Abstract
Cancer immunotherapy has been significantly effective on multiple cancers; however, there are still a distinct number of non-responding patients and various immune-related adverse events in responding patients. It is known that heterogeneity of intestinal microbiota may lead to different outcomes of therapy. Previous studies have reported that intestinal microbiota is probably attributed to influence the efficacy of cancer immunotherapy. Some intestinal bacteria could synergize with immune checkpoint blockade agents and optimize the immune response against multiple cancers. Therefore, understanding the roles of intestinal microbiota could help to improve the clinical efficacy of cancer immunotherapy. In this review, we first introduced the close relationships between intestinal microbiota and intestinal immune system. Then, we described the emerging evidences that intestinal microbiota responses to cancer immunotherapy. Finally, we briefly reviewed the technical development on intestinal microbiota research.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
In recent years, cancer immunotherapy has become very successful against distinct metastatic malignancies [1,2,3], due in great part to the clinical success of immune checkpoint blockade and chimeric antigen receptor (CAR)-modified T cell therapy. Immune checkpoint inhibitors (ICIs) have improved the survival of cancer patients. The ipilimumab and tremelimumab blocking cytotoxic T lymphocyte-associated antigen-4 (CTLA-4) have been evaluated in the treatment of melanoma [4], prostate [5], lung [6], and pancreatic [7] carcinomas and have demonstrated an overall survival benefit in cancer patients [8]. However, the functions of cytotoxic T cells are inhibited in the tumor microenvironment, which conduces to cancer cell immune evasion [9]. In recent years, blockade of programmed cell death 1 (PD-1, nivolumab and pembrolizumab) and its ligand programmed death ligand (PD-L1) performs higher response rates and more prolonged overall survival than blockade of CTLA-4 (ipilimumab) [10, 11]. In addition, an exciting new approach CAR-T cell therapy in the fight against cancer, which refines the design of CARs and improves the cellular manufacturing processes with the purpose of delivering safe and efficacious therapeutic T cells, is bringing the promising therapeutic platform for more cancer patients.
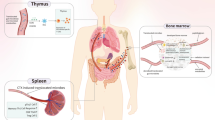
Unfortunately, the beneficial effects of these immunotherapy strategies are seen only in a subgroup of patients [12], and these ICIs treatments are closely associated with various immune-related adverse events. The most common immune toxicities include colitis, diarrhea, thyroid dysfunction, dermatologic event, liver disorder, and lung disorder [13]. Although these side effects have become relieved to some extent by using corticosteroid therapy, new effective indicators of response and toxicity are necessary to improve the compliance to immunotherapy. The recent flurry of scientific studies on the effects of intestinal microbiota in response to cancer immunotherapy opens up an entirely novel approach to the treatment of cancer diseases [14,15,16]. The intestine faces constant challenges from food antigens, pathogens, and commensals and has to make appropriate responses precisely and quickly. The intestinal microbiota is important for human metabolism such as production of short-chain fatty acids, essential vitamins, and amino acids [17]. Intestinal microbiota is also a key for the development of the mucosal immune system [18]. Therefore, intestinal microbiota appears a very promising area of research in modulating the immune system and finding an impact in anti-tumor immunotherapy. The purpose of this review is to help us better understand the role of intestinal microbiota and improve the efficacy in cancer immunotherapy by the regulation of intestinal microbiota.
Intestinal microbiota and intestinal immune system
The intestinal microbiota plays essential roles in modulating the intestinal immune response to keep intestinal immune homeostasis [19]. Intestinal microbiota is capable to module host physiology and/or nutritional status and influences not only the intestine but also distant organs [20].
