Abstract
Most of the heritability in frontotemporal dementia (FTD) is accounted for by autosomal dominant hexanucleotide expansion in the chromosome 9 open reading frame 72 (C9orf72), pathogenic/likely pathogenic variants in progranulin (GRN), and microtubule-associated protein tau (MAPT) genes. Until now, there has been no systematic analysis of these genes in the Serbian population. Herein, we assessed the frequency of the C9orf72 expansion, pathogenic/likely pathogenic variants in GRN and MAPT in a well-characterized group of 472 subjects (FTD, Alzheimer’s disease - AD, mild cognitive impairment - MCI, and unspecified dementia - UnD), recruited in the Memory Center, Neurology Clinic, University Clinical Center of Serbia. The C9orf72 repeat expansion was detected in 6.98% of FTD cases (13.46% familial; 2.6% sporadic). In the UnD subgroup, C9orf72 repeat expansions were detected in 4.08% (8% familial) individuals. Pathogenic variants in the GRN were found in 2.85% of familial FTD cases. Interestingly, no MAPT pathogenic/likely pathogenic variants were detected, suggesting possible geographical specificity. Our findings highlight the importance of wider implementation of genetic testing in neurological and psychiatric practice managing patients with cognitive-behavioral and motor symptoms.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Frontotemporal dementia (FTD) is associated with frontal and temporal lobe degeneration that results in progressive personality/behavior changes and impairment of language functions [1]. FTD is a highly heritable group of neurodegenerative disorders, with around 40% of patients with a strong family history [2].
The most common pathogenic/likely pathogenic variants with autosomal dominant inheritance are found in the chromosome 9 open reading frame 72 (C9orf72), progranulin (GRN), and microtubule-associated protein tau (MAPT) genes each causing between ∼ 5 and 10% of all FTD [3], with geographical variability presented in some case series like the predominance of GRN pathogenic/likely pathogenic variants in Northern Italy and the Basque country [4, 5]. C9orf72 hexanucleotide expansion seems to be the most common global cause of genetic FTD, followed by GRN and MAPT genes. These pathogenic/likely pathogenic variants account for 20–30% of the familial and 5–10% of sporadic FTD cases [3]. In Caucasian/Western populations, pathogenic/likely pathogenic variants in C9orf72, GRN, and MAPT could be responsible for 8–25%, 5–22%, and 5–15%, cases respectively [6], whereas in other genes are found in less than 5% patients with FTD [7].
Alzheimer’s disease (AD) and FTD may share clinical features in early stages, making it difficult to differentiate between these two diseases. Hexanucleotide repeat expansion in C9orf72 was reported either within clinically diagnosed AD patients or pathologically confirmed AD cases [8, 9]. In the study that included asymptomatic and symptomatic individuals with a family history of FTD, the pathogenic/likely pathogenic variants in one of the genes (C9orf72, GRN, or MAPT) were reported, with the bvFTD as the most common finding. Further, in C9orf72 expansion carriers, the behavioral (n = 3) and cognitive (n = 6) mild cognitive impairment (MCI) variants are found. In MAPT carriers, the bvFTD showed high frequency, followed by the MCI behavior variant, while GRN carriers had more diverse phenotypes in FTD and AD dementia spectrum phenotypes [10].
Until now, no comprehensive analyses of pathogenic/likely pathogenic variants in the major three genes have been conducted in the Serbian population. In the present study, we screened C9orf72, GRN, and MAPT in the cohort with cognitive-behavioral disorders (FTD, MCI, AD, and unspecified dementia -UnD) recruited at the Memory Center of the Neurology Clinic in Belgrade, University Clinical Centre of Serbia.
