Abstract
Perampanel approved by FDA in 2012 is a first-in-class antiepileptic drug which inhibits α-amino-3-hydroxy-5-methyl-4-isoxazolepropionic acid (AMPA) receptor currents. It is markedly more active than many of its close analogs, and the reasons for this activity difference are not quite clear. Recent crystallographic studies allowed the authors to identify the location of its binding site. Unfortunately, the resolution is low, and the detailed description of perampanel binding mode is still in part speculative. Here we provide a detailed DFT-level conformational analysis of perampanel in a vacuum and in the solvents, mimicking the protein environment, followed by quantum theory of atoms in molecules (QTAIM), non-covalent interactions (NCI), and natural bond orbital (NBO) analyses. The findings indicate the electrostatic nature of the intramolecular interactions which contribute to energy differences of the conformations in a vacuum whereas the increase of dielectric constant leads to the energy equalization of conformations. Based on these results, the docking study was performed to investigate possible binding modes of perampanel and its close analogs in AMPA receptors. The influence of the pyridine nitrogen and cyano group position was explained based on the results of conformational analysis and molecular docking. These findings may contribute to the design of novel antiepileptic drugs and the development of novel approaches to treat neurodegenerative diseases and major depressive disorder.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Ionotropic glutamate receptors are membrane-embedded tetrameric proteins acting as ligand-gated ion channels involved in the process of fast synaptic transmission. Their natural regulation plays an important role in the processes of learning and memory formation [1]. Blocking of these receptors leads to the decrease in synaptic transmission of glutamatergic synapses and may be used in the treatment of many CNS disorders such as epilepsy [2], Parkinson’s disease [3], and neuropathic pain [4]. The channel blockers may also possess neuroprotective function due to the ability of some glutamate receptor subtypes to transport calcium ions through the membrane causing the increased risk of the excitotoxicity-associated processes and the neuronal cell death [5]. Beside the inhibition, the stimulation of ionotropic glutamate receptors via positive allosteric modulators (PAMs) is also a subject of scientific interest since it can be used in the development of cognitive enhancers [1]. Among the glutamate receptors types, α-amino-3-hydroxy-5-methyl-4-isoxazolepropionic acid (AMPA) receptors are considered to be the most appropriate for this purpose due to the reduced ability for the most abundant AMPA receptor subtypes in the brain to permeate calcium ions [6].
AMPA subtype of ionotropic glutamate receptors is a perspective target for small molecule drugs, and AMPA receptor antagonists were shown to have anticonvulsant activity and thus being suitable for epilepsy treatment [7]. The first FDA-approved anticonvulsant, which acts through AMPA receptor antagonism is perampanel [8]. Its successful clinical use [9] will stimulate the development of the other AMPA receptor antagonists applied to epilepsy treatment. It is important to note that certain close analogs of perampanel [8] demonstrate the so-called activity cliffs, i.e., sharp activity decay in comparison with perampanel. The reasons explaining this effect have not been reported yet. Thorough conformational analysis of perampanel and the related structures accompanied by the estimation of the rotation potential energy surfaces (PES) might be useful in understanding perampanel biological behavior. Comparison of this compound with its structurally close but less active analogs may help to determine the features which are crucial for biological activity. It should be noted that the robust conformational analysis can be helpful to investigate interactions observed in protein-ligand complexes [10, 11].
The development of the novel drugs of this class is facilitated by the release of the X-ray structures of AMPA receptor complexes with several noncompetitive antagonists including perampanel [12]. Unfortunately, due to the relatively low resolution, some atomistic details of ligand binding need refinement before using in the structure-based analog design. Nevertheless, a possible mechanism of perampanel action was revealed [12, 13]. Figure 1 (left) demonstrates the location of perampanel binding site while the right part of it reveals the fact that perampanel binding pocket is accommodated by the M3 helix in the open-channel form (Fig. 1 (right)).
Molecular modeling approaches have been extensively used to develop novel ligands of glutamate receptors [14,15,16,17]. However, the perampanel binding site was not investigated due to high uncertainty of its location. Recently published crystallographic data [12] allowed researchers to fill this gap. The goal of the current study is to rationalize these data and refine the perampanel position in the binding site using quantum chemical and molecular modeling approaches.
In the current work, perampanel (compound 1) and five of its structurally close but less active analogs (compounds 2–6, Fig. 2) were investigated by means of density functional theory (DFT) methods together with quantum theory of atoms in molecules (QTAIM), natural bond orbital (NBO), and non-covalent interaction (NCI) analyses applied. The shape and height of the internal rotation barriers were estimated to assess the conformational flexibility of the investigated molecules. Based on the results of quantum chemical calculations, a robust docking study using the X-ray structure of full-length AMPA receptor [12] was carried out for perampanel and its analogs.
