Abstract
Adult diffuse gliomas form a heterogeneous group of tumors of the central nervous system that vary greatly in histology and prognosis. A significant advance during the last decade has been the identification of a set of genetic lesions that correlate well with histology and clinical outcome in diffuse gliomas. Most characteristic driver mutations consist of isocitrate dehydrogenase 1 (IDH1) and IDH2, and H3 histone family member 3A, which are strongly associated with DNA and histone methylation patterns. A well-characterized DNA methylation aberration is on the O6-methylguanine-DNA methyltransferase promoter. This aberration is associated with an improved response to the DNA alkylating agent, temozolomide. Methylation alterations are used for classification or treatment decisions of diffuse gliomas. This supports the importance of considering epigenomic aberrations in the pathogenesis of gliomas. Recent DNA methylation analyses revealed a small group of IDH mutant diffuse gliomas exhibiting decreased DNA hypermethylation resulting in substantial unfavorable prognosis comparable to glioblastoma. Thus, DNA methylation patterns may become a new standard that replaces the conventional grading system based on histological diagnosis. In this review, we summarize recent developments regarding the contributions of methylation patterns to the pathogenesis of adult diffuse glioma, the interactions between methylation patterns and driver mutations, and potential epigenomic targeted therapies.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Diffuse gliomas, the most frequent primary brain tumors, are a heterogeneous group of brain tumors with distinct histological and clinical features. Classification and tumor grading of diffuse gliomas were originally defined by histological diagnosis [1]. However, this classification strategy is subjective and does not reflect intratumoral heterogeneity and interobserver variation of the histological diagnosis, suggesting that it cannot reliably guide patient care [2, 3]. The advent of microarray and next-generation sequencing enabled genome-wide genomic and transcriptomic sequencing and has revealed several genetic alterations that clearly classify diffuse gliomas into discrete subtypes with characteristic molecular and clinical features. This knowledge led to the adoption of an integrated diagnosis with molecular information in the World Health Organization (WHO) Classification of Tumors of the Central Nervous System (CNS), revised in 2016 [4].
Mutations of isocitrate dehydrogenase 1 (IDH1) and IDH2 occur in the majority of lower grade gliomas (LGGs) [5, 6]. IDH mutant LGGs are subclassified into two groups based on the presence (oligodendroglioma) or absence (astrocytoma) of chromosome 1p and 19q co-deletion (1p19q) [7, 8]. Glioblastomas (GBM) are also classified based on the IDH mutation. IDH mutant GBM has genetic alterations and an age distribution similar to LGGs, and largely consists of secondary GBM that originates from a preexisting LGG [5, 9]. IDH mutations lead to the production of 2-hydroxyglutarate (2-HG) that inhibits the activity of histone and DNA demethylases, resulting in the hypermethylation of DNA and histones that drive the disease phenotype.
Other diffuse gliomas frequently have mutations in genes that encode chromatin-regulating enzymes. H3 histone family member 3A (H3F3A) and histone cluster 1 H3 family member b/c (HIST1H3B/C) encode the H3.3 and H3.1 histone H3 variants, respectively. H3K27M, which results from replacement of the 27th lysine (K) residue of these genes by methionine (M), is a characteristic mutation for pediatric midline high-grade gliomas, whereas H3G34R/V, which results from replacement of the 34th glycine (G) residue by arginine (R) or valine (V), is found frequently in hemispheric high-grade gliomas in children and young adults [10,11,12]. The H3F3A mutation inhibits methyltransferases resulting in aberrant genome-wide DNA methylation patterns.
These driver gene mutations alter genome-wide or focal DNA methylation patterns, and clearly classify diffuse gliomas into distinct subtypes with characteristic clinical features, including age distribution, tumor location, and prognosis. This suggests the importance of epigenomic changes in the initiation and progression of diffuse gliomas. Several classification schemes based on genome-wide DNA methylation were proposed and could provide further refinement in each defined subgroup of diffuse gliomas. Here, we review recent developments regarding the contributions of DNA methylation patterns to the pathogenesis of adult diffuse glioma, the interactions between methylation patterns and driver mutations, and potential epigenomic targeted therapies, including inhibitors of DNA methylation and IDH mutant enzymes.
DNA methylation
One of the most commonly studied epigenetic alterations in malignant tumors is DNA methylation. DNA methylation is the covalent transfer of a methyl group to the 5′ position of the cytosine ring, primarily at a cytosine-phosphate-guanine (CpG) dinucleotide, resulting in 5-methylcytosine. The DNA methylation status results from the action of methyltransferases and demethyltransferases. DNA methyltransferase 1 (DNMT1) is responsible for maintenance of the DNA methylation pattern after DNA replication, whereas DNMT3A, DNMT3B, and DNMT3L are responsible for de novo methylation [13,14,15]. The ten-eleven translocation family of enzymes 1–3 are involved in DNA demethylation by conversion of 5-methylcytosine to 5-hydroxymethylcytosine (Fig. 1) [16]. CpG islands are short interspersed DNA sequences that deviate significantly from the average genomic pattern by an elevated G + C base composition, and are located preferentially at gene promoters [17].
