Abstract
Context
The specialised family of triggering receptors expressed on myeloid cells (TREMs) plays a pivotal role in causing neurodegenerative disorders and activating microglial anti-inflammatory responses. Nasu-Hakola disease (NHD), a rare autosomal recessive disorder, has been associated with mutations in TREM2, which is also responsible for raising the risk of Alzheimer’s disease (AD). Herein, we have made an endeavour to differentiate the confirmed pathogenic variants in TREM2 extra-cellular domain (ECD) linked with NHD and AD using mutation-induced fold stability change (∆∆G), with the computation of 12distinct structure-based methods through saturation mutagenesis. Correlation analysis between relative solvent accessibility (RSA) and ∆∆G expresses the discrete distributive behaviour of mutants associated with TREM2 in AD (R2 = 0.061) and NHD (R2 = 0.601). Our findings put an emphasis on W50 and V126 as major players in maintaining V-like domain in TREM2. Interestingly, we discern that both of them interact with a common residue Y108, which is dissolved upon mutation. This Y108 could have structural or functional role for TREM2 which can be an ideal candidate for further study. Furthermore, the residual interaction network highlights the importance of R47 and R62 in maintaining the CDR loops that are crucial for ligand binding. Future studies using biophysical characterisation of ligand interactions in TREM2-ECD would be helpful for the development of novel therapeutics for AD and NHD.
Methods
ConSurf algorithm and ENDscript were used to determine the position and conservation of each residue in the wild-type ECD of TREM2. The mutation-induced fold stability change (∆∆G) of confirmed pathogenic mutants associated with NHD and AD was estimated using 12 state-of-the-art structure-based protein stability tools. Furthermore, we also computed the effect of random mutation on these sites using computational saturation mutagenesis. Linear regression analysis was performed using mutants ∆∆G and RSA through GraphPad software. In addition, a comprehensive non-bonded residual interaction network (RIN) of wild type and its mutants of TREM2-ECD was enumerated using RING3.0.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The innate immune receptor TREM2 (triggering receptor expressed on myeloid cells) is a transmembrane glycoprotein having V-immunoglobulin extra-cellular domains (ECDs) and cytoplasmic tails, located on human chromosome 6p21.1, expressed in myeloid cells and tissue-specific macrophages [1,2,3,4]. Expression of TREM2 is augmented by anti-inflammatory molecules in vitro; however, in vivo expression is upregulated under pathological conditions [5, 6]. A variety range of microglial immune functions, comprising of chemotaxis, phagocytosis, autophagy, survival and proliferation, proinflammatory cytokine production, and lipid metabolism, are controlled by TREM2 [1, 7,8,9,10,11,12]. Overall TREM2 is comprised of an intracellular domain (ICD), transmembrane domain (TMD) segment, and an ECD which binds to the lipopolysaccharide (LPS), lipoteichoic acid (LTAs) [13], and high- and low-density lipoproteins and apolipoproteins [14]. Through its ectodomain, the TM protein binds ligands for the activation of intercellular signalling pathways controlling the innate immune responses [15]. Signalling of TREM2 involves binding to the DAP12 (DNAX-activating protein of 12 kDa), further activating SYK (Spleen Tyrosine Kinase) tyrosine kinase [16]. Stimulation of DAP12 pathway encourages hydrolysis of PIP2 into IP3 at the cell membrane, making it possible for AD-associated variants to impair the activation of DAP12, and inhibition of PLCℽ2 activities will enhance the PIP2 level [17]. As a result, TREM2’s ability to guard against AD will be diminished [18]. Cleaved ECD gives rise to the soluble form of TREM2 (sTREM2), which supervises the interaction between neurons and the surrounding microenvironment [17]. The sTREM2 level in cerebral spinal fluid (CSF) is said to be a well-founded analyst for Alzheimer’s disease (AD) [19]. This CSF sTREM2 has also been studied as a biomarker of microglia activation in the setting of neuroinflammatory and neurodegenerative disorders. Since CSF sTREM2 provides information on the timing of microglia activation in relation to Aβ deposition, tau aggregation, neurodegeneration, and the onset of clinical symptoms, it has been the subject of extensive research in AD [20, 21], whereas plasma sTREM2 has not been found to be a suitable biomarker for the determination of status of TREM2 atypical variants or anticipated AD [22]. Genome wide association studies (GWAS) have shown TREM2 loses function as a consequence of appearance of homozygous and heterozygous variants linked to neurodegenerative disorders, e.g. AD, frontotemporal dementia (FTD), Nasu-Hakola disease (NHD), and Parkinson’s disease (PD) [5, 23,24,25,26]. The most common variant of AD in TREM2 is located near the putative ligand-interacting region (PLIS), whereas the variants associated with NHD are grossly damaging by folding, truncating, or unfolding [23, 24].
Construction of neurofibrillary tangles and neuroinflammation with built-up amyloid-beta (aβ) plaque and gradual memory loss categorise the neurodegenerative disorder AD [27, 28]. AD, characterised by a slow progressive loss of cognitive function, is the most common form of dementia among the aged population [29]. Progression of AD in CNS encompasses aβ plaque and neurofibrillary tangles of hyperphosphorylated tau protein concerned to synapse loss, neuronal death eventually leading to cognitive decline [27, 30]. TREM2 acts as a neuroprotector by suppressing the amyloid beta diffusion with the regulation of microglial activation around amyloid plaques during the early and mid-term AD [8, 31, 32]. The risk factor of AD is enhanced by several folds with the reduction of the microglial phagocytosis of amyloid plaque [31, 33]. Mutation of TREM2 associated with AD takes place on its ectodomain encoded by exon2 which is said to be linked with production and function of sTREM2 which further instigate AD development [24, 34,35,36]. R47H (Arg47His) variant is the most common form of AD reported to triple the risk with 23% of faster dementia rate, by disrupting the ligand-receptor interactions [35, 37, 38]. Another mutation associated with late-onset AD is R62H (Arg62His) that inhibits immune responses and reduces microglial activation [39, 40]. Other than the above two variants, there are other TREM2 mutations associated with AD which can be listed as D87N and H157Y [23, 41]. Pathogenic variant of NHD Q33X is said to be a risk modifier for AD [36].
Polycystic lipomenbranous ostedysplasia with sclerosing leukoembrancephalopathy (PLOSL), popularly known as NHD, is an early onset dementia and continual bone fracture due to polycystic osseous lesions, a rare autosomal disorder with the occurrence of frontal lobe dysfunction eventually leading to death before the fifth decade of life [42,43,44]. Neuropathological discoveries also indicate corporation of myelin loss and loss of neurons, astrocyte proliferation, and activation of microglia in cerebral white matter and basal ganglia [45]. Biallelic mutations in two genes TREM2 or TYRO protein tyrosine kinase binding protein (TYROBP) bring about the heterogeneous syndrome NHD [43]. NHD is divided into 4 phases, namely latent, osseous, early neurological, and late neurological [14, 46]. TREM2 is a crucial player for NHD pathogenesis involved with the differentiation of myeloid progenitors concerning dendritic cells, osteoclasts, and microglia having deleterious effects on its own function due to mutation [23, 42]. Mutation W50C associated with NHD was first seen in a patient of Greek exhibiting personality change at the age of 30 [44], where tryptophan-cysteine substitution is said to be harmful [44]. Q33X variant linked to NHD, in a homozygous state, leads a premature stop codon in place of glutamine. W44X and W78X were first derived from a Bolivian and Swedish families respectively in a homozygous state [43]. The variant V126G [45] hints protein misfolding [47]. AlthoughV126G exhibits poor cell-surface expression and is incapable of N-linked glycosylation in the Golgi, it may be subject to a different O-linked glycosylation pathway [48]. Variants D134G [43] and E14X [49] are also said to be associated with NHD (Data retrieved from http://www.alzoforum.org/mutations). TREM2 variant associated with NHD in heterozygous carriers may confer increased AD risk [23, 39, 50].
