Abstract
Context
In cellular environments, the reduction of disulfide bonds is pivotal for protein folding and synthesis. However, the intricate enzymatic mechanisms governing this process remain poorly understood. This study addresses this gap by investigating a disulfide bridge reduction reaction, serving as a model for comprehending electron and proton transfer in biological systems. Six potential mechanisms for reducing the dimethyl disulfide (DMDS) bridge through electron and proton capture were explored. Thermodynamic and kinetic analyses elucidated the sequence of proton and electron addition. MD-PMM, a method that combines molecular dynamics simulations and quantum-chemical calculations, was employed to compute the redox potential of the mechanism. This research provides valuable insights into the mechanisms and redox potentials involved in disulfide bridge reduction within proteins, offering an understanding of phenomena that are challenging to explore experimentally.
Methods
All calculations used the Gaussian 09 software package at the MP2/6–311 + g(d,p) theory level. Visualization of the molecular orbitals and electron densities was conducted using Gaussview6. Molecular dynamics simulations were performed using GROMACS with the CHARMM36 force field. The PyMM program (Python Program for QM/MM Simulations Based on the Perturbed Matrix Method) is used to apply the Perturbed Matrix Method to MD simulations.
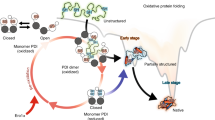
Graphical Abstract

Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
“Life is nothing more than an electron looking for a place to rest,” a citation by Hungarian Nobel laureate Albert Szent–Györgyi perfectly describes the importance of electron transfer within molecules. This phenomenon plays a crucial role in various biological processes, such as the function of photosystems I and II, which act as oxidoreductases that catalyze reactions within the respiratory chain. The redox potential (E°) of a molecule is a key measure of its ability to transfer electrons, offering valuable insight into its electronic behavior. Because this property is similar to the adiabatic ionization potential, many calculations have been performed to estimate it [1]. However, progress in this field has been limited by technical challenges such as identifying suitable reference reactions and maintaining equilibrium between redox species and concentration effects [2, 3]. On the other hand, computational approaches have become promising tools for studying chemical processes associated with redox reactions [1, 4, 5]. They have applications in electrochemical studies, including corrosion analysis [6] and water treatment [7, 8], as well as in biochemical contexts, such as determining the redox potential of enzymes such as lysyl oxidase-like 2 (LOXL2) [9] and proteins such as plastocyanin and rusticyanin [10].
Thioredoxins (Trxs) are proteins that have various functions, serving as electron and proton donors for the reduction of disulfide bridges in target proteins [11, 12]. The electronic structure of the valence shell, with a s2p4 configuration, makes these elements highly active in redox mechanisms. Trx, present in all cell types, has a Cys-S–S-Cys redox center, which is crucial for controlling numerous essential cellular processes, such as promoting cell growth, inhibiting apoptosis, and modulating inflammation [13]. In the reduced state, Trx is active, whereas it becomes inactive when oxidized [14].
In particular, the Trx system is frequently overexpressed in cancer cells and is associated with tumor growth by promoting malignant development through various mechanisms, including genetic rearrangements, gene amplification, loss of growth control, and resistance to therapies [15, 16]. In addition, the reduced form of Trx1 inhibits apoptosis when it binds to the apoptosis signal-regulating kinase (ASK-1) [17]. This critical role of the thioredoxin system in tumor progression underscores the need to effectively target this system to develop anticancer therapies [15]. One promising therapeutic target is the inhibition of disulfide bond reduction within oxidized Trx [18]. Therefore, the molecular details of this reduction mechanism need to be fully understood to advance the development of strategies aimed at selectively targeting this protein [19].
Complete reduction of the disulfide bond involves the addition of two electrons and two protons, resulting in the formation of two thiols and the breaking of the sulfur-sulfur bond. Experimental and theoretical methodologies have been employed to elucidate this mechanism in simple and complex systems [20].
Disulfide bonds have been the subject of numerous experimental studies that have focused mainly on detecting the presence or absence of these bonds but have not conclusively determined the point at which the disulfide bond is broken [21,22,23,24]. Recent research has focused on the mechanism of dimethyl disulfide (DMDS) reduction by phosphines [25].
Previous experimental work has suggested that the irreversible addition of two successive electrons to a sulfur–sulfur bond leads to the formation of an intermediate radical and is crucial for bond cleavage [26]. Another proposed mechanism involves electron capture at protonated sites [27]. Other suggestions are that sulfur–sulfur bond rupture occurs exclusively by protonation, indicating that proton addition is essential for the bond to be cleaved [28].
The addition of a single electron to a disulfide bond is the primary reaction most often chosen in theoretical investigations to study redox reactions using quantum chemical methods [20, 29, 30]. This reaction produces a radical anion that carries a two-center-three-electron (2c-3e) chemical half-bond between the sulfur atoms [31].
Another possible product state of the reduction reaction is a zwitterionic compound, resulting from the addition of an electron and a proton. Such findings suggest that the addition of a proton to the radical anion does not further stabilize the reduced state, as spontaneous bond elongation is observed by optimizations with quantum mechanical (QM) methods and hybrid quantum mechanics and molecular mechanics (QM/MM) methods [20].
For a comprehensive study and its implications for further research, we explored six reaction pathways for the two-electron, two-proton reduction of the disulfide bond in the dimethyl disulfide (DMDS) molecule. Thermodynamics and kinetics are key factors in determining the viability of a chemical reaction [32] and therefore play a crucial role in the evaluation of these reaction pathways. In addition to electron and proton affinities, the thermodynamics of electron and proton transfer reactions are characterized by redox potentials (E°) and pKa values. These two parameters serve as indicators of the free energy (ΔG°). In addition, the kinetics of these transfer reactions can be correlated with their thermochemistry using principles derived from Marcus theory [33, 34].
Computational methods
Electronic structure and thermochemistry
In this study, we employed two highly accurate ab initio methods, the second-order Møller–Plesset perturbation method (MP2), and coupled-cluster with single, double, and perturbative triple excitations (CCSD(T)), using the 6–311 + g(d,p) and aug-cc-pvtz basis sets, respectively. These combinations are well known for their extensive coverage and have previously yielded significant and noteworthy insights into disulfide studies [30].
Nine optimized structures were obtained by performing a relaxed scan of the potential energy surface (PES) along the CSSC dihedral angle to identify the optimal conformations in both the gas and solvated phases. Subsequently, vibrational frequencies were calculated at the same level of theory to ensure the absence of imaginary frequencies and to estimate thermochemical parameters.
Note that the energies labeled as CCSD(T) energies denote single-point energies computed at the CCSD(T)/aug-cc-pVTZ theoretical level, using geometries optimized through MP2/6–311 + G(d,p) calculations.
