Abstract
The study aims to characterise the species identification and antimicrobial susceptibility testing (AST) results of Nocardial isolates from adult patients across major public hospitals in Queensland, Australia, over a 15-year period. A multi-centre retrospective observational study of Nocardia sp. isolates was conducted from 7 major public hospitals in Queensland, Australia, over a 15-year period. Clinical samples from patients aged ≥ 18 years that isolated Nocardia sp. were included. Demographic and clinical data were collected, along with species identification and AST results. Overall, 484 Nocardia sp. were isolated. Most patients were male (297, 61%) with a mean (IQR) age of 60 (51–75) and a median (IQR) Charlson Comorbidity Index of 4 (2–6). Of these, 239 (49%) patients were immunosuppressed. Organisms were most frequently isolated from sputum (174, 36%), and superficial swabs (102, 21%). Patients presented with pulmonary infections (165, 35%) and superficial skin and soft tissue infections (87, 18%) most commonly. One hundred (21%) isolates were deemed pulmonary colonisation and were not treated. Of the speciated organisms, N. nova complex was the most common (93, 19%), followed by N. farcinica complex (79, 16%). Organisms were reliably susceptible to linezolid (240/245, 98%), amikacin (455/470, 97%), and trimethoprim/sulfamethoxazole (459/476, 96%), but less so to imipenem (243/472, 51%) and ceftriaxone (261/448, 58%). This is the largest Australian description of Nocardia sp. to date. Given antimicrobials are often commenced prior to AST results and sometimes even speciation, characterisation of local species and antibiogram data is important to guide empiric choices and local guidelines.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Nocardia, a ubiquitous environmental Gram-positive actinomycete, is a rare cause of human disease, but infections are frequently associated with high morbidity and mortality [1]. The worldwide incidence of Nocardia sp. infections has been increasing, largely due to an improvement in detection and identification methods, but also due to the increasing number of immunocompromised patients [2]. A move from biochemical to molecular identification methods has improved species discrimination but resulted in significant taxonomic change and revision of our understanding of this important pathogen [3]. Species previously grouped together were recognised to be distinct with correspondingly dissimilar drug susceptibility patterns [3].
Nocardiosis typically presents as one of three clinical syndromes: cutaneous infection, which often occurs in immunocompetent individuals after direct inoculation and may present as either skin and soft tissue infection (SSTI), deeper bone and joint infections such as osteomyelitis and septic arthritis, pulmonary infection, and disseminated disease which can include bloodstream infection and central nervous system (CNS) involvement [4]. Disseminated disease is usually seen in immunocompromised patients — typically in those with impaired cellular immunity — and is associated with a mortality rate of up to 40% [5].
Treatment is made more difficult by the frequent delays in diagnosis and, even when the diagnosis is established, limited treatment options for the isolated Nocardia species [6]. While intraspecies antimicrobial susceptibility is predictable for individual species, there is significant variability between species, and marked differences in their geographical distribution [7]. It is therefore essential to define locally prevailing Nocardia species precisely to inform optimal empiric antibiotic selection.
In the last decade, there have been 3 reviews of nocardiosis in the different states of Australia, but given the varying geography and climates across the country, these are not necessarily generalisable. Two studies — one in New South Wales and a single-site study in Victoria — examined 270 and 68 isolates respectively; these were published in 2020 and 2018 respectively [8, 9]. A 2016 study conducted in the Northern Territory, which is more similar to Queensland in its climate, examined 61 isolates [10]. However, the last reported study of nocardiosis in Queensland was published in 1992 and examined only 102 patients [11]. This study represents the largest Australian series of Nocardia sp. isolates to date.
Hence, this study aimed to describe the species and antimicrobial susceptibility testing (AST) results of Nocardia isolates identified in adults treated at major public hospitals in Queensland, Australia, over a 15-year period. It was hoped that this data might be used to inform the optimal selection of empirical therapies of patients diagnosed with Nocardiosis in the region.
