Abstract
We analysed mutations in katG, inhA and rpoB genes, and isoniazid phenotypic resistance levels in Mycobacterium tuberculosis isolates from drug-resistant TB patients from São Paulo state, Brazil. Isolates resistant to the critical concentration of isoniazid in MGIT (0.1 µg/mL) were screened for mutations in katG 315 codon, inhA promoter region and rpoB RRDR by MTBDRplus assay and subjected to determination of isoniazid resistance levels by MGIT 960. Discordances were resolved by Sanger sequencing. Among the 203 isolates studied, 109 (54%) were isoniazid-monoresistant, 47 (23%) MDR, 29 (14%) polydrug-resistant, 12 (6%) pre-XDR and 6 (3%) XDR. MTBDRplus detected isoniazid mutations in 75% (153/203) of the isolates. Sequencing of the entire katG and inhA genes revealed mutations in 18/50 wild-type isolates by MTBDRplus (10 with novel mutations), resulting in a total of 32/203 (16%) isolates with no mutations detected. 81/83 (98%) isolates with katG 315 mutations alone had intermediate resistance. Of the 66 isolates with inhA C-15T mutation alone, 51 (77%) showed low-level, 14 (21%) intermediate and 1 (2%) high-level resistance. 5/6 (83%) isolates with mutations in both katG and inhA had high-level resistance. Inferred mutations corresponded to 22% (16/73) of all mutations found in rpoB. Mutations detected in katG regions other than codon 315 in this study might be potential new isoniazid resistance markers and could explain phenotypic resistance in some isolates without katG and inhA classic mutations. In our setting, 16% of isoniazid-resistant isolates, some with high-level resistance, presented no mutations either in katG or inhA.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Drug resistance in Mycobacterium tuberculosis is due mainly to mutations in its genome. More than 95% of rifampicin (RIF) resistance conferring mutations occur in an 81-bp region (codons 507–533) of rpoB gene, known as the RIF resistance determining region (RRDR) [1]. As for isoniazid (INH) resistance, the most common associated mutations occur in codon 315 of katG and, in the promoter region of inhA, mostly at position − 15 [2]. Globally, 64% of phenotypic INH resistance is associated with katG 315 mutations, while mutations in inhA − 15 are observed in 19% of INH-resistant isolates [3, 4].
katG gene mutations have been associated with at least moderate- to high-level INH resistance, above the usual tested concentrations of 0.2 and 1 µg/mL in solid medium and 0.1 and 0.4 µg/mL in MGIT 960 liquid medium. katG S315T, the most frequent mutation found in this gene, is associated with minimum inhibitory concentrations (MICs) ranging from 2 to > 10 µg/mL in MGIT 960, while inhA promoter mutations have been associated with low-level resistance (0.1 < MIC ≤ 1 µg/mL) in MGIT 960 [5].
Combinations of mutations in katG 315 and inhA promoter are known to confer high-level resistance (MIC > 10 µg/mL) [6]. However, previous studies have shown phenotypic heterogeneity of INH resistance-associated mutations, with a wide range of MICs observed among INH-resistant isolates [6,7,8].
A mutation leading to a decrease in drug susceptibility should not always exclude an anti-tuberculosis (TB) drug from a treatment regimen, since low-level resistance does not necessarily imply clinical resistance [5]. This is the case for INH, which can still be effective at elevated doses in patients infected by M. tuberculosis strains with low-level resistance, i.e. strains presenting specific inhA promoter mutations without katG mutations [4, 9]. On the other hand, isolates with katG 315 mutations only, particularly the high-confidence S315T and S315N, should be reported as high-level INH-resistant regardless of their MIC, so that INH is not used in the treatment [9]. Thus, patients harbouring isolates with mutations in inhA promoter only might be beneficiated by a treatment regimen with double dose of INH (600 mg) [10].
Here, we investigated genetic mutations in rpoB, katG and inhA genes of M. tuberculosis clinical isolates with different drug resistance profiles and associated katG and inhA mutations with INH phenotypic levels of resistance.
Methods
Setting and study population
The state of São Paulo is the most populous in Brazil, presenting a TB incidence of 40.8/100,000 in 2018 [11]. Adolfo Lutz Institute is the reference laboratory for São Paulo, receiving annually 8000 isolates of mycobacteria for species identification and drug susceptibility testing (DST). M. tuberculosis isolates from patients under risk of drug-resistant TB are subjected to DST by MGIT 960 (Becton Dickinson, MD, USA). These patients include smear-positive cases after 2 months on TB treatment, contacts of drug-resistant TB patients, retreatment cases, immunosuppressed persons, alcohol/illicit drugs abusers, healthcare workers, homeless individuals, indigenous, immigrants, inmates and inpatients.