Mutant isocitrate dehydrogenase (IDH)-induced methyltransferase of histones and DNA. Wild-type IDH converts isocitrate into α-ketoglutarate (α-KG), producing NADPH. Mutant IDH converts α-KG into 2-hydroxyglutarate (2-HG), consuming NADPH. 2-HG inhibits the activity of histone lysine demethylase and ten-eleven translocation enzymes (DNA demethylases), resulting in the hypermethylation of histones and DNA. Inhibition of DNA methyltransferase and mutant IDH results in demethylation of DNA and/or histones. wtIDH wild-type IDH, mutIDH mutant IDH, 3me tri-methylation, 5mC 5-methylcytosine, 5hmC 5-hydroxymethylcytosine, 5-AZA 5-azacytidine, DEC 5-aza-2′-decitabine
One of the most important characteristics of cancer is the decrease in global DNA methylation (demethylation) affecting intergenic regions, DNA repetitive sequences, and gene bodies, including regulatory sequences. By contrast, hypermethylation of CpG islands in promoter regions is also a frequent phenomenon [18]. Such hypermethylation is an important mechanism of inducing transcriptional silencing of tumor suppressor and DNA repair genes, and may be a critical step during tumor formation [19, 20]. In GBM, aberrant promoter DNA methylation patterns of genes involved in key biological pathways have been reported. For example, the retinoblastoma, receptor tyrosine kinase (RTK)/phosphoinositide 3-kinase (PI3K), p53, and WNT pathways are affected by hypermethylation of CpG island promoters [21,22,23,24,25,26,27].
Glioma CpG island methylator phenotype (G-CIMP) and IDH mutation
Originally, the CIMP was defined by the genome-wide hypermethylation of CpG islands, first described in the context of colorectal cancers [28]. The concept of CIMP quickly became the focus of several cancer studies, including gliomas. A large fraction of gliomas exhibit substantial DNA hypermethylation and have been termed as G-CIMP [29, 30]. CIMP-positive tumors carry distinct clinicopathological and molecular features; the G-CIMP-positive subtype is closely associated with mutations of IDH1 and IDH2, and an improved prognosis [30].
Somatic mutations in IDH are observed in a wide spectrum of human cancers, most commonly diffuse gliomas, myeloid malignancies, chondrosarcomas, and intrahepatic cholangiocarcinomas [5, 31,32,33]. IDH is an important metabolic enzyme in the tricarboxylic acid cycle. Wild-type IDH catalyzes the oxidative decarboxylation of isocitrate into α-ketoglutarate (α-KG), which is associated with the regulation of histone and DNA demethylation in normal cells (Fig. 1). IDH gene mutations are drivers for the development of gliomas and map to specific arginine residues within the catalytic pocket of the enzyme; mutations in IDH1 and IDH2 occur mostly at arginines 132 and 172, respectively [34, 35]. Mutant IDH converts α-KG to the so-called “oncometabolite” 2-HG, which is a competitive inhibitor of the protein family of α-KG-dependent dioxygenases, including histone lysine demethylase and the ten-eleven translocation family enzymes involved in DNA demethylation (Fig. 1) [36, 37]. Because αKG regulates histone and DNA demethylation, inhibition of α-KG by 2-HG results in cellular hypermethylation. Mutant IDH changes the cellular redox environment by altering the ratio of NADPH to NADP+, and is also involved with hypoxia inducible factor-1α regulatory proteins and nucleic acid metabolism [38]. Furthermore, IDH mutant gliomas display a characteristic tumor immune microenvironment exhibiting a lower number of tumor-infiltrating lymphocytes and perturbation of nuclear factor of activated T cells transcriptional activity and polyamine biosynthesis, resulting in suppression of T cell activity [39].
O6-Methylguanine-DNA methyltransferase (MGMT) promoter methylation
MGMT is a DNA repair protein involved in cellular defenses against mutagenesis and toxicity from alkylating agents, including temozolomide. Within diffuse gliomas, most well-characterized DNA methylation aberrations are not only G-CIMP, but also MGMT promoter DNA methylation. Patients harboring gliomas with MGMT promoter DNA methylation demonstrate increased overall survival and time to progression of the disease after chemotherapy and/or radiotherapy [40, 41]. Approximately 80% of IDH mutant secondary high-grade gliomas (anaplastic astrocytoma or GBM) have MGMT methylation, suggesting a strong correlation between the two [42]. Some studies have used the MGMT methylation status as a stratification tool in clinical trials [43]. Although there is a strong correlation between IDH mutation and MGMT methylation in gliomas, the role of MGMT methylation status on the benefit from temozolomide therapy in IDH mutant glioma is less clear [44]. IDH mutant grade II and III diffuse gliomas (LGGs), in contrast to GBM, usually carry two copies of chromosome 10 on which MGMT resides (10q26.3). Thus, MGMT may not be silenced completely, resulting in resistance of MGMT to temozolomide therapy [45].