In electron transfer reactions, in which species with different electron numbers are compared, we assessed spin contamination via open-shell calculations. The observed S^2 value was approximately 0.77, which slightly deviated from the expected value of 0.75 but remained within the acceptable range (< 10%) for spin contamination. For solvent effects, we used the Implicit Integral Equation Formalism Polarizable Continuum Model (IEFPCM) within Gaussian 09, considering water as the solvent. Thermochemical parameters, including zero-point energy(ZPE) correction, were computed under standard conditions of room temperature (295.15 K) and pressure (1 atm). To simplify the calculations and comparative analyses, the reactants and products were assumed to have equal concentrations of 1 M. The free energy of the proton in water was derived from experimental data found in the literature, which is typically approximately − 262.4 kcal/mol [35]. The bond dissociation energy is the energy required to break a covalent bond and form two radical fragments through a homolytic break, each retaining one electron from the original shared pair. This energy is obtained using the enthalpy change according to Eq. (1) [36].
The equilibrium constants of the reactions were estimated using the following equation:
where \(\Delta G^\circ\) is the reaction Gibbs free energy, R is the ideal gas constant, and T is the temperature in Kelvin.
Energy barriers
The analysis of the DMDS reduction mechanism involves evaluating the energy barrier during transitions between different species. Marcus theory [33] was employed for this purpose, utilizing Eq. (3) to estimate the activation energy.
The theory relies on the Born–Oppenheimer approximation, which separates the movement of electrons and nuclei by assuming that changes in the nuclear configurations of the system occur infinitely more slowly than the quantum transfer of electrons. Consequently, electron transfer dynamics are controlled by nuclear dynamics, which include both intramolecular and solvent reorganizations [37].
The energy required to reach the electron transfer point is denoted as λ. This energy can be arbitrarily divided into two parts as Eq. (4) [37]:
- \({\lambda }_{{\text{out}}}\):
-
(outer) characterizes external environment (solvent) fluctuations, essentially polarization.
- \({\lambda }_{\text{in}}\):
-
(inner) represents the internal contribution, i.e., deformations of angles or bonds in the molecules.
Electron transfer
The reorganization energy of each electron transfer is obtained by adopting the method reported by Buda [38].
In this strategy, the internal and solvent contributions (also known as the inner and outer spheres respectively) are not separated but rather calculated as a single parameter (Eq. (5)).
where \(E\left(i/f\right)\) and \(E\left(i/i\right)\) are the energy of the initial state in the optimized geometry of the final state and the energy of the initial state in its optimized geometry, respectively. Vertical electronic transitions were calculated by considering nonequilibrium solvation [38]. This process was carried out in two steps. First, a single-point energy calculation was performed for the initial state using the keyword “noneq = write” in the PCM input section to record nonequilibrium solvation information based on this initial state. Next, the energy calculation was performed by reading the solvation information from the checkpoint file of the first step using the “noneq = read” keyword. The reorganization energy values are obtained as follows:
The value of \({\lambda }_{ET}\) is the simple arithmetic mean between the two values of \({\lambda }_{\to }\) and \({\lambda }_{\leftarrow }\).
Proton transfer
Since the proton is a quantum particle, the proton transfer (PT) reaction can also be described by the Marcus equations [39]. Consequently, the PT reorganization energy can be obtained from Eq. (4).
\({\lambda }_{\text{in}}\) is defined as the energy barrier \(\Delta {G}^{\#}\) when the free energy of the reaction is null \(\Delta {G}^{^\circ }=0\) [34]. It is estimated by calculating the initial state (before proton transfer) in the optimized geometry of the protonated final state.
\({\lambda }_{{\text{out}}}\) is obtained according to Eq. (8) [39].
where:
- \({\epsilon }_{^\circ }\):
-
Permittivity under vacuum (8.85 × 10–12 C2/N·m2).
- \({\epsilon }_{op}\):
-
Optical constant of water (1.78).
- \({\epsilon }_{s}\):
-
Static dielectric constant of water (80.0).
- \(a\):
-
The radius of the solute cavity was calculated using the VOLUME method implemented in Gaussian 09, which estimates the volume within a certain density contour (1 × 10−3 electrons/Bohr3) using Monte Carlo integration.
- \({\mu }_{IS}\):
-
Dipole moment of the initial state (D).
- \({\mu }_{FS}\):
-
Dipole moment of the final state (D).
Reaction rates
In many enzymatic reactions, sequential coupled transfer (1e−/1H+) (Fig. 1) depends on the stability of the intermediate state, thereby affecting the overall reaction rate. The rate constants were established by employing the Eyring equation (Eq. (9)) [39].
\(\Delta {G}^{\#}\) represents the activation energy, \({K}_{b}\) is Boltzmann’s constant, and \(h\) is Planck’s constant. If the intermediate state has a short lifetime (indicated by a low value of \({k}_{-1}\)), there may not be enough time for the second transfer (\({k}_{2}\)) to occur, and the reaction may not proceed efficiently. Moreover, electron transfer can occur either before or after protonation, leading to corresponding rates of \({k}_{ep}\) and \({k}_{pe}\), respectively. The overall rate constant for this reaction is given by:
In general, one of the electron–proton (ep) or proton–electron (pe) mechanisms may dominate, depending on the individual reaction rates [40]. The model by Hammes–Schiffer and coworkers responds well to these scenarios [40]:
If the reverse process of step 1 is faster than that of step 2, \({k}_{-1}\gg {k}_{2},\) then:
If the inverse process of step 1 is slower than that of step 2, \({k}_{-1}\ll {k}_{2}\), in this case:
where \(x\) = \({k}_{ep}\) or \({k}_{pe}\)
Energy decomposition analysis (EDA)
The binding energy \({\Delta E}_{{\text{bind}}}\) is the energy required to form the AB system from the isolated fragments A and B (Eq. (13)) [41]:
where \(E\left({AB}^{AB}\right)\), \(E\left({A}^{A}\right)\), and \(E\left({B}^{B}\right)\) represent the energy of the AB system and the energies of fragments A and B, respectively, each in its optimized geometry. In the EDA formalism [41], \({\Delta E}_{{\text{bind}}}\) is decomposed into interaction energy (Eq. (17)) and preparation energy as follows [42]:
Here, \(E\left({A}^{0,AB}\right)\) denotes the energy of fragment A calculated in the geometry of the AB system with a given electronic configuration (\({A}^{0}\)), which may not correspond to that of the ground state for the isolated fragment. Thus defined, the preparation energy will be the energy required to deform the fragment geometries from their isolated forms to the geometries in the AB interaction system. LMOEDA was used as implemented in the Gamess software. In this method, the total interaction energy is decomposed into electrostatic energy, exchange energy, repulsion energy, polarization energy, and dispersion energy [43]:
QM/MM calculations
For QM/MM calculations, the MD-PMM method (molecular dynamics simulations combined with the perturbed matrix method) is applied within the framework of the atom-based expansion approximation, which is capable of providing accurate estimates of thermodynamic properties at a limited computational cost [44, 45]. In this approach, the entire system is divided into two groups: the quantum center (QC), which is the region of the system where the quantum process under study occurs and is treated at the quantum mechanics level. The remaining part of the system (also called the environment), the solvent in our case, is treated as a time-dependent electrostatic perturbation, obtained through MD simulations.