Study design
This was a prospective study including all M. tuberculosis isolates referred to Adolfo Lutz Institute between January and December 2016, and resistant to 0.1 μg/mL of INH (along or not with other drugs) by MGIT 960 DST. Only one isolate per patient was included, except for patients with isolates showing different mutations along the study period.
Routine drug susceptibility testing
DST for first-line drugs was performed on MGIT 960 using SIRE kit (Becton Dickinson, MD, USA). Critical concentrations tested were 1 μg/mL, 0.1 μg/mL, 1 μg/mL and 5 μg/mL of streptomycin, INH, RIF and ethambutol, respectively. The isolates were also tested against second-line drugs. Critical concentrations were 1 µg/mL of amikacin, 2.5 µg/mL of capreomycin and kanamycin, 2 µg/mL of ofloxacin and 1 µg/mL of levofloxacin [12,13,14]. Isolates were classified as INH-monoresistant when they presented INH resistance only; as polydrug-resistant when they were resistant to INH and other first-line drug(s), except RIF; as MDR when they presented resistance at least to INH and RIF; as pre-extensively drug-resistant (pre-XDR) when they were MDR and resistant to at least one of injectable second-line drugs or fluoroquinolones; and as extensively drug-resistant (XDR) when they were MDR and resistant to both injectables and fluoroquinolones. On January 2021, the WHO updated the definitions of both pre-XDR- and XDR-TB, which classify pre-XDR-TB as the disease caused by an MDR or RIF-resistant strain which is also resistant to a fluoroquinolone, and XDR-TB as TB caused by an MDR or RIF-resistant strain which is also resistant to a fluoroquinolone and bedaquiline and/or linezolid (https://www.who.int/news/item/27-01-2021-who-announces-updated-definitions-of-extensively-drug-resistant-tuberculosis). Here, we applied the previous definitions of pre-XDR- and XDR-TB, because this study was performed before these changes.
Determination of INH resistance levels
The INH-resistant isolates at 0.1 µg/mL were tested by MGIT 960 against 1 µg/mL, 3 µg/mL and 10 µg/mL of INH [6, 13, 15]. The tests were incubated into MGIT instrument according to Rüsch-Gerdes et al. (2006) [16]. Isolates with no growth at 1 µg/mL were considered low-level-resistant (0.1 < MIC ≤ 1 µg/mL), while the ones growing at 1 µg/mL or at both 1 µg/mL and 3 µg/mL, but not at 10 µg/mL (1 < MIC ≤ 3 µg/mL or 3 < MIC ≤ 10 µg/mL) were considered having intermediate resistance, and the ones growing at all concentrations were considered high-level-resistant (MIC>10 µg/mL) [6, 15]. Despite the clinical breakpoint of 0.4 µg/mL of INH recommended by the Clinical & Laboratory Standards Institute (CLSI) for definition of low-level resistance, existing data on correlation of inhA promoter mutations and INH MICs are not enough to support this clinical breakpoint, according to the WHO [9]. Therefore, based on previous studies on MGIT 960 liquid medium, we decided to adopt the concentration of 1 µg/mL as a breakpoint for low-level INH resistance [6, 7, 15].
DNA extraction
DNA of the isolates was extracted by thermal lysis. One milliliter of the culture in MGIT was incubated at 95 °C for 20 min and frozen at −20 °C twice [17]. The lysates were kept at −20 °C until MTBDRplus and Sanger sequencing were performed.