DNA methylation-based classification of diffuse gliomas
The cancer methylome consists of not only somatically acquired changes in DNA methylation, but also characteristics retaining some traits of the cell of origin. This enables one to determine the origin of metastasis of unknown primary cancers [46, 47]. Furthermore, the DNA methylation profile of cancers can be used to subclassify CNS tumors, that were considered previously to be homogeneous diseases, into discrete subtypes with characteristic molecular and clinical features [48, 49]. Recently, Capper et al. defined 82 methylation-based CNS tumor classes, encompassing new and known tumor groups, using tissue samples from more than 2800 individuals with cancer [50]. The results of this classification scheme suggested that methylation profiling may offer an avenue for expanding and improving CNS tumor diagnoses.
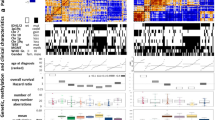
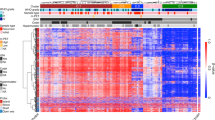
Ceccarelli et al. performed a large genome and methylome analyses using over 1000 diffuse gliomas from The Cancer Genome Atlas dataset and found seven discrete subtypes with distinct molecular and clinical features (Table 1) [51]. This DNA methylation-based classification recapitulated the aforementioned diffuse glioma classification based on IDH mutations and 1p19q co-deletion. In IDH wild-type tumors, gliomas were further classified into four subgroups: classic-like, mesenchymal-like, LGm6-GBM, and pilocytic astrocytoma (PA)-like. Most classic-like and mesenchymal-like tumors belonged to “classical” and “mesenchymal” gene expression-based subgroups, respectively, which were described by Verhaak et al. [52]. The PA-like subgroups contained a larger fraction of LGG compared to “non-PA-like” IDH-wild-type clusters, including classic-like, mesenchymal-like, and LGGm6-GBM tumors. The PA-like subgroup exhibited distinct clinical features including markedly longer survival and younger age, and was characterized by a relatively low frequency of typical GBM genetic alterations. By contrast, non-PA-like IDH-wild-type tumors carried a poor prognosis and frequent GBM-like genetic alterations such as epidermal growth factor receptor amplification, telomerase reverse transcriptase promoter mutation, and chromosome seven gain and ten loss. Thus, most non-PA-like IDH-wild-type tumors, regardless of histological diagnosis, are classified into WHO grade IV according to cIMPACT-NOW update 3 [53]. In IDH mutated and non-1p19q-co-deleted tumors (astrocytoma IDH-mut), they are further classified into two subgroups based on the extent of DNA methylation: G-CIMP-high and -low. The G-CIMP-low subgroup exhibits a substantial unfavorable prognosis comparable to GBM, and carries genetic abnormalities in cell cycle and RTK/PI3K genes such as CDKN2A, CDK4, and PIK3R1. These have been reported to be genetic alterations with unfavorable prognostic value in astrocytoma IDH-mut [54].
The updated WHO classification of CNS tumors represents a shift in tumor diagnosis by integrating histological and molecular findings [4]. However, in LGGs, the grading system based on histological criteria remains mainly focused on tumor proliferation activity. Regarding proliferative activity, there are no strict criteria for delineating mitotic index cut-off values to distinguish between grade II and III tumors. This could result in substantial interobserver variability in determining the grade [2, 3]. Some studies reported that, according to the WHO classification revised in 2016, the role of tumor grade in patient survival was not well-characterized in LGG, after accounting for the IDH mutation status and 1p19q co-deletion [34, 55, 56]. However, each LGG subgroup classified based on IDH mutation and 1p19q co-deletion has a highly variable clinical course. Based on these results, a more objective grading system using defined diffuse glioma subgroups is required. In diffuse gliomas, the G-CIMP-low subgroup may be the new standard that replaces the conventional grading system based on histological diagnosis.
DNA hypomethylation and tumor malignancy
To investigate the temporal dynamics of methylation patterns of G-CIMP, de Souza et al. compared DNA methylation profiles using primary and recurrent samples of LGGs from 77 patients (200 tumors). The results showed that some primary G-CIMP-high cases exhibited a demethylation pattern after disease recurrence that was observed primarily in G-CIMP-low tumors. This result suggested that the G-CIMP-high subgroup was a predecessor to the G-CIMP-low subgroup [51, 57]. A dramatic loss of DNA methylation during the progression and/or recurrence of IDH mutant LGGs was also reported in other papers [58, 59].