Methods
Ethics approval
Research was conducted in accordance with the Declaration of Helsinki and national and institutional standards. Ethical approval was provided by the Metro South Health Human Research Ethics Committee (LNR/2019/QMS/48872). As the data was retrospective and de-identified, the Committee waived the requirement for informed consent. Site-specific approval was obtained for each participating hospital.
Data collection
This retrospective multi-centre study was conducted at 7 large public hospitals in Queensland, Australia, a state of 4.7 million people spread across 1.9 million km2 (supp Fig. 1). Climate is variable across the state with arid regions in west, tropical in the north, and subtropical in the south.
Clinical samples from patients aged ≥ 18 years collected between 1st January 2004 and 31st December 2018 that isolated Nocardia sp. in culture were included in the analysis. All isolates were initially identified in these hospitals’ microbiology laboratories and were then referred to a state-wide tertiary reference laboratory (Queensland Mycobacterium Reference Laboratory) for genus confirmation, speciation, and antimicrobial susceptibility testing. Episodes where the same organism was isolated from the same patient within 12 months were excluded, as were specimens in which the site of collection was not documented. Demographic, clinical, and laboratory data were collected to determine the site of infection, the Nocardia species and AST.
SSTI was defined as cellulitis or cutaneous abscess. Bone and joint infection included osteomyelitis and septic arthritis. Disseminated disease was bloodstream infection or ≥ 2 discontiguous sites of involvement (not including multiple SSTI sites). Central nervous system (CNS) infection included meningitis and cerebral abscesses. Pulmonary disease included abscesses, nodular disease, and pneumonia. Pulmonary colonisation was defined as any Nocardia spp. isolated from pulmonary samples where patients recovered without treatment. Patients were classified as immunosuppressed if they had a current history of malignancy, chemotherapy, transplantation, human immunodeficiency virus, or were taking any systemic immunosuppressive medications (oral corticosteroids at any dose, calcineurin inhibitors, mammalian target of rapamycin inhibitors, or antimetabolites). Comorbidity was quantified using the Charlson Comorbidity Index.
Isolates and antimicrobial susceptibility testing
Primary isolation of putative Nocardia sp. was conducted by local microbiology laboratories using standard media applicable to the specimen type. In some cases, prolonged incubation was used to enhance detection. Isolates were referred to the reference laboratory where identification and antimicrobial susceptibility testing was performed. The oldest isolates in this study were confirmed phenotypically as Nocardia sp. and speciated using patterns of susceptibility. AST was performed using disc susceptibility testing as advocated by Wallace et al. and recorded as susceptible, intermediate, or resistant [12]. MIC data was not available for these isolates. Identification of newer isolates was first attempted by mass spectrometry. Where possible, species were grouped into complexes as defined by Conville et al. [3].
Evolution of methods saw the introduction of 16S rDNA, Sanger sequencing for species identification (2007), and broth microdilution (BMD) for AST using the commercial Sensititre assay (Trek Diagnostics/ThermoFisher) (2011). Despite 16S rDNA sequencing, with reference matching to NCI BLAST, some isolates failed to meet criteria for species reporting (≥ 99.6%) match as per Clinical and Laboratory Standards Institute (CLSI) recommendations [13] and were reported to genus level only. Methodology and interpretation of BMD minimum inhibitory concentrations were determined using CLSI standards [13]. Due to differences in AST methodology and reporting over the time period and laboratories, the numbers of isolates tested for each antibiotic varied.
Data management and analysis
Data were entered into an electronic database (RedCAP) and analysed using statistical software (Rstudio). Descriptive statistics were performed and are presented as the absolute number (%), mean ± standard deviation or median (interquartile range) as appropriate. Analysis for trend was conducted using an extension of the Wilcoxon rank-sum test [14].