Intestinal barrier
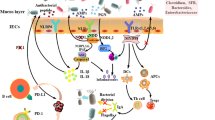
Usually, the spatial interactions between intestinal microbiota and intestinal immune system can be divided into three functional layers. Facing to the intestinal lumen, the first layer, rich in mucus, can also be divided into another two sublayers: the outer sublayer and the inner layer. The outer sublayer is very abundant in microbiota, while the inner layer has high concentration of bactericidal antimicrobial peptides (AMPs) and secretory IgA. The second layer is made up of a monolayer of intestinal epithelial cells (IECs), which are mainly composed by goblet cells, absorptive enterocytes, and enteroendocrine cells, paneth cells, M cells, and so on [21, 22]. IECs play a key role in separating the internal body organs from the outside environment by the formation of tight junctions and secretion of mucus and AMPs [21]. IECs express pattern recognition receptors (PRRs), including nod-like receptors (NLRs) and toll-like receptors (TLRs) [23]. The production of some types of AMPs, like angiogenin-4, regenerating islet-derived protein 3γ (REGIIIγ), and regenerating islet-derived protein 3β (REGIIIβ), is influenced by commensal microbes in a toll-like receptor-dependent way [21]. In IECs layer, paneth cells are the leading producer of AMPs [24]. The M cells are a very important cell type, because these cells work directly with the immune system [21]. The third layer is formed by lamina propria (LP) and mesentery. Microbe-associated molecular patterns from colonizing bacteria are sensed by PRRs or dendritic cells (DCs) that activate T and B cells in isolated lymphoid follicles. DCs that capture antigens through IECs or from LP migrate to mesenteric lymph node to induce the differentiation of effector T cells [25]. The interactions of intestinal microbiota, intestinal epithelium, and mucosal immune system lead to a local and systemic homeostasis.
Interactions of intestinal microbiota and intestinal immune system
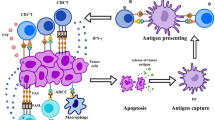
The gastrointestinal tract is inhabited by various microbes including commensal bacteria and pathogenic bacteria. Usually, commensal bacteria are beneficial for the host, while pathogenic bacteria are able to cause problems, such as intestinal inflammation and invasiveness. The intestinal immune system shapes the intestinal microbiota composition, and the latter regulates the intestinal immune system responses [26]. In a symbiosis context, microbe-associated molecular patterns constantly stimulate IECs to secrete some immunological mediators, such as IL-33, IL-25, and tumor growth factor-β (TGF-β), which induce the development of tolerogenic macrophages and tolerogenic DCs [25, 27]. And tolerogenic DCs could produce TGF-β and retinoic acids that activate the development of T regulatory cells (Tregs). Therefore, the intestinal immune system associated with intestinal microbiota could establish and maintain an anti-inflammatory environment through tregs, macrophages, and tolerogenic DCs. In a dysbiosis context, the pathogenic bacteria overcome commensal bacteria and disrupt the regulated anti-inflammatory environment. This unstable state can induce IECs and activate dendritic cells and macrophages to secrete inflammatory cytokines (IL-1β, Il-6, IL-12, and IL-23). These cytokines stimulate the development of TH1 cells and TH17 cells leading to chronic inflammation [27]. Furthermore, intestinal microbiota could produce high levels of endotoxins, which will cause systemic inflammation once in the bloodstream, resulting in the progression of many human diseases.