Mechanisms by which DNA hypomethylation is associated with tumor malignancy involve altered cis-regulatory elements as well as promoter hypomethylation that lead to transcriptional upregulation of genes [58]. CpG sites are found not only in gene promoters, but also in gene bodies or cis-regulatory elements such as enhancers, silencers, and insulators [60, 61]. These regulatory elements contain binding sites for transcription factors and act to increase or decrease transcription. Interplay within the protein-DNA complex forms a three-dimensional folded chromatin loop called a topology associated domain (TAD). TADs are mediated by insulator proteins containing CCCTC-binding factors (CTCFs). The chromatin structure is composed of loops, and CTCFs block communication between enhancers and promoters in intergenic regions (Fig. 2). In IDH mutant gliomas, hypermethylation at CTCF binding sites reduces the CTCF binding capacity. This leads to aberrant enhancer–gene interactions and the upregulation of oncogenes such as platelet-derived growth factor receptor-α in glioblastoma [62]. It is speculated that the G-CIMP-low subgroup exhibits a loss of genome-wide DNA methylation, including CTCF binding sites, influencing chromatin architecture by disrupting insulator binding [63].
Chromatin disorganization associated with DNA methylation in CCCTC-binding factor (CTCF) binding sites. a Topology associated domains (TADs) are mediated by insulator CTCF proteins that block the interaction between enhancers and promoters. b DNA methylation within CTCF binding sites reduces the capacity of CTCF binding. This leads to aberrant enhancer-gene interactions and the upregulation of oncogenes
DNA methylation targeted therapies
Genetic and epigenetic alterations, including mutations of IDH and H3F3A, are associated with tumor initiation and progression in diffuse gliomas. Furthermore, tumor cells routinely use the epigenomic process to escape from chemotherapy and host immune responses. Hence, a growing emphasis of recent drug discovery efforts has been on targeting the epigenome, including DNA methylation [64].
There are two classes of drugs targeting the epigenome: broad reprogrammers and drugs targeting focal regions (targeted therapy). Broad reprogrammers include DNA methylation inhibitors (DNMTi), histone deacetylase inhibitors, and bromodomain and extra-terminal motif protein inhibitors. Epigenomic targeted therapies include inhibitors of enhancer of zeste 2 polycomb repressive complex 2 (EZH2) and IDH. EZH2 targets within the polycomb repressor complex, which H3K27M inhibits [65].
DNMTi promote global DNA demethylation in a dose-dependent manner by depleting or degrading DNA methyltransferases, resulting in large-scale changes in gene expression. However, they generally did not tend to affect cancer-specific gene expression [60, 66]. DNMTi, such as 5-azacytidine and 5-aza-2′-decitabine, are effective against hematologic neoplasms and are approved by the United States Food and Drug Administration (FDA) for treating the myelodysplastic syndrome, which can progress to a rapidly growing cancer of bone marrow cells called acute myeloid leukemia (AML) [67, 68]. However, the effects of DNMTi are diverse and generally have a slow onset. Additionally, low-dose 5-azacytidine or 5-aza-2′-decitabine treatment can induce long-lasting decreases in self-renewal and tumorigenicity of tumor-initiating cells with minimal cytotoxic effects [69]. These results suggest mechanisms other than the inactivation of tumor-suppressor genes and the activation of crucial oncogenes must exist by which DNMTi can target cancer. For example, DNMTi induce a cell-autonomous immune activation response by permitting the expression of endogenous retroviruses that were silenced previously by DNA methylation [70, 71]. This antiviral response may underlie some of the antitumor activity of these drugs. A next generation DNA hypomethylating agent, guadecitabine, inhibits DNMT with better pharmacodynamic characteristics; a phase 3 study is currently being conducted in AML to delineate its effectiveness [72]. As for diffuse gliomas, a phase 1 study for 5-azacytidine is underway for patients with hematologic or solid tumor malignancies, including GBM.