Therefore, the Hamiltonian operator \(\widehat{H}\) of the quantum center (QC) integrated into the perturbing environment is as follows:
where \(\widehat{H}^\circ\) is the unperturbed electronic Hamiltonian provided by the gas-phase QM calculation, and \(\widehat{V}\) is the perturbation operator for the electrostatic effect of the environment. In this study, the entire molecules were chosen as QCs, and their unperturbance properties (electronic energies, dipole moments, and ESP charges) were evaluated at the MP2/6–311 + G(d,p) or CCSD(T)/aug-cc-pvtz level of theory.
All molecular dynamics (MD) simulations were conducted using GROMACS 5.1.2 software, employing the CHARMM36 force field in the NVT ensemble (constant particle number, volume, and temperature) at 300 K. The simulations were carried out for a duration of 40 ns with a time step of 2 fs. Topologies were generated using the Ante Chamber Python Parser (acpype) interface [46], and counterions were included to neutralize the total charge of the charged systems. To replicate experimental conditions, the volume of the cubic simulation box was selected to mimic the isobaric solute insertion in either liquid water or N,N-dimethylformamide (DMF) at infinite dilutions [47]. Long-range interactions were treated using the Ewald summation technique with an accuracy of 0.0001. All bond lengths were constrained using the LINCS algorithm, and periodic boundary conditions were applied in all directions to prevent edge effects. A simple point charge water model (SPC216) solvent model was utilized for water solvent. For simulations involving DMF solvent, we first ran a 40 ns simulation of 587 molecules of DMF in a cubic box with a length of 4.245 nm. This was done to mimic the experimental density of DMF solvent.
The acidity constants for the protonation reactions are given by Eq. (18) [48] at 25 °C in aqueous solution.
The terms in the above equation are as follows:
- \(\Delta A\):
-
Helmholtz free energy obtained from the MD-PMM calculation.
- \(\Delta {\mu }_{{H}^{+}}\):
-
The experimental solvation-free energy of a proton. We used a value of − 1112.5 kJ/mol, which indicates that the standard potential of the hydrogen electrode is E°(SHE) = 4.28 [49].
- \(\Delta {u}_{el}^{0}\):
-
Energy difference of the unperturbed electronic ground state of QC (in the gas phase) calculated during deprotonation.
- \({K}_{b}Tln\left({K}_{b}T{\rho }^{\ominus }P\right)\):
-
Correction for the standard state change from 1 atm to 1 mol L − 1 (at 298 K).
- \({G}_{{AH}^{+}\to A+{H}_{\text{gas}}^{+}}^{{\text{gaz}}}\):
-
Estimation of the resulting gas phase free energy using the following approximation:
where
- \(\Delta {A}_{v}^{{\text{gas}}}\):
-
is the vibrational contribution to the gas-phase deprotonation-free energy, and
- \({\mu }_{{H}^{+}}^{{\text{gas}}}\):
-
is the standard chemical potential of the proton in the gas phase.
Results and discussion
Intermediates and reaction pathways
Six possible reaction paths have been proposed to study the reduction of the disulfide bridge in dimethyl disulfide (DMDS). This reaction occurs by capturing two electrons and two protons (Fig. 2).
Each pathway in Fig. 2 involves three intermediates before reaching the two thiols. We therefore distinguished seven intermediates, including three anionic species (B, C, F); three cationic species (D, G, H); and one possibly zwitterionic compound (E).
From Table 1, we can identify three distinct classes of intermediates, classified according to their disulfide bond type. The first class comprises species (D and G) with covalent bonds (2c-2e), indicating that protonation alone has no major influence on bond elongation. The SS bond distance increases by 0.97% and 4.32%, respectively. The second class is composed of species characterized by a hemibond (B and H); two centers and three electron bonds are noted (2c-3e), where the electronic distribution between the two sulfur atoms is equally balanced. They correspond to elongations of 33.5% and 35.4%, respectively. Finally, the C, E, and F species have no disulfide bonds in their molecular structure. Vibration frequencies vary in correlation with intersulfide distances. A higher vibration frequency indicates a higher energetic vibration and therefore corresponds to a relatively strong and rigid bond, particularly for species A, D, and G. On the other hand, a very low vibration frequency is associated with species C, E, and I.
These results suggest that disulfide bond cleavage necessarily involves the addition of at least one electron and one proton, or two successive electrons.
A comparison of the dihedral angles in the gas phase with those in solution (Table 2) reveals the effect of the solvent on the reaction, manifested mainly in the orientation of the dihedral angle, which varies by at most 20°. In the presence of the solvent, protonations lead to more open angles compared to those formed upon reduction. The dihedral angle around the disulfide bond is indeed the degree of freedom that undergoes the most changes during reduction [50].
Bond dissociation energies (BDEs) and reduction trends
The bond dissociation energies (Table 3 and Fig. 3) enable the identification of trends in the disulfide bond strength during dimethyl disulfide reduction.
Investigation of the BDE variation of disulfide through reduction pathways starting with protonation (c) or with electron uptake (d). “E” and “P” denote electron transfer and proton transfer reactions, respectively. For instance, the EPPE pathway involves a sequence of electron–proton–proton–electron transfer reactions
The initial step produces the first anionic intermediate (B) with a BDE dissociation energy ~ 6 times smaller than that of the neutral molecule (A). This energy is further reduced by 8.53 kcal/mol after protonation. Moreover, the successive addition of two electrons leads to a negative value, indicating an unstable intermediate (C).
The protonation of the system (E) increases the BDE by 22.7 kcal/mol, compared to B, which indicates a stronger bond. Therefore, disulfide bond rupture may be less favored after this step. However, we note that the protonation of DMDS leading to D causes a slight weakening of the bond studied.
This trend in dissociation energy suggested that the disulfide bond became easier to break as electrons were added; protonation had a slight impact.
Electronic and energy parameters
Electron affinities
Electron uptake can be measured by electron affinity (EA), which represents the amount of energy released when a gas-phase system captures an electron. The higher the EA is, the greater the energy released and the more favorable the reaction.
Figure 4 illustrates the adiabatic electron affinity (AEA) and vertical electron affinity (VEA) of each reduction step. These values are calculated as the difference in total energies between (i) the optimized initial and anionic states and (ii) the initial and anionic states in the geometry of the initial molecule. First, an electron can be added to a neutral system (A), after a protonation step (D), or to a doubly protonated system (G). Then, the second electron can either be captured by (E) molecule or added as a final step to the cationic system (H). We note that the presence of protons leads to a greater affinity for electrons. In the case of the first reduction, this increases from 0.009 eV (AEA of the neutral molecule) to 6.48 eV with a single proton and to 12.74 eV with two protons, which is twice that of the AEA of the once-protonated molecule.