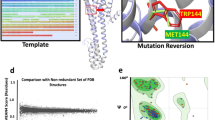
The structure of perampanel (compound 1) and its analogs (compounds 2–6). Bold capital letters denote the nomenclature of aromatic cycles. Colored bonds define the dihedral angles D1, D2, and D3 used for conformational analysis. Activity data is presented on the right [8]
Methods
The conformational space of molecules 1–6 was investigated using DFT methods as implemented in ORCA quantum chemistry program (version 4.1.0) [18, 19]. The initial conformer geometries were found by optimization using the MMFF94s force field [20], as implemented in Avogadro program [21]. The geometry optimizations and the relaxed PES scans were performed at the B3LYP/6-31G* level [22,23,24]. The PES scans were obtained by systematic sampling (with 5° step size) of the dihedral angles D1–D3, defined in Fig. 2. The results obtained were verified by calculations with basis set extension and application of other hybrid GGA functional: B3LYP/6-311G* and PBE0/cc-pVTZ methods [25,26,27]. The polarizable continuum model (PCM) was applied in its density-based form for solvent (SMD) [28] for all calculations concerning the solvation effects. In all cases, RIJCOSX [29] and RIJONX [30] approximations for 4-electron integrals had been applied with the corresponding auxiliary basis sets generated automatically [31]. Natural bond orbital (NBO) [32] analysis was carried out in JANPA program [33]. In attempt to characterize intramolecular interactions, the calculations within the framework of the quantum theory of atoms in molecules (QTAIM) [34] and non-covalent interactions (NCI) [35] analyses were performed in the Multiwfn program [36]. Electrostatic potential map (ESP) was generated using the ORCA package.
For docking studies, a crystal structure of the AMPA receptor (namely, rat homotetrameric GluA2 subtype AMPA receptor) was taken from Protein Data Bank (PDB code 5L1F) [12, 37].
Docking of molecule 1 in the Autodock Vina [38] program was performed with rigid receptor geometry and flexible ligand. Initial ligand geometry was taken from the gas phase B3LYP/6-31G* level geometry optimizations. The box of size 30 × 30 × 30 Å was centered near the crystallographic position of the ligand amide nitrogen located on cycle C. Exhaustiveness parameter was set to 16 to achieve better conformational sampling. One hundred runs each resulting in 10 best poses were performed independently.
Other two docking programs (FRED [39,40,41] and Rosetta [42]) require a ligand conformer library for conformational sampling. Two different libraries were utilized for molecule 1. The first one (Lib1) consisted of the equilibrium geometries of molecule 1 obtained in the gas phase B3LYP/6-31G* level calculations with the degenerated conformers removed. The second one (Lib2) was composed of the non-equilibrium geometries of molecule 1 resulted from the 8 relaxed PES scans with respect to cycle B conducted for each possible minimum energy orientations of cycles A and D. As for molecules 2–6, conformer libraries were composed of their equilibrium geometries similar to Lib1 for molecule 1.
Molecular docking using the FRED program, which is a part of the OpenEye package, was carried out both for Lib1 and Lib2 conformer libraries constructed for molecule 1. Search space was derived from the structure of the protein-ligand complex using Pdb2Receptor utility from the OpenEye package [41]. Docking resolution parameter was switched to “high,” which sets rotational and translational step sizes to 1 Å. Top 100 poses were obtained for both types of conformer libraries.
Molecular docking using the Rosetta program was performed with full ligand and receptor flexibility. It required the receptor structure to be relaxed with regard to the Rosetta energy function prior to docking to avoid inaccuracies of binding mode prediction related to unfavorable protein sidechain conformations in the X-ray structure [42]. The relaxation was conducted using Relax application from the Rosetta package [43]. Sidechain flexibility was allowed for all residues while the backbone was kept rigid. Total of 900 structures were generated, and the best one was chosen according to the Rosetta “total score” function.
The docking itself was performed using the Rosetta Ligand Dock application [42] according to the recommendations given by the program developers [44]. Sidechain conformational sampling parameters were set as recommended. Backbone minimization procedure was enabled with no harmonic restraints applied. The backbone flexibility and arbitrary sidechain rotations were allowed for residues near the ligand position (default cutoffs are 6 Å and 7 Å, respectively) [45]. The search space covered the crystallographic position of the ligand with allowed arbitrary translations up to 5 Å. “soft” repulsive and “old” electrostatic potentials were applied for pose generation step as recommended. Restrained minimization of ligand geometry was enabled for molecules 2–6 as well as for molecule 1 with Lib1 conformer library. Five thousand poses were generated in total independently. As for the docking of molecule 1 with Lib2 conformer library, ligand geometry was not optimized and the number of poses to generate was increased to 20,000 to achieve more comprehensive conformational sampling. As recommended, top 5% of poses were selected based on “total score” and the best part of the obtained poses were chosen according to the “interface delta” score which represents the change of the “total score” upon ligand binding [44]. The visualizations were obtained using Visual Molecular Dynamics (VMD) program of version 1.9.4a8 [46].
Results and discussion
Equilibrium geometries and internal interactions
The search for the reasons underlying the activity cliffs reported for the compounds 1–6 [8] was started with the investigation of the equilibrium geometries of these molecules.
To clarify the description of the conformational space of molecules 1–6, we introduced three dihedral angles: D1, D2, and D3 (Fig. 2). Each conformer in this paper will be described by the code constructed out of three numbers related to the D1, D2, and D3 dihedral angles and indicating the quadrant in which their values lie (for example, a conformer with code 111 has D1, D2, and D3 lying in the 1st quadrant, for more information, see Fig. S1).
The quantum chemical study of molecules 1–6 had started with geometry optimizations in the gas phase at the B3LYP/6-31G* level. The results of the above calculations are presented in Figs. 3 and 4 (left). One may observe that the values of the dihedral angle D2 for molecules 1, 5, and 6 in equilibrium geometries lie close to 0° or 180° reflecting the increased planarity of the B-C moiety. It should also be noted that some conformers of these molecules result in structural degeneration with the dihedral angle D2, changing its quarter passing through the 0° or 180° marks during geometry optimization.