Over the years, neurodegenerative diseases have been a concern to the scientific community. There are unanswered questions how mutation of a single protein can cause two separate diseases related to dementia. To infer how the pathogenic mutation associated with AD and NHD affects the structure and function of ECD of TREM2 using 12 structure-based state-of-the-art computational protein folding stability (∆∆G) tools. Present study is focused on confirmed pathogenic variants such as W50C, W44X, W78X, and V126G are associated with NHD while R47H, R62H, and D87N are increasing the risk of AD onset. Q33X is found to be associated both in AD and NHD. All these mutants are accumulated in the ECD region of TREM2 which is important for the ligand interaction and signalling. Protein stability assays also decipher the changes in RSA and residual depth in WT and mutant variants. Further changes in the residual depth and RSA correlation were determined and the results were corroborated with thermodynamic stability (∆∆G) of the above mutant variants. Mutants associated with AD are usually present on the surface of the TREM2 while the residues associated with NHD development are located in buried region of the same protein. Moreover, previous mutational analysis reported that NHD mutations are disrupting the structure than variants AD. We performed a correlation analysis between solvent accessibility and thermodynamic stability of the protein in both the disease conditions which revealed that NHD has a strong correlation while AD does not show any significance which could be the one of the differential etiology of these two diseases. Missense mutations at conserved or important sites frequently impair the interaction in the native condition and result in reduction of protein stability, solubility, and signalling etc. [51]. In most of the confirmed pathogenic mutants, we observed a loss of contacts (non-bonded contacts) which disrupt the local fold of protein and lead to affect the functionality of TREM2. Further in vitro studies and experimental structural data of mutants can establish the effects of mutants in two different disease conditions.
Materials and methods
Collection of TREM2 variants
Pathogenic mutants of TREM2 linked to AD and NHD were assembled from database of Alzoforum (http://www.alzoforum.org/mutations). Present study has considered ECD of TREM2 protein with its 5 mutant variants with confirmed pathogenicity for NHD and 4 variants for AD as listed in Table S1. To carry our research forward, we retrieved a crystal structure of wild type TREM2 (5UD7) with resolution of 2.20 Å from RCSB-PDB [52].
Conservation analysis TREM2
To determine the position and their conservation of critical residues in TREM2 protein, we employed ConSurf algorithm [53]. It is a web-based tool that helps to find out the evolutionary information of each amino acid residues and also highlights structurally and functionally important regions in the protein. Crystal structure of wild-type TREM2 (5UD7-A chain) was taken as input for ConSurf. Then the outcome was obtained with the function, position, and evolutionary conservation with color-coded values. Furthermore, we proceeded for ENDscript [54] for generating a detailed interactive representation of wild-type structure of TRME2 ECD (5UD7) to identify conservation zones. Resulted outcome was studied to see the conserved and variable residues highlighted in different colour shades. The results also showed the accessible and buried regions as well as the hydrophobic and hydrophilic sites.
Electrostatic surface potential mapping
The electrostatic potential surfaces of TREM2 protein (5UD7) were computed using the APBS-Electrostatics plug-in tool of PyMOL2.5 [55]. The linearised Poisson-Boltzmann (PB) equation with a bulk solvent radius of 1.4 Å and a dielectric constant of 78 was used to compute the solvent-accessible surface area of the same protein. The electrostatic positive and negative surfaces TREM2 were shown by using a contour (kT/e) value of 1.
Protein folding stability prediction
Protein stability is one of the long-term goals of protein structure analysis [56] which is essential for improvement of the effectiveness of rational protein design. To achieve this end, it is favoured to study how mutations effect protein stability further leading to the development of several methods for prediction of stability change produced by substitution of residues [57]. A total twelve freely available structure-based protein stability prediction web servers were chosen to compute protein stability (∆∆G) upon single point mutation. Selected methods utilise a range of algorithms for estimating the change in protein stability. PoPMuSiC [58] uses combined statistical potential to for ∆∆G prediction whereas CUPSAT [59] applies atomic potential from chemical characteristics and torsion potentials derived empirically. Methods like mCSM [60] and I-Mutant3.0 [61] use machine learning methods. Web-based server FireProt [62] implements the same thresholds used by FoldX [63] and Rosetta [62]. Table 1 provides a detailed insight into the tools used in the study. For I-Mutant3.0, NeEMO [64] user can define the parameters, where 25 °C temperature was considered and pH was 7. But most of the tools run on default frame works. SDM was considered to calculate residual depth and RSA values for wild type and mutants as well. In addition, to know the effect of random mutation on protein stability, we employed computational saturation mutagenesis of the selected pathogenic mutants.
Association of RSA with ∆∆G
To see whether RSA affects the stability and function of protein, we executed the correlation analysis on computationally calculated ∆∆G with RSA. All the tools in the study provided with similar RSA values to WT and the variants. Therefore, we opted for SDM tool for the RSA calculations in all variants [66]. Correlation analysis between RSA and ∆∆G was performed by using GraphPad 6.0 software by taking ∆∆G as the independent variable, while RSA was dependable variable.
Residual interaction networks
Each mutant was developed by using the mutagenesis wizard of PyMOL2.5. For a better knowledge about the interaction, we generated the residual interaction network (RIN) for the wild type protein and then further for each generated mutant using RING3.0 [71], a web-based platform which provides a detailed non-covalent molecular interaction based upon geometrical parameters. The determined results were then analysed to decipher the changes occurred between the residual interactions of mutants as compared to wild type.
Results and discussion
Transmembrane receptor TREM2 is found on microglia and plays a significant role in neuroinflammation. Variants of the TREM2 gene were discovered by GWAS, which is said to be increasing the chance of developing AD by 2 to 4 times. Several research using mouse models have shown a connection between TREM2 gene alterations and their impact on tau pathology and amyloid load in AD-affected brain tissue [72]. Similarly, NHD is a progressive and fatal neurodegenerative disease. Patients with NHD typically develop symptoms such as cognitive decline, behavioural changes, movement disorders, and bone fractures at early life around 30 s–40 s. Few mutations in TREM2 are also reported to increase the chances of disease development. Molecular basis of pathogenicity due to missense mutations are under active area of research. Previous studies emphasise on mutant variants for the development of two distinct diseases but how they are differentiating in the etiologies are yet to be revealed. Here, the mutants of TREM2 with confirmed pathogenicity were taken for study to find out their role in protein stability, RSA, and in functionality. Using experimental structural information, computational exploration of protein folding stability offers a clear knowledge of protein function and related mutation in these two pathological conditions, which may aid in development of novel therapeutics.
Distribution of confirmed pathogenic variants within ECD of TREM2
Variants such as W50C, W44X, W78X, and V126G are associated with NHD while R47H, R62H, and D87N are increasing the risk of AD onset. Q33X is found to be associated both in AD and NHD. Furthermore, R47H and R62H mutation are well-studied mutants that increase the risk for the AD development. R47 has an important role in maintaining structural integrity of complementarity-determining region 2 (CDR2) and putative positive ligand-interacting surface. The R47H mutation of TREM2 disrupt the above interacting surface which significantly affect the functionality by altering ligand binding [73]. Similarly, other variants are also found to pose greater risk for the onset of above diseases but their molecular basis of disease development has not been explored. Experimental studies on above variants would be longer and tedious procedures. Hence, computational analysis would add insights for the molecular basis of disease in mutant variants.