Detection of mutations
All the isolates were subjected to GenoType MTBDRplus v. 2.0 (Hain Lifescience, Nehren, Germany) for detection of mutations in katG 315 codon, in inhA promoter region, and in rpoB RRDR, as described elsewhere [18, 19]. Sanger sequencing was performed for isolates with discordant results between DST and MTBDRplus, such as those with inferred mutations (wild type and mutant probes absent) and those which were phenotypically resistant to INH but presented no mutations by MTBDRplus. The inhA promoter was amplified and sequenced with primers inhA-1 and inhA-2 (positions −168 to 80 in reference to start codon) [20]. The whole inhA and katG genes were sequenced with primers inhA3 and inhA4 (positions −13 to 379), inhA3F and inhA5R (positions 381 to 840), and forward and reverse primers katG-P4, katG-P5, katG-P6, katG-P7 and katG-P8 (positions −135 to 2202 of katG plus 431 nucleotides after the end of the gene) [21]. Primers rpoB-1 and rpoB-2 were used to amplify and sequence a 350-bp fragment of rpoB encompassing the RRDR [22]. Each PCR reaction included 12.5 μL of PrimeSTAR Max DNA Polymerase (Takara Bio, Shiga, Japan), 5 pmol of primers for inhA promoter and katG, 10 pmol of primers for inhA coding region and rpoB, 2 μL of DNA template and PCR grade water for a final volume of 25 μL. Amplification comprised 30 cycles of 98 °C for 10 s, 55 °C for 5 s and 72 °C for 20 s. Amplicons were purified with ExoSAP-it (Affymetrix, CA, USA) and sequenced with ABI 3130×L Genetic Analyzer and the BigDye Terminator version 3.1 Kit (Applied Biosystems, CA, USA). Sequences were analysed using BioEdit v7.2.5 software and the web-based MUBII-TB-DB and BLAST tools [19].
Nucleotide sequence accession numbers
The sequences with novel mutations found in katG and inhA were deposited in GenBank (http://www.ncbi.nlm.nih.gov/) under the following accession numbers: MW809517-MW809528 and MW815499.
Data analysis
Fisher’s exact test was used for comparisons between proportions. The significance threshold was set at 0.05. Statistical analyses were performed using the web-based OpenEpi program [23].
Results
Phenotypic resistance to first- and second-line drugs
There were 203 isolates resistant to INH at 0.1 µg/mL by MGIT 960, in the study period. Most of the isolates were obtained from sputum (n = 194, 95%), followed by bronchoalveolar lavage (n = 4, 2%), tracheal aspirate (n = 2, 1%), pleural fluid (n = 2, 1%) and bone tissue (n = 1, 1%). These isolates belonged to 200 patients, as 3 patients had 2 isolates presenting different mutations (2 polydrug-resistant isolates from one patient, and 2 MDR isolates from the other two patients). Among the 203 isolates, 109 (54%) were INH-monoresistant, 47 (23%) MDR, 29 (14%) polydrug-resistant, 12 (6%) pre-XDR (all of them resistant to fluoroquinolones) and 6 (3%) XDR.
INH resistance levels
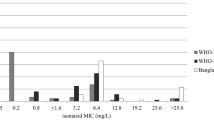
Among the 203 isolates tested, 75 (37%) were INH low-level-resistant (0.1 < MIC ≤ 1 µg/mL), 107 (53%) showed intermediate resistance [34 (17%) with 1 < MIC ≤ 3 µg/mL, and 73 (36%) with 3 < MIC ≤ 10 µg/mL], and 21 (10%) were high-level-resistant (MIC > 10 µg/mL). Most of the 75 isolates with low-level resistance were INH-monoresistant (88%, n = 66) (Fig. 1). Interestingly, INH-monoresistant isolates accounted for 33% (n = 7) of high-level-resistant isolates (Fig. 1).
Mutations detected by MTBDRplus
Regarding INH-associated mutations, of the 203 isolates tested, 82 (40%) had mutations only in katG 315: 79 (39%) with S315T1 (AGC > ACC), 2 (1%) with S315T2 (AGC > ACA) and 1 with an inferred mutation. Sixty-six isolates (33%) showed mutations only in inhA, all with C-15T. Mutations in both genes were detected in 5 (3%) isolates: katG S315T1 + inhA inferred (n = 2), katG inferred + inhA C-15T (n = 2) and katG S315T1 + inhA C-15T (n = 1) (Fig. 2). A considerable proportion of INH-resistant isolates (25%, n = 50) presented no mutations in any of these targets.
Figure 2 shows the distribution of katG 315 and inhA promoter mutations according to the phenotypic resistance of the isolates. While the katG S315T mutation alone was found mainly among INH-monoresistant (n = 22, 27%), MDR (n = 29, 36%) and polydrug-resistant (n = 19, 24%) isolates, the majority of the isolates (n = 54, 82%) with only inhA C-15T mutation were INH-monoresistant. Most of the isolates showing no mutations either in katG 315 and inhA promoter were INH-monoresistant (n = 32, 64%); however, 18% (n = 9) and 14% (n = 7) of the isolates without katG 315 and inhA promoter mutations were MDR and polydrug-resistant, respectively. Absence of these mutations was also detected in 1 pre-XDR and 1 XDR isolates.