Materials and methods
Participants
The present study was conducted as a part of the project on dementia genetics from January 2011 to January 2021. Patients with various cognitive-behavioral presentations were prospectively recruited at the Memory Center, Neurology Clinic, University Clinical Center of Serbia (UCCS). The consecutive subjects (n = 472) consisted of 129 patients with FTD, 176 with AD, 118 with MCI, and 49 with UnD. The patients were referred from specialized dementia inpatient and outpatient units, covering dementia cases from all over Serbia. The patients were unrelated, except for two sisters in the AD group, and one FTD patient had a sister with MCI. The group of healthy controls (n = 96) without a history of dementia, parkinsonism, and motor neuron disease were tested as well for the C9orf72 expansion.
The three major FTD genes (C9orf72, GRN, and MAPT) were screened in 265 (83 in FTD, 80 in AD, 22 in UnD, and 80 in MCI) patients. Sequencing analysis of GRN and MAPT genes was performed in 271 (84 in FTD, 84 in AD, 23 in UnD, and 80 in MCI) and 276 patients (88 in FTD, 82 in AD, 23 in UnD, and 83 in MCI), respectively. All recruited patients (n = 472) were screened for the C9orf72 expansion.
The participants underwent detailed clinical and cognitive-behavioral examination and imaging protocols: computerized tomography, magnetic resonance imaging (MRI), and FDG positron emission tomography (PET) imaging. The diagnosis of AD was made on the existing criteria [11]. The cerebrospinal fluid (CSF) AD biomarkers (amyloid-beta, tau, and phospho-tau) were determined for the patients who gave consent for lumbar puncture (∼ 74%). The enrolled patients with FTD met the research criteria for behavioral variant FTD (bvFTD), primary progressive aphasia (PPA), Semantic dementia (SD), and progressive nonfluent aphasia (PNFA) following the criteria: (a) International Behavioural Variant FTD Criteria Consortium [12]; and (b) developed by an international group of PPA investigators [13]. The STROBE checklist for the present study was completed based on the STROBE cohort guidelines [14].
The Goldman scores were used to determine the family histories. Each patient was assigned a score from 1 to 4 with an adjustment that split Goldman category 3 into 3 and 3.5 [15]. Score 1 is assigned to autosomal dominant inheritance with at least 3 people in 2 generations affected with FTLD, ALS, CBD, or PSP, with one person being a first-degree relative of the other two. Category 2 means a family aggregation if there were at least three relatives with dementia or ALS but do not meet the criteria for autosomal dominant inheritance. A score of 3 is assigned if there was a single affected first-degree family member with dementia or ALS with the age of onset before 65y and 3.5 for the onset above 65y. Category 4 was assigned when there was no contributory family history or with an unknown family history. Medical history records were obtained for affected family members whenever possible.
Standard protocol approvals, registrations, and patient consents
The study was approved by the Ethics Committee of the Faculty of Medicine, University of Belgrade, and Ethics from the UCCS. A neurologist took the necessary clinical information after obtaining informed written consent from the patients and their participating family members. Whenever a patient was not capable of consenting, a legally authorized representative was included. For all patients, information regarding the age of onset, disease duration, family history, and clinical features were obtained from patients and caregiver reports.
Genetic analyses
After obtaining the informed consent, blood samples were drawn from all patients, and DNA was extracted using standard protocols. The first coding exon, as well as the exons 9–13 of MAPT gene (transcript NM_001377265.1), and all exons of GRN gene (transcript NM_002087.4) with their surrounding regions (up to 30 base pairs), were sequenced using Sanger sequencing. All the variants were classified according to the American College of Medical Genetics and Genomics and the Association for Molecular Pathology Guidelines [16]. The variants were searched through the ClinVar database (https://www.ncbi.nlm.nih.gov/clinvar/) [17]. Minor allele frequency in the non–Finnish European population was obtained from the Genome Aggregation Database (gnomAD (v.3.1.1), http://gnomad.broadinstitute.org/) [18] and pathogenicity prediction was performed using in silico prediction software SIFT (https://sift.bii.a-star.edu.sg) [19], PolyPhen-2 (http://genetics.bwh.harvard.edu/pph2/) [20] and CADD (https://cadd.gs.washington.edu) [21]. We applied a stringent pathogenicity cut-off of 20 for the CADD score. Sizing of hexanucleotide repeats in the C9orf72 gene was performed by fragment analysis on ABI 3500 Genetic Analyzer (Applied Biosystems, Foster City, CA, USA) using the two-step protocol. First, normal-size alleles were genotyped. In the second step, for the homozygous samples, we performed repeat-primed PCR (RP-PCR) with previously published primers by Renton et al., 2011 [22]. To exclude false negative results, additional RP-PCR with primers by DeJesus-Hernandez et al., 2011 [23] was performed, as recommended by Rollinson et al., 2015 [24]. Pathogenic cut-off size was 30 repeats [22]. All the expansion carriers were confirmed with Southern blot.