Structural and functional conservation of important residues in TREM2
TREM2 is 230 amino acid long protein with three different regions: an N-terminal matured ectodomain (ECD, residues 19–174), a membrane-spanning region (residues 175–195), and a C-terminal cytosolic tail (residues 196–230). The residue 1–18 amino acids act as signalling peptide for the above protein. The structure of ECD domain, spanning from 19 to 134 is taken for the above study (PDB: 5UD7). It is comprised of eight Beta strands (A–F) and three important complementarity-determining regions (CDR loops), namely CDR1 (P37–R47), CDR2 (T66–R76), and CDR3 (H144–E117). These regulator loops are associated with ligand interaction. Recent investigations emphasised the deleterious effect of mutations on these regions [74]. To identify the functionally relevant regions in the TREM2 ectodomain as well as to analyse the evolutionary conservation at each amino acid site, this study employed a web-based tool ConSurf [53]. ConSurf analysis (as shown in Fig. 1A and Table S2) depicts that the critical residues are mostly falling in the conserved regions and few important residues are sited at variable regions in the ECD. The variants those are highlighted in Fig. 1C are considered in the study. Residues such as Q33, W44, R47, and D87 are predicted to be at conserved region in the protein whileW50 and V126 are highly conserved in TREM2 protein family (Fig. 1A). Residues such as W78 and R62 are surface exposed and present at variable regions of C and C″ beta strands but they are well associated with risk of developing NHD and AD respectively. Furthermore, Q33 is sited on the surface of the second Beta strand at N-terminal regions (Fig. 1B); mutation at this position causes loss of TREM2 expression [75]. Similarly, other surface exposed residues such as W44 and R47 are present in the CDR1 loop region, responsible for ligand interaction. Another important residue D87 is also present on the surface of D beta strand (Fig. 1B), while W78 is buried in the Ig domain C″ strand. Substitutions at these residues result into lack of lipid binding to TREM2 and expression of truncated protein respectively [43]. Likewise, W50 residue is in the core of the protein in C′ strand and actively participating in the maintenance of structural integrity of TREM2 protein. Upon mutation, V126 is buried and presents at the V-like Ig domain on G strand, predicted to have a function in the maintenance of hydrophobic region in the protein [47]. All the residues are present in the positively charged hydrophilic region except V126, which is in hydrophobic region illustrated in the electrostatic potential surface analysis (Fig. 1D).
Molecular architecture, position, and conservation of each amino acid at ectodomain (19–129) in TREM2. (A) ConSurf analysis depicts evolutionary conserved amino acids and they are colour coded. Residues those are membrane exposed or inside the core of the protein are separately demarcated as e and b respectively. Conserved structural and functional residues are also marked as s and f respectively. (B) Structure of TREM 2 ectodomain comprising of 9 beta strands and 3 CDR regions. (C) Mutants those included in the present study are shown in stick representation. (D) Electrostatic potential surface map of TREM2 ECD computed using APBS-plugin of PyMOL2.5 [55]
To confirm our ConSurf analysis, we further proceeded for ENDscript [54], a programme that produces images of aligned sequences with secondary structure data. Result displayed Q33, W44, R47, and R62 to be present at the variable regions, where Q33, W44, and R62 are highly accessible and R47 is intermediate (Fig. 2). Variant W50 is structurally important, highly conserved and buried residue likewise V126 lies in the buried and conserved region of the protein. W78 and D87 are present at the conserved regions which are accessible (Fig. 2).
Detailed analysis of secondary structure, evolutionary conservation, and properties of amino acid residues in TREM2 via structure-based sequence alignment inTREM2 family homologous to 5UD7.On top, secondary structure constituents are shown: β-strands with arrows, turns with TT letters, and helices with squiggles. In the sequencing block, conserved residues are written in red colour. Below bar represents the accessibility of 5UD7 where blue denotes for accessible, cyan for intermediate, and white shows the buried regions. On the second bar, pink represents hydrophobic and cyan hydrophilic
Estimation of thermodynamic stability
Protein stability can be further defined as the difference in energy between the folded and unfolded state of the protein [76]. It has been insightful for increasing the resistance against high temperature and harsh solution [77, 78] reducing immunogenicity due to local and global unfolding and subsequent aggregation [79,80,81,82], designing of novel functions and for directed evolution [83, 84]. In this study, apart from computing the thermodynamic stability of each confirmed pathogenic mutants, we also used computational saturation mutagenesis to estimate the protein folding stability as a test case. It is a technique where at the given position, single codon is replaced with all possible amino acids [85]. The methods use algorithms based on machine leaning approaches, energy-based, and knowledge-based methods to predict the thermodynamic stability of protein. As input, we have used the variants of TREM2 on its ectodomain associated with AD and NHD. The difference in free energy change between WT and mutant protein is defined as the thermodynamic stability, ∆∆G = ∆GMutant − ∆GWild type. For most of the tools, the calculated thermodynamic stability is said to be stabilizing if ∆∆G > 0 and destabilizing if ∆∆G < 0. For tools such as PoPMuSiC [58], NeEMO [64], PremPS, Rosetta, and FoldX [63], ∆∆G < 0 is said to be stabilizing, where ∆∆G > 0 is destabilizing. The predicted ∆∆G values with other parameters for all the tools are shown in Table S3. Web-based tool Fireprot [86] produces ∆∆G for both Rosetta [87] and FoldX [63], so we have considered results separately for FoldX and Rosetta. The predicted ∆∆G for all the tools is shown in Fig. 3 and Table S3 for the variants of both AD and NHD. All of the mutant variants have different G values, and most of them contribute to the destabilisation in TREM2. When the AD and NHD groups are compared, the higher destabilising impact is shown in the NHD conditions. Comparing the ∆∆G values of five variants, saturation mutagenesis at W50 and V126 are detrimental as the above values are affecting protein stability. Substitution at W50 and V126 is having deleterious effect both on structural and functional aspect of TREM2 protein [44]. W50 is present on the C′ β strand and as per ConSurf analysis; it is a highly conserved, buried residue having a structural role. It is located adjacent to C51 residue that responsible for the maintenance of Ig V-like domain in the protein [88]. Thus, substitution in the targeted position also affecting the interaction with neighbouring residue and probably affect the downstream signalling. Likewise, ConSurf predicts that V126 is a buried residue with high evolutionary conservation. It is also responsible for the responsible for the maintenance of hydrophobic core in Ig-like domain. Upon substitution, the above residue might cause disruption of the hydrophobic core [47]. MD simulation analysis deciphered that upon mutation to W50C and V126G results into higher residual mobility and H-bonding pattern in CDR1 region. V126G mutation results into the reduced H-bonding between R47 and T66 while increased the interaction between R47 and N68 that might alter the conformational change in CDR loops [89]. From AD pathological group, R47 and R62 positions were more pronounced with changes in the ∆∆G (Fig. 3). Both are present in the CDR1 and CDR2 regions respectively. These two regions are important for ligand binding and downstream signalling [89]. Experimental study on R47H has shown a change in the conformation and affecting ligand interaction significantly with phosphatidylserine, phosphatidyl ethanolamine, and sphingolipids [73, 90] while MD simulation results imply R62H mutation affects the binding loop of TREM2 at a low extent. Furthermore, Yeh et al. found that there is reduction in the binding ability of TREM2 with CLU/APOJ and APOE when both R47H and R62H mutations were present [91].
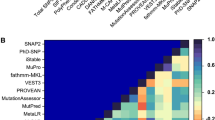
Representation of calculated ∆∆G with respect to different pathogenic mutants in Alzheimer’s (AD) and Nasu-Hakola disease (NHD) by saturation mutagenesis in different structure-based tools. R47H, R62H, and D87N are included in AD while W44X, W50C, W78X, and V126G are under NHD. Q33X is present in both AD and NHD
Q33X is found to be involved in both the diseases along with frontotemporal dementia [92] has a moderate effect on protein stability. Likely, D87 which is a surface exposed residue associated with AD risk has exhibited a destabilisation effect on TREM2 in saturated mutagenesis. To find out how this mutation is affecting protein stability, we included RSA and residual depth for further insights. Generated relative solvent accessibility values were studied to see if there is any difference between the two disorders’ RSA values and we could see that NHD mutants W50C and V126G have RSA values of 0.3% and 0% respectively according to all the tools except I-Mutant3.0 and Pop Music, and all the AD variants have an RSA value more than 20%. Details about the calculated RSA are provided in Table S4.