Recent studies have focused on the interactions between the intestinal microbiota and the immune system. Masahata et al. found that IgA-secreting cells were closely associated with the maintenance of intestinal microbial homeostasis and contributed to shaping the healthy intestinal microbial community, indicating that the development of immune system had a close accordance with intestinal microbiota [28]. Atarashi et al. demonstrated that a mixture of Clostridia strains from the human intestinal microbiota was able to induce the accumulation of Tregs and IL-10 production in intestine [29]. Cording et al. concluded that intestinal microbial stimulus locally influenced the Treg proliferation and systemically affected conventional CD4+ T cells [30]. The intestinal microbiota undoubtedly influences the regulatory cells; however, the mechanisms induced by microbes influencing the development of Tregs remain unknown. Obata et al. [31] explored the changes in IL-2 expressing CD4+ T cells and FoxP3+ Treg cells by inoculating germ-free mice with commensal microbiota. The results found that changes of IL-2+ CD4+ T cells were different from Treg cell expansion, suggesting that commensal microbiota stimulated the development of the Tregs in an IL-2-dependent manner. In addition, some studies tried to identify the metabolites of intestinal microbiota to influence the immune system and regulate homeostasis. Smith et al. [32] found that germ-free mice had a series of immunological problems, and the abundance of three types of short-chain fatty acids (SCFAs: acetic acid, butyric acid, and propionic acid) was significantly decreased. After treating the germ-free mice with SCFAs for 3 weeks, these mice showed an increasing in frequency and number of colonic Tregs. Therefore, SCFAs metabolized by intestinal microbiota played a key role in maintaining homeostasis through Tregs. The result was also confirmed by the study of Furusawa et al. [33], who found that the luminal concentrations of SCFAs were positively correlated with the number of regulatory cells in the colon.
As the conclusions described above, intestinal microbiota and immune system interact continuously to maintain a complex dynamic equilibrium for host health. A complete understanding of the relationship between intestinal microbiota and intestinal immunity is very important for the treatment of human many diseases.
Intestinal microbiota influences the efficacy of cancer immunotherapy
Intestinal microbiota has ascended to prominence as important modulators of host immunity and has made the possibility of influencing the outcome of cancer immunotherapy. Table 1 lists some researches about intestinal microbiota in response to cancer immunotherapy. In this following, we summarized some studies about intestinal microbiota influencing the efficacy of cancer immunotherapy.
Intestinal microbiota in response to CTLA-4-based immunotherapy
Different from cytotoxic therapies, ICIs regulate tumor changes via enhancing host immune activation. Antibodies targeting CTLA-4 have been successfully used as cancer immunotherapy. Ipilimumab is a fully human monoclonal antibody directed against CTLA-4, approved as the first drug for improving the overall survival of patients with metastatic melanoma [8]. Previous studies have addressed the role of intestinal microbiota in immunomodulatory effects of CTLA-4 blockade [34]. Marie Vétizou et al. found tumors in antibiotic-treated or germ-free mice did not respond to CTLA-4 blockade; however, this defect was overcome by gavage with Bacteroides fragilis, by adoptive transfer of Bacteroides fragilis-specific T cells, or by immunization with Bacteroides fragilis polysaccharides [34]. This study indicates that Bacteroidales plays an important role in the immunostimulatory effects of CTLA-4 blockade. Chaput et al. reported that Faecalibacterium and other Firmicutes were closely associated with beneficial clinical response, but a higher representation of Bacteroides genus in metastatic melanoma patients had a poor response to anti-CTLA-4 treatment [35]. The conclusion was inconsistent with the results of Marie Vétizou trial in mouse models [34]. The discrepancy is mainly ascribed to different models. Apart from this, it is difficult to exclude other microbes interfering results in mouse experiment. Therefore, further studies should be carried out to evaluate the effect of CTLA-4-based immunotherapy on intestinal microbiota.