(Left) Conformer energies for molecules 1–6 in the gas phase obtained at the B3LYP/6-31G* level. Degenerated conformers are not shown. Drop lines are drawn only for molecule 2 since it is the only one having the full set of conformers (from 111 to 442). (Right) Conformer energies for molecule 1 in solvents with different ε (B3LYP/6-31G* calculations in SMD solvent model). One may notice the stabilization of certain conformations of molecules 1–4 in the gas phase caused by AB-attraction which is not observed in the PCM calculations. The numeric values of energy differences of conformations can be found in Supporting Materials (Tables S1, S2)
The geometry optimizations were followed by the investigation of the intramolecular interactions in equilibrium geometries. The analysis of non-covalent interactions (NCI) demonstrated that each of the lowest energy conformers of molecules 1–4 has the van der Waals contact between the cycle A ortho-cyano group and the cycle B aromatic hydrogen (Fig. 5b). The interaction behind this phenomenon was called AB-attraction. The NBO and QTAIM analyses along with the electrostatic potential map (ESP) investigation suggested the electrostatic nature of this interaction (Fig. 5a, for results of the NBO and QTAIM studies, see Table S3A). NCI analysis also found a weaker interaction of similar nature between cycles A and D hereafter referred to as AD-attraction (Fig. 5c, for analysis, see Table S3B). AB- and AD-attractions along with repulsion between the cycle C carbonyl oxygen and the cycle A ortho-cyano group define the conformational energy landscapes of molecules 1–4. These interactions are absent in the case of molecules 5 and 6 lacking the cyano group in the ortho-position of cycle A. It explains the low values of the relative conformational energies observed for these molecules in the gas phase B3LYP/6-31G* level calculations (Fig. 4 (left)).
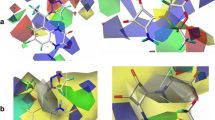
a The ESP surface of the molecule 1 (conformer 141), contour value −0.5 a.u.; blue and red colors represent negative and positive ESP values, respectively. b, c The NCI surfaces illustrating AB- (b) and AD- (c) attractions in molecule 1, conformers 141 and 241, respectively. The NCI isosurfaces were built with reduced density gradient (RDG) of 0.7, and the color palette was scaled from − 0.035 a.u. < sign(λ2)ρ < 0.02 a.u. so that the red and blue regions represent repulsive and attractive interactions, respectively, and the green ones represent van der Waals contacts
As far as the nature of the dominant internal interactions for molecules 1–6 in the gas phase was identified, their conformational space in the non-vacuum medium should be considered next. The electrostatic nature of the major intramolecular interactions implies the decrease of the relative conformational energies in the environment with dielectric constant value ε > 1. The approving calculations at the B3LYP/6-31G* level with the polarizable continuum model (PCM, namely, SMD) applied were performed only for molecule 1. To cover the relevant range of the ε values, three solvents had been chosen: chloroform (ε = 4.9), acetone (ε = 20.7), and water (ε = 80.4). The results of the PCM model calculations for molecule 1 indeed demonstrate a sharp decrease in energy differences of conformations even for small ε values as compared to ε = 1 (Fig. 4 (right)).
It should be noted that D1, D2, and D3 values in equilibrium geometries remained nearly the same for different solvents (mean difference below 7° for each angle and solvent). To conclude, the results of PCM calculations suggest that in the non-vacuum environment, the occupancies of different conformational states of molecule 1 are practically equal. The above conclusion may be extrapolated onto molecules 2–6 as well due to the similar nature of the internal interactions.
The results of the performed quantum chemical calculations suggest that the internal interactions have little to no effect in explaining the biological activity of molecules 1–6 due to partial screening in the non-vacuum environment. However, the geometric features such as different degree of planarity of the B-C moiety might still underlie the reported activity cliffs [8].
Internal rotation
At the next step of the search for an explanation of the reported activity cliffs, the conformational flexibility of molecules 1–6 was considered. The rotational potential energy surfaces (PESs) of their aromatic cycles were investigated by performing the set of relaxed PES scans with respect to the dihedral angles D1, D2, and D3.
Cycle D having a similar structure for all the considered molecules was investigated only for compound 1. Rotation profiles of cycle A were calculated for molecules 1, 5, and 6 representing three different positions of the cyano group. Cycle B scans were performed for each molecule. In the case of compound 1, PES scans were also conducted for all possible minimum energy orientations of cycles A and B.
As expected, the rotational profiles of cycles A and B having two ortho-substituents exhibit four energy maxima of moderate height at 0°, 90°, 180°, and 270° (Fig. 6a). The 0° and 180° maxima are caused by the steric repulsion of the ortho-substituents while the maxima at 90° and 270° result from the loss of conjugation between the two aromatic π-systems.
a–c Rotation profiles (normalized to their global minima) of cycle A for molecules 1, 5, and 6 and cycle B for molecules 2–4 (a), cycle B for molecule 1 (with and without AB-attraction, starting conformer is specified in parentheses) and molecules 5–6 (b), and cycle D for molecule 1 (c). d Cycle B rotation profiles for molecule 1 with all eight possible orientations of cycles A and D (normalized to the minimum energy conformer)
The similar picture is observed for cycle D (Fig. 6c) though the energy maxima at 0° and 180° are markedly higher than those at 90° and 270°. It may reflect the lower degree of π-conjugation between the aromatic systems of cycles C and D (for an approximate analysis, see Table S4, Fig. S2).