Statistical analyses
Statistical analyses were performed with the Statistica program (version 12). The potential differences between groups were assessed with the Pearson chi2 test, and Fischer’s exact test was used when the expected values were < 5 in more than 20% of the contingency cells. A p-value of < 0.05 was considered significant. In the correlation analysis of the repeat size and the age of onset in the FTD group, we have used the maximum value of the obtained repeats. The analysis was performed using the Spearman correlation coefficient.
Results
All subjects originated from the population of Serbia. Demographic and clinical data of each group (FTD, AD, MCI, and UnD) are presented in Table 1., and in the Supplementary file text (Supplementary material). In the FTD group there were 84 patients with bvFTD, 5 with FTD-ALS, 34 with PNFA, and 6 with SD. To our knowledge, parental consanguinity of either degree was not present in our cohort.
In 265 subjects tested for C9orf72, GRN, and MAPT, 2.64% had a pathogenic variant in C9orf72 (n = 6) and in GRN (n = 1), whereas in the group of 472, we found almost the same frequency 2.54% (n = 12).
C9orf72 gene
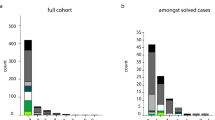
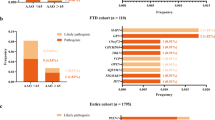
Nine patients with repeat expansions were from the FTD group (9/129; 6.98%), and two patients belonged to the UnD group (2/49; 4.08%). We did not detect any presence of the pathogenic C9orf72 repeat expansion in MCI or AD subjects. The distribution of the C9orf72 alleles is presented in Supplementary Fig. 1 and Supplementary file text. The most frequent alleles in patients and the control group were in the wild-type range with 2, 5, and 8 repeats. In addition, one FTD and one AD patient were carriers of the intermediate number of repeats (24 and 28 repeats, respectively) in C9orf72 (Supplementary Fig. 2a&b, and Supplementary file text).
C9orf72 repeats expansion in FTD
Demographic and clinical characteristics of C9orf72 carriers in the FTD group
In the FTD group (n = 129), among 52 positive family history and 77 sporadic, there was no statistically significant difference in the age of onset (p = 0.55). Nine patients (6.98%; 5 females, 4 males) were C9orf72 repeat expansion heterozygous carriers with increasing frequency (13.46%) within the group with a positive family history, and in 2.6% of sporadic cases. A significant difference was shown for the age of onset between the C9orf72 carriers and non-carriers (p = 0.007), indicating that heterozygous carriers of the C9orf72 expansion in the FTD group were about 7 years younger at the onset than FTD patients without the expansion (mean age 50.56 ± 8.25 vs. 57.63 ± 7.47). There was no difference in the age of onset (p = 0.756) nor in disease duration (p = 0.379) between the female and male carriers. Details of demographic and clinical characteristics and imaging findings are presented in Table 2. The typical clinical phenotypes varied between C9orf72 repeat expansion carriers; 7 patients with bvFTD and 2 with FTD-ALS overlapping presentation. Detailed descriptions of patients’ neurological signs and behavioral symptoms are given in Table 2.