Intestinal microbiota in response to PD-1/PD-L1-based immunotherapy
Immune checkpoint inhibitors (ICIs) targeting the PD-1/PD-L1 lead to sustained clinical responses in cancer patients [40]. Routy et al. found that abnormal intestinal microbial composition caused primary resistance to ICIs [37]. The relative abundance of Akkermansia muciniphila significantly affected the clinical responses to ICIs, proved by oral supplementation with Akkermansia muciniphila for non-responders to restore the efficacy of PD-1 blockade [37]. Mastson et al. analyzed fecal samples from metastatic melanoma patients before anti-PD-1 immunotherapy based on 16S rRNA sequencing, metagenomic shotgun sequencing, and quantitative polymerase chain [38]. The results suggested that commensal microbiome including Bifidobacterium longum, Collinsella aerofaciens, and Enterococcus faecium could have important impacts on anti-tumor immunity. Frankel et al. showed that melanoma patients who responded to ICIs were enriched with Bacteroides caccae using metagenomic shotgun sequencing method [36]. Wargo et al. also found that intestinal microbial diversity and composition in metastatic melanoma patients that responded to the anti-PD-1 therapy were significantly different from that in non-responding patients [39]. The responders had higher alpha diversity and relative abundance of Ruminococcaceae bacteria, and the non-responders had lower diversity and higher relative abundance of Bacteroidales [39]. In addition, Sivan et al. explored melanoma growth in mice harboring distinct commensal microbiota and found Bifidobacterium could promote anti-tumor immunity and facilitate anti-PD-L1 efficacy [16]. Above all these findings, intestinal microbiota could improve PD-1 and PD-L1 immunotherapy by regulating the immune response. Therefore, keeping the intestinal microbiota healthy could help cancer patients improve therapeutic efficacy.
Furthermore, CAR-T cell therapy has the remarkable potential to become promising therapeutic platform especially for a few cancer patients with hematologic tumors in recent years. However, adoptive T cell therapy is still in its infancy, and a number of challenges need to be considered to provide safe and reliable cellular products. Up to now, there is no related study about intestinal microbiota in response to the efficacy of CAR-T cell therapy.
Immune-related adverse events involved in intestinal microbiota
Blockades of CTLA-4 and PD-1 often lead to immune-related adverse events that are mostly exposed to intestinal microbiota [41]. During CTLA-4 and PD-1 blockade treatment, intestinal epithelial cell injury results in the loss of integrity of intestinal barrier. Some commensal bacteria such as Enterococcus hirae can influence systemic inflammation by destroying intestinal barrier into secondary immune organs even tumor bed [42]. Marie Vétizou and colleagues identified that two species from Bacteroidales and Burkholderiales order significantly reduced the histopathological colitis associated with anti-CTLA-4 therapy in mice model, which was a common, high-risk, immune-related adverse event [34]. Krista Dubin et al. found that the Bacteroidales species were closely linked with decreased incidence of colitis in patients with metastatic melanoma who have undergo ipilimumab treatment [43]. However, it also reported that certain Bacteroidales species in the gut were correlated with colitis [44]. One convincing reason is that the anatomical structures of gastrointestinal tract and intestinal wall linings in human and mouse are significantly different [45]. Another reason is that many intestine-colonizing microbes in mice are not found in humans [46]. Additionally, individual diet and lifestyle contribute to the intestinal microbiota in response to immunotherapy [47, 48].
New techniques on intestinal microbiota research
Much of our current understanding of interactions between intestinal microbiota and immune system has been mainly acquired from studies of germ-free animals. However, the composition of human and animal intestinal microbiota can be defined from polymorphisms of bacterial genes. Therefore, technological advances play great roles in facilitating studies of complex intestinal microbiota and their functions. The DNA sequencing technology (next-generation sequencing, metagenomic) improvements have identified potential functions of microbes in human gut [49, 50]. However, a vast majority still have no known functions, reflecting the great diversity and biochemical potential of microbiota remaining to be discovered. The mRNA sequencing (metatranscriptomic), revealing which genes are expressed by specific organisms from spatial and temporal scales, has offered a wealth of knowledge about the expression of microbial genes in human gut [51]. The nucleic acid sequencing has helped to explore and understand microbial phylogenetic and functional compositions in human intestinal microbiomes [49, 52]; however, it is also desirable to know which proteins (metaproteomic) and metabolites (metabolomic) play key roles in performing special functions. These proteins and metabolites produced by microbes were measured by mass spectrometry, which had high sensitivity, resolution, and throughput in providing metaproteomic or metabolomic measurements. Furthermore, additional technologies such as gas-phase ion mobility spectrometry and liquid chromatography separations are also being used to identify the more proteins and metabolites.