In general, each protonation generates an increase of approximately 6 eV in the AEA, which translates into an increase in the exothermicity of approximately 141 kcal/mol.
We confirmed this result by examining the gas-phase free energies corresponding to the first and second reductions for the singly and doubly protonated states (Table 4).
The electron uptake by the doubly protonated state releases approximately 141 kcal/mol more energy than the electron uptake by a singly protonated system. This excess energy is remarkably close to the proton affinity of the sulfur atom (158.9 kcal/mol) [51].
The negative value of VEA underlines the importance of geometrical relaxation for the initial capture of the first electron by the neutral dimethyl disulfide, without which the odd electron is spontaneously repelled [50].
Figure 5 illustrates the predicted geometry changes (relaxation energy) obtained by subtracting the VEA from the AEA. The most significant change in geometry occurs when an electron is added to a protonated disulfide (39.42 kcal/mol). In contrast, the lowest relaxation energy characterizes electron capture by the intermediate (E) (6.58 kcal/mol), indicating that (E) and (F) have similar geometries.
Proton affinities
Gas-phase proton affinity (PA) is defined as the opposite of the enthalpy variation associated with the gas-phase reaction between a proton and a molecule [52]. It thus measures the tendency of a molecule to react with a proton to form its conjugate acid [53]. It is also worth mentioning the concept of gas-phase basicity (GB), which is related to the free energy of the aforementioned reaction 52 (Table 5).
The highest proton affinities, 14.57 and 14.90 eV, correspond to negatively charged species B and F, respectively. On the other hand, neutral systems A and E have relatively intermediate proton affinities of 8.32 eV (\({PA}_{{\text{exp}}}=8.45\) eV and \({GB}_{{\text{exp}}}=8.12\) eV [54] and 9.10 eV, respectively.
To better understand how protonation occurs energetically and structurally, we need a solid, physical description of the process. By virtue of quantum mechanics, which treats all the constituents of a quantum system as a whole, we can break down a protonation reaction (composed of a molecule and a proton in a vacuum) into three distinct and successive stages [53]: (a) ionization of the molecule, (b) attachment of the ejected electron to the incident proton and formation of the hydrogen atom, and (c) creation of a new bond between the molecular cation radical and hydrogen. The first step involves a cost in the form of the ionization energy required for protonation. However, this expense is offset by the electronic affinity of the proton (13.6 eV), which significantly contributes to the overall exothermicity of the process. Protonation is therefore possible when the ionization energy (IP) of the molecule is lower than the electronic affinity of the proton EA(H+) = 13.6 eV [52]. Furthermore, this process is more favorable when the ionization energy is low, thus reducing the associated energy costs. In this context, the protonation of species (D) is unfavorable, as its ionization potential is 14.38 eV. The latter has an electron affinity twice that of the proton and is therefore more likely to capture an electron.
According to Scheme (a) of Fig. 6, compound (B) has a high affinity for accepting a proton at a very low energy cost (0.01 eV). Regarding the second protonation, the anionic molecule (F) has the greatest proton affinity and lowest ionization energy. Therefore, we suggest that the protonation steps cannot occur successively. First, a proton is added to the anionic molecule (B). After a reduction step, another protonation occurs to form the two-thiol compound (I).
Thermodynamic and kinetic considerations for pathway selection
To reduce the number of potential reaction paths, we followed the strategy proposed by Galano [55]. This method is primarily based on evaluating the thermochemistry of all reaction pathways through their Gibbs free energies of reaction (∆G). Then, the data were used to identify the endergonic (∆G > 0) and exergonic (∆G < 0) paths, and we later considered only the exergonic routes for further kinetic analysis. This simplified strategy is based on the assumption that the endergonic pathways are reversible and that the products formed will therefore not be observed. However, it is important to note that these pathways can still be important if their products react rapidly. In this case, they should be included in the kinetic study.
Thermodynamically favorable pathways
Figure 7 shows the energy diagram for each potential reaction pathway. In each scenario, the electron transfer steps exhibit a decrease in energy, indicating that they release more energy than protonation reactions, which conversely show an increase in energy.
These findings show that DMDS protonation is energetically unfavorable. The EPEP and EEPP routes, on the other hand, may be favorable because they avoid high-energy intermediates. These observations suggest that DMDS reduction is initiated by an electron capture step to form the (DMSD) ion, thus ruling out the PEPE, PEEP, and PPEE pathways. Furthermore, the second protonation in the EPPE and PEPE pathways is marked by a high energy of 63 kcal/mol, making it highly unfavorable. One can conclude that the free energy diagram suggests two possible pathways, both starting with an electron capture step, namely, the EEPP and EPEP pathways. It is crucial to complete this study by examining the direction of the reactions to provide a more complete description of the stability of the intermediates formed. The equilibrium constants Ke for each elementary reaction are presented in the following section (Table 6). High values of Ke (> 1) indicate that the reactions strongly favor the formation of products. In other words, at equilibrium, the concentration of the products will be significantly greater than that of the reactants [56]. However, the low Ke values (< 1) associated with the protonation of systems A and E suggest that these two reactions occur in the reverse direction, favoring the reformation of reactants.
Although our initial results indicated that protonation is an endothermic reaction, it is important to consider the potential influence of explicit solvent–solute interactions, which are not fully accounted for in the continuum approach to solvation. To explore this further, we conducted a quantum mechanical/molecular mechanical (QM/MM) calculation using the MD-PMM approach outlined in the methodology section. Our findings revealed that even in the presence of explicit aqueous solvent molecules, the protonation of DMDS remained highly unfavorable (\(\Delta {G}^{0}\) = 71.87 kcal/mol).
Reaction kinetics and rate constants
The simplest approach to integrate kinetics into theoretical methodologies is likely through the calculation of reaction barriers \(\Delta {G}^{\#}\) [57]. To determine whether it is necessary to include the PEPE and PEEP pathways in the kinetic analysis, we evaluated the rate of the D + 1e reaction after the initial endogenous protonation step. This process exhibits rapid kinetics, with a rate constant K = 1.9 × 1011 s−1. Consequently, both pathways appear to be particularly crucial in this study. The parameters required to apply Eq. (3) to the reduction and protonation reactions are outlined in Tables 7 and 8, respectively.
In each reaction, the driving force (− ∆G°) is greater than the total reorganization energy (λ). Thus, the process occurs in the inverted region of Marcus. This explains the high energy barrier of the E + 1e− → F reaction. In this region, electron transfer reactions slow down when they become highly thermodynamically favorable, a counterintuitive relationship between kinetics and thermodynamics [58].