Results
Demographics
Over the 15-year period, 484 Nocardia sp. isolates were identified in clinical samples at the study hospitals (supp Fig. 1). Most patients were male (297, 61%), with a median age of 66 (IQR 51–75), and a median (IQR) Charlson Comorbidity Index of 4 (2–6). Almost half (239, 49%) of patients were immunosuppressed, most commonly due to steroid use (supp Fig. 2). There was no trend towards increasing number of Nocardia cases over time (p = 0.21) (supp Fig. 5).
Site of infection
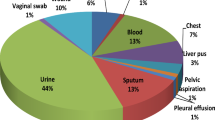
Organisms were isolated from a variety of samples, most commonly sputum (174, 36%), and superficial swabs (102, 21%) (Table 1). This corresponded to 165 (34%) pulmonary infections and 87 (18%) skin and soft tissue infections (Table 2; supp Fig. 3): the two leading clinical syndromes observed. 100 (21%) sputum isolates were thought to represent colonisation.
Speciation
Of the isolates that were able to be identified to species level, N. nova complex was the most common (93, 19%), followed by N. farcinica complex (79, 16%) (supp Fig. 4). Among patients with pulmonary infection, N. cyriacigeorgica complex was the most common isolate (39, 24%) followed by N. nova complex (34, 21%). Of those deemed to be colonised, N nova complex was most common (32, 32%), followed by N. farcinica complex (21, 21%) (Table 2). In bronchial washings, N. cyriacigeorgica complex was most frequently isolated (20, 22%), and in sputum, this was N. nova complex (47, 27%) (Table 1).
N. brasiliensis was the predominant organism (51, 59%) isolated from those with skin and soft tissue infections (SSTI) followed by N. nova (9, 10%) (Table 2). This corresponded to the predominant isolation of N. brasiliensis from superficial swabs (47, 46%) (Table 1). Identified species causing disseminated disease were mostly N. farcinica complex (15, 27%) (Table 2).
Significant numbers of isolates were not able to be identified to the species level (Tables 1–2). There was no trend towards improved speciation of isolates over time (p = 0.06) (supp Fig. 4). Despite the 12 month limit imposed between each episode, 29 samples from 11 patients grew the same organism more than 12 months apart. These were largely deemed pulmonary colonisation (21, 72%).
Antimicrobial susceptibility testing results
Pooled sensitivities of all Nocardia sp. isolated demonstrated that the isolates were most frequently susceptible to linezolid (240/245, 98%), followed by amikacin (455/470, 97%), then trimethoprim/sulfamethoxazole (TMP/SXT; 459/476, 96%). Conversely, only 51% of the isolates were susceptible to imipenem (243/472), and ceftriaxone (261/448, 58%) (Table 3). Linezolid susceptibility was only available for 50% of the isolates due to changes in AST reporting over the study period.
N. nova complex, the predominant isolate in this dataset, was reliably susceptible to TMP/SXT (93/93, 100%), amikacin (93/93, 100%), linezolid (40/40, 100%), and imipenem (90/93, 97%), but less so to ceftriaxone (60/88, 68%) (supp Fig. 6). N. farcinica complex, the second most commonly speciated isolate, was similarly susceptible to TMP/SXT (72/77, 94%), amikacin (74/76, 95%), and linezolid (30/31, 97%), but less so to ceftriaxone (10/73, 14%) and imipenem (31/76, 41%) (supp Fig. 7). Furthermore, species-specific AST breakdowns are presented in supplementary Figs. 8–11.
Of the isolates unable to be identified to species level, the majority were once again susceptible to linezolid (51/51, 100%), TMP/SXT (74/77, 96%), and amikacin (74/76, 99%), whereas susceptibility to ceftriaxone (39/75, 52%) and imipenem (47/77, 61%) was more variable.