As for RIF-associated mutations, most of the isolates (64%, n = 130/203) did not present any mutation in rpoB (all of them were phenotypically RIF-susceptible, 105 INH-monoresistant and 25 polydrug-resistant). Among the 73 isolates with rpoB mutations, the most prevalent was S531L mutation, which occurred alone in 51 (70%) isolates: 37 MDR, 11 pre-XDR and 3 XDR. Four (6%) isolates presented D516V mutation (2 MDR and 2 XDR), and 1 (1%) pre-XDR isolate had H526Y mutation. One (1%) MDR isolate presented both S531L and D516V mutations. rpoB inferred mutations occurred in 16 (22%) isolates, of which 4 were INH-monoresistant, 4 polydrug-resistant, 7 MDR and 1 pre-XDR.
Mutations detected by Sanger sequencing
Fifty-five isolates were subjected to katG and inhA sequencing (Table 1). Among the 50 isolates with no mutations detected either in katG 315 and inhA promoter by MTBDRplus, 18 (36%) presented mutations by sequencing, while 32 (64%) continued without mutations. Point mutations were the most prevalent in katG and occurred in isolates with different levels of resistance to INH. Four isolates presented heteroresistance (defined as the coexistence of susceptible and resistant organisms to a drug in the same M. tuberculosis population): 1 with katG S315R mutation (which was not detected by MTBDRplus) and 3 with mutations in other codons. The 3 isolates with deletions presented high levels of resistance: A122G − 3 < MIC ≤ 10 µg/mL, R463L + A478R − MIC > 10 µg/mL and K590_None − MIC > 10 µg/mL. The latter isolate presented also a silent mutation in inhA (G76G) (Table 1).
Sequencing of the 2 isolates with katG S315T and inhA promoter inferred mutation revealed had inhA G-17 T mutation, and both showed high-level INH resistance. Of the 3 isolates with katG inferred mutations, 2 had S315N mutation (one presented heteroresistance and the other presented also the katG A506T mutation), and the other had S315G mutation alone (Table 1).
rpoB RRDR sequencing was performed for the 16 isolates with inferred mutations by MTBDRplus (Table 2). The most frequently mutated codon was 526, with 6 isolates presenting mutations in this target, 5 of them phenotypically RIF-susceptible (2 with H526L, 2 with H526N and 1 with H526S).
INH resistance levels and mutations
Considering the results from MTBDRplus and sequencing together, we assessed the association between INH resistance levels and mutations in katG and inhA for the 203 isolates (Table 3) (Fig. 3). Overall, 99 (49%) isolates presented mutations in katG only, of which 83 (84%), occurred in 315. All these 83 isolates showed intermediate- to high-level INH resistance. Among the 16 isolates with mutations in katG codons other than the 315, only 4 had low-level resistance. Of the 66 (33%) isolates with inhA C-15T mutation only, 51 (77%) showed low-level resistance, 14 (21%) had intermediate resistance and 1 (2%) showed high-level resistance. Among the 6 isolates with mutations in both katG and inhA, 5 (83%) were high-level INH-resistant, while 1 (17%) was intermediate. Even after sequencing of entire katG and inhA, no mutations were detected in 32/203 (16%) isolates, of which 20 were low-level-resistant, 6 intermediate and 6 high-level INH-resistant.
Discussion
We investigated mutations related to INH and RIF resistance in 203 M. tuberculosis clinical isolates with different drug resistance profiles and associated mutations found in katG and inhA with INH phenotypic levels of resistance. MTBDRplus kit was able to identify INH resistance-associated mutations in 75% (153/203) of the isolates. Still, 25% (50/203) of the isolates phenotypically resistant to INH would be diagnosed by the test as not having mutations, which means that resistance was not detected. Analysing the entire katG and inhA genes by sequencing enhanced the molecular diagnosis of INH resistance, as mutations in other codons of katG were found in 17/50 isolates (plus one S315R mutant), resulting in a total of 32/203 (16%) isolates without mutations in these targets.