Positive family history (n = 7) was present significantly more in patients harboring C9orf72 repeat expansion than in patients without (p = 0.018). FTD-ALS overlapping phenotype was present in five individuals (3.88%) within the FTD group. Overlapping FTD-ALS clinical presentation was more common in expansion carriers relative to patients without (22.22% vs. 2.5%, respectively).
The maximum number of C9orf72 repeats obtained among all FTD patients was ∼ 742. The correlation analysis of the number of expanded repeats and patients’ age of onset did not show significance (p > 0.05).
C9orf72 carriers within the UnD group
In the group with UnD (n = 49; 25 positive family history, 24 sporadic), 2 female patients (4.08%) carried the C9orf72 repeat expansion. Within the positive family history, both expansion carriers (8%) had a Goldman score category 3. One patient had a family member (aunt) with ALS at an age before 65y, and the other with a score 3.5 had a mother with dementia (aged over 65y). The clinical picture is described in Table 2 with various cognitive-motor problems that included stuttering, agrammatism, apraxia, walking and postural stability disorders, dysphagia, dysarthria, and cerebellar ataxia. The maximum number of the obtained repeats with Southern blot, for these two patients, was ∼ 590 repeats.
GRN variants
Pathogenic GRN variant (c.1252 C > T; p.R418*) was identified in one out of 271 screened patients (Table 2). Based on the in silico prediction software CADD, it has a score of 27.5. The variant is also present in the ClinVar database and characterized as pathogenic by multiple sources. The patient was a female aged 67y, with first clinical symptoms in the language domain that appeared several years before, with difficulties finding the right word when speaking or starting a conversation. Also, she exhibited hesitant speech, and preferred listening. The symptoms worsened as time passed, with pauses in speech, speaking in shorter sentences, and grammatical errors. She was diagnosed with a PNFA variant of FTD. The patient’s mother was diagnosed with dementia at the age of 64y, while the patient’s sister had a more severe clinical picture diagnosed at the age of 52y.
MAPT variants
Screening of selected MAPT exons in 276 patients did not reveal proven pathogenic variants in our cohort. We identified only 1 benign variant in one patient (Excel file in Supplementary material).
Discussion
This report presents the first analysis of the frequency of pathogenic variants in the three major FTD genes (C9orf72, GRN, and MAPT) within the Serbian population from the Memory Center. The study included individuals with neurodegenerative conditions such as AD, FTD, MCI, and UnD. Among the 472 individuals tested, pathogenic variants were found in 12 cases, accounting for a frequency of 2.64%. Specifically, pathogenic variants were identified in C9orf72 (11 cases) and GRN (1 case), while in MAPT, there were no pathogenic variants.
In total, 7.75% of the FTD group had pathogenic variants in the C9orf72 (n = 9) and GRN (n = 1) genes. These results are lower when compared to findings from Greece, Germany, Turkey, and Sweden [25,26,27,28]. The frequency of pathogenic variants with a positive family history was highest in our FTD cases (15.38%) and lower in sporadic cases (2.60%). A recent study in the Turkish cohort revealed a 25% rate of pathogenic variants in the selected genes among familial cases [27]. Similarly, the Greek study reported a 23.9% rate in familial cases and only one sporadic case with a likely pathogenic variant in MAPT [25]. Overall, pathogenic variants in GRN, MAPT, and hexanucleotide repeat expansions in C9orf72 can be present in 60% of familial FTD cases [29].
In this study, the C9orf72 repeat expansion was detected in 13.46% of familial FTD cases and 2.60% of sporadic FTD cases, which is relatively lower compared to other studies. Published data on the frequency of the C9orf72 repeat expansion in FTD cohorts from Western Europe shows a possible north-south descending gradient. The reported frequency reaches up to 28.7% in familial and up to 6% in sporadic FTD cases [30, 31]. The highest mutation rates were observed in Scandinavian familial FTD cases, such as Sweden (26.5%) [28] and Finland (29.33%) [32], while Italy and Portugal had the lower rates (6.09% and 6.62%, respectively). Germany registered the lowest frequency at 4.82% [32]. Our findings show that the frequency of C9orf72 repeat expansion in sporadic cases was below 3%, consistent with previously published data [31]. In our study, the presence of a positive family history was significantly more frequent among expansion carriers compared to those without the expansion, aligning with previously published data [33].