In WT, when Q33 is substituted with polar amino acid D or N, the residual depth is remaining same with increases RSA value. On the contrary, when it is switched to non-polar amino acid such as A and S, there is a decrease in both RSA and residual depth. When W44 is substituted with polar amino acids, thermodynamic stability of protein is significantly affected. Polar substitution such as Nand T causes increase in residual depth with concomitant decrease in RSA. Substitution with non-polar amino acid causes increase RSA with unchanged residual depth which often lead to destabilise the protein structure [93]. The protein stability value is highly affected when W50 is mutated to any polar amino acids. Irrespective of polar or non-polar amino acids, RSA of all the variants is increased, signifying the destabilisation upon mutation and establishing the crucial role of W50 in TREM2 protein. W78X is a nonpolar variant, when the substitutions are polar residues; there is increase in the RSA percentage with decrease in the residual depth. Similarly, R62H, D87N, and R47H variants are exhibiting increase in RSA with decrease or minimal change in residual depth across all variants. At V126 position, we could not observe any changes in the RSA. But residual depth and stability are affected with both polar and non-polar amino acid substitution (Table S5). Although all mutations are affecting protein stability, the analysis of residual depth and alteration in RSA indicate that buried or hydrophobic residues are mostly affected. The residues such as R62H, R47H, and D87N are surface residues and all of them are following a similar trend while W44X, W78X, W50C, and V126G are exhibiting maximum variations in RSA and residual depth. They are generally inducing the risk for NHD and could have a structural role in TREM2 protein. Furthermore, correlation of RSA and ∆∆G in various mutant variants will clarify the role.
Correlation between relative solvent accessibility (RSA) and the predicted ∆∆G
The main purpose of our exploration is to investigate how AD differ from NHD despite mutation taking place on the same protein TREM2, using computational and statistical approaches.
Here, we hypothesise that relative solvent accessibility would affect thermodynamic stability of protein that leads to neurodegenerative disorders. The differential correlation values would reveal the molecular basis of pathogenicity in two varied conditions. The relative solvent accessibility (RSA) of each residue was calculated by using SDM on the basis of the wild-type structure as input. The classification of residues is based on the RSA values. The residues with RSA < 20% were classified as buried and exposed residues are considered when RSA value is exceeding 20%. Then the graph was plotted in GraphPad Prism 6.0, where RSA was taken as a dependable while ∆∆G was independent factor. Then simple regression analysis is deciphering that the correlation is existing between these two factors in case of NHD (R2 = 0.60), while, in case of AD, the correlation was not significant (R2 = 0.06) (Fig. 4). Our analysis is well corroborated with the previous RSA and residual depth analysis. This result also supported by the in vitro experimental data suggesting that most of the NHD variants are buried, less exposed to the solvent and Alzheimer’s disease variants are on the surface being available to the solvent more [47]. Furthermore, evaluating the interaction pattern after mutation would offer a clear picture on the functionality of those critical residues.
Linear regression analysis between relative solubility accessibility (RSA) and ∆∆G in different mutant variants of AD and NHD. The plot was developed using simple regression analysis where ∆∆G and RSA are independent and dependent variables respectively. (A) AD is showing a minimal correlation. (B) A positive correlation is established in NHD
Residual network analysis in WT and mutant conformations
A web-based tool RING3.0 [71] was employed to decipher the interaction networks among the residues present in WT. Upon mutation to those residues cause changes in the interaction pattern (Fig. 5). All interactions in WT and alternation with mutation are shown in Table 2. In TREM 2 (ectodomain 19–134), these residues were found to be interacted with each other via 67 no of hydrogen bonds, 3 pi–pi interaction, 1 disulphide interaction, and 75 number of Van der Waal’s interactions. By zooming out specific residues in WT as well as in mutant variants, Q33 interacts with T94 via van der Waals interaction, but upon mutation to A33 resulted into loss of that contact. Q33 is present on the surface area B-β strand which is connecting to CDR1 loop in TREM2, responsible for ligand interaction. After a mutation, the lack of contact might change the aforementioned protein’s structure and function, which could lead to AD, NHD, or FTD. W44 is an exposed hydrophobic residue present at the CDR1 region in WT TREM2 protein, exhibiting pi–pi interaction with W70, present in the CDR2 region. A hydrogen bond is observed with M41 residue of same CDR1 region. Upon mutation to A41, the hydrogen bond is persisting with M41 but pi–pi interaction with W70 is demolished. The CDR1 and CDR2 regions are interconnected by different atomic interaction for structural integrity and ligand interaction [74]. The missing pi-pi network might disturb the structure and function which can be deciphered via structural analysis as well as classical molecular dynamics simulations. W50 is a conserved and buried residue located at C′ strand in TREM2. Mutational results show that RSA and residual depth are getting highly disturbed on substitution to C. Looking into the interaction pattern, W50 form some pi–pi interaction and 3 van der Waals bonds with Y108 while 2 van der Waals interaction with I95 positions. It interacts with V63 via 1 van der Waals and 1 H-bond, and V64 is in contact with W50 via 2 H-bonds. Upon mutation to C50, hydrogen bonds are remained same as in WT while there is loss of van der Waals contacts. Likewise, pi–pi interaction between C50 and Y108 is missing. Previous molecular dynamics simulation-based literature on structural dynamic behaviour of TREM2 protein has shown a minimal change occurring with the above mutation [74]. But it stands out as the greater risk variants for NHD and as C is a smaller and polar residue could have a role in structural integrity which is to be understood in further studies.
Residue interaction networks (RIN) displaying the various non-bonded interactions between all residues in WT and mutants of TREM2 ECD associated with AD and NHD. All interactions such as van der Waals interaction, H-bond, di-sulphide bond, and pi-pi interactions are shown with a distance threshold (Å) for H-bond 3.5, ionic 4, π-cation 5, π-π stacking 6.5, disulphide 2.5, and van der Waals 0.5. Q33, R62, and W78 have been exempted from the figure due to the complete dissolve of bonds
V126 is essential for maintaining the hydrophobic core in TREM2, which forms van der Waals contacts with L32 and Y108 and interacts with F24 and G106 residues by one and two hydrogen bonds respectively. Upon substitution to G leads to the loss of contacts between Y108, F24, and L32 but the interaction with G106 remains unchanged.
It is noteworthy that W50 and V126 are interacting with Y108 by two varied interaction patterns. Upon mutation, the bonds with Y108 are lost. This Y108 could be another important residue for the structure or functionality of TREM2, as it is a core residue present in the Ig domain along with F24, D104, and A105 [47]. W78 is a hydrophobic core residue present at C″ strand, interacting with A83, S81, and T82 via van der Waals networks. Upon mutations to alanine, all the interactions are lost.
D87 is polar amino acid and presents on the surface of D beta strand in TREM2. It makes contacts with G90, G91, and T92. Upon mutation to N87 which is again a polar amino acid, the interaction pattern remains the same. But how it affects the stability or causing disease effect is yet to be revealed. R62H and R47H are important risk variants for the development of AD. Both the residues are polar amino acids present on the surface of TREM2. R62 is on the C″ strand while R47 is present on the CDR1 region. Mutation on histidine is relatively a smaller amino acid having imidazole group instead of guanidino group in arginine. R62 has a van der Waals contact with A49 but upon the substitution with H62 causes the loss of above interaction. R47 has also one hydrogen bond with leu113, and on missense mutation with H47, the above interaction is lost. But few van der Waals contacts: N68, L69, Y70, and H67 due to the substitution of H are established. This interaction might change the structure of CDR1 loop which is shown in previous literature [35].
Conclusion
Most challenging aspect in neurodegenerative disorders has been identification of drug target and the molecular mechanism of the diseases. Prediction of protein stability helps in protein engineering and for the development of immunotherapeutic agent for drug discovery as well. Owing to limited confirmed pathogenic mutant data, we envisioned saturation mutagenesis technique, which can evaluate the effects of all probable substitutions (random mutations) at a certain protein residue position.
This study utilises structure-based computational approaches that deliver a range of ∆∆G for considered 8 mutants of TREM2 with known pathogenicity on its ECD. Observation based on ∆∆G score reveals most of the mutants to be destabilizing. Variants associated with NHD show shorter RSA values implying them to be hydrophobic in nature and being present in the core regions. Furthermore, while equated with residual depth, we see a decrease in RSA when depth is increased. Due to their presence in the enfolded regions, NHD variants are found to be indulged in structural alteration of TREM2, whereas AD variants are not affecting structural integrity rather they are influencing the ligand binding as most of them are surface exposed. The linear regression analysis also supports the hypothesis of RSA playing a role in the stability of TREM2 in case of NHD, where ∆∆G is found to be correlated with RSA.