Metagenomic, metatranscriptomic, mass spectrometry-based metaproteomic and metabolomic gas-phase ion mobility spectrometry and liquid chromatography separations have offered a deeper understanding of composition and function of microbiomes. However, there are still many challenges to be addressed, such as extraction of biomolecules from complex environmental samples (human gut), assembly of complete genomes, statistical and mathematical models to integrate the data, and sufficient storage and analysis options for meta-data to provide meaningful biological insights.
Conclusions
Mounting evidences indicate that the intestinal microbiota influences cancer patients in response to immunotherapy, including the therapy efficacy and side effects. Intestinal microbiota probably becomes a novel biomarker of immune response. However, it still requires extensive studies, development, and testing, especially applying all acquired knowledge to transfer from mice models to human beings. Furthermore, the technological and computational improvements will contribute to a better understanding of how cancer immunotherapy affects microbial functions and facilitate human health strategy improvement. Summing up, regulating the intestinal microbiota probably helps to improve tumor control, augment immune responses, and enhance the efficacy of immunotherapy in the demanding fight against cancer.
Abbreviations
- AMPs:
-
Antimicrobial peptides
- CAR:
-
Chimeric antigen receptor
- CTLA-4:
-
Cytotoxic T lymphocyte-associated antigen-4
- DCs:
-
Dendritic cells
- ICIs:
-
Immune checkpoint inhibitors
- LP:
-
Lamina propria
- mAbs:
-
Monoclonal antibodies
- NLRs:
-
Nod-like receptors
- PRRs:
-
Pattern recognition receptors
- PD-1:
-
Programmed cell death 1
- PD-L1:
-
Programmed cell death ligand 1
- REGIIIγ:
-
Regenerating islet-derived protein 3γ
- REGIIIβ:
-
Regenerating islet-derived protein 3β
- SCFA:
-
Short-chain fatty acids
- TLRs:
-
Toll-like receptors
- Tregs:
-
T regulatory cells
- TGF-β:
-
Tumor growth factor-β
References
Restifo NP, Dudley ME, Rosenberg SA (2012) Adoptive immunotherapy for cancer: harnessing the T cell response. Nat Rev Immunol 12(4):269
Brahmer J, Reckamp KL, Baas P, Crinò L, Eberhardt WEE, Poddubskaya E, Antonia S, Pluzanski A, Vokes EE, Holgado E (2015) Nivolumab versus docetaxel in advanced squamous-cell non-small-cell lung cancer. N Engl J Med 373(17):123–135
Shalapour S, Lin XJ, Bastian IN, Brain J, Burt AD, Aksenov AA, Vrbanac AF, Li W, Perkins A, Matsutani T (2017) Erratum: inflammation-induced IgA+ cells dismantle anti-liver cancer immunity. Nature 551(7680):340–345
Ribas A, Kefford R, Marshall MA, Punt CJ, Haanen JB, Marmol M, Garbe C, Gogas H, Schachter J, Linette G (2013) Phase III randomized clinical trial comparing tremelimumab with standard-of-care chemotherapy in patients with advanced melanoma. J Clin Oncol 31(5):616–622
Kwon ED, Drake CG, Scher HI, Fizazi K, Bossi A, Aj VDE, Krainer M, Houede N, Santos R, Mahammedi H (2014) Ipilimumab versus placebo after radiotherapy in patients with metastatic castration-resistant prostate cancer that had progressed after docetaxel chemotherapy (CA184-043): a multicentre, randomised, double-blind, phase 3 trial. Lancet Oncol 15(10):700–712