As mentioned previously, IDH mutations are drivers in diffuse gliomas through 2HG production, leading to DNA and histone hypermethylation. Because the effects of 2HG on chromatin and cell differentiation are largely reversible, IDH mutant enzyme inhibiting agents may be useful for treating IDH mutant malignancies [73]. First-generation IDH mutant enzyme inhibitors demonstrated activity in AML [74, 75]. Recently, the IDH1 inhibitor, ivosidenib (AG-120), and the mutant IDH2 inhibitor, enasidenib (AG-221), induced clinical responses in phase 1/2 trials and were approved by the FDA for patients with relapsed/refractory IDH mutant AML [76, 77]. In diffuse gliomas, the mutant IDH1 inhibitor induced near-complete 2HG inhibition in vitro and in vivo, but not all IDH mutant glioma cell lines were sensitive to these inhibitors, possibly because of no appreciable changes in genome-wide DNA methylation [78, 79]. Some basic studies suggested that inhibition of mutant IDH may induce proliferation of IDH mutant glioma cell lines in vitro [78]. Decreased DNA hypermethylation was associated with the malignant phenotype and decreased overall survival in an astrocytoma IDH mutant, as mentioned previously. Genome-wide or focal demethylation with DNMTi or IDH mutant inhibiting drugs could inactivate tumor-suppressor genes, activate oncogenes, and/or promote demethylation of MGMT, which could lead to resistance to the alkylating agent, temozolomide, in GBM. Moreover, while mutation of IDH initiates gliomagenesis, and is retained upon glioma recurrence, mutant IDH and 2HG might not be required for clonal expansion at tumor recurrence [80]. This raises questions about its importance for tumor maintenance, and the suitability of targeting IDH mutants for treatment. By contrast, one clinical study showed the effectiveness of IDH inhibitors. Specifically, AG-120 monotherapy was associated with a favorable safety profile and prolonged stable disease in a previously treated non-enhancing IDH1 mutant glioma patient population [81]. Furthermore, additional IDH inhibitors were designed and have entered clinical trials. These include the IDH1-mutant inhibitors IDH-305, DS-1001b, and BAY-1436032, and a pan inhibitor of mutant IDH1 and IDH2 enzymes, vorasidenib (AG-881), which fully penetrates the blood–brain barrier. The results of these clinical trials are eagerly awaited.
Conclusion
Diffuse gliomas are increasingly understood to involve epigenetic alterations in addition to genetic modifications. Driver mutations in epigenetic regulator genes have clarified the etiology of diffuse gliomas and defined their molecular subtypes. The advent of genome-wide analysis technologies has revealed comprehensive DNA methylation patterns, as well as genomic and transcriptomic alterations in diffuse gliomas. DNA methylation-based classification schemes not only confirmed the aforementioned genetic subtypes according to IDH mutations and 1p19q co-deletion, but also identified the characteristic subtype “G-CIMP-low”, which exhibited decreased DNA hypermethylation and poor prognosis compared to its G-CIMP-high counterpart. Longitudinal studies suggested that the G-CIMP-low subgroup was a successor to the G-CIMP-high subgroup. It is expected that an analysis of the effect of demethylating specific genes will improve our understanding of the pathogenesis of glioma, regardless of its histological grade.
Researchers are rapidly developing drugs targeting the epigenome, including DNA methylation and mutant IDH, for treating IDH mutant tumors. While not all IDH mutant glioma cell lines were sensitive to these inhibitors, IDH inhibitors prolonged stable disease in patients with non-enhancing IDH1 mutant glioma. In general, non-enhancement in magnetic resonance imaging is a characteristic feature of LGG, not GBM. LGG accounts for 94 and 40% of G-CIMP-high and -low, respectively (Table 1). Thus, the extent of DNA methylation is possibly associated with whether or not the IDH inhibitor is effective.
Although this review focuses on aberrations in DNA methylation, additional epigenetic alterations contribute to the pathogenesis of glioma. These alterations include aberrant epigenetic regulation and altered histone modification patterns. A thorough examination of epigenetic alterations may reveal novel avenues of treatment, consequently increasing survival rates in patients with diffuse gliomas.