Additionally, inter-residual network interaction highlights the non-bonded interactions in WT and mutant, and how a missense mutation is leading structural alteration of protein. A single hydrogen or pi–pi interaction or a van der Waals interaction is crucial for the WT functionality. Loss of such interaction could result in alteration in the structural integrity of the protein. Our observed residual interaction network displays that the inter-residual interaction is mostly lost upon mutation, but R47H has shown establishment of new contacts altering the arrangement of CDR2 loop region and downstream β-strand C″, important for phospholipid interaction. This residue could be an important aspect for ligand binding which needs to be studied further to provide a clear understanding to develop novel therapeutics for neurodegenerative disorders. W50C and V126G display maximum destabilizing effect upon mutational analysis; they have shown complete loss of interaction with single polar amino acid substitution. We also find both the residues are interacting with Y108 residues in WT which is lost upon mutation. The residue Y108 might be playing a crucial role in the functionality of the protein and needs to be studied in the future.
Abbreviations
- AD:
-
Alzheimer’s disease
- Aβ:
-
Amyloid beta
- CDR:
-
Complementarity-determining regions
- ECD:
-
Extra-cellular domain
- NHD:
-
Nasu-Hakola disease
- RIN:
-
Residue interaction network
- RSA:
-
Relative solvent accessibility
- TREM:
-
Triggering receptors expressed on myeloid
References
Takahashi K, Rochford CDP, Neumann H (2005) Clearance of apoptotic neurons without inflammation by microglial triggering receptor expressed on myeloid cells-2. J Exp Med 201:647–657. https://doi.org/10.1084/jem.20041611
Bouchon A, Dietrich J, Colonna M (2000) Cutting edge: inflammatory responses can be triggered by TREM-1, a novel receptor expressed on neutrophils and monocytes. J Immunol 164:4991–4995. https://doi.org/10.4049/jimmunol.164.10.4991
Allcock RJN, Barrow AD, Forbes S, Beck S, Trowsdale J (2003) The human TREM gene cluster at 6p21.1 encodes both activating and inhibitory single lgV domain receptors and includes NKp44. Eur J Immunol 33:567–577. https://doi.org/10.1002/immu.200310033
Gratuze M, Leyns CEG, Holtzman DM (2018) New insights into the role of TREM2 in Alzheimer’s disease. Mol Neurodegener 13. https://doi.org/10.1186/s13024-018-0298-9
Borroni B, Ferrari F, Galimberti D, Nacmias B, Barone C, Bagnoli S, Fenoglio C, Piaceri I, Archetti S, Bonvicini C, Gennarelli M, Turla M, Scarpini E, Sorbi S, Padovani A (2014) Heterozygous TREM2 mutations in frontotemporal dementia. Neurobiol Aging 35(934):e7-934.e10. https://doi.org/10.1016/j.neurobiolaging.2013.09.017
Lue LF, Schmitz CT, Serrano G, Sue LI, Beach TG, Walker DG (2015) TREM2 protein expression changes correlate with Alzheimer’s disease neurodegenerative pathologies in post-mortem temporal cortices. Brain Pathol 25:469–480. https://doi.org/10.1111/bpa.12190
Koth LL, Cambier CJ, Ellwanger A, Solon M, Hou L, Lanier LL, Abram CL, Hamerman JA, Woodruff PG (2010) DAP12 is required for macrophage recruitment to the lung in response to cigarette smoke and chemotaxis toward CCL2. J Immunol 184:6522–6528. https://doi.org/10.4049/jimmunol.0901171
Wang Y, Ulland TK, Ulrich JD, Song W, Tzaferis JA, Hole JT, Yuan P, Mahan TE, Shi Y, Gilfillan S, Cella M, Grutzendler J, DeMattos RB, Cirrito JR, Holtzman DM, Colonna M (2016) TREM2-mediated early microglial response limits diffusion and toxicity of amyloid plaques. J Exp Med 213:667–675. https://doi.org/10.1084/jem.20151948
Otero K, Shinohara M, Zhao H, Cella M, Gilfillan S, Colucci A, Faccio R, Ross FP, Teitelbaum SL, Takayanagi H, Colonna M (2012) TREM2 and β-catenin regulate bone homeostasis by controlling the rate of osteoclastogenesis. J Immunol 188:2612–2621. https://doi.org/10.4049/jimmunol.1102836
Carlyle WC, Mcclain JB, Tzafriri AR, Bailey L, Brett G, Markham PM, Stanley JRL, Edelman ER, Sciences CB, Technologies E, Park LT (2015) TREM2 maintains microglial metabolic fitness in Alzheimer’s disease. Cell 162:561–567
Andreone BJ, Przybyla L, Llapashtica C, Rana A, Davis SS, van Lengerich B, Lin K, Shi J, Mei Y, Astarita G, Di Paolo G, Sandmann T, Monroe KM, Lewcock JW (2020) Alzheimer’s-associated PLCγ2 is a signaling node required for both TREM2 function and the inflammatory response in human microglia. Nat Neurosci 23:927–938. https://doi.org/10.1038/s41593-020-0650-6
Bouchon A, Hernández-Munain C, Cella M, Colonna M (2001) A DAP12-mediated pathway regulates expression of CC chemokine receptor 7 and maturation of human dendritic cells. J Exp Med 194:1111–1122. https://doi.org/10.1084/jem.194.8.1111
Kawabori M, Kacimi R, Kauppinen T, Calosing C, Kim JY, Hsieh CL, Nakamura MC, Yenari MA (2015) Triggering receptor expressed on myeloid cells 2 (TREM2) deficiency attenuates phagocytic activities of microglia and exacerbates ischemic damage in experimental stroke. J Neurosci 35:3384–3396. https://doi.org/10.1523/JNEUROSCI.2620-14.2015
Hakola HP (1972) Neuropsychiatric and genetic aspects of a new hereditary disease characterized by progressive dementia and lipomembranous polycystic osteodysplasia. Acta Psychiatr Scand Suppl 232:1–173
Lue LF, Schmitz C, Walker DG (2015) What happens to microglial TREM2 in Alzheimer’s disease: immunoregulatory turned into immunopathogenic? Neuroscience 302:138–150. https://doi.org/10.1016/j.neuroscience.2014.09.050
Deczkowska A, Weiner A, Amit I (2020) The physiology, pathology, and potential therapeutic applications of the TREM2 signaling pathway. Cell 181:1207–1217. https://doi.org/10.1016/j.cell.2020.05.003
Wunderlich P, Glebov K, Kemmerling N, Tien NT, Neumann H, Walter J (2013) Sequential proteolytic processing of the triggering receptor expressed on myeloid cells-2 (TREM2) protein by ectodomain shedding and γ-secretase- dependent intramembranous cleavage. J Biol Chem 288:33027–33036. https://doi.org/10.1074/jbc.M113.517540
Takalo M, Wittrahm R, Wefers B, Parhizkar S, Jokivarsi K, Kuulasmaa T, Mäkinen P, Martiskainen H, Wurst W, Xiang X, Marttinen M, Poutiainen P, Haapasalo A, Hiltunen M, Haass C (2020) The Alzheimer’s disease-associated protective Plcγ2-P522R variant promotes immune functions. Mol Neurodegener 15. https://doi.org/10.1186/s13024-020-00402-7
Molinuevo JL, Ayton S, Batrla R, Bednar MM, Bittner T, Cummings J, Fagan AM, Hampel H, Mielke MM, Mikulskis A, O’bryant S, Scheltens P, Sevigny J, Shaw LM, Soares HD, Tong G, Trojanowski JQ, Zetterberg H, Blennow K (2018) Current state of Alzheimer’s fluid biomarkers. Acta Neuropathol. 136:821–853. https://doi.org/10.1007/s00401-018-1932-x