We excluded the PPEE and EEPP routes from this study. The former was discarded due to its unfavorable energetics, whereas the latter involves a second electron transfer with a significantly high barrier of 133.10 kcal/mol, making its kinetics disadvantageous. Figure 8 illustrates the reaction pathways included in the kinetic analysis. The horizontal and vertical arrows indicate the reduction and protonation steps, respectively. The activation energy for each step is shown above the corresponding arrow in the unit of kcal/mol.
Reduction of the neutral molecule occurs without any major energy barrier (0.12 kcal/mol). This means that the reaction is close to the transition point between the normal regime and the inverse regime, as defined in Marcus’ theory. This point corresponds to the moment when electron transfer is the fastest [59]. In comparison, electron transfer after the protonation step is accompanied by a low barrier of 2.06 kcal/mol. On the other hand, protonation of neutral DMDS requires a higher activation energy than protonation following the reduction step. Consequently, the protonation reaction becomes much more favorable after the reduction step. Moreover, the activation energy for this initial protonation closely matches the dissociation energy (BDE = 9.6 kcal/mol) of the (2C-3e) bond within the B species. This indicates that the energy liberated upon bond cleavage contributes significantly to driving the reaction forward. To simplify our investigation, we divided the four steps of electron and proton transfer into two distinct stages: first, the transfer of one electron and one proton (e1,p1) in the first square in Fig. 8 and then the transfer of another electron and a second proton (e2,p2) in the second square in Fig. 8. Initially, the reaction proceeds via two parallel pathways, each comprising two sequential steps, labeled ep or pe (see Fig. 1). The overall rate is dictated by the faster pathway, whereas within each pathway, the rate is constrained by the slower step [40, 60].
Table 9 provides an overview of the calculated rate constants \({k}_{pe}\) and \({k}_{ep}\) corresponding to reactions A → D → E and A → B → E, respectively. The activation energy for the reverse process is calculated as the deviation between the energy barrier in the forward direction and the reaction energy:
The significant value of \({k}_{-1}\) in the A → D → E pathway confirms the reversibility of proton transfer. Indeed, deprotonation can occur before the electron is captured. Given the dominance of the \({k}_{ep}\) term in the overall rate constant k = \({k}_{ep}\) + \({k}_{pe}\) [40], we can deduce that in the reaction studied, the system first captures the electron and then attracts the proton. Applying the same approaches to the second electron and proton pair, we obtain \({k^{\prime}}_{pe}\) = 2.93 × 10–167 s-1 and \({k^{\prime}}_{ep}\) = 1.08 × 10–35 s-1 for the paths E → C → I and E → F → I, respectively. Clearly, the (p2,e2) path is very unfavorable because of the substantial energy barriers it involves. Our results suggest that the most favorable path from a thermodynamic and kinetic perspective is the electron–proton–electron–proton sequence (Fig. 9). Consequently, DMDS reduction is initiated by electron capture, which is a quasi-spontaneous reaction. Next, the anionic molecule attracts a proton to form a stable (E) intermediate. As the proton approaches (S….H \(\simeq\) 1.8 A°), the (2C-3e) bond is broken, providing the energy to form the new S–H bond. The third step (second reduction) is considered the critical kinetic step due to its relatively high activation energy, resulting in a slow reaction rate. It should be noted that the formation and cleavage of disulfide are known to be ultimately thermodynamically controlled, and the kinetics of this process are slow [61].
Energy decomposition analysis (EDA)
Each system in the chosen reaction pathway (electron–proton–electron–proton) is subdivided into two monomers, as illustrated in Fig. 10.
This approach aims to assess the binding energy between fragments, allowing detailed analysis of interactions within the reaction mechanism.
We observe a similarity between trends in the evolution of \({E}_{{\text{prep}}}\) and the activation energy (Table 10), where system F corresponds to both the highest activation energy (see Table 7) and preparation energy. Furthermore, the positive values of \({E}_{{\text{prep}}}\) in systems F and I imply that both sulfur monomers are more stable in their isolated geometries. Next, we focus on the interaction energy, for which we employ EDA analysis to decompose it into electrostatic, exchange, polarization, repulsion, and dispersion energy components (Fig. 11).
Similar results can be observed for the anionic intermediates (B and F) and in the neutral systems (E and I). This may be because protonation primarily reduces the orbital overlap between the two monomers, consequently affecting the exchange and repulsion energies. In the case of anionic systems, the exchange term mainly contributes to the stabilization energy. With the exception of neutral DMDS, the contribution of polarization energy in all species is less than that of electrostatic energy, indicating a more electrostatic than covalent nature of disulfide bonds. Notably, the minimum polarization energy of − 0.94 kcal/mol in the E system underlines the cleavage of the covalent disulfide bond. After the second electron is captured, the electrostatic energy increases considerably due to a higher dipole moment from E to F, with 3.03 Debye and 8.90 Debye, respectively. In addition, the disulfide distance is reduced by 8.73%, favoring greater orbital overlap and, consequently, an increase in exchange energy, although this is accompanied by a significant destabilization of the \({E}_{{\text{prep}}}\).
Natural bond orbitale (NBO) analysis
The NBO analysis reveals some interesting details, particularly regarding the nature of the 2c-3e bond (B), where one electron is shared between the two sulfur atoms (0.5e-: 0.5e-) and is located mainly in the bonding orbital (BD) (Fig. 12). Remarkably, the energy level of this bonding orbital is closely aligned with that of the half-filled nonbonding doublets (LP(3)) newly formed on the two sulfur atoms. Upon protonation (E system), a significant electronic rearrangement occurs to allow the formation of a new S–H bond (BD). Next, the unprotonated sulfur captures the second electron on the LP(3) orbital to form the F system. The LUMO orbital in the E system is more stable than that in the neutral system, which explains the trend observed in electron affinity (see Fig. 4). The major interaction within system F involves a charge transfer from the occupied LP3(S1) orbital to the vacant BD* (SH) orbital, which indicates the formation of a hydrogen bond. This interaction produces a stabilization energy of 19.44 kcal/mol. As a result, the length of the SS bond is shortened by 0.65 Å, and the overall stability of the system is enhanced. The sulfur acceptor is at a distance of 2.33 Å from the hydrogen, which agrees with the conventional threshold for identifying hydrogen bonds involving sulfur (2.8 Å) [62]. Notably, hydrogen bonds with thiolate as an acceptor form even shorter bonds (~ 2.34 Å) [63]. Furthermore, the length of the hydrogen bonds identified between cysteine (a donor) and half cysteine (derived from the removal of a hydrogen atom from the thiol group of cysteine) (an acceptor) is 2.29 Å [63].
Noncovalent interaction (NCI)
The noncovalent interaction (NCI) method characterizes noncovalent interactions in three-dimensional space by identifying peaks in the reduced density gradient at low electron densities. In general, weak van der Waals interactions appear at very low densities (ρ < 0.01 a.u.), while stronger hydrogen bonds appear at higher density values (0.01 < ρ < 0.06 a.u.). To distinguish between attractive and repulsive interactions, the NCI index uses the second eigenvalue of the electron density Hessian matrix (λ2). λ2 < 0 indicates bonding interactions, while λ2 > 0 indicates repulsive interactions. Consequently, plotting the reduced density gradient as a function of the sign (λ2)ρ effectively distinguishes between attractive and repulsive noncovalent interactions (Fig. 13).