References
Louis DN, Ohgaki H, Wiestler OD et al (2007) The 2007 WHO classification of tumours of the central nervous system. Acta Neuropathol 114:97–109
Aldape K, Simmons ML, Davis RL et al (2000) Discrepancies in diagnoses of neuroepithelial neoplasms: the San Francisco Bay Area Adult Glioma Study. Cancer 88:2342–2349
van den Bent MJ (2010) Interobserver variation of the histopathological diagnosis in clinical trials on glioma: a clinician’s perspective. Acta Neuropathol 120:297–304
Louis DN, Perry A, Reifenberger G et al (2016) The 2016 World Health Organization classification of tumors of the central nervous system: a summary. Acta Neuropathol 131:803–820
Yan H, Parsons DW, Jin G et al (2009) IDH1 and IDH2 mutations in gliomas. N Engl J Med 360:765–773
Ichimura K (2012) Molecular pathogenesis of IDH mutations in gliomas. Brain Tumor Pathol 29:131–139
Cairncross JG, Ueki K, Zlatescu MC et al (1998) Specific genetic predictors of chemotherapeutic response and survival in patients with anaplastic oligodendrogliomas. J Natl Cancer Inst 90:1473–1479
Yamamichi A, Ohka F, Aoki K et al (2018) Immunohistochemical ATRX expression is not a surrogate for 1p19q codeletion. Brain Tumor Pathol 35:106–113
Parsons DW, Jones S, Zhang X et al (2008) An integrated genomic analysis of human glioblastoma multiforme. Science 321:1807–1812
Schwartzentruber J, Korshunov A, Liu XY et al (2012) Driver mutations in histone H3.3 and chromatin remodelling genes in paediatric glioblastoma. Nature 482:226–231
Wu G, Broniscer A, McEachron TA et al (2012) Somatic histone H3 alterations in pediatric diffuse intrinsic pontine gliomas and non-brainstem glioblastomas. Nat Genet 44:251–253
Wu G, Diaz AK, Paugh BS et al (2014) The genomic landscape of diffuse intrinsic pontine glioma and pediatric non-brainstem high-grade glioma. Nat Genet 46:444–450
Hermann A, Gowher H, Jeltsch A (2004) Biochemistry and biology of mammalian DNA methyltransferases. Cell Mol Life Sci 61:2571–2587
Quina AS, Buschbeck M, Di Croce L (2006) Chromatin structure and epigenetics. Biochem Pharmacol 72:1563–1569
Goll MG, Bestor TH (2005) Eukaryotic cytosine methyltransferases. Annu Rev Biochem 74:481–514
Huang Y, Rao A (2014) Connections between TET proteins and aberrant DNA modification in cancer. Trends Genet 30:464–474
Bird AP (1986) CpG-rich islands and the function of DNA methylation. Nature 321:209–213
Esteller M (2008) Epigenetics in cancer. N Engl J Med 358:1148–1159
Esteller M (2007) Epigenetic gene silencing in cancer: the DNA hypermethylome. Hum Mol Genet 16(Spec No 1):R50–R59
Berdasco M, Esteller M (2010) Aberrant epigenetic landscape in cancer: how cellular identity goes awry. Dev Cell 19:698–711
Costello JF, Berger MS, Huang HS et al (1996) Silencing of p16/CDKN2 expression in human gliomas by methylation and chromatin condensation. Cancer Res 56:2405–2410
Watanabe T, Yokoo H, Yokoo M et al (2001) Concurrent inactivation of RB1 and TP53 pathways in anaplastic oligodendrogliomas. J Neuropathol Exp Neurol 60:1181–1189
Nakamura M, Yonekawa Y, Kleihues P et al (2001) Promoter hypermethylation of the RB1 gene in glioblastomas. Lab Invest 81:77–82
Bello MJ, Rey JA (2006) The p53/Mdm2/p14(ARF) cell cycle control pathway genes may be inactivated by genetic and epigenetic mechanisms in gliomas. Cancer Genet Cytogen 164:172–173
Amatya VJ, Naumann U, Weller M et al (2005) TP53 promoter methylation in human gliomas. Acta neuropathologica 110:178–184
Lambiv WL, Vassallo I, Delorenzi M et al (2011) The Wnt inhibitory factor 1 (WIF1) is targeted in glioblastoma and has a tumor suppressing function potentially by induction of senescence. Neuro-Oncology 13:736–747
Gotze S, Wolter M, Reifenberger G et al (2010) Frequent promoter hypermethylation of Wnt pathway inhibitor genes in malignant astrocytic gliomas. Int J Cancer 126:2584–2593
Toyota M, Ahuja N, Ohe-Toyota M et al (1999) CpG island methylator phenotype in colorectal cancer. Proc Natl Acad Sci USA 96:8681–8686
Noushmehr H, Weisenberger DJ, Diefes K et al (2010) Identification of a CpG island methylator phenotype that defines a distinct subgroup of glioma. Cancer Cell 17:510–522
Turcan S, Rohle D, Goenka A et al (2012) IDH1 mutation is sufficient to establish the glioma hypermethylator phenotype. Nature 483:479–483
Mardis ER, Ding L, Dooling DJ et al (2009) Recurring mutations found by sequencing an acute myeloid leukemia genome. N Engl J Med 361:1058–1066