In contrast, the lack of one of ortho-substituents for molecules 1, 5, and 6 possessing 2-pyridyl moiety as cycle B leads to elimination of the potential energy maxima near 0° and 180° from the cycle B rotation profiles (Fig. 6b, d). Consequently, the areas of energetically favorable D2 values shift towards 0° and 180° for 2-pyridyl moiety as compared to 3-pyridyl, 4-pyridyl, and phenyl (compounds 2–4) ones. In addition, this feature explains the cases of structural degeneration observed during geometry optimization. As it was shown by the NBO analysis, no significant covalent interaction takes place between the cycle B nitrogen and the cycle C hydrogens when they approach each other during the rotation of cycle B (Fig. S3). Thus, a reasonable hypothesis is the increased planarity of the B-C moiety underlies the higher potency of perampanel as compared to compounds 2–4. It should be noted that in the crystallographic ligand conformation, the value of D2 is 130° which is not favorable for molecule 1 (1.5 kcal/mol from the nearest gas phase minimum).
In order to assess the applicability of the B3LYP/6-31G* method to the PES investigation in the gas phase, we recalculated one of the cycle B rotation profiles of molecule 1 at the B3LYP/6-311G** and PBE0/cc-pVTZ levels. These calculations completely reproduced the B3LYP/6-31G* results (Fig. S4).
To conclude, investigation of the PESs of molecules 1–6 demonstrated moderate conformational flexibility of their aromatic cycles. It also revealed higher planarity of the B-C moiety in molecules 1, 5, and 6 as compared to molecules 2–4. We hypothesize this feature may underlie the sharp activity decrease reported for the latter ones [8].
Docking study
The results of the quantum chemical investigation of molecules 1–6 demonstrated that in non-vacuum environment, the relative energies of their conformers are equal within 1 kcal/mol. Therefore, the protein-ligand interactions rather than the features of the ligand conformational space contribute more to the structure-activity relationships of the studied molecules. However, the difference in conformational behavior of cycle B observed in the gas phase B3LYP/6-31G* level calculations may underlie the lower antagonist activity of molecules 2–4 as compared to compound 1. It should also be noted that molecules 1–6 are composed of three aromatic cycles of quite similar shape and contain no functional groups for which strong specific and directional interactions may be expected. This fact complicates the identification of their unique modes of binding to the target protein.
Recently, Yelshanskaya et al. [12] reported the structure of the full-length rat homotetrameric GluA2 subtype AMPA receptor in complex with molecule 1 determined by the X-ray crystallography. The publication of this structure allowed to identify the approximate position of the ligand in the binding pocket and therefore opened the possibility for computational investigation of the protein-ligand interactions for molecule 1. However, the reported structure is characterized by modest resolution (4 Å) and high B-factor values in the ligand binding region (~ 200 Å2) which may reflect either the uncertainty of the atomic positions caused by thermal motion or the superposition of a set of conformations which restrict its application in the structure-based ligand design.
As it was observed in the X-ray structure [12], molecule 1 binds to the target protein in a flexible linker region between its ligand-binding domain (LBD) and transmembrane domain (TMD). The structures of the two other AMPA receptor noncompetitive antagonists bound at the same site (CP-465022 and GYKI-53655, Fig. 7) with the target protein [12] indicated the noticeable variations of the binding pocket shape depending upon the ligand structure. These variations involve the position change for both sidechain and backbone atoms of the neighboring amino acids.
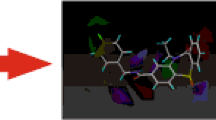
X-ray structures of the GluA2 subtype AMPA receptor in complex with its noncompetitive antagonists: perampanel (a), GYKI-53655 (b), and CP-465022 (c). Ligand structures are transparent for a more clear illustration of the binding pocket flexibility. Dashed lines represent plausible hydrogen bonds. Traditional residue numbering for AMPA receptors (i.e., without the signal peptide) is used. PDB codes: 5L1F, 5L1H, and 5L1E for a, b, and c, respectively
In order to improve our understanding of the binding of molecule 1 to the AMPA receptor, the comprehensive docking study was performed. At the first step, we attempted to reproduce the binding mode of molecule 1 observed in the X-ray structure by means of rigid molecular docking using the FRED and Autodock Vina programs. The latter one was used by Yelshanskaya et al. [12] for reportedly successful verification of the crystallographically identified ligand position.
In our Autodock Vina–based study, all 100 runs resulted in the same best-scored pose markedly different from the one observed in the X-ray structure. The crystallographic pose was scored only for the 4th or 3rd one (in 5 of 100 runs) of 10 best poses. In the FRED-based study, for both Lib1 and Lib2 conformer libraries, the crystallographic ligand position was scored only at no earlier than on the 63rd position out of 100.
Since neither the FRED nor Autodock Vina programs reproduced the crystallographically identified ligand orientation, we attempted to suggest a better alternative for the computed binding mode of molecule 1 by performing the robust molecular docking study. As it was already mentioned, the crystal structures of the GluA2 subtype AMPA receptor in complex with different noncompetitive antagonists (Fig. 7) demonstrated high flexibility of the residues composing the binding site. Therefore, the reliable docking study should be conducted with regard to the conformational flexibility of amino acids surrounding the ligand, which is naturally taken into account by the Rosetta software. The first docking study was aimed at finding a new binding mode of molecule 1 and utilized Lib1 conformer library. Then, the second docking study with Lib2 library was carried out to investigate, which of the cycle B orientations are preferred for binding to the receptor. It is important to note that the ligand conformational energy distribution does not affect the docking results in the Rosetta program.