In our group of FTD expansion carriers, the average age of onset was around 50 years, which is lower than the previously reported 57 years [30]. The youngest carrier in our cohort was 36 years old, and the oldest was 64 years old at the age of onset, fitting within the expected range of 30 to 76.3 years [30, 31, 34].
The predominant clinical presentation in our C9orf72 expansion-positive patients was the bvFTD phenotype (7 out of 9 cases), consistent with previous reports [30, 31, 35, 36]. In this study, the FTD-ALS phenotype strongly predicted a pathogenic C9orf72 mutation, observed in 2 out of 5 cases.
In our UnD cases with a positive family history, we had a relatively high frequency (8%) of C9orf72 repeat expansion carriers. This finding aligns with previous research, as C9orf72 expansions have been detected in various neurodegenerative and psychiatric disorders [37]. Additionally, rare cases of C9orf72 expansions associated with ataxia syndrome are reported in the literature [38], which explains the occurrence of ataxia in one of our UnD patients.
In this study, we have identified only one GRN pathogenic variant in the FTD group, found in a 65-year-old female patient with a PNFA phenotype and a positive family history. Previous reports indicate that clinical presentations of patients with GRN pathogenic variants include PNFA, progressive mixed aphasia, and social-executive disorder [39]observed in 24% of such patients [40]. The relative frequency of GRN variants in FTD is 4.8% overall and 12.8% in familial forms. Notably, 3.2% of apparently sporadic FTD patients carried a GRN pathogenic or likely pathogenic variant, suggesting the possibility of de novo mutations or incomplete penetrance [41]. Therefore, more systematic genetic testing should be considered, even in patients without an apparent family history of FTD [41].
The current study has several limitations. First, we were unable to confirm the diagnosis for many deceased family members and lacked pathohistological diagnoses. However, all subjects were evaluated using recent clinical criteria [12, 13] and utilized available CSF biomarkers and FDG PET for AD diagnosis. Challenges such as limited clinical and family history, misdiagnoses, and psychiatric disorders mimicking FTD can make it difficult to accurately estimate a disease frequency [42]. Our study focused on patients primarily under 65 years old, but recent findings suggest FTD incidence peaks at 71 years [43].
The study confirms that C9orf72 repeat expansion is the most common genetic cause of FTD in Serbia, although slightly lower when compared to European averages. Genetic testing for this expansion is recommended for patients with this neurodegenerative disease. It is important to consider testing patients with UnD based on the diverse clinical similarities. Further research is needed to explore geographic and ethnic implications. Our study did not find pathogenic variants in MAPT, possibly due to targeted analysis or the rarity of these variants in our population. Healthcare professionals should be aware of the genetic risk in patients with FTD and UnD. Genetic counseling is advised for patients and families with a detailed family history.