Amary MF, Bacsi K, Maggiani F et al (2011) IDH1 and IDH2 mutations are frequent events in central chondrosarcoma and central and periosteal chondromas but not in other mesenchymal tumours. J Pathol 224:334–343
Farshidfar F, Zheng S, Gingras MC et al (2017) Integrative genomic analysis of cholangiocarcinoma identifies distinct IDH-mutant molecular profiles. Cell Rep 19:2878–2880
Suzuki H, Aoki K, Chiba K et al (2015) Mutational landscape and clonal architecture in grade II and III gliomas. Nat Genet 47:458–468
Hartmann C, Meyer J, Balss J et al (2009) Type and frequency of IDH1 and IDH2 mutations are related to astrocytic and oligodendroglial differentiation and age: a study of 1,010 diffuse gliomas. Acta Neuropathol 118:469–474
Dang L, White DW, Gross S et al (2010) Cancer-associated IDH1 mutations produce 2-hydroxyglutarate. Nature 465:966
Ward PS, Patel J, Wise DR et al (2010) The common feature of leukemia-associated IDH1 and IDH2 mutations is a neomorphic enzyme activity converting alpha-ketoglutarate to 2-hydroxyglutarate. Cancer Cell 17:225–234
Clark O, Yen K, Mellinghoff IK (2016) Molecular pathways: isocitrate dehydrogenase mutations in cancer. Clin Cancer Res 22:1837–1842
Bunse L, Pusch S, Bunse T et al (2018) Suppression of antitumor T cell immunity by the oncometabolite (R)-2-hydroxyglutarate. Nat Med 24:1192–1203
Hegi ME, Diserens AC, Gorlia T et al (2005) MGMT gene silencing and benefit from temozolomide in glioblastoma. N Engl J Med 352:997–1003
Stupp R, Mason WP, van den Bent MJ et al (2005) Radiotherapy plus concomitant and adjuvant temozolomide for glioblastoma. N Engl J Med 352:987–996
Juratli TA, Kirsch M, Geiger K et al (2012) The prognostic value of IDH mutations and MGMT promoter status in secondary high-grade gliomas. J Neurooncol 110:325–333
Stupp R, Hegi ME, Gorlia T et al (2014) Cilengitide combined with standard treatment for patients with newly diagnosed glioblastoma with methylated MGMT promoter (CENTRIC EORTC 26071–22072 study): a multicentre, randomised, open-label, phase 3 trial. Lancet Oncol 15:1100–1108
Baumert BG, Hegi ME, van den Bent MJ et al (2016) Temozolomide chemotherapy versus radiotherapy in high-risk low-grade glioma (EORTC 22033–26033): a randomised, open-label, phase 3 intergroup study. Lancet Oncol 17:1521–1532
Bady P, Delorenzi M, Hegi ME (2016) Sensitivity analysis of the MGMT-STP27 model and impact of genetic and epigenetic context to predict the MGMT methylation status in gliomas and other tumors. J Mol Diagn 18:350–361
Fernandez AF, Assenov Y, Martin-Subero JI et al (2012) A DNA methylation fingerprint of 1628 human samples. Genome Res 22:407–419
Moran S, Martinez-Cardus A, Sayols S et al (2016) Epigenetic profiling to classify cancer of unknown primary: a multicentre, retrospective analysis. Lancet Oncol 17:1386–1395
Sturm D, Orr BA, Toprak UH et al (2016) New brain tumor entities emerge from molecular classification of CNS-PNETs. Cell 164:1060–1072
Sahm F, Schrimpf D, Stichel D et al (2017) DNA methylation-based classification and grading system for meningioma: a multicentre, retrospective analysis. Lancet Oncol 18:682–694
Capper D, Jones DTW, Sill M et al (2018) DNA methylation-based classification of central nervous system tumours. Nature 555:469–474
Ceccarelli M, Barthel FP, Malta TM et al (2016) Molecular profiling reveals biologically discrete subsets and pathways of progression in diffuse glioma. Cell 164:550–563
Verhaak RG, Hoadley KA, Purdom E et al (2010) Integrated genomic analysis identifies clinically relevant subtypes of glioblastoma characterized by abnormalities in PDGFRA, IDH1, EGFR, and NF1. Cancer Cell 17:98–110
Brat DJ, Aldape K, Colman H et al (2018) cIMPACT-NOW update 3: recommended diagnostic criteria for “Diffuse astrocytic glioma, IDH-wildtype, with molecular features of glioblastoma, WHO grade IV”. Acta Neuropathol 136:805–810
Aoki K, Nakamura H, Suzuki H et al (2018) Prognostic relevance of genetic alterations in diffuse lower-grade gliomas. Neuro Oncol 20:66–77
Cancer Genome Atlas Research N, Brat DJ, Verhaak RG et al (2015) Comprehensive, integrative genomic analysis of diffuse lower-grade gliomas. N Engl J Med 372:2481–2498
Olar A, Wani KM, Alfaro-Munoz KD et al (2015) IDH mutation status and role of WHO grade and mitotic index in overall survival in grade II-III diffuse gliomas. Acta Neuropathol 129:585–596