Both docking studies suggested nearly identical binding modes for molecule 1 (Fig. 8b, c). Moreover, the binding modes are markedly different from the one found in the X-ray structure. In the suggested poses, cycle B of molecule 1 was placed near the apex of the M3 transmembrane helix which is thought to be responsible for the ion channel gating [1]. This event is accompanied by the conformational change in the Leu620 and Leu624 residues belonging to the M3 helix where the former one is a part of the highly conservative SYTANLAAF motif. In the Lib2-based study, cycle B nitrogen was found to form a hydrogen bond with Ser615 from the SYTANLAAF motif of the adjacent subunit. This interaction may be important for the stabilization of the pyridine nitrogen in a hydrophobic environment. Other aromatic cycles of the ligand are also placed into the hydrophobic areas in both best-scored poses. Cycle A was found to form a π-stacking interaction with Phe623 residue from the SYTANLAAF motif. Its cyano group was exposed to the solvent and did not form a hydrogen bond with Ser516, which was observed in the X-ray structure. Cycle D was placed near the pre-M1 helix in both docking studies yet in a different orientation.
Binding modes of molecule 1 to the GluA2 subtype AMPA receptor found in the X-ray structure (a), Lib1-based docking study (b), and Lib2-based docking study with a deep variation of cycle B orientation (c). Traditional residue numbering for AMPA receptors (i.e., without the signal peptide) is used. Dashed lines represent plausible hydrogen bonds
As for the cycle B orientations favorable for binding to the target protein, the results of the Lib2-based study suggested that conformers with nearly planar B-C moiety geometry are preferred regardless the ligand orientation in the binding pocket (Fig. S5). In the best-scored poses of both docking runs, values of the D2 dihedral angle lay within 10° from 0 to 180° which is only energetically favorable if cycle B is represented by 2-pyridyl moiety as it was shown by the quantum chemical calculations.
To complete the description of the docking results, it should be pointed out that in the both docking studies, high variability of the ligand position and orientation was observed among the 10 best-scored poses. It may reflect the uncertainty of the ligand binding mode caused by the similarity of its aromatic cycles. It is also consistent with high B-factor values reported for the X-ray structure. The conformational flexibility of the neighboring amino acid sidechains was also observed but it never led to a considerable change of the binding pocket shape (except those of Leu620 and Leu624). The backbone flexibility was found to be insignificant.
The binding modes obtained using the Rosetta software were evaluated by the rigid docking in the FRED and Autodock Vina programs. During this process, molecule 1 was docked to the receptor structure taken from the Rosetta results. Both the FRED and Autodock Vina programs produced best-scored poses closely resembling the ones obtained by the Rosetta software with a slightly different orientation of cycle D.
To further evaluate the obtained poses, we employed the mutagenesis data reported by Yelshanskaya et al. [12] along with the 2Fo-Fc electron density deposited to the PDB [37]. The results of the alanine scan mutagenesis study [12] are in agreement with the binding mode proposed here. The residues reported to be insignificant for ligand affinity (i.e., substitution of which with alanine leads to a less than twofold IC50 increase: Ser510, Phe515, Leu518, Asp519, Tyr523, and Ser788) do not have any contacts with molecule 1 in both of the suggested binding modes. The influence of Phe623, Pro520, and Phe517 onto the perampanel binding (202.6-, 14.5-, and 2.4-fold IC50 increase, respectively) can be explained by more or less tight hydrophobic contacts with ligand molecule present in both X-ray structure and docking results.
However, the interpretation of the Ser615, Asn791, and Ser516 influence onto the ligand IC50 is different between the crystallographic pose and docking results. First, the influence of the SYTANLAAF member Ser615 (14.0-fold IC50 increase) finds better explanation in the pose obtained in Lib2-based docking study, where this residue forms a hydrogen bond with the ligand cycle B nitrogen, which is implausible in the crystal structure (N…O distances are 3.4 Å and 5.4 Å, respectively). Second, a steric contact between Asn791 (7.2-fold IC50 increase) and the ligand molecule is found to be closer in the crystallographic pose. However, the hydrogen bond with the cycle B nitrogen suggested by Yelshanskaya et al. [12] is doubtful (N…NH2 distance is 4.8 Å). Third, the influence of Ser516 (4.5-fold IC50 increase), which is explained by a hydrogen bond formation with ligand cyano group in the crystal structure (N…O distance is 2.9 Å), finds alternative interpretation in the docking results. Here, this influence is explained by a structural role of this residue arising from the hydrogen bond formation with carbonyl oxygen of an upstream pre-M1 amino acid Gly513 (O…O distance is 2.5 Å, Fig. 8). In that case, S516A mutation would lead to a looser pre-M1 segment packing, therefore expanding the binding site and decreasing the perampanel affinity. Unfortunately, there are no mutagenesis data regarding Leu624 influence onto the ligand binding while the Leu620 mutation to alanine was reported to result into a nonfunctional receptor [12].
Regarding the 2Fo-Fc electron density available in the PDB [37], the crystallographically identified binding mode lays in a better accordance with experimental data, as compared to the docking results (Fig. 9). At the 1σ level (left), cycles C and B of the ligand along with major parts of cycles A and D are covered by electron density in the crystallographic pose. As for the binding mode suggested by docking, the whole cycle D and a major part of cycle A are covered by the electron density while cycle B and nearly a half of cycle C are not. However, 2Fo-Fc electron density having a resolution of 4.0 Å fails to cover almost all amino acid sidechains surrounding the ligand except those belonging to the pre-M1 helix. At the 2σ level (right), only cycles C and B are covered for the crystallographic ligand position. As for the docking results, electron density covers about a half of cycle D, the carbonyl oxygen of cycle C, and the cyano group of cycle A. Therefore, the experimental 2Fo-Fc density demonstrates poorer coverage of the ligand position resulted from the docking study as compared to the crystallographic pose. However, this may result from the thermal motion of cycles B and C and, most probably, from the flexibility of cycle B orientation. Crystallographic experiments including derivatives of molecule 1 with cyano group substituted by Br or I atoms could be useful for understanding the exact ligand position in the binding pocket.