Lynch TJ, Bondarenko I, Luft A, Serwatowski P, Barlesi F, Chacko R, Sebastian M, Neal J, Lu H, Cuillerot JM (2012) Ipilimumab in combination with paclitaxel and carboplatin as first-line treatment in stage IIIB/IV non-small-cell lung cancer: results from a randomized, double-blind, multicenter phase II study. J Clin Oncol 30(17):2046–2054
Aglietta M, Barone C, Sawyer MB, Moore MJ, Jr MW, Bagalà C, Colombi F, Cagnazzo C, Gioeni L, Wang E (2014) A phase I dose escalation trial of tremelimumab (CP-675,206) in combination with gemcitabine in chemotherapy-naive patients with metastatic pancreatic cancer. Ann Oncol 25(9):1750
Hodi FS, O’Day SJ, Mcdermott DF, Weber RW, Sosman JA, Haanen JB, Gonzalez R, Robert C, Schadendorf D, Hassel JC (2016) Improved survival with ipilimumab in patients with metastatic melanoma. N Engl J Med 363(8):711–723
Zou W (2005) Immunosuppressive networks in the tumour environment and their therapeutic relevance. Nat Rev Cancer 5(4):263–274
Larkin J, Chiarionsileni V, Gonzalez R, Grob JJ, Cowey CL, Lao CD, Schadendorf D, Dummer R, Smylie M, Rutkowski P (2015) Combined nivolumab and ipilimumab or monotherapy in previously untreated melanoma. N Engl J Med 373(1):23
Robert C, Schachter J, Long GV, Arance A, Grob JJ, Mortier L, Daud A, Carlino MS, Mcneil C, Lotem M (2015) Pembrolizumab versus ipilimumab in advanced melanoma. N Engl J Med 372(26):2521–2532
Postow MA, Callahan MK, Wolchok JD (2015) Immune checkpoint blockade in cancer therapy. J Clin Oncol 33(17):1974–1982
Weber JS, Kähler KC, Hauschild A (2012) Management of immune-related adverse events and kinetics of response with ipilimumab. J Clin Oncol 30(21):2691–2697
Peled JU, Devlin SM, Staffas A, Lumish M, Khanin R, Littmann ER, Ling L, Kosuri S, Maloy M, Slingerland JB (2017) Intestinal microbiota and relapse after hematopoietic-cell transplantation. J Clin Oncol 35(15):JCO2016703348
Iida N, Dzutsev A, Stewart CA, Smith L, Bouladoux N, Weingarten RA, Molina DA, Salcedo R, Back T, Cramer S (2013) Commensal bacteria control cancer response to therapy by modulating the tumor microenvironment. Science 342(6161):967–970
Sivan A, Corrales L, Hubert N, Williams JB, Aquinomichaels K, Earley ZM, Benyamin FW, Lei YM, Jabri B, Alegre ML (2015) Commensal Bifidobacterium promotes antitumor immunity and facilitates anti-PD-L1 efficacy. Science 350(6264):1084–1089
Brestoff JR, Artis D (2013) Commensal bacteria at the interface of host metabolism and the immune system. Nat Immunol 14(7):676
Randall TD, Mebius RE (2014) The development and function of mucosal lymphoid tissues: a balancing act with micro-organisms. Mucosal Immunol 7(3):455
Mortha A, Chudnovskiy A, Hashimoto D, Bogunovic M, Spencer SP, Belkaid Y, Merad M (2014) Microbiota-dependent crosstalk between macrophages and ILC3 promotes intestinal homeostasis. Science 343(6178):1249288
Nicholson JK, Holmes E, Kinross J, Burcelin R, Gibson G, Jia W, Pettersson S (2012) Host-gut microbiota metabolic interactions. Science 336(6086):1262–1267
Goto Y, Ivanov II (2013) Intestinal epithelial cells as mediators of the commensal–host immune crosstalk. Immunol Cell Biol 91(3):204–214
Collins SM, Surette M, Bercik P (2012) The interplay between the intestinal microbiota and the brain. Nat Rev Microbiol 10(11):735