We observed that low-level INH resistance was detected more frequently in INH-monoresistant isolates (66/109, 61%) than in MDR (4/47, 9%, p < 0.0001) and polydrug-resistant (5/29, 17%, p = 0.0002) isolates. Considering MTBDRplus results only, frequencies of katG S315T mutation in MDR (29/47, 62%, p < 0.0001), and polydrug-resistant (19/29, 66%, p < 0.0001) isolates were significantly higher than in INH-monoresistant isolates (22/109, 20%). The inhA C-15T mutation was also more frequent in INH-monoresistant isolates (54/109, 50%) than in MDR (6/47, 13%, p < 0.0001) and polydrug-resistant (2/29, 7%, p < 0.0001) isolates. Marttila et al. (1998) and Hazbón et al. (2006) also found association between S315T mutation and MDR isolates [24, 25].
Mutations in katG and inhA are the most frequently encountered in INH-resistant isolates [26]. Among the 171 isolates with any katG and/or inhA mutation detected by MTBDRplus and sequencing in our study, 99 (58%) had mutations in katG alone, while 66 (39%) had only the C-15T inhA mutation, and six (3%) had mutations in both genes.
In this study, mutations in katG 315 were associated with an intermediate level of INH resistance, presented by 81/83 (98%) isolates with mutations in this region only. Regarding mutations in inhA promoter alone, 51/66 (77%) isolates with C-15T substitution showed low-level resistance to INH. Cambau et al. (2015) found 93% of isolates presenting katG S315T mutation with intermediate-level resistance (3 ≤ MIC ≤ 10 µg/mL) and 83% of isolates with inhA promoter mutations having low-level resistance (0.1 < MIC ≤ 1 µg/mL) [15]. Lempens et al. (2018) found that 83% of isolates with katG S315T mutation had intermediate resistance (3.2 ≤ MIC ≤ 12.8 µg/mL), while isolates with inhA C-15T showed low-level (0.8 ≤ MIC ≤ 1.6 µg/mL) and intermediate resistance [8]. In our study, 14 (21%) and 1 (2%) of the 66 isolates with inhA C-15T mutation alone had respectively intermediate- and high-level resistance.
Concurrent mutations in katG and inhA have been associated with high-level INH resistance. In our study, 5/6 (83%) isolates with mutations in both genes presented INH MIC > 10 µg/mL, while only 1 isolate had intermediate resistance. Similar results were found by Lempens et al. (2018), showing that molecular tests such as MTBDRplus are accurate in identifying isolates with high-level INH resistance which cannot be overcome by increasing INH dose in the treatment [8].
In our study, isolates presenting mutations in katG regions other than the 315 codon (n = 17) showed varying levels of INH resistance. To the best of our knowledge, 10 of these isolates presented novel mutations (P232T, D381N, W438R, A478R, I497T, K590_None, N660D, W668Y, G680R and W728G), of which 8 showed high-level resistance. Previous studies also detected different degrees of INH resistance in isolates with mutations in katG other than in 315 [8, 27, 28]. The only isolate with a mutation in the coding region of inhA (G76G) had also two mutations in katG, and high-level resistance.
The great variability of mutations we encountered along katG, including isolates harbouring more than one mutation, might be explained by the fact that this gene is located in a highly variable region of the genome containing repetitive DNA sequences, which favours the instability of the region and the high frequency of mutations [29]. katG point mutations were more frequent than deletions in our study as previously reported elsewhere [27,28,29].
The WHO has just released a catalogue of > 17,000 M. tuberculosis mutations and their association with drug resistance [30]. The 10 novel katG mutations we found are not present in this catalogue, which comprises isolates from 41 different countries. Except for the katG R463L mutation (which is considered a polymorphism and is not associated with drug resistance), the other mutations we found in katG regions outside the 315 codon that are present in the WHO catalogue have a very low frequency, so that their association with INH resistance is classified as uncertain. The scarcity of information on these katG mutations in the catalogue reinforces the importance of this work, as more data on association of INH MIC levels and infrequent mutations are needed.
The current CLSI clinical breakpoint to define INH low-level resistance (0.4 µg/mL) does not correspond to the upper end of the MIC distribution of inhA promoter mutants, hindering the adoption of such concentration to decide if the patient can be treated with high-level INH or not. According to Ghodousi et al. (2019) results on MIC distributions of inhA promoter mutants, a clinical breakpoint of 1 or 2 µg/mL would be more suitable than 0.4 µg/mL to phenotypically separate isolates between low- and high-level INH-resistant [31].