Filipello F, Goldsbury C, You SF, Locca A, Karch CM, Piccio L (2022) Soluble TREM2: innocent bystander or active player in neurological diseases? Neurobiol Dis 165:105630. https://doi.org/10.1016/j.nbd.2022.105630
Suárez‐Calvet M, Kleinberger G, Araque Caballero MÁ, Brendel M, Rominger A, Alcolea D, Fortea J, Lleó A, Blesa R, Gispert JD, Sánchez‐Valle R (2016) sTREM-2 cerebrospinal fluid levels are a potential biomarker for microglia activity in early-stage Alzheimer's disease and associate with neuronal injury markers. EMBO Mol Med 8: 466–476. https://doi.org/10.15252/emmm.201506123
Rodriguez-Vieitez E, Ashton NJ (2021) Plasma sTREM2: a potential marker of cerebrovascular injury in neurodegenerative disorders. Brain 144:3283–3285. https://doi.org/10.1093/brain/awab399
Guerreiro R, Wojtas A, Bras J, Carrasquillo M, Rogaeva E, Majounie E, Cruchaga C, Sassi C, Kauwe JSK, Younkin S, Hazrati L, Collinge J, Pocock J, Lashley T, Williams J, Lambert J-C, Amouyel P, Goate A, Rademakers R, Morgan K, Powell J, St. George-Hyslop P, Singleton A, Hardy J (2013) TREM2 variants in Alzheimer’s disease. N Engl J Med 368:117–127. https://doi.org/10.1056/nejmoa1211851
Jonsson T, Stefansson H, Steinberg S, Jonsdottir I, Jonsson PV, Snaedal J, Bjornsson S, Huttenlocher J, Levey AI, Lah JJ, Rujescu D (2013) Variant of TREM2 associated with the risk of Alzheimer’s disease. N Engl J Med 368:107–116. https://doi.org/10.1056/NEJMoa1211103
Rayaprolu S, Mullen B, Baker M, Lynch T, Finger E, Seeley WW, Hatanpaa KJ, Lomen-Hoerth C, Kertesz A, Bigio EH, Lippa C (2013) TREM2 in neurodegeneration: evidence for association of the p. R47H variant with frontotemporal dementia and Parkinson’s disease. Mol Neurodegener 8:1–5. https://doi.org/10.1186/1750-1326-8-19
Cuyvers E, Bettens K, Philtjens S, Van Langenhove T, Gijselinck I, van der Zee J, Engelborghs S, Vandenbulcke M, Van Dongen J, Geerts N, Maes G, Mattheijssens M, Peeters K, Cras P, Vandenberghe R, De Deyn PP, Van Broeckhoven C, Cruts M, Sleegers K (2014) Investigating the role of rare heterozygous TREM2 variants in Alzheimer’s disease and frontotemporal dementia. Neurobiol Aging 35(726):e11-726.e19. https://doi.org/10.1016/j.neurobiolaging.2013.09.009
Holtzman DM, Morris JC, Goate AM (2011) Alzheimer’s disease: the challenge of the second century. Sci Transl Med 3. https://doi.org/10.1126/scitranslmed.3002369
Hardy J, Selkoe DJ (2002) The amyloid hypothesis of Alzheimer’s disease: progress and problems on the road to therapeutics. Science (80-. ) 297:353–356. https://doi.org/10.1126/science.1072994
Area-Gomez E, Schon EA (2017) Alzheimer disease. Adv Exp Med Biol 997:149–156. https://doi.org/10.1007/978-981-10-4567-7_11
Huang et al (2002) Alzheimer mechanisms and therapeutic strategies. Science (80-. ) 148:1204–1222
Schoepp NG, Khorosheva EM, Schlappi TS, Curtis MS, Humphries RM, Angeles L, Annex B, Angeles L, Hindler JA, Angeles L, Annex B, Angeles L, Ismagilov RF (2017) TREM2 haplodeficiency in mice and humans impairs the microglia barrier function leading to decreased amyloid compaction and severe axonal dystrophy. Neuron 55:9557–9561
Keren-Shaul H, Spinrad A, Weiner A, Matcovitch-Natan O, Dvir-Szternfeld R, Ulland TK, David E, Baruch K, Lara-Astaiso D, Toth B, Itzkovitz S, Colonna M, Schwartz M, Amit I (2017) A unique microglia type associated with restricting development of Alzheimer’s disease. Cell 169:1276-1290.e17. https://doi.org/10.1016/j.cell.2017.05.018
Condello C, Yuan P, Schain A, Grutzendler J (2015) Microglia constitute a barrier that prevents neurotoxic protofibrillar Aβ42 hotspots around plaques. Nat Commun 6. https://doi.org/10.1038/ncomms7176
Colonna M, Wang Y (2016) TREM2 variants: new keys to decipher Alzheimer disease pathogenesis. Nat Rev Neurosci 17:201–207. https://doi.org/10.1038/nrn.2016.7
Sudom A, Talreja S, Danao J, Bragg E, Kegel R, Min X, Richardson J, Zhang Z, Sharkov N, Marcora E, Thibault S, Bradley J, Wood S, Lim AC, Chen H, Wang S, Foltz IN, Sambashivan S, Wang Z (2018) Molecular basis for the loss-of-function effects of the Alzheimer’s disease-associated R47H variant of the immune receptor TREM2. J Biol Chem 293:12634–12646. https://doi.org/10.1074/jbc.RA118.00235
Cheng-Hathaway PJ, Reed-Geaghan EG, Jay TR, Casali BT, Bemiller SM, Puntambekar SS, Von Saucken VE, Williams RY, Karlo JC, Moutinho M, Xu G, Ransohoff RM, Lamb BT, Landreth GE (2018) The Trem2 R47H variant confers loss-of-function-like phenotypes in Alzheimer’s disease. Mol Neurodegener 13. https://doi.org/10.1186/s13024-018-0262-8
Luis EO, Ortega-Cubero S, Lamet I, Razquin C, Cruchaga C, Benitez BA, Lorenzo E, Irigoyen J, Pastor MA, Pastor P, Initiative Alzheimer’s Disease Neuroimaging, (ADNI), (2014) Frontobasal gray matter loss is associated with the TREM2 p. R47H variant. Neurobiol Aging 35:2681–2690. https://doi.org/10.1016/j.neurobiolaging.2014.06.007
Del-Aguila JL, Fernández MV, Schindler S, Ibanez L, Deming Y, Ma S, Saef B, Black K, Budde J, Norton J, Chasse R, Harari O, Goate A, Xiong C, Morris JC, Cruchaga C (2018) Assessment of the genetic architecture of Alzheimer’s disease risk in rate of memory decline. J Alzheimer’s Dis 62:745–756. https://doi.org/10.3233/JAD-170834
Jin SC, Benitez BA, Karch CM, Cooper B, Skorupa T, Carrell D, Norton JB, Hsu S, Harari O, Cai Y, Bertelsen S, Goate AM, Cruchaga C (2014) Coding variants in TREM2 increase risk for Alzheimer’s disease. Hum Mol Genet 23:5838–5846. https://doi.org/10.1093/hmg/ddu277
Sims R, Badarinarayan N, Raybould R, Heilmann-Heimbach S, Vronskaya M, Hoffmann P (2017) Rare coding variants in PLCG2, ABI3 and TREM2 implicate microglial-mediated innate immunity in Alzheimer’s disease. Nat Genet 49:1373–1384
Jiang T, Hou J-K, Gao Q, Yu J-T, Zhou J-S, Zhao H-D, Zhang Y-D (2016) TREM2 p.H157Y variant and the risk of Alzheimer’s disease: a meta-analysis involving 14,510 subjects. Curr Neurovasc Res 13:318–320. https://doi.org/10.2174/1567202613666160808095530
Paloneva J, Kestilä M, Wu J, Salminen A, Böhling T, Ruotsalainen V, Hakola P, Bakker AB, Phillips JH, Pekkarinen P, Lanier LL (2000) Loss-of-function mutations in TYROBP (DAP12) result in a presenile dementia with bone cysts. Nat Genet 25:357–361. https://doi.org/10.1038/77153
Paloneva J, Manninen T, Christman G, Hovanes K, Mandelin J, Adolfsson R, Bianchin M, Bird T, Miranda R, Salmaggi A, Tranebjærg L, Konttinen Y, Peltonen L (2002) Mutations in two genes encoding different subunits of a receptor signaling complex result in an identical disease phenotype. Am J Hum Genet 71:656–662. https://doi.org/10.1086/342259