Figure 13 shows isosurfaces generated by Multiwfn 3.8 with an isovalue of s(r) = 0.05 a.u., visualized using VMD 1.9.4. In system (B), the blue isosurface indicates a strong interaction between sulfur atoms with some covalent character. Conversely, systems (E) and (I) show only green surfaces, indicating weak interactions. The presence of weak hydrogen bonding in system F is clearly evident in the NCI analysis, where a light blue surface appears between the hydrogen and acceptor sulfur, accompanied by a peak in the reduced density gradient at < 0.02 a.u.
Electrostatic potential iso-surfaces provide insight into changes in charge distribution during dimethyl disulfide reduction (Fig. 14). Notably, upon the introduction of the first unpaired electron, a large negative region is generated around the sulfur atoms. Then, when a proton is introduced, this negative charge is neutralized and localized around one of the sulfur atoms. Next, the second electron is added to the unprotonated sulfur, to create a highly polarized system, as shown by NBO analysis. Finally, the resulting thiols (system I) show nonequivalent charges. This discrepancy suggests that the thiol with the most positively charged hydrogen is more likely to undergo initial deprotonation. This process leads to the formation of reactive thiolate ion, which is crucial for nucleophilic attack in thiol–disulfide exchange reactions [61].
Redox potential calculation
In our scientific approach, our primary objective was to determine the reduction potential of dimethyl disulfide (DMDS) by 2 electrons and 2 protons in aqueous media. However, our efforts were limited by the lack of experimental data in the literature, and the only value available corresponds to the first reduction in dimethylformamide (DMF) solvent [64]. To ensure consistency with this reference, we first calculated the reduction potential corresponding to the first step in a DMF solvent.
For now, in our simulation, the solvent has been treated with the IEFPCM model which provides a continuous approach to solvation, considering the solvent as a homogeneous medium surrounding the solute molecule. However, as the continuum approach does not include actual solvent molecules in the calculation, it neglects specific short-range solute–solvent interactions such as hydrogen bonding and van der Waals effets or the specific structure of the solvent around the solute [65]. Hydrogen bonding, in particular, has proved important for the accurate calculation of redox potentials [66]. Indeed, Kılıç and Ensing predict a direct correlation between the number of hydrogen bonds accepting or donating substituents of a compound and its redox potential [67].
In this respect, we use the molecular dynamics-perturbed matrix method (MD-PMM) [44] approach for a quantum mechanical/molecular mechanical (QM/MM) calculation (see section on calculation methods).
Dimethyl disulfide is taken as the quantum center (QC) and analyzed using two different quantum methods for comparative purposes. Conversely, the solvent is considered to be the perturbing environment and is subjected to molecular dynamics simulations (further details are provided in the methodology section). The redox potential was calculated using the following equations:
Here, \(\Delta A\) is the variation in the Helmholtz free energy calculated via the PyMM program [68, 72] considering both oxidized and reduced ensembles. \({E}_{j}\) represents the liquid junction potential, \({E^\circ }_{SHE}=4.28V\) represents the standard potential of the hydrogen electrode, and \(E^\circ \left(\frac{SCE}{SHE}\right)=0.24V\) is the potential of the saturated calomel electrode (SCE) relative to the standard hydrogen electrode (SHE) [49].
The combination of CCSD(T) (for QC) and MD (for solvent) gave the redox potential in the best agreement with the experimental value (Table 11). Therefore, this model is suitable for calculating reduction potentials and acidity constants in aqueous environments. E° is calculated using Eq. (15) (without the Ej correction). The pKa values were estimated according to the methodology described by Zanetti–Polzi et al. [48]. Finally, Eq. (17) is used to determine the reduction potential for the overall reaction [69]:
As expected, the reduction potential of DMDS in water (Table 12) is less negative than that of the nonaqueous DMF solvent due to the higher dielectric constant of water (80.2) compared to that of DMF (36.7) [70].
With regard to electron uptake reactions [71], the \(C{H}_{3}SHSC{H}_{3}\) system, with a less negative reduction potential for the couple, is a more powerful oxidant than neutral DMDS in the \(C{H}_{3}SSC{H}_{3}/\text{[}C{H}_{3}SSC{H}_{3}{]}^{-}\) redox couple \(\text{[}C{H}_{3}SHSC{H}_{3}{]}^{0}\) is easier to reduce than \(C{H}_{3}SSC{H}_{3}\). It is, however, accompanied by a higher energy barrier, as seen previously (Table 7). The same pattern is observed in protonation reactions, where the \({pK}_{a}^{2}\) value is lower than \({pK}_{a}^{1}\), indicating that the second protonation is expected to occur more easily; however, this is associated with a higher activation energy (Table 8). This phenomenon is a hallmark of the inverted Marcus region, where most exoergic reactions systematically proceed more slowly [72].
Conclusion
As disulfide bond reduction is a crucial reaction in many biochemical processes, it is important to understand its detailed mechanism and to identify the precise stage at which the disulfide bridge is broken.
By studying the six possible \({2e}^{-}\)/\({2H}^{+}\) reduction pathways from thermodynamic and kinetic perspectives, this work identified the most favorable reaction route. This pathway begins with the uptake of a first electron to form a metastable anionic radical with a (2S-3e) bond, followed by a protonation reaction that causes the (S–S)bond to break definitively. Next, a second electron is captured by the unprotonated sulfur, creating an anionic intermediate with a relatively strong hydrogen bond. This is the kinetically determinant step. Last, a second protonation is required to form the two distinct thiols. The rate constants show an inverted dependence on the driving forces (\(\Delta {G}^{0}\)). This observation of an inverted region may enable new strategies to understand and control the reaction.
The MD‒PMM method is used to calculate the redox potential of the first DMDS reduction, giving a value of − 1.98 V. This theoretical result agrees well with the experimental value reported in the literature (− 1.88 V). In addition, this approach enables us to determine the redox potentials (E°) and acidity constants (pKa) of the elementary reactions in aqueous solution. The redox potential for the overall reduction of DMDS was calculated to be 0.32 V.
In summary, this study represents the first attempt to predict disulfide reduction potentials through the involvement of two electrons and two protons. This highlights the importance of computational methodologies in exploring and elucidating phenomena that pose challenges to experimental investigation. In addition, this study provides new perspectives by providing essential details on the mechanism underlying disulfide bridge reduction, thus offering valuable guidance for experimental investigation and discovery.
Data availability
The data that support the findings of this study are available from the corresponding author upon reasonable request.