Binding modes of molecule 1 found in the X-ray structure (blue) and resulted from the docking study (orange) superposed with 2Fo-Fc electron density available in PDB [37] (transparent, blue). Contour values 1.0σ (left) and 2.0σ (right). Amino acid sidechains from the docking results are not shown
To sum up, the above results allowed us to suggest an alternative binding mode for molecule 1 and to determine which of the cycle B orientations are favorable for binding to the target protein. These results also suggested the conformational changes at the apex of the M3 transmembrane helix and the formation of the intersubunit hydrogen bond with Ser615, which may be important for the antagonist activity. Given this knowledge, we can now investigate the binding modes of the less active compounds 2–6 and use the obtained results to explain the reported activity cliffs [8].
The binding modes of molecules 2–6 were determined by means of the fully flexible molecular docking performed with the Rosetta software. The best-scored poses determined for molecules 2–6 were found to be markedly different from that of molecule 1. According to the docking results, Leu620 and Leu624 conformational transitions leading to the formation of the hydrophobic cavity near the SYTANLAAF motif located at the apex of the M3 helix occur only in the case of molecules 1 and 2. In the best-scored poses of other molecules, this cavity is not present. Here, only two hydrophobic cavities are present: one located near the pre-M1 helix and another located between the Phe623 sidechain and Pro520.
In molecules with isomeric position of cycle B nitrogen, location of pyridine moiety is strongly affected by the availability of hydrogen bond donors needed to solvate the pyridine nitrogen in a hydrophobic environment. For the 3-pyridyl isomer (molecule 2) the intersubunit hydrogen bond with Ser615 cannot be formed and the solvation is achieved by the interaction with pre-M1 member Ser516 in the best-scored pose. This apparently decreases the stability of the complex since the cavity near the pre-M1 helix remains unoccupied and the hydrophobic pocket near the SYTANLAAF motif is filled with cycle D having inappropriate orientation (according to results of the Lib2-based docking). As for the 4-pyridyl analog (molecule 3), neither Ser615 nor Ser516 can provide a hydrogen bond necessary for the pyridine nitrogen solvation. In the best-scored pose of this molecule, cycle B resides under the Phe623 sidechain with pyridine nitrogen forming a hydrogen bond with Ser785 from the water-accessible linker segment above the M4 helix of the adjacent subunit. Cycle A occupies the hydrophobic pocket between the pre-M1 and M3 helices with no neighboring hydrogen bond donor to solvate its cyano group. Further decrease in stability is caused by the cycle D placement into the solvent-accessible linker region above the M4 helix.
As for the molecule 4, where cycle B is represented by a phenyl moiety, cycles B and D are held in the binding pocket by nondirectional hydrophobic interactions. The lack of planarity in the B-C moiety probably prevents cycle B from entering the cavity near the SYTANLAAF motif. Cycle A cyano group forms a hydrogen bond with Ser516, and the cavity near the pre-M1 helix remains unoccupied.
In molecules with isomeric cyano group position, the major determinant of the binding mode is the steric compatibility of cycle A to the available hydrophobic pockets. Thus, the most appropriate hydrophobic pocket for 3-cyanophenyl moiety of molecule 5 is located near the pre-M1 helix, as observed in the best-scored pose. This position of cycle A is however incompatible with cycle B occupation of the neighboring hydrophobic pockets, and this moiety stays in the water-exposed area above the M4 helix. Contraversely, the hydrophobic pocket near the pre-M1 helix is incapable of accommodating the 4-cyanophenyl cycle of molecule 6. In the best-scored pose, this moiety resides under the Phe623 sidechain with its cyano group placed into the water-accessible area above the M4 helix of the adjacent subunit. The cyano group forms a hydrogen bond with M3 residue Asn619. In that pose, cycle D stays in the solvent-exposed area above the M4 helix and cycle B resides in the pocket near the pre-M1 segment with no hydrogen bond solvating its pyridine nitrogen.
Similar to molecule 1, the ligand position of molecules 2–6 in the binding pocket varied significantly among the 10 best-scored poses. The shape of the binding pocket itself was also subjected to slight alterations, though its overall geometry remained nearly the same except for the Leu620 and Leu624 conformational changes. Along with these residues, conformational alterations were also seen for Ser516, Phe517, and Asp519 from the pre-M1 helix, Phe623 from the SYTANLAAF motif, and Asn791 from the apex of the M4 transmembrane helix.
To conclude, the results of the performed docking studies allowed us to propose an alternative binding mode for molecule 1. According to the suggested mode, the binding of perampanel is accompanied by the Leu620 and Leu624 conformational changes and the intersubunit hydrogen bond formation, which allows the 2-pyridyl moiety of molecule 1 to occupy the hydrophobic cavity near the SYTANLAAF motif. The results of the docking study also suggested that cycle B orientation preferable for binding to the target protein can only be achieved if this cycle is represented by 2-pyridyl moiety. Docking of molecules 2–6 allowed us to suggest the reasons underlying the reported activity cliffs. On the one hand, modifications of cycle B make this moiety incapable of occupying the hydrophobic cavity near the SYTANLAAF motif. On the other hand, cycle A modifications lead to a significant changes in the binding modes governed mainly by the availability of hydrophobic pockets capable of accommodating this moiety.