Dardiotis E, Siokas V, Pantazi E, Dardioti M, Rikos D, Xiromerisiou G, Markou A, Papadimitriou D, Speletas M, Hadjigeorgiou GM (2017) A novel mutation in TREM2 gene causing Nasu-Hakola disease and review of the literature. Neurobiol Aging 53(194):e13-194.e22. https://doi.org/10.1016/j.neurobiolaging.2017.01.015
Klünemann HH, Ridha BH, Magy L, Wherrett JR, Hemelsoet DM, Keen RW, De Bleecker JL, Rossor MN, Marienhagen J, Klein HE, Peltonen L, Paloneva J (2005) The genetic causes of basal ganglia calcification, dementia, and bone cysts: DAP12 and TREM2. Neurology 64:1502–1507. https://doi.org/10.1212/01.WNL.0000160304.00003.CA
Paloneva J, Autti T, Raininko R, Partanen J, Salonen O, Puranen M, Hakola P, Haltia M (2001) CNS manifestations of Nasu-Hakola disease: a frontal dementia with bone cysts. Neurology 56:1552–1558. https://doi.org/10.1212/wnl.56.11.1552
Kober DL, Alexander-Brett JM, Karch CM, Cruchaga C, Colonna M, Holtzman MJ, Brett TJ (2016) Neurodegenerative disease mutations in TREM2 reveal a functional surface and distinct loss-of-function mechanisms. Elife 5. https://doi.org/10.7554/eLife.20391
Sirkis DW, Aparicio RE, Schekman R (2017) Neurodegeneration-associated mutant TREM2 proteins abortively cycle between the ER and ER–Golgi intermediate compartment. Mol Biol Cell 28:2723–2733. https://doi.org/10.1091/mbc.e17-06-0423
Paloneva J, Mandelin J, Kiialainen A, Böhling T, Prudlo J, Hakola P, Haltia M, Konttinen YT, Peltonen L (2003) DAP12/TREM2 deficiency results in impaired osteoclast differentiation and osteoporotic features. J Exp Med 198:669–675. https://doi.org/10.1084/jem.20030027
Song W, Hooli B, Mullin K, Jin SC, Cella M, Ulland TK, Wang Y, Tanzi RE, Colonna M (2017) Alzheimer’s disease-associated TREM2 variants exhibit either decreased or increased ligand-dependent activation. Alzheimer’s Dement 13:381–387. https://doi.org/10.1016/j.jalz.2016.07.004
Tang N, Dehury B, Kepp KP (2019) Computing the pathogenicity of Alzheimer’s disease presenilin 1 mutations. J Chem Inf Model 59:858–870. https://doi.org/10.1021/acs.jcim.8b00896
Berman PEBHM, Westbrook J, Feng Z, Gilliland G, Bhat TN, Weissig H, Shindyalov IN, Berman HM, Westbrook J, Feng Z, Gilliland G, Bhat TN, Weissig H, Shindyalov IN, Bourne PE (2000) The Protein Data Bank nucleic acids research, 28: 235–242. Protein Data Bank Nucleic Acids Res 28(2000):235–242
Ashkenazy H, Abadi S, Martz E, Chay O, Mayrose I, Pupko T, Ben-Tal N (2016) ConSurf 2016: an improved methodology to estimate and visualize evolutionary conservation in macromolecules. Nucleic Acids Res 44(2016):W344–W350. https://doi.org/10.1093/NAR/GKW408
Gouet P, Robert X, Courcelle E (2003) ESPript/ENDscript: extracting and rendering sequence and 3D information from atomic structures of proteins. Nucleic Acids Res 31:3320–3323. https://doi.org/10.1093/nar/gkg556
Schrödinger L (n.d.) The PyMOL Molecular Graphics, Version 2.5.2
Daggett V, Fersht AR (2003) Is there a unifying mechanism for protein folding? Trends Biochem Sci 28:18–25. https://doi.org/10.1016/S0968-0004(02)00012-9
Gromiha MM, Huang L-T (2011) Machine learning algorithms for predicting protein folding rates and stability of mutant proteins: comparison with statistical methods. Curr Protein Pept Sci 12:490–502. https://doi.org/10.2174/138920311796957630
Dehouck Y, Kwasigroch JM, Gilis D, Rooman M (2011) PoPMuSiC 2.1: a web server for the estimation of protein stability changes upon mutation and sequence optimality. BMC Bioinformatics 12. https://doi.org/10.1186/1471-2105-12-151
Parthiban V, Gromiha MM, Schomburg D (2006) CUPSAT: prediction of protein stability upon point mutations. Nucleic Acids Res 34. https://doi.org/10.1093/nar/gkl190
Pires DEV, Ascher DB, Blundell TL (2014) MCSM: predicting the effects of mutations in proteins using graph-based signatures. Bioinformatics 30:335–342. https://doi.org/10.1093/bioinformatics/btt691
Capriotti E, Fariselli P, Casadio R (2005) I-Mutant2.0: predicting stability changes upon mutation from the protein sequence or structure. Nucleic Acids Res 33. https://doi.org/10.1093/nar/gki375
Musil M, Stourac J, Bendl J, Brezovsky J, Prokop Z, Zendulka J, Martinek T, Bednar D, Damborsky J (2017) FireProt: web server for automated design of thermostable proteins. Nucleic Acids Res 45:W393–W399. https://doi.org/10.1093/nar/gkx285
Schymkowitz J, Borg J, Stricher F, Nys R, Rousseau F, Serrano L (2005) The FoldX web server: an online force field. Nucleic Acids Research 33:W382–W388. https://doi.org/10.1093/nar/gki387
Giollo M, Martin AJM, Walsh I, Ferrari C, Tosatto SCE (2014) NeEMO: a method using residue interaction networks to improve prediction of protein stability upon mutation. BMC Genomics 15:1–11. https://doi.org/10.1186/1471-2164-15-S4-S7
Pucci F, Bourgeas R, Rooman M (2016) Predicting protein thermal stability changes upon point mutations using statistical potentials: introducing HoTMuSiC. Sci Rep 6. https://doi.org/10.1038/srep23257
Worth CL, Preissner R, Blundell TL (2011) SDM - a server for predicting effects of mutations on protein stability and malfunction. Nucleic Acids Res 39. https://doi.org/10.1093/nar/gkr363
Pires DEV, Ascher DB, Blundell TL (2014) DUET: a server for predicting effects of mutations on protein stability using an integrated computational approach. Nucleic Acids Res 42. https://doi.org/10.1093/nar/gku411
Rodrigues CHM, Pires DEV, Ascher DB (2018) DynaMut: predicting the impact of mutations on protein conformation, flexibility and stability. Nucleic Acids Res 46:W350–W355. https://doi.org/10.1093/nar/gky300
Chen Y, Lu H, Zhang N, Zhu Z, Wang S, Li M (2020) PremPS: predicting the impact of missense mutations on protein stability. PLoS Comput Biol 16:1–22. https://doi.org/10.1371/journal.pcbi.1008543
Montanucci L, Capriotti E, Birolo G, Benevenuta S, Pancotti C, Lal D, Fariselli P (2022) DDGun: an untrained predictor of protein stability changes upon amino acid variants. Nucleic Acids Res 50:W222–W227. https://doi.org/10.1093/nar/gkac325
Clementel D, Del Conte A, Monzon AM, Camagni GF, Minervini G, Piovesan D, Tosatto SCE (2022) RING 3.0: fast generation of probabilistic residue interaction networks from structural ensembles. Nucleic Acids Res. 50:W651–W656. https://doi.org/10.1093/nar/gkac365