References
Onyike CU, Diehl-Schmid J (2013) The epidemiology of frontotemporal dementia. Int Rev Psychiatry 25(2):130–137
Rohrer JD, Guerreiro R, Vandrovcova J, Uphill J, Reiman D, Beck J et al (2009) The heritability and genetics of frontotemporal lobar degeneration. Neurology 73(18):1451–1456
Greaves CV, Rohrer JD (2019) An update on genetic frontotemporal dementia. J Neurol 266(8):2075–2086
Borroni B, Bonvicini C, Galimberti D, Tremolizzo L, Papetti A, Archetti S et al (2011) Founder effect and estimation of the age of the Progranulin Thr272fs mutation in 14 Italian pedigrees with frontotemporal lobar degeneration. Neurobiol Aging 32(3):555e1–555e8
Barandiaran M, Estanga A, Moreno F, Indakoetxea B, Alzualde A, Balluerka N et al (2012) Neuropsychological features of asymptomatic c.709-1G > A progranulin mutation carriers. J Int Neuropsychological Society: JINS 18(6):1086–1090
Le Ber I (2013) Genetics of frontotemporal lobar degeneration: an up-date and diagnosis algorithm. Rev Neurol 169(10):811–819
Seelaar H, Rohrer JD, Pijnenburg YA, Fox NC, van Swieten JC (2011) Clinical, genetic and pathological heterogeneity of frontotemporal dementia: a review. J Neurol Neurosurg Psychiatry 82(5):476–486
Cacace R, Van Cauwenberghe C, Bettens K, Gijselinck I, van der Zee J, Engelborghs S et al (2013) C9orf72 G4C2 repeat expansions in Alzheimer’s disease and mild cognitive impairment. Neurobiol Aging 34(6):1712e1–1712e7
Kohli MA, John-Williams K, Rajbhandary R, Naj A, Whitehead P, Hamilton K et al (2013) Repeat expansions in the C9ORF72 gene contribute to Alzheimer’s disease in caucasians. Neurobiol Aging 34(5):1519e5–151912
Ramos EM, Dokuru DR, Van Berlo V, Wojta K, Wang Q, Huang AY et al (2020) Genetic screening of a large series of north American sporadic and familial frontotemporal dementia cases. Alzheimer’s Dement J Alzheimer’s Assoc 16(1):118–130
McKhann GM, Knopman DS, Chertkow H, Hyman BT, Jack CR Jr., Kawas CH et al (2011) The diagnosis of dementia due to Alzheimer’s disease: recommendations from the National Institute on Aging-Alzheimer’s Association workgroups on diagnostic guidelines for Alzheimer’s disease. Alzheimer’s Dement J Alzheimer’s Assoc 7(3):263–269
Rascovsky K, Hodges JR, Knopman D, Mendez MF, Kramer JH, Neuhaus J et al (2011) Sensitivity of revised diagnostic criteria for the behavioural variant of frontotemporal dementia. Brain 134(Pt 9):2456–2477
Gorno-Tempini ML, Hillis AE, Weintraub S, Kertesz A, Mendez M, Cappa SF et al (2011) Classification of primary progressive aphasia and its variants. Neurology 76(11):1006–1014
von Elm E, Altman DG, Egger M, Pocock SJ, Gotzsche PC, Vandenbroucke JP et al (2008) The strengthening the reporting of Observational studies in Epidemiology (STROBE) statement: guidelines for reporting observational studies. J Clin Epidemiol 61(4):344–349
Beck J, Rohrer JD, Campbell T, Isaacs A, Morrison KE, Goodall EF et al (2008) A distinct clinical, neuropsychological and radiological phenotype is associated with progranulin gene mutations in a large UK series. Brain 131(Pt 3):706–720
Richards S, Aziz N, Bale S, Bick D, Das S, Gastier-Foster J et al (2015) Standards and guidelines for the interpretation of sequence variants: a joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet Medicine: Official J Am Coll Med Genet 17(5):405–424
Landrum MJ, Lee JM, Benson M, Brown GR, Chao C, Chitipiralla S et al (2018) ClinVar: improving access to variant interpretations and supporting evidence. Nucleic Acids Res 46(D1):D1062–D7
Karczewski KJ, Francioli LC, Tiao G, Cummings BB, Alfoldi J, Wang Q et al (2020) The mutational constraint spectrum quantified from variation in 141,456 humans. Nature 581(7809):434–443
Sim NL, Kumar P, Hu J, Henikoff S, Schneider G, Ng PC (2012) SIFT web server: predicting effects of amino acid substitutions on proteins. Nucleic Acids Res. ;40(Web Server issue):W452–W457