Discussion
Since the advent of molecular techniques, taxonomy of Nocardia sp. has expanded rapidly and many new species have been identified [3]. These have been grouped into complexes based on genotypic and phenotypic similarity, but clinical correlation based on these complexes is still nebulous [3]. Despite the use of 16S rDNA sequencing, a large proportion of the isolates were still only able to be identified down to the genus level (80, 17%) and the percentage of isolates identified only to species level did not decrease over time. This is likely in part due to lack of research in the field and limited availability of sequencing data in the NCBI database. Thus, species identification remains difficult despite progression in molecular techniques. This has the potential to delay de-escalation or optimal antimicrobial prescribing, given the variation in susceptibility patterns over the different species.
Species distribution of our isolates differed from previous international studies highlighting the geographical variation and environmental requirements for different Nocardia sp. Of the isolates identified to species level in our study, N. nova complex was the most commonly isolated, followed by N. farcinica complex, N. cyriacigeorgica complex, then N. brasiliensis. The high proportion of N. farcinica complex isolates in our study is of concern given this species is known to be particularly resistant and is increasingly recognised as a nosocomial pathogen [15]. Hence, awareness of Nocardia sp. as a pathogen remains important, especially given increases in comorbidity amongst patients, and degree of immunosuppression with the advent of more immunologically challenging donor-recipient matches and expansion of novel treatment of malignancies. Our study did also contain a large proportion of N. brasiliensis species (62, 13%) which is in keeping with its role as major SSTI pathogen as well as previous observations that this species is more frequently found in tropical or subtropical climates [16].
In contrast, a Japanese study found a predominance of N. farcinica complex, followed by N. cyriacigeorgica complex and N. brasiliensis [17]. Studies conducted in Spain and USA found N. cyriacigeorgica complex followed by N. nova complex were the most prevalent species [18, 19]. A French study of 793 isolates found a clear predominance of N. farcinica and abscessus complex [20]. While climate is likely to play a large role, part of the difference could be due to the over-representation of pulmonary isolates in our study which could reflect persistent colonisation despite treatment.
Other studies have described a change in the species distribution in keeping with changing patient and environmental factors: in particular, Lebeaux et al. documented an increase in the prevalence of N. farcinica complex over time, which they postulated was secondary to an increase in infections amongst solid organ transplant recipients [15, 20]. A similar time trend was not observed in our study.
Australian therapeutic guidelines recommend trimethoprim/sulfamethoxazole (TMP/SXT) for cutaneous disease, TMP/SXT with either ceftriaxone or linezolid for pulmonary disease, and TMP/SXT and linezolid, with amikacin, imipenem, or meropenem for severe or disseminated disease [21]. Most of our isolates were susceptible to most of these antimicrobials with the notable exceptions of ceftriaxone and imipenem. The high rate of resistance to ceftriaxone is likely contributed by the proportion of N. farcinica complex in our dataset, which has high rates of resistance to beta-lactam antibiotics [22]. This is of concern given that ceftriaxone is the recommended backbone of therapy for pulmonary disease according to our local guidelines [16, 21]. One study previously reported increasing TMP/SXT resistance which was suggested to be plasmid mediated, but our isolates had low rates of resistance (< 5%), as consistent with other recent studies [17, 20, 23,24,25].
In Australian guidelines, meropenem is recommended in the initial treatment of severe or disseminated disease, but our local laboratories are unable to test specifically for it. Clinicians sometimes use imipenem results to extrapolate meropenem susceptibility, but this is not recommended by our reference laboratory as the validity of this is questionable [26]. In addition, the high rates of resistance to imipenem found here suggest that amikacin, which is recommended as an alternative, may be more effective, and should be considered as first-line in disseminated disease over meropenem [21]. However, the use of amikacin may be limited by its significant side-effect profile. For example, pre-existing renal impairment may be a contraindication in vulnerable transplant patients, as would vestibular and ototoxicity in patients with CNS disease.