Li Z, Del-Aguila JL, Dube U, Budde J, Martinez R, Black K, Xiao Q, Cairns NJ, Network The Dominantly Inherited Alzheimer, (DIAN), Dougherty JD, Lee J-M, Morris JC, Bateman RJ, Karch CM, Cruchaga C, Harari O, (2018) Genetic variants associated with Alzheimer’s disease confer different cerebral cortex cell-type population structure. Nucleic Acids Res 10:1–19. https://doi.org/10.1186/s13073-018-0551-4
Sudom A, Talreja S, Danao J, Bragg E, Kegel R, Min X, Richardson J, Zhang Z, Sharkov N, Marcora E, Thibault S, Bradley J, Wood S, Lim A-C, Chen H, Wang S, Foltz IN, Sambashivan S, Wang Z (2018) Molecular basis for the loss-of-function effects of the Alzheimer’s disease–associated R47H variant of the immune receptor TREM2. J Biol Chem 293:12634–12646. https://doi.org/10.1074/jbc.ra118.002352
Dash R, Choi HJ, Moon IS (2020) Mechanistic insights into the deleterious roles of Nasu-Hakola disease associated TREM2 variants. Sci Rep 10. https://doi.org/10.1038/s41598-020-60561-x
Guerreiro RJ, Lohmann E, Brás JM, Gibbs JR, Rohrer JD, Gurunlian N, Dursun B, Bilgic B, Hanagasi H, Gurvit H, Emre M, Singleton A, Hardy J (2012) Using exome sequencing to reveal mutations in TREM2 presenting as a frontotemporal dementia–like syndrome without bone involvement. JAMA Neurol 1. https://doi.org/10.1001/archneurol.2013.579
Ahmad F (2022) Protein stability [determination] problems. Front Mol Biosci 9. https://doi.org/10.3389/fmolb.2022.880358
Bommarius AS, Paye MF (2013) Stabilizing biocatalysts. Chem Soc Rev 42:6534–6565. https://doi.org/10.1039/c3cs60137d
Magliery TJ (2015) Protein stability: computation, sequence statistics, and new experimental methods. Curr Opin Struct Biol 33:161–168. https://doi.org/10.1016/j.sbi.2015.09.002
Sauerborn M, Brinks V, Jiskoot W, Schellekens H (2010) Immunological mechanism underlying the immune response to recombinant human protein therapeutics. Trends Pharmacol Sci 31:53–59. https://doi.org/10.1016/j.tips.2009.11.001
Manning MC, Chou DK, Murphy BM, Payne RW, Katayama DS (2010) Stability of protein pharmaceuticals: an update. Pharm Res 27:544–575. https://doi.org/10.1007/s11095-009-0045-6
Boulet-Audet M, Byrne B, Kazarian SG (2014) High-throughput thermal stability analysis of a monoclonal antibody by attenuated total reflection FT-IR spectroscopic imaging. Anal Chem 86:9786–9793. https://doi.org/10.1021/ac502529q
Uversky V, Breydo L, Redington J (2017) When good goes awry: the aggregation of protein therapeutics. Protein Pept Lett 24:340–347. https://doi.org/10.2174/0929866524666170209153421
Bloom JD, Labthavikul ST, Otey CR, Arnold FH (2006) Protein stability promotes evolvability. Proc Natl Acad Sci U S A 103:5869–5874. https://doi.org/10.1073/pnas.0510098103
Nobuhiko T, Tawfik DS (2009) Stability effects of mutations and protein evolvability. Curr Opin Struct Biol 19:596–604
Reetz MT, Carballeira JD (2007) Iterative saturation mutagenesis (ISM) for rapid directed evolution of functional enzymes. Nat Protoc 2:891–903. https://doi.org/10.1038/nprot.2007.72
Stourac J, Dubrava J, Musil M, Horackova J, Damborsky J, Mazurenko S, Bednar D (2021) FireProtDB: database of manually curated protein stability data. Nucleic Acids Res 49:D319–D324. https://doi.org/10.1093/nar/gkaa981
Kellogg EH, Leaver-Fay A, Baker D (2011) Role of conformational sampling in computing mutation-induced changes in protein structure and stability. Proteins Struct Funct Bioinforma 79:830–838. https://doi.org/10.1002/prot.22921
Reifschneider A, Robinson S, van Lengerich B, Gnörich J, Logan T, Heindl S, Vogt MA, Weidinger E, Riedl L, Wind K, Zatcepin A, Pesämaa I, Haberl S, Nuscher B, Kleinberger G, Klimmt J, Götzl JK, Liesz A, Bürger K, Brendel M, Levin J, Diehl‐Schmid J, Suh J, Di Paolo G, Lewcock JW, Monroe KM, Paquet D, Capell A, Haass C (2022) Loss of TREM2 rescues hyperactivation of microglia, but not lysosomal deficits and neurotoxicity in models of progranulin deficiency. EMBO J 41. https://doi.org/10.15252/embj.2021109108
Dash R, Munni YA, Mitra S, Choi HJ, Jahan SI, Chowdhury A, Jang TJ, Moon IS (2022) Dynamic insights into the effects of nonsynonymous polymorphisms (nsSNPs) on loss of TREM2 function. Sci Rep 12. https://doi.org/10.1038/s41598-022-13120-5
Hall-Roberts H, Agarwal D, Obst J, Smith TB, Monzón-Sandoval J, Di Daniel E, Webber C, James WS, Mead E, Davis JB, Cowley SA (2020) TREM2 Alzheimer’s variant R47H causes similar transcriptional dysregulation to knockout, yet only subtle functional phenotypes in human iPSC-derived macrophages. Alzheimer’s Res Ther 12:1–27. https://doi.org/10.1186/s13195-020-00709-z
Yeh FL, Wang Y, Tom I, Gonzalez LC, Sheng M (2016) TREM2 binds to apolipoproteins, including APOE and CLU/APOJ, and thereby facilitates uptake of amyloid-beta by microglia. Neuron 91:328–340. https://doi.org/10.1016/j.neuron.2016.06.015
Patel D, Mez J, Vardarajan BN, Staley L, Chung J, Zhang X, Farrell JJ, Rynkiewicz MJ, Cannon-Albright LA, Teerlink CC, Stevens J, Corcoran C, Gonzalez Murcia JD, Lopez OL, Mayeux R, Haines JL, Pericak-Vance MA, Schellenberg G, Kauwe JSK, Lunetta KL, Farrer LA (2019) Association of rare coding mutations with Alzheimer disease and other dementias among adults of European ancestry. JAMA Netw Open 2(3):e191350. https://doi.org/10.1001/jamanetworkopen.2019.1350
Strub C, Alies C, Lougarre A, Ladurantie C, Czaplicki J, Fournier D (2004) Mutation of exposed hydrophobic amino acids to arginine to increase protein stability. BMC Biochem 5:1–6. https://doi.org/10.1186/1471-2091-5-9
Acknowledgements
The authors acknowledge the support from ICMR-RMRC, Bhubaneswar, for computational facility. The authors also acknowledge DHR for providing Young Scientist grant to BD. The authors also acknowledge ICMR for providing ICMR-Centenary Postdoctoral Fellowship to SP.
Author information
Authors and Affiliations
Contributions
Preety Sthutika Swain: methodology, software investigation, data curation, visualisation, writing — original draft.
Sunita Panda: data curation, methodology, review and editing.
Sanghamitra Pati: writing — review and editing.
Budheswar Dehury: conceptualisation, methodology, review and editing, and supervision.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
The data supporting this manuscript is listed in the “Supplementary information” section.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Swain, P.S., Panda, S., Pati, S. et al. Computational saturation mutagenesis to explore the effect of pathogenic mutations on extra-cellular domains of TREM2 associated with Alzheimer’s and Nasu-Hakola disease. J Mol Model 29, 360 (2023). https://doi.org/10.1007/s00894-023-05770-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00894-023-05770-7