Change history
31 May 2024
A Correction to this paper has been published: https://doi.org/10.1007/s00894-024-05998-x
References
Arumugam K, Becker U (2014) Computational redox potential predictions: Applications to inorganic and organic aqueous complexes, and complexes adsorbed to mineral surfaces. Minerals 4:345–387
Xiao Z, La Fontaine S, Bush AI et al (2019) Molecular mechanisms of glutaredoxin enzymes: Versatile hubs for thiol–disulfide exchange between protein thiols and glutathione. J Mol Biol 431:158–177
Hogg PJ (2003) Disulfide bonds as switches for protein function. Trends Biochem Sci 28:210–214
Matsui T, Kitagawa Y, Okumura M et al (2015) Accurate standard hydrogen electrode potential and applications to the redox potentials of vitamin C and NAD/NADH. J Phys Chem A 119:369–376
Chen CG, Nardi AN, Amadei A et al (2022) Theoretical modeling of redox potentials of biomolecules. Molecules 27:1077
Stenlid JH, Dos Santos EC, Bagger A et al (2020) Electrochemical interface during corrosion of copper in anoxic sulfide-containing groundwater—A computational study. J Phys Chem C 124:469–481
Zuo K, Garcia-Segura S, Cerrón-Calle GA et al (2023) Electrified water treatment: Fundamentals and roles of electrode materials. Nat Rev Mater 8:472–490
Tentscher PR, Lee M, Von Gunten U (2019) Micropollutant oxidation studied by quantum chemical computations: Methodology and applications to thermodynamics, kinetics, and reaction mechanisms. Acc Chem Res 52:605–614
Lin L, Zou H, Li W, Xu LY, Li EM, Dong G (2021) Redox Potentials of Disulfide Bonds in LOXL2 Studied by Nonequilibrium Alchemical Simulation. Front Chem 9:797036
Olsson MHM, Hong G, Warshel A (2003) Frozen density functional free energy simulations of redox proteins: Computational studies of the reduction potential of plastocyanin and rusticyanin. J Am Chem Soc 125:5025–5039
Tron AE, Bertoncini CW, Chan RL et al (2002) Redox regulation of plant homeodomain transcription factors. J Biol Chem 277:34800–34807
Trotter EW, Grant CM (2003) Non-reciprocal regulation of the redox state of the glutathione–glutaredoxin and thioredoxin systems. EMBO Rep 4:184–188
Fernandes PA, Ramos MJ (2004) Theoretical insights into the mechanism for thiol/disulfide exchange. Chem Eur J 10:257–266
Arnér ESJ, Holmgren A (2000) Physiological functions of thioredoxin and thioredoxin reductase. Eur J Biochem 267:6102–6109
Zhang J, Duan D, Osama A, Fang J (2021) Natural molecules targeting thioredoxin system and their therapeutic potential. Antioxid Redox Signal 34(14):1083–1107
Arnér ESJ, Holmgren A (2006) The thioredoxin system in cancer. Semin Cancer Biol 16:420–426
Kylarova S, Kosek D, Petrvalska O et al (2016) Cysteine residues mediate high-affinity binding of thioredoxin to ASK1. FEBS J 283:3821–3838
Xu J, Fang J (2021) How can we improve the design of small molecules to target thioredoxin reductase for treating cancer? Expert Opin Drug Discov 16:331–333
Le Moan N (2007) Approches globales de l'état redox du résidu cystéine (Doctoral dissertation, Université Paris Sud-Paris XI)
Baldus IB, Gräter F (2012) Mechanical force can fine-tune redox potentials of disulfide bonds. Biophys J 102:622–629
Bustanji Y, Samorì B (2002) The mechanical properties of human angiostatin can be modulated by means of its disulfide bonds: A single-molecule force-spectroscopy study. Angew Chem 114:1616–1618
Carl P, Kwok CH, Manderson G et al (2001) Forced unfolding modulated by disulfide bonds in the Ig domains of a cell adhesion molecule. Proc Natl Acad Sci 98:1565–1570
Bhasin N, Carl P, Harper S et al (2004) Chemistry on a single protein, vascular cell adhesion molecule-1, during forced unfolding *. J Biol Chem 279:45865–45874
Li H, Fernandez JM (2003) Mechanical design of the first proximal Ig domain of human cardiac titin revealed by single molecule force spectroscopy. J Mol Biol 334:75–86
Mthembu SN, Sharma A, Albericio F et al (2020) Breaking a couple: disulfide reducing agents. ChemBioChem 21:1947–1954
Johnson DL, Polyak SW, Wallace JC et al (2003) Probing the stability of the disulfide radical intermediate of thioredoxin using direct electrochemistry. Int J Pept Res Ther 10:495–500. https://doi.org/10.1007/s10989-004-2410-y
Ganisl B, Breuker K (2012) Does electron capture dissociation cleave protein disulfide bonds? ChemistryOpen 1:260–268
Houée-Levin C (2002) Determination of redox properties of protein disulfide bonds by radiolytic methods. Methods Enzymol 353:35–44
Dumont É, Loos P-F, Assfeld X (2008) Effect of ring strain on disulfide electron attachment. Chem Phys Lett 458:276–280
Dumont É, Laurent AD, Assfeld X et al (2011) Performances of recently-proposed functionals for describing disulfide radical anions and similar systems. Chem Phys Lett 501:245–251
Dumont É, Loos P-F, Assfeld X (2008) Factors governing electron capture by small disulfide loops in two-cysteine peptides. J Phys Chem B 112:13661–13669
Mayer JM, Rhile IJ (2004) Thermodynamics and kinetics of proton-coupled electron transfer: stepwise vs. concerted pathways. Biochimica et Biophysica Acta (BBA)-Bioenergetics 1655:51–58
Marcus RA (1968) Theoretical relations among rate constants, barriers, and Broensted slopes of chemical reactions. J Phys Chem 72:891–899
Silverman DN (2000) Marcus rate theory applied to enzymatic proton transfer. Biochimica et Biophysica Acta (BBA)-Bioenergetics 1458(1):88–103
Walton JC (2017) Radical-enhanced acidity: Why bicarbonate, carboxyl, hydroperoxyl, and related radicals are so acidic. J Phys Chem A 121:7761–7767
Ouellette RJ, Rawn JD (2015) 3-Introduction to organic reaction mechanisms. In: Ouellette RJ, Rawn JD (eds) Organic Chemistry Study Guide. Elsevier, Boston, pp 31–46
Hajj, V.: Transfert couplé électron/proton et coupure de liaison dans des systèmes bio-inspirés.