Our study is limited by its retrospective nature, the high proportion of isolates not able to speciated, and the absence of MIC data with the older isolates, where susceptibilities were recorded categorically (i.e. susceptible, intermediate, and resistant, without specific MIC data due to the use of disc diffusion AST). In addition, changes in speciation and AST methods over the timeframe could reduce reliability of identification and AST. Due to the retrospective nature of this data, it was not possible to discern the different methods of testing, nor was it possible to retest the isolates. Our study also had a predominance of pulmonary isolates, which could also reflect persistent colonisation and ease of sampling. Definitions of immunosuppression included steroid use at any dose, which could overestimate the number of patients who were deemed immunosuppressed. However, acknowledging these limitations, this is still the largest Australian study of Nocardia sp. to date and provides understanding of the local milieu and antimicrobial susceptibility profiles. Given the wide geographical variation in species, this is important to understand in order to guide empiric treatment recommendations, especially given ongoing difficulty in identifying isolates to species level despite advances in molecular technology.
In light of the antimicrobial susceptibility profiles, review of local guidelines should be considered, especially with regards to empiric ceftriaxone and meropenem. Investigation into clinical treatment and outcomes in these settings is necessary to help guide this. Further work should also be undertaken to enable accurate identification and susceptibility testing, for example, through expansion of sequencing databases. It is likely that evolution of mass spectrometry techniques as well as the application of Next Generation Sequencing techniques will allow greater accuracy of species identification as well as potential resistance prediction.
Data availability
The datasets generated during and/or analysed during the current study are not publicly available due to patient confidentiality and privacy reasons, but may be available from the corresponding author on reasonable request.
References
Wilson JW (2012) Nocardiosis: updates and clinical overview. Mayo Clin Proc 87(4):403–407
Restrepo A, Clark NM (2019) Nocardia infections in solid organ transplantation: guidelines from the Infectious Diseases Community of Practice of the American Society of Transplantation. Clin Transplant 33(9):e13509
Conville PS, Brown-Elliott BA, Smith T, Zelazny AM (2018) The complexities of Nocardia taxonomy and identification. J Clin Microbiol 56(1):e01419–e01417
Saubolle MA, Sussland D (2003) Nocardiosis: review of clinical and laboratory experience. J Clin Microbiol 41(10):4497–4501
Ambrosioni J, Lew D, Garbino J (2010) Nocardiosis: updated clinical review and experience at a tertiary center. Infection 38(2):89–97
Brown-Elliott BA, Wallace RJ Jr. (2017) In vitro susceptibility testing of Tedizolid against isolates of Nocardia. Antimicrob Agents Chemother 61(12):e01537–17
Uhde KB, Pathak S, McCullum I Jr, Jannat-Khah DP, Shadomy SV, Dykewicz CA, Clark TA, Smith TL, Brown JM (2010) Antimicrobial-resistant nocardia isolates, United States, 1995–2004. Clin Infect Dis 51(12):1445–1448
Tan YE, Chen SCA, Halliday CL (2020) Antimicrobial susceptibility profiles and species distribution of medically relevant Nocardia species: results from a large tertiary laboratory in Australia. J Glob Antimicrob Resist 20:110–117
Paige EK, Spelman D (2019) Nocardiosis: 7-year experience at an Australian tertiary hospital. Intern Med J 49(3):373–379
McGuinness SL, Whiting SE, Baird R, Currie BJ, Ralph AP, Anstey NM, Price RN, Davis JS, Tong SY (2016) Nocardiosis in the Tropical Northern Territory of Australia, 1997-2014. Open Forum Infect Dis 3(4):ofw208