Conclusions
According to the results of quantum chemical calculations for a series of perampanel analogs, the intramolecular interactions are too weak to solely explain the conformational equilibrium in the protein medium (ε = 4–5). Although the gas phase calculations demonstrate significant stabilization of certain conformations by AB-attraction, this effect can be generally neglected for the considered environment due to the partial screening in a non-vacuum medium. However, the rotation profiles around the bond connecting cycles B and C demonstrate the increased planarity of the B-C moiety in the case of the 2-pyridyl substituent as compared to 3-pyridyl (2), 4-pyridyl (3), and phenyl (4), which, we believe, contributes to a better accommodation of the ligand in the binding site. The results of the robust docking study shed the light on the origin of the activity cliffs observed for a set of molecules (1–6). The analysis of the alternative binding mode of perampanel proposed here reveals several observations, which are helpful in explaining the reported activity cliffs. First, the planar geometry of the B-C moiety was suggested to be preferred for binding to the AMPA receptors. Second, an intersubunit hydrogen bond was proposed, which is necessary for the stabilization of a pyridine nitrogen in a hydrophobic environment. This hydrogen bond can be formed only for 2-pyridyl moiety while the 3-pyridyl and 4-pyridyl perampanel isomers should pay the desolvation penalty upon binding. Finally, the conformational transitions of two leucine residues located near the presumed ion channel gating mechanism were suggested on the basis of the results of the docking study. The affinity decrease upon the change of the cyano group position was suggested to be caused by the alterations of the geometry of the corresponding aromatic cycle. For 4-cyano isomer (6), this cycle cannot be properly placed into the binding site due to the steric ligand-receptor non-complementarity, while for 3-cyano isomer (5), this cycle is tightly buried in the hydrophobic pocket near the pre-M1 helix. In both cases, the 2-pyridyl moiety cannot reach the gating region. This rationale may be useful for the design of novel noncompetitive AMPA receptor antagonists. The performed study yielded the alternative hypothesis of the perampanel binding mode which is consistent with computational results described above and demonstrates the necessity of a more thorough experimental investigation of the perampanel binding to AMPA receptors.
References
Traynelis SF, Wollmuth LP, McBain CJ, Menniti FS, Vance KM, Ogden KK, Hansen KB, Yuan H, Myers SJ, Dingledine R (2010) Glutamate receptor ion channels: structure, regulation, and function. Pharmacol Rev 62:405–496
Swanson GT (2009) Targeting AMPA and kainate receptors in neurological disease: therapies on the horizon? Neuropsychopharmacology 34:249–250
Aarsland D, Ballard C, Walker Z, Bostrom F, Alves G, Kossakowski K, Leroi I, Pozo-Rodriguez F, Minthon L, Londos E (2009) Memantine in patients with Parkinson’s disease dementia or dementia with Lewy bodies: a double-blind, placebo-controlled, multicentre trial. Lancet Neurol 8:613–618
Petrenko AB, Yamakura T, Baba H, Shimoji K (2003) The role of N-methyl-D-aspartate (NMDA) receptors in pain: a review. Anesth Analg 97:1108–1116
Dong XX, Wang Y, Qin ZH (2009) Molecular mechanisms of excitotoxicity and their relevance to pathogenesis of neurodegenerative diseases. Acta Pharmacol Sin 30:379–387
Burnashev N, Monyer H, Seeburg PH, Sakmann B (1992) Divalent ion permeability of AMPA receptor channels is dominated by the edited form of a single subunit. Neuron 8:189–198
Meldrum BS (2000) Glutamate as a neurotransmitter in the brain: review of physiology and pathology. J Nutr 130:1007S–1015S
Hibi S, Ueno K, Nagato S, Kawano K, Ito K, Norimine Y, Takenaka O, Hanada T, Yonaga M (2012) Discovery of 2-(2-oxo-1-phenyl-5-pyridin-2-yl-1,2-dihydropyridin-3-yl)benzonitrile (perampanel): a novel, noncompetitive α-amino-3-hydroxy-5-methyl-4-isoxazolepropanoic acid (AMPA) receptor antagonist. J Med Chem 55:10584–10600
Frampton JE (2015) Perampanel: a review in drug-resistant epilepsy. Drugs 75:1657–1668
Silla JM, Silva DR, Freitas MP (2017) Theoretical study on the conformational bioeffect of the fluorination of acetylcholine. Mol Inform 36:1700084
Horvath D, Marcou G, Varnek A (2018) Monitoring of the conformational space of dipeptides by generative topographic mapping. Mol Inform 36:1700115
Yelshanskaya MV, Singh AK, Sampson JM, Narangoda C, Kurnikova M, Sobolevsky AI (2016) Structural bases of noncompetitive inhibition of AMPA-subtype ionotropic glutamate receptors by antiepileptic drugs. Neuron 91:1305–1315
Twomey EC, Yelshanskaya MV, Grassucci RA, Frank J, Sobolevsky AI (2017) Channel opening and gating mechanism in AMPA-subtype glutamate receptors. Nature 549:60–65
Barygin OI, Grishin EV, Tikhonov DB (2011) Argiotoxin in the closed AMPA receptor channel: experimental and modeling study. Biochemistry 50:8213–8220
Elhallaoui M, Laguerre M, Carpy A, Ouazzani FC (2002) Molecular modeling of noncompetitive antagonists of the NMDA receptor: proposal of a pharmacophore and a description of the interaction mode. J Mol Model 8:65–72