Our data show that either isolates with katG 315 mutations only or with inhA C-15T mutation alone have overlapping MICs that fall into the intermediate category. Similar results were found by Lempens et al. (2018), showing that inhA promoter mutations do not always indicate very low, and katG mutations do not necessarily cause very high levels of INH resistance levels [8]. As a consequence, the Global Laboratory Initiative recommends that, once a katG 315 mutation is detected by LPA, the result should be reported as high-level INH resistance, while an isolate with an inhA promoter mutation should be reported as “at least” low-level-resistant (http://www.stoptb.org/wg/gli/assets/documents/LPA_test_web_ready.pdf). Further research is necessary to elucidate whether high-dose INH contributes to successful outcomes in patients with low- or intermediate-level resistance, caused by a katG or inhA mutation alone.
All RIF-resistant isolates by MGIT 960 had rpoB mutations detected by MTBDRplus. Among the 73 isolates with rpoB mutations, 22% (n = 16) showed inferred mutations (50% of them were phenotypically susceptible to RIF at 1.0 µg/mL). Such borderline mutations (mutations which are detected as RIF-susceptible by phenotypic DST) are related to a higher risk of treatment failure or relapse when the standard first-line regimen with RIF is used in the treatment [32, 33]. If isolates presenting borderline mutations are misdiagnosed as RIF-susceptible and treatment with RIF is inadequately maintained, RIF resistance might spread in the community. Therefore, the WHO has just released a report in which the RIF critical concentration in MGIT decreased to 0.5 µg/mL, so that misclassification of these mutations can be reduced [9].
This study has limitations. We sequenced entire katG and inhA genes only for isolates with MTBDRplus wild-type results, so we might have missed mutations in other regions of these genes in isolates with katG and/or inhA mutations found by MTBDRplus. Moreover, we did not investigate other genes known to be related with INH resistance, such as those that regulate katG expression (furA-katG and sigI intergenic region), ahpC, oxyR, kasA and ndh, and genes involved in efflux pump mechanisms [26, 29]. Mutations in these genetic regions might explain INH resistance in isolates with wild-type results for both katG and inhA whole genes in our study. We also did not assess RIF resistance level determination for the inferred rpoB mutations.
Conclusions
We have shown that mutations in katG 315 are associated with intermediate- to high-level resistance to INH, while inhA C-15T is related to low- and also intermediate-level resistance in M. tuberculosis clinical isolates. We have described 10 novel mutations in different regions of katG in isolates with varying levels of resistance to INH. Such mutations might be potential new INH resistance markers and could explain INH resistance in isolates without classic mutations in katG 315 and inhA promoter region. In our setting, 16% of INH-resistant isolates, some with high-level resistance, did not show mutations in either katG or inhA genes.
Data availability
The datasets generated and analysed during the current study are available from the corresponding author on reasonable request.
Code availability
Not applicable.
References
Ramaswamy S, Musser JM (1998) Molecular genetic basis of antimicrobial agent resistance in Mycobacterium tuberculosis: 1998 update. Tuber Lung Dis 79:3–29
Domínguez J, Böttger EC, Cirillo D, Cobelens F, Eisenach KD, Gagneux S et al (2016) Clinical implications of molecular drug resistance testing for Mycobacterium tuberculosis: a TBNET/RESIST-TB consensus statement. Int J Tuberc Lung Dis 20(1):24–42
Seifert M, Catanzaro D, Catanzaro A, Rodwell TC (2015) Genetic mutations associated with isoniazid resistance in Mycobacterium tuberculosis: a systematic review. PLoS ONE 10(3):e0119628
Dean AS, Zignol M, Cabibbe AM, Falzon D, Glaziou P, Cirillo DM et al (2020) Prevalence and genetic profiles of isoniazid resistance in tuberculosis patients: a multicountry analysis of cross-sectional data. PLoS Med 17:e1003008