Georghiou PR, Blacklock ZM (1992) Infection with Nocardia species in Queensland A review of 102 clinical isolates. Med J Aust 156(10):692–697
Wallace RJ Jr, Steele LC, Sumter G, Smith JM (1988) Antimicrobial susceptibility patterns of Nocardia asteroides. Antimicrob Agents Chemother 32(12):1776–1779
CLSI (2018) Interpretive criteria for identification of bacteria and fungi by targeted DNA sequencing. CLSI guideline MM18 Wayne, PA: Clinical and Laboratory Standards Institute
Cuzick J (1985) A Wilcoxon-type test for trend. Stat Med 4(1):87–90
Schaal KP, Lee H-J (1992) Actinomycete infections in humans — a review. Gene 115(1):201–211
Brown-Elliott BA, Brown JM, Conville PS, Wallace RJ Jr (2006) Clinical and laboratory features of the Nocardia spp. based on current molecular taxonomy. Clin Microbiol Rev 19(2):259–282
Toyokawa M, Ohana N, Ueda A, Imai M, Tanno D, Honda M, Takano Y, Ohashi K, Saito K, Shimura H (2021) Identification and antimicrobial susceptibility profiles of Nocardia species clinically isolated in Japan. Sci Rep 11(1):16742
Valdezate S, Garrido N, Carrasco G, Medina-Pascual MJ, Villalón P, Navarro AM, Saéz-Nieto JA (2017) Epidemiology and susceptibility to antimicrobial agents of the main Nocardia species in Spain. J Antimicrob Chemother 72(3):754–761
Schlaberg R, Fisher MA, Hanson KE (2014) Susceptibility profiles of Nocardia isolates based on current taxonomy. Antimicrob Agents Chemother 58(2):795–800
Lebeaux D, Bergeron E, Berthet J, Djadi-Prat J, Mouniee D, Boiron P, Lortholary O, Rodriguez-Nava V (2019) Antibiotic susceptibility testing and species identification of Nocardia isolates: a retrospective analysis of data from a French expert laboratory, 2010–2015. Clin Microbiol Infect 25(4):489–495
Therapeutic Guidelines. Melbourne: therapeutic guidelines limited. https://www.tg.org.au. Accessed Sept 2022
Wallace RJ Jr, Tsukamura M, Brown BA, Brown J, Steingrube VA, Zhang YS, Nash DR (1990) Cefotaxime-resistant Nocardia asteroides strains are isolates of the controversial species Nocardia farcinica. J Clin Microbiol 28(12):2726–2732
Valdezate S, Garrido N, Carrasco G, Villalón P, Medina-Pascual MJ, Saéz-Nieto JA (2015) Resistance gene pool to co-trimoxazole in non-susceptible Nocardia strains. Front Microbiol 6:376
Lai CC, Liu WL, Ko WC, Chen YH, Tang HJ, Huang YT, Hsueh PR (2011) Antimicrobial-resistant nocardia isolates, Taiwan, 1998–2009. Clin Infect Dis 52(6):833–835
Wang H, Zhu Y, Cui Q, Wu W, Li G, Chen D, Xiang L, Qu J, Shi D, Lu B (2022) Epidemiology and antimicrobial resistance profiles of the Nocardia species in China, 2009 to 2021. Microbiol Spectr 10(2):e0156021
Brown-Elliott BA, Killingley J, Vasireddy S, Bridge L, Wallace RJ Jr (2016) In vitro comparison of Ertapenem, Meropenem, and Imipenem against isolates of rapidly growing mycobacteria and Nocardia by use of broth nicrodilution and Etest. J Clin Microbiol 54(6):1586–1592
Acknowledgements
The authors acknowledge the QMRL and staff who conducted microbiological identification and susceptibility testing on these isolates.
Author information
Authors and Affiliations
Contributions
All authors contributed to the study conception, design, and data collection. Material preparation was performed by Evan Bursle and Andrew Henderson. Analysis was performed by Beatrice Sim, Luke Aaron, Evan Bursle, and Andrew Henderson. The first draft of the manuscript was written by Beatrice Sim, and all authors commented on previous versions of the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Consent to participate
Not applicable.
Consent for publication
Not applicable.
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Sim, B.Z., Aaron, L., Coulter, C. et al. A multi-centre retrospective study of Nocardia speciation and antimicrobial susceptibility in Queensland, Australia. Eur J Clin Microbiol Infect Dis 42, 339–345 (2023). https://doi.org/10.1007/s10096-022-04542-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10096-022-04542-0