Abstract
We evaluated fungal and bacterial diversity in an established moss carpet on King George Island, Antarctica, affected by ‘fairy ring’ disease using metabarcoding. A total of 127 fungal and 706 bacterial taxa were assigned. Ascomycota dominated the fungal assemblages, followed by Basidiomycota, Rozellomycota, Chytridiomycota, Mortierellomycota and Monoblepharomycota. The fungal community displayed high indices of diversity, richness and dominance, which increased from healthy through infected to dead moss samples. A range of fungal taxa were more abundant in dead rather than healthy or fairy ring moss samples. Bacterial diversity and richness were greatest in healthy moss and least within the infected fairy ring. The dominant prokaryotic phyla were Actinobacteriota, Proteobacteria, Bacteroidota and Cyanobacteria. Cyanophyceae sp., whilst consistently dominant, were less abundant in fairy ring samples. Our data confirmed the presence and abundance of a range of plant pathogenic fungi, supporting the hypothesis that the disease is linked with multiple fungal taxa. Further studies are required to characterise the interactions between plant pathogenic fungi and their host Antarctic mosses. Monitoring the dynamics of mutualist, phytopathogenic and decomposer microorganisms associated with moss carpets may provide bioindicators of moss health.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Antarctic vegetation is dominated by bryophytes, with 116 species currently recognised representing cosmopolitan, endemic and bipolar taxa (Ochyra et al. 2008; Câmara et al. 2019). Mosses may form extensive carpets in some parts of Antarctica, particularly in the maritime Antarctic, contributing to the greatest development of ‘fellfield’ communities globally and providing habitats and ameliorating Antarctica’s extreme environmental conditions for contained microbial and invertebrate communities (Smith 1984; de Carvalho et al. 2019; Prather et al. 2019). Well established Antarctic moss carpets may act as "sentinels" sensitive to environmental changes, particularly in temperature and hydration, across the Antarctic Peninsula region (Prather et al. 2019). Moss carpet health has been a subject of research attention since the early years of Antarctic terrestrial research (Robinson et al. 2018). One of the most frequently reported concerns relating to moss health is that of attack by initially unidentified organism(s) resulting in the formation of a concentric ring (‘fairy ring’) visible on the surface of the carpet which eventually results in the death of the moss (Wilson 1951; Racovitza 1959; Hawksworth 1973; Longton 1973; Fenton 1983; Ochyra et al. 2008; Tojo et al. 2012; Pawłowska et al. 2017). Most recently, Rosa et al. (2020a) recorded the development of fairy rings on previously unreported moss species from new locations in the western Antarctic Peninsula region, suggesting that the disease is more widespread in maritime Antarctica than previously believed and may be increasing in prevalence.
The majority of studies Antarctic moss fairy rings have considered that fungi are the cause of the disease. However, there is no consensus about which species is/are the phytopathogenic agent(s) causing the disease. Fenton et al. (1983) was the first to propose Coleroa turfosorum, Bryosphaeria megaspora and Epibryon chorisodontii (Ascomycota), recovered from infected mosses on Signy Island, as the causative agent. Tojo et al. (2012) proposed Pythium polare (Oomycota) to be the species affecting Sanionia uncinata on King George Island. Pawłowska et al. (2017) recovered and proposed Psychronectria hyperantarctica as the phytopathogenic fungus causing fairy rings, in line with previous work by Putzke and Pereira (2012). Most recently, Rosa et al. (2020a) reported that fairy rings host multiple fungal taxa, which might therefore act in consortium in causing the disease.
Despite the continent’s typically extreme conditions, Antarctic fungi represent a diverse eukaryote microbial group, including symbionts, decomposers and opportunistic taxa, amongst which are phytopathogenic taxa (Rosa et al. 2019). Globally, approximately 300 species from 80 genera of Ascomycota are known to parasitize mosses or liverworts (Döbbeler 1997). Earlier studies of the cause of fairy ring disease relied on culturing approaches and direct morphological identification. Rosa et al. (2020a) was the first study to use molecular tools to identify fungi potentially involved. DNA metabarcoding using high-throughput sequencing (HTS) is increasingly recognised as an important tool in investigating fungal diversity in various Antarctic ecosystems (Rosa et al. 2020b, c; Rosa et al. 2021). Therefore, in the present study, we evaluated fungal and bacterial diversity associated with different stages of the development of fairy ring disease in well established moss carpets on King George Island, South Shetland Islands, maritime Antarctic.
Methods
Moss carpet sampling and identification
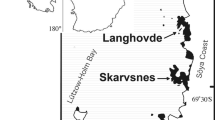
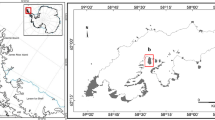
Fungal and bacterial occurrence and diversity were investigated across different fairy ring disease stages in a well-established moss carpet on the Keller Peninsula, King George Island, South Shetland Islands (maritime Antarctica; Fig. 1) during the austral summer of 2019/20. Three moss samples (each approximately 4 cm diameter) from each visible stage of the disease, defined as healthy, fairy ring (infected) and dead (Fig. 2), were obtained. The samples were immediately stored in sterilized whirl pack bags and frozen at −20 °C until further use. The moss carpet was formed by the species Sanionia uncinata (Hedw.) Loeske, with identification confirmed based on macro- and micromorphological characteristics with reference to Ochyra et al. (2008). All moss specimens are deposited in the University of Brasília Herbarium (UB).
Satellite images a, b and c (obtained in Google Earth Pro, 2019) indicating where the moss samples were obtained. a Antarctic continent with the north-west Antarctic Peninsula and South Shetland Islands inside the red rectangle, b Antarctic Peninsula with King George Island inside the red rectangle, c King George Island with the Keller Peninsula inside the red rectangle, d aerial view of the well established moss carpet (total area 530 m2) from which samples were obtained on the Keller Peninsula, close to the