Lavelle EC, Murphy C, O’Neill LAJ, Creagh EM (2009) The role of TLRs, NLRs, and RLRs in mucosal innate immunity and homeostasis. Mucosal Immunol 3(1):17
Cani PD, Everard A, Duparc T (2013) Gut microbiota, enteroendocrine functions and metabolism. Curr Opin Pharmacol 13(6):935–940
Kamada N, Seo SU, Chen GY, Núñez G (2013) Role of the gut microbiota in immunity and inflammatory disease. Nat Rev Immunol 13(5):321–335
Round JL, Mazmanian SK (2009) The gut microbiome shapes intestinal immune responses during health and disease. Nat Rev Immunol 9(5):313–323
Maynard CL, Elson CO, Hatton RD, Weaver CT (2012) Reciprocal interactions of the intestinal microbiota and immune system. Nature 489(7415):231–241
Masahata K, Umemoto E, Kayama H, Kotani M, Nakamura S, Kurakawa T, Kikuta J, Gotoh K, Motooka D, Sato S (2014) Generation of colonic IgA-secreting cells in the caecal patch. Nat Commun 5(4):3704
Atarashi K, Tanoue T, Oshima K, Suda W, Nagano Y, Nishikawa H, Fukuda S, Saito T, Narushima S, Hase K (2013) Treg induction by a rationally selected mixture of Clostridia strains from the human microbiota. Nature 500(7461):232
Cording S, Fleissner D, Heimesaat MM, Bereswill S, Loddenkemper C, Uematsu S, Akira S, Hamann A, Huehn J (2013) Commensal microbiota drive proliferation of conventional and Foxp3(+) regulatory CD4(+) T cells in mesenteric lymph nodes and Peyer’s patches. Eur J Microbiol Immunol 3(1):1
Obata Y, Furusawa Y, Endo TA, Sharif J, Takahashi D, Atarashi K, Nakayama M, Onawa S, Fujimura Y, Takahashi M (2014) The epigenetic regulator Uhrf1 facilitates the proliferation and maturation of colonic regulatory T cells. Nat Immunol 15(6):571
Smith PM, Howitt MR, Panikov N, Michaud M, Gallini CA, Bohloolyy M, Glickman JN, Garrett WS (2013) The microbial metabolites, short chain fatty acids, regulate colonic Treg cell homeostasis. Science 341(6145):569–573
Uetake C, Takahashi D, Topping DL, Miyauchi E, Nakato G, Koseki H, Ohno H, Clarke JM, Kikuchi J, Kato K (2013) Commensal microbe-derived butyrate induces the differentiation of colonic regulatory T cells. Nature 504(7480):446
Vétizou M, Pitt JM, Daillère R, Lepage P, Waldschmitt N, Flament C, Rusakiewicz S, Routy B, Roberti MP, Duong CP (2015) Anticancer immunotherapy by CTLA-4 blockade relies on the gut microbiota. Science 350(6264):1079
Chaput N, Lepage P, Coutzac C, Soularue E, Le RK, Monot C, Boselli L, Routier E, Cassard L, Collins M (2017) Baseline gut microbiota predicts clinical response and colitis in metastatic melanoma patients treated with ipilimumab. Ann Oncol 28(6):1368
Frankel AE, Coughlin LA, Kim J, Froehlich TW, Yang X, Frenkel EP, Koh AY (2017) Metagenomic shotgun sequencing and unbiased metabolomic profiling identify specific human gut microbiota and metabolites associated with immune checkpoint therapy efficacy in melanoma patients. Neoplasia 19(10):848
Routy B, Le CE, Derosa L, Cpm D, Alou MT, Daillè R, Fluckiger A, Messaoudene M, Rauber C, Roberti MP (2018) Gut microbiome influences efficacy of PD-1-based immunotherapy against epithelial tumors. Science 359(6371):91
Matson V, Fessler J, Bao R, Chongsuwat T, Zha Y, Alegre ML, Luke JJ, Gajewski TF (2018) The commensal microbiome is associated with anti-PD-1 efficacy in metastatic melanoma patients. Science 359(6371):104