de Souza CF, Sabedot TS, Malta TM et al (2018) A distinct DNA methylation shift in a subset of glioma CpG island methylator phenotypes during tumor recurrence. Cell Rep 23:637–651
Mazor T, Pankov A, Johnson BE et al (2015) DNA methylation and somatic mutations converge on the cell cycle and define similar evolutionary histories in brain tumors. Cancer Cell 28:307–317
Bai H, Harmanci AS, Erson-Omay EZ et al (2016) Integrated genomic characterization of IDH1-mutant glioma malignant progression. Nat Genet 48:59–66
Yang X, Han H, De Carvalho DD et al (2014) Gene body methylation can alter gene expression and is a therapeutic target in cancer. Cancer Cell 26:577–590
Singer M, Kosti I, Pachter L et al (2015) A diverse epigenetic landscape at human exons with implication for expression. Nucleic Acids Res 43:3498–3508
Flavahan WA, Drier Y, Liau BB et al (2016) Insulator dysfunction and oncogene activation in IDH mutant gliomas. Nature 529:110–114
Malta TM, de Souza CF, Sabedot TS et al (2018) Glioma CpG island methylator phenotype (G-CIMP): biological and clinical implications. Neuro Oncol 20:608–620
Jones PA, Issa JP, Baylin S (2016) Targeting the cancer epigenome for therapy. Nat Rev Genet 17:630–641
Lewis PW, Muller MM, Koletsky MS et al (2013) Inhibition of PRC2 activity by a gain-of-function H3 mutation found in pediatric glioblastoma. Science 340:857–861
Mund C, Brueckner B, Lyko F (2006) Reactivation of epigenetically silenced genes by DNA methyltransferase inhibitors basic concepts and clinical applications. Epigenetics-Us 1:7–13
Issa JP (2005) Optimizing therapy with methylation inhibitors in myelodysplastic syndromes: dose, duration, and patient selection. Nat Clin Pract Oncol 2(Suppl 1):S24–S29
Kaminskas E, Farrell A, Abraham S et al (2005) Approval summary: azacitidine for treatment of myelodysplastic syndrome subtypes. Clin Cancer Res 11:3604–3608
Tsai HC, Li H, Van Neste L et al (2012) Transient low doses of DNA-demethylating agents exert durable antitumor effects on hematological and epithelial tumor cells. Cancer Cell 21:430–446
Chiappinelli KB, Strissel PL, Desrichard A et al (2017) Inhibiting DNA methylation causes an interferon response in cancer via dsRNA including endogenous retroviruses. Cell 169:361
Roulois D, Loo Yau H, Singhania R et al (2015) DNA-demethylating agents target colorectal cancer cells by inducing viral mimicry by endogenous transcripts. Cell 162:961–973
Kantarjian HM, Roboz GJ, Kropf PL et al (2017) Guadecitabine (SGI-110) in treatment-naive patients with acute myeloid leukaemia: phase 2 results from a multicentre, randomised, phase 1/2 trial. Lancet Oncol 18:1317–1326
Losman JA, Looper RE, Koivunen P et al (2013) R)-2-hydroxyglutarate is sufficient to promote leukemogenesis and its effects are reversible. Science 339:1621–1625
Stein EM (2015) IDH2 inhibition in AML: finally progress? Best Pract Res Cl Ha 28:112–115
Yen K, Travins J, Wang F et al (2017) AG-221, a first-in-class therapy targeting acute myeloid leukemia harboring oncogenic IDH2 mutations. Cancer Discov 7:478–493
Stein EM, DiNardo CD, Pollyea DA et al (2017) Enasidenib in mutant IDH2 relapsed or refractory acute myeloid leukemia. Blood 130:722–731
DiNardo CD, Stein EM, de Botton S et al (2018) Durable remissions with ivosidenib in IDH1-mutated relapsed or refractory AML. New Engl J Med 378:2386–2398
Rohle D, Popovici-Muller J, Palaskas N et al (2013) An inhibitor of mutant IDH1 delays growth and promotes differentiation of glioma cells. Science 340:626–630
Tateishi K, Wakimoto H, Iafrate AJ et al (2015) Extreme vulnerability of IDH1 mutant cancers to NAD + depletion. Cancer Cell 28:773–784
Mazor T, Chesnelong C, Pankov A et al (2017) Clonal expansion and epigenetic reprogramming following deletion or amplification of mutant IDH1. Proc Natl Acad Sci USA 114:10743–10748
Mellinghoff IK, Touat M, Maher E et al (2017) Ag-120, a first-in-class mutant Idh1 inhibitor in patients with recurrent or progressive Idh1 mutant glioma: updated results from the phase 1 non-enhancing glioma population. Neuro-Oncology 19:10–11
Acknowledgements
We would like to thank Editage (http://www.editage.jp) for English language editing.
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Aoki, K., Natsume, A. Overview of DNA methylation in adult diffuse gliomas. Brain Tumor Pathol 36, 84–91 (2019). https://doi.org/10.1007/s10014-019-00339-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10014-019-00339-w