Böttger EC (2011) Drug resistance in Mycobacterium tuberculosis: molecular mechanisms and laboratory susceptibility testing. In: Donald PR, van Helden PD (eds) Progress in respiratory research. Karger, Basel, pp 1–17
Kambli P, Ajbani K, Sadani M, Nikam C, Shetty A, Udwadia Z et al (2015) Defining multidrug-resistant tuberculosis: correlating GenoType MTBDRplus assay results with minimum inhibitory concentrations. Diagn Microbiol Infect Dis 82(1):49–53
Springer B, Calligaris-Maibach R, Ritter C, Böttger E (2008) Tuberculosis drug resistance in an area of low endemicity in 2004 to 2006: semiquantitative drug susceptibility testing and genotyping. J Clin Microbiol 46(12):4064–4067
Lempens P, Meehan CJ, Vandelannoote K, Fissette K, de Rijk P, Van Deun A et al (2018) Isoniazid resistance levels of Mycobacterium tuberculosis can largely be predicted by high-confidence resistance-conferring mutations. Sci Rep 8(1):3246
World Health Organization (2021) Technical report on critical concentrations for drug susceptibility testing of isoniazid and the rifamycins (rifampicin, rifabutin and rifapentine). https://apps.who.int/iris/rest/bitstreams/1330649/retrieve. Accessed 10 Mar 2021
World Health Organization (2016) Treatment guidelines for drug-resistant tuberculosis. http://www.who.int/tb/areas-of-work/drug-resistant-tb/MDRTBguidelines2016.pdf. Accessed 10 Mar 2021
Center of Epidemiologic Surveillance, state of São Paulo (2019) Tuberculose no Estado de SP: Situação e desafios em tempos de crisehttp://www.saude.sp.gov.br/resources/cve-centro-de-vigilancia-epidemiologica/areas-de-vigilancia/tuberculose/eventos/tb19_forum_laedisantoscomoestamos.pdf. Accessed 10 Mar 2021
Rodrigues C, Jani J, Shenai S, Thakkar P, Siddiqi S, Mehta A (2008) Drug susceptibility testing of Mycobacterium tuberculosis against second-line drugs using the BACTEC MGIT 960 system. Int J Tuberc Lung Dis 12:1449e55
Adami AG, Gallo JF, Pinhata JMW, Martins MC, Giampaglia CMS, Oliveira RS (2017) Modified protocol for drug susceptibility testing of MGIT cultures of Mycobacterium tuberculosis by the MGIT 960. Diagn Microbiol Infect Dis 87:108–111
Gallo JF, Pinhata JMW, Saraceni CP, Oliveira RS (2017) Evaluation of the BACTEC MGIT 960 system and the resazurin microtiter assay for susceptibility testing of Mycobacterium tuberculosis to second-line drugs. J Microbiol Methods 139:168e71
Cambau E, Viveiros M, Machado D, Raskine L, Ritter C, Tortoli E et al (2015) Revisiting susceptibility testing in MDR-TB by a standardized quantitative phenotypic assessment in a European multicentre study. J Antimicrob Chemother 70:686–696
Rüsch-Gerdes S, Pfyffer GE, Casal M, Chadwick M, Siddiqi S (2006) Multicenter laboratory validation of the BACTEC MGIT 960 technique for testing susceptibilities of Mycobacterium tuberculosis to classical second-line drugs and newer antimicrobials. J Clin Microbiol 44(3):688–692
Matsui T, Pinhata JMW, Rabello MCDS, Brandão AP, Ferrazoli L, Leão SC et al (2020) Frequency of first and second-line drug resistance-associated mutations among resistant Mycobacterium tuberculosis clinical isolates from São Paulo Brazil. Mem Inst Oswaldo Cruz 115:e200055
Crudu V, Stratan E, Romancenco E, Allerheiligen V, Hillemann A, Moraru N (2012) First evaluation of an improved assay for molecular genetic detection of tuberculosis as well as rifampin and isoniazid resistances. J Clin Microbiol 50(4):1264–1269
Brandão AP, Pinhata JMW, Oliveira RS, Galesi VMN, Caiaffa-Filho HH, Ferrazoli L (2019) Speeding up the diagnosis of multidrug-resistant tuberculosis in a high-burden region with the use of a commercial line probe assay. J Bras Pneumol 45(2):e20180128
Perdigão J, Macedo R, João I, Fernandes E, Brum L, Portugal I (2008) Multidrug-resistant tuberculosis in Lisbon, Portugal: a molecular epidemiological perspective. Microb Drug Resist 14(2):133–143
Machado D, Perdigão J, Ramos J, Couto I, Portugal I, Ritter C et al (2013) High-level resistance to isoniazid and ethionamide in multidrug-resistant Mycobacterium tuberculosis of the Lisboa family is associated with inhA double mutations. J Antimicrob Chemother 68(8):1728–1732
Portugal I, Maia S, Moniz-Pereira J (1999) Discrimination of multidrug-resistant Mycobacterium tuberculosis IS6110 fingerprint subclusters by rpoB gene mutation analysis. J Clin Microbiol 37(9):3022–3024