Buda M (2013) On calculating reorganization energies for electrochemical reactions using density functional theory and continuum solvation models. Electrochim Acta 113:536–549
Martínez AG, Gómez PC, de la Moya S et al (2021) Revealing the mechanism of the water autoprotolysis on the basis of Marcus theory and TD-DFT methodology. J Mol Liq 324:115092
Hammes-Schiffer S, Stuchebrukhov AA (2010) Theory of coupled electron and proton transfer reactions. Chem Rev 110:6939–6960
Gimferrer M, Danés S, Andrada DM et al (2023) Merging the energy decomposition analysis with the interacting quantum atoms approach. J Chem Theory Comput 19:3469–3485
Bickelhaupt FM, Baerends EJ (2000) Kohn‐Sham density functional theory: predicting and understanding chemistry. Rev Comput Chem 1–86. https://doi.org/10.1002/9780470125922.ch1
Su P, Li H (2009) Energy decomposition analysis of covalent bonds and intermolecular interactions. J Chem Phys 131:014102
Zanetti-Polzi L, Galdo SD, Daidone I et al (2018) Extending the perturbed matrix method beyond the dipolar approximation: Comparison of different levels of theory. Phys Chem Chem Phys 20:24369–24378
De Sciscio ML, D’Annibale V, D’Abramo M (2022) Theoretical evaluation of sulfur-based reactions as a model for biological antioxidant defense. Int J Mol Sci 23:14515
Sousa da Silva AW, Vranken WF (2012) ACPYPE-Antechamber python parser interface. BMC Res Notes 5:367. https://doi.org/10.1186/1756-0500-5-367
Del Galdo S, Marracino P, D’Abramo M, Amadei A (2015) In silico characterization of protein partial molecular volumes and hydration shells. Phys Chem Chem Phys 17(46):31270–31277
Zanetti-Polzi L, Daidone I, Amadei A (2020) Fully atomistic multiscale approach for pKa prediction. J Phys Chem B 124:4712–4722
Ho J, Coote ML, Cramer CJ, Truhlar DG (2015) Theoretical calculation of reduction potentials. Organic electrochemistry 5:229–259
Laurent A (2010) Etude de phénomènes électroniques de macromolécules à l'aide de méthodes hybrides QM-MM (Doctoral dissertation, Université Henri Poincaré-Nancy 1)
Lemmon EW, McLinden MO, Friend DG, Linstrom P, Mallard W (2011) NIST chemistry webbook. NIST stand ref database (69):20899
Moser A, Range K, York DM (2010) Accurate proton affinity and gas-phase basicity values for molecules important in biocatalysis. J Phys Chem B 114:13911
Maksić ZB, Kovačević B, Vianello R (2012) Advances in determining the absolute proton affinities of neutral organic molecules in the gas phase and their interpretation: A theoretical account. Chem Rev 112:5240–5270
Hunter EPL, Lias SG (1998) Evaluated gas phase basicities and proton affinities of molecules: An update. J Phys Chem Ref Data 27:413–656
Galano, A. (2015). Free radicals induced oxidative stress at a molecular level: The current status, challenges and perspectives of computational chemistry based protocols. J Mex Chem Soc 59(4):231–262
Narayan GM, Valles A, Venegas F, Yi J, Narayan M (2020) Learnings from the relation between the number of forward and reverse reactions (transfer cycles) required to converge to equilibrium and the ratio of the forward to the reverse rate constants in simple chemical reactions. ACS Omega 6(1):38–45
Parada GA, Goldsmith ZK, Kolmar S et al (2019) Concerted proton-electron transfer reactions in the Marcus inverted region. Science
Rosspeintner A, Lang B, Vauthey E (2013) Ultrafast photochemistry in liquids. Annu Rev Phys Chem 64:247–271
Stuchebrukhov AA (2003) Electron transfer reactions coupled to proton translocation: Cytochrome oxidase, proton pumps, and biological energy transduction. J Theor Comput Chem 02:91–118
Maksić ZB, Vianello R (2007) Physical origin of chemical phenomena: Interpretation of acidity, basicity, and hydride affinity by trichotomy paradigm. Pure Appl Chem 79:1003–1021
Yi MC, Khosla C (2016) Thiol–disulfide exchange reactions in the mammalian extracellular environment. Annu Rev Chem Biomol Eng 7:197–222
van Bergen LAH, Alonso M, Palló A et al (2016) Revisiting sulfur H-bonds in proteins: The example of peroxiredoxin AhpE. Sci Rep 6:30369
Zhou P, Tian F, Lv F et al (2009) Geometric characteristics of hydrogen bonds involving sulfur atoms in proteins. Proteins 76:151–163
Antonello S, Benassi R, Gavioli G et al (2002) Theoretical and electrochemical analysis of dissociative electron transfers proceeding through formation of loose radical anion species: Reduction of symmetrical and unsymmetrical disulfides. J Am Chem Soc 124:7529–7538
Sterling CM, Bjornsson R (2018) Multistep explicit solvation protocol for calculation of redox potentials. J Chem Theory Comput 15(1):52–67
Wang LP, Van Voorhis T (2012) A polarizable QM/MM explicit solvent model for computational electrochemistry in water. J Chem Theory Comput 8(2):610–617
Kılıç M, Ensing B (2019) Microscopic picture of the solvent reorganization during electron transfer to flavin in water. J Phys Chem B 123(46):9751–9761
Chen CG, Nardi AN, Amadei A et al (2023) PyMM: An open-source python program for QM/MM simulations based on the perturbed matrix method. J Chem Theory Comput 19:33–41
Quinone. 1 e– and 2 e–/2 H+ reduction potentials: Identification and analysis of deviations from systematic scaling relationships | Journal of the American Chemical Society Available from: https://doi.org/10.1021/jacs.6b05797
Coordination sphere versus protein environment as determinants of electronic and functional properties of iron-sulfur proteins | SpringerLink Available from: https://doi.org/10.1007/3-540-62888-6_5
Wardman P (1989) Reduction potentials of one-electron couples involving free radicals in aqueous solution. J Phys Chem Ref Data 18:1637–1755
Andrieux CP, Gamby J, Hapiot P et al (2003) Evidence for inverted region behavior in proton transfer to carbanions. J Am Chem Soc 125:10119–10124
Acknowledgements
The authors would like to thank the laboratory of Theory-Modeling-simulation, LPCTUMR CNRS UL 7565 Université de Lorraine 1. All calculations have been done using the local computing resources of the LPCT laboratory-UMR CNRS Computer Center. The authors are most appreciative of the assistance of Pr. Marco D’Abramo (Dept. of Chemistry, Sapienza University of Rome) who initiated Lina Ould Mohamed for QM/MM and Molecular Dynamics (MD) methods.
Author information
Authors and Affiliations
Contributions
O. , A., As. wrote the main manuscript text and O. and G. prepared tables and figures . All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Ethical approval
Not applicable.
Competing interests
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
The original online version of this article was revised due to a small detail in the first affiliation that was overlooked.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Ould Mohamed, L., Abtouche, S., Ghoualem, Z. et al. Unraveling redox pathways of the disulfide bond in dimethyl disulfide: Ab initio modeling. J Mol Model 30, 180 (2024). https://doi.org/10.1007/s00894-024-05963-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00894-024-05963-8