Gopalakrishnan V, Spencer CN, Nezi L, Reuben A, Andrews MC, Karpinets TV, Prieto PA, Vicente D, Hoffman K, Wei SC (2018) Gut microbiome modulates response to anti-PD-1 immunotherapy in melanoma patients. Science 359(6371):97
Pardoll D (2015) Cancer and the immune system: basic concepts and targets for intervention. Semin Oncol 42(4):523–538
Beck KE, Blansfield JA, Tran KQ, Feldman AL, Hughes MS, Royal RE, Kammula US, Topalian SL, Sherry RM, Kleiner D (2006) Enterocolitis in patients with cancer after antibody blockade of cytotoxic T-lymphocyte-associated antigen 4. J Clin Oncol 24(15):2283–2289
Mao K, Baptista AP, Tamoutounour S, Zhuang L, Bouladoux N, Martins AJ, Huang Y, Gerner MY, Belkaid Y, Germain RN (2018) Innate and adaptive lymphocytes sequentially shape the gut microbiota and lipid metabolism. Nature 554 (7691)
Dubin K, Callahan MK, Ren B, Khanin R, Viale A, Ling L, No D, Gobourne A, Littmann E, Huttenhower C (2016) Intestinal microbiome analyses identify melanoma patients at risk for checkpoint-blockade-induced colitis. Nat Commun 7:10391
Lucke K, Miehlke S, Jacobs E, Schuppler M (2006) Prevalence of Bacteroides and Prevotella spp. in ulcerative colitis. J Med Microbiol 55(5):617–624
Nguyen TLA, Vieira-Silva S, Liston A, Raes J (2015) How informative is the mouse for human gut microbiota research? Dis Model Mech 8(1):1–16
Ley RE, Bäckhed F, Turnbaugh P, Lozupone CA, Knight RD, Gordon JI (2005) Obesity alters gut microbial ecology. Proc Natl Acad Sci U S A 102(31):11070–11075
Graf D, Cagno RD, Fåk F, Flint HJ, Nyman M, Saarela M, Watzl B (2015) Contribution of diet to the composition of the human gut microbiota. Biochem J 26(1):477–480
Conlon M, Bird A (2015) The impact of diet and lifestyle on gut microbiota and human health. Nutrients 7(1):17–44
Lamendella R, Verberkmoes N, Jansson JK (2012) ‘Omics’ of the mammalian gut – new insights into function. Curr Opin Biotechnol 23(3):491–500
Willing BP, Dicksved J, Halfvarson J, Andersson AF, Lucio M, Zheng Z, Järnerot G, Tysk C, Jansson JK, Engstrand L (2011) A pyrosequencing study in twins shows that gastrointestinal microbial profiles vary with inflammatory bowel disease phenotypes. Gastroenterology 139(6):1844–1854.e1
Zoetendal EG, Raes J, Bogert BVD, Arumugam M, Booijink CC, Troost FJ, Bork P, Wels M, Vos WMD, Kleerebezem M (2012) The human small intestinal microbiota is driven by rapid uptake and conversion of simple carbohydrates. ISME J 6(7):1415–1426
Lozupone CA, Stombaugh JI, Gordon JI, Jansson JK, Knight R (2012) Diversity, stability and resilience of the human gut microbiota. Nature 489(7415):220–230
Acknowledgements
The assistance of the staff is gratefully appreciated.
Funding
This study was supported by the funding from Project funded by China Postdoctoral Science Foundation (2016M602094), Qingdao Application Research Project (2016047), and Qingdao People’s Livelihood Science and Technology Program (16-6-2-3-nsh).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare that they have no competing interests.
Rights and permissions
About this article
Cite this article
Cong, J., Zhang, X. Roles of intestinal microbiota in response to cancer immunotherapy. Eur J Clin Microbiol Infect Dis 37, 2235–2240 (2018). https://doi.org/10.1007/s10096-018-3374-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10096-018-3374-8