Dean AG, Sullivan KM, Soe MM. OpenEpi: Open Source Epidemiologic Statistics for Public Health. http://www.openepi.com/Menu/OE_Menu.htm. Accessed 10 Mar 2021
Marttila HJ, Soini H, Eerola E, Vyshnevskaya E, Vyshnevskiy BI, Otten TF et al (1998) A Ser315Thr substitution in katG is predominant in genetically heterogeneous multidrug-resistant Mycobacterium tuberculosis isolates originating from the St. Petersburg area in Russia. Antimicrob Agents Chemother 42(9):2443–2445
Hazbón MH, Brimacombe M, Bobadilla del Valle M, Cavatore M, Guerrero MI, Varma-Basil M et al (2006) Population genetics study of isoniazid resistance mutations and evolution of multidrug-resistant Mycobacterium tuberculosis. Antimicrob Agents Chemother 50(8):2640–2649
Vilchèze C, Jacobs WR Jr (2014) Resistance to isoniazid and ethionamide in Mycobacterium tuberculosis: genes, mutations, and causalities. Microbiol Spectr 2(4):MGM2-0014–2013
Cardoso RF, Cooksey RC, Morlock GP, Barco P, Cecon L, Forestiero F et al (2004) Screening and characterization of mutations in isoniazid-resistant Mycobacterium tuberculosis isolates obtained in Brazil. Antimicrob Agents Chemother 48(9):3373–3381
Kandler JL, Mercante AD, Dalton TL, Ezewudo MN, Cowan LS, Burns SP et al (2018) Validation of novel Mycobacterium tuberculosis isoniazid resistance mutations not detectable by common molecular tests. Antimicrob Agents Chemother 62(10):e00974-e1018
Unissa AN, Subbian S, Hanna LE, Selvakumar N (2016) Overview on mechanisms of isoniazid action and resistance in Mycobacterium tuberculosis. Infect Genet Evol 45:474–492
World Health Organization (2021) Catalogue of mutations in Mycobacterium tuberculosis complex and their association with drug resistance. https://www.who.int/publications/i/item/9789240028173. Accessed 28 June 2021
Ghodousi A, Tagliani E, Karunaratne E, Niemann S, Perera J, Köser CU et al (2019) Isoniazid resistance in Mycobacterium tuberculosis is a heterogeneous phenotype composed of overlapping MIC distributions with different underlying resistance mechanisms. Antimicrob Agents Chemother 63:e00092-e119
Miotto P, Tessema B, Tagliani E, Chindelevitch L, Starks AM, Emerson C et al (2017) A standardised method for interpreting the association between mutations and phenotypic drug resistance in Mycobacterium tuberculosis. Eur Respir J 50(6):1701354
Van Deun A, Decroo T, Aung KJM, Hossain MA, Gumusboga M, De Rijk WB, et al (2021) Mycobacterium tuberculosis borderline rpoB mutations: emerging from the unknown. Eur Respir J. https://doi.org/10.1183/13993003.00783-2021
Acknowledgements
The authors thank the technical assistance and support provided by the staff of Núcleo de Tuberculose e Micobacterioses and Centro de Bacteriologia of IAL, as well as the TB laboratory network from São Paulo for the valuable partnership.
Funding
This work was supported in part by the Fundação de Amparo à Pesquisa do Estado de São Paulo (FAPESP) [grants numbers 2012/51756–5 and 2017/16082–7].
Author information
Authors and Affiliations
Contributions
JMWP and RSO conceived the study and designed the experiments; JMWP and FFM performed and analysed MTBDRplus tests; JMWP, APB and MCSR performed and analysed gene sequencing; JMWP collected laboratory data; JMWP and RSO analysed the data; JMWP wrote the draft; and JMWP, RSO, APB and LF reviewed and edited the draft. All authors read and approved the final manuscript and agreed to be responsible for all aspects of the work.
Corresponding author
Ethics declarations
Ethics approval
This study was approved by Adolfo Lutz Institute Ethics Committee (Plataforma Brasil no. 1.473.145).
Consent to participate
Not applicable.
Consent for publication
Not applicable.
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Pinhata, J.M.W., Brandao, A.P., Mendes, F.d.F. et al. Correlating genetic mutations with isoniazid phenotypic levels of resistance in Mycobacterium tuberculosis isolates from patients with drug-resistant tuberculosis in a high burden setting. Eur J Clin Microbiol Infect Dis 40, 2551–2561 (2021). https://doi.org/10.1007/s10096-021-04316-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10096-021-04316-0