Karlov DS, Lavrov MI, Palyulin VA (2016) Pharmacophore analysis of positive allosteric modulators of AMPA receptors. Russ Chem Bull 65:581–587
Karlov DS, Lavrov MI, Palyulin VA, Zefirov NS (2018) MM-GBSA and MM-PBSA performance in activity evaluation of AMPA receptor positive allosteric modulators. J Biomol Struct Dyn 36:2508–2516
Neese F (2012) The ORCA program system. Wiley Interdiscip Rev Comput Mol Sci 2:73–78
Neese F (2018) Software update: the ORCA program system, version 4.0. Wiley Interdiscip Rev Comput Mol Sci 8:e1327
Halgren TA (1999) MMFF VI. MMFF94s option for energy minimization studies. J Comput Chem 20:720–729
Hanwell MD, Curtis DE, Lonie DC, Vandermeersch T, Zurek E, Hutchison GR (2012) Avogadro: an advanced semantic chemical editor, visualization, and analysis platform. J Cheminform 4:17
Becke AD (1996) Density-functional thermochemistry. IV. A new dynamical correlation functional and implications for exact-exchange mixing. J Chem Phys 104:1040–1046
Hertwig RH, Koch W (1997) On the parameterization of the local correlation functional. What is Becke-3-LYP? Chem Phys Lett 268:345–351
Hehre WJ, Ditchfield R, Pople JA (1972) Self-consistent molecular orbital methods. 12. Further extensions of Gaussian-type basis sets for use in molecular-orbital studies of organic-molecules. J Chem Phys 56:2257–2261
Krishnan R, Binkley JS, Seeger R, Pople JA (1980) Self-consistent molecular orbital methods. XX. A basis set for correlated wave functions. J Chem Phys 72:650–654
Perdew JP, Ernzerhof M, Burke K (1996) Rationale for mixing exact exchange with density functional approximations. J Chem Phys 105:9982–9985
Dunning TH (1989) Gaussian basis sets for use in correlated molecular calculations. I. The atoms boron through neon and hydrogen. J Chem Phys 90:1007–1023
Marenich AV, Cramer CJ, Truhlar DG (2009) Universal solvation model based on solute electron density and on a continuum model of the solvent defined by the bulk dielectric constant and atomic surface tensions. J Phys Chem B 113:6378–6396
Neese F, Wennmohs F, Hansen A, Becker U (2009) Efficient, approximate and parallel Hartree-Fock and hybrid DFT calculations. A “chain-of-spheres” algorithm for the Hartree-Fock exchange. Chem Phys 35:98–109
Neese F, Olbrich G (2002) Efficient use of the resolution of the identity approximation in time-dependent density functional calculations with hybrid density functionals. Chem Phys Lett 362:170–178
Stoychev GL, Auer AA, Neese F (2017) Automatic generation of auxiliary basis sets. J Chem Theory Comput 13:554–562
Weinhold F, Landis CR (2001) Natural bond orbitals and extensions of localized bonding concepts. Chem Educ Res Pract Eur 2:91–104
Nikolaienko TY, Bulavin LA, Hovorun DM (2014) JANPA: an open source cross-platform implementation of the Natural Population Analysis on the Java platform. Comput Theor Chem 1050:15–22
Bader RFW (1991) A quantum theory of molecular structure and its applications. Chem Rev 91:893–928
Johnson ER, Keinan S, Mori-Sánchez P, Contreras-García J, Cohen AJ, Yang W (2010) Revealing noncovalent interactions. J Am Chem Soc 132:6498–6506
Lu T, Chen F (2012) Multiwfn: a multifunctional wavefunction analyzer. J Comput Chem 33:580–592
Berman HM, Westbrook J, Feng Z, Gilliland G, Bhat TN, Weissig H, Shindyalov IN, Bourne PE (2000) The Protein Data Bank. Nucleic Acids Res 28:235–242
Trott O, Olson AJ (2010) AutoDock Vina: improving the speed and accuracy of docking with a new scoring function, efficient optimization and multithreading. J Comput Chem 32:455–461
OEDOCKING 3.3.0.3: OpenEye Scientific Software, Santa Fe, NM. http://www.eyesopen.com. Accessed 01.09.2019
McGann M (2011) FRED pose prediction and virtual screening accuracy. J Chem Inf Model 51:578–596
McGann M (2012) FRED and HYBRID docking performance on standardized datasets. J Comput Aided Mol Des 26:897–906
Kaufmann KW, Meiler J (2012) Using RosettaLigand for small molecule docking into comparative models. PLoS One 7:e50769
Nivón LG, Moretti R, Baker D (2013) A Pareto-optimal refinement method for protein design scaffolds. PLoS One 8:e59004
Rosetta Ligand Dock Application. https://www.rosettacommons.org/docs/latest/application_documentation/docking/ligand-dock. Accessed 01.09.2019
Davis IW, Baker D (2009) RosettaLigand docking with full ligand and receptor flexibility. J Mol Biol 385:381–392
Humphrey W, Dalke A, Schulten K (1996) VMD: visual molecular dynamics. J Mol Graph 14:33–38
Acknowledgments
The academic license for OpenEye software was kindly provided by OpenEye Scientific Software Inc. to Dr. Vladimir A. Palyulin laboratory.
Funding
This work was supported by the Russian Science Foundation under Grant No. 18-75-00077.
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
ESM 1
(DOCX 507 kb)
Rights and permissions
About this article
Cite this article
Guseynov, AA.D., Pisarev, S.A., Shulga, D.A. et al. Computational characterization of the glutamate receptor antagonist perampanel and its close analogs: density functional exploration of conformational space and molecular docking study. J Mol Model 25, 312 (2019). https://doi.org/10.1007/s00894-019-4188-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00894-019-4188-z