Adzhubei IA, Schmidt S, Peshkin L, Ramensky VE, Gerasimova A, Bork P et al (2010) A method and server for predicting damaging missense mutations. Nat Methods 7(4):248–249
Rentzsch P, Witten D, Cooper GM, Shendure J, Kircher M (2019) CADD: predicting the deleteriousness of variants throughout the human genome. Nucleic Acids Res 47(D1):D886–D94
Renton AE, Majounie E, Waite A, Simon-Sanchez J, Rollinson S, Gibbs JR et al (2011) A hexanucleotide repeat expansion in C9ORF72 is the cause of chromosome 9p21-linked ALS-FTD. Neuron 72(2):257–268
DeJesus-Hernandez M, Mackenzie IR, Boeve BF, Boxer AL, Baker M, Rutherford NJ et al (2011) Expanded GGGGCC hexanucleotide repeat in noncoding region of C9ORF72 causes chromosome 9p-linked FTD and ALS. Neuron 72(2):245–256
Rollinson S, Bennion Callister J, Young K, Ryan SJ, Druyeh R, Rohrer JD et al (2015) Small deletion in C9orf72 hides a proportion of expansion carriers in FTLD. Neurobiol Aging 36(3):1601e1–1601e5
Ramos EM, Koros C, Dokuru DR, Van Berlo V, Kroupis C, Wojta K et al (2019) Frontotemporal dementia spectrum: first genetic screen in a Greek cohort. Neurobiol Aging 75:224e1e8
Wagner M, Lorenz G, Volk AE, Brunet T, Edbauer D, Berutti R et al (2021) Clinico-genetic findings in 509 frontotemporal dementia patients. Mol Psychiatry 26(10):5824–5832
Guven G, Lohmann E, Bras J, Gibbs JR, Gurvit H, Bilgic B et al (2016) Mutation frequency of the Major Frontotemporal Dementia Genes, MAPT, GRN and C9ORF72 in a Turkish cohort of Dementia patients. PLoS ONE 11(9):e0162592
Oijerstedt L, Chiang HH, Bjorkstrom J, Forsell C, Lilius L, Lindstrom AK et al (2019) Confirmation of high frequency of C9orf72 mutations in patients with frontotemporal dementia from Sweden. Neurobiol Aging 84:241 e21- e25
Olszewska DA, Lonergan R, Fallon EM, Lynch T (2016) Genetics of Frontotemporal Dementia. Curr Neurol Neurosci Rep 16(12):107
Majounie E, Renton AE, Mok K, Dopper EGP, Waite A, Rollinson S et al (2012) Frequency of the C9orf72 hexanucleotide repeat expansion in patients with amyotrophic lateral sclerosis and frontotemporal dementia: a cross-sectional study. Lancet Neurol 11(4):323–330
Simon-Sanchez J, Dopper EG, Cohn-Hokke PE, Hukema RK, Nicolaou N, Seelaar H et al (2012) The clinical and pathological phenotype of C9ORF72 hexanucleotide repeat expansions. Brain 135(Pt 3):723–735
van der Zee J, Gijselinck I, Dillen L, Van Langenhove T, Theuns J, Engelborghs S et al (2013) A pan-european study of the C9orf72 repeat associated with FTLD: geographic prevalence, genomic instability, and intermediate repeats. Hum Mutat 34(2):363–373
Snowden JS, Rollinson S, Thompson JC, Harris JM, Stopford CL, Richardson AM et al (2012) Distinct clinical and pathological characteristics of frontotemporal dementia associated with C9ORF72 mutations. Brain 135(Pt 3):693–708
Gijselinck I, Van Langenhove T, van der Zee J, Sleegers K, Philtjens S, Kleinberger G et al (2012) A C9orf72 promoter repeat expansion in a Flanders-Belgian cohort with disorders of the frontotemporal lobar degeneration-amyotrophic lateral sclerosis spectrum: a gene identification study. Lancet Neurol 11(1):54–65
Kartanou C, Karadima G, Koutsis G, Breza M, Papageorgiou SG, Paraskevas GP et al (2017) Screening for the C9ORF72 repeat expansion in a Greek frontotemporal dementia cohort. Amyotroph Lateral Scler Frontotemporal Degeneration 19(1–2):152–154
Rohrer JD, Isaacs AM, Mizielinska S, Mead S, Lashley T, Wray S et al (2015) C9orf72 expansions in frontotemporal dementia and amyotrophic lateral sclerosis. Lancet Neurol 14(3):291–301
Marogianni C, Rikos D, Provatas A, Dadouli K, Ntellas P, Tsitsi P et al (2019) The role of C9orf72 in neurodegenerative disorders: a systematic review, an updated meta-analysis, and the creation of an online database. Neurobiol Aging. ;84:238 e25- e34.
Corcia P, Vourc’h P, Guennoc AM, Del Mar Amador M, Blasco H, Andres C et al (2016) Pure cerebellar ataxia linked to large C9orf72 repeat expansion. Amyotroph Lateral Scler Frontotemporal Degeneration 17(3–4):301–303
Van Deerlin VM, Wood EM, Moore P, Yuan W, Forman MS, Clark CM et al (2007) Clinical, genetic, and pathologic characteristics of patients with frontotemporal dementia and progranulin mutations. Arch Neurol 64(8):1148–1153
Gass J, Cannon A, Mackenzie IR, Boeve B, Baker M, Adamson J et al (2006) Mutations in progranulin are a major cause of ubiquitin-positive frontotemporal lobar degeneration. Hum Mol Genet 15(20):2988–3001
Le Ber I, van der Zee J, Hannequin D, Gijselinck I, Campion D, Puel M et al (2007) Progranulin null mutations in both sporadic and familial frontotemporal dementia. Hum Mutat 28(9):846–855
Benussi A, Padovani A, Borroni B (2015) Phenotypic heterogeneity of monogenic Frontotemporal Dementia. Front Aging Neurosci. ;7
Logroscino G, Piccininni M, Graff C, Hardiman O, Ludolph AC, Moreno F et al (2023) Incidence of syndromes Associated with Frontotemporal Lobar Degeneration in 9 European countries. JAMA Neurol 80(3):279
Acknowledgements
We thank the patients and caregivers who agreed to participate in the study.
Funding
This study was supported by the Ministry of Education and Science of the Republic of Serbia (grant no. 200110).
Author information
Authors and Affiliations
Contributions
Elka Stefanova, MD, PhD: Conceptualization; Data curation; Formal analysis; Investigation; Resources; Project Administration; Visualization; Writing Drafting/revision of the manuscript. Ana Marjanović, PhD: Conceptualization; Data curation; Investigation; Methodology; Visualization; Writing Drafting/revision of the manuscript. Valerija Dobričić, PhD: Conceptualization; Investigation; Methodology; Visualization; Writing Drafting/revision of the manuscript. Gorana Mandić Stojmenović, MD, PhD: Resources. Tanja Stojković, MD, PhD: Resources. Marija Branković, PhD: Investigation; Methodology. Maksim Šarčević, MD: Resources. Ivana Novaković, MD, PhD: Conceptualization; Resources; Funding Acquisition; Writing Drafting/revision of the manuscript. Vladimir S Kostić: Conceptualization of the study, Funding Acquisition; Writing Drafting/revision of the manuscript.
Corresponding author
Ethics declarations
Data sharing statement
Anonymized data not published within this article will be made available by request from any qualified investigator.
Conflict of interest
The authors declare no conflict of interests.
Consent statement
All of the participants provided written informed consent to participate in the study.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Stefanova, E., Marjanović, A., Dobričić, V. et al. Frequency of C9orf72, GRN, and MAPT pathogenic variants in patients recruited at the Belgrade Memory Center. Neurogenetics 25, 193–200 (2024). https://doi.org/10.1007/s10048-024-00766-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10048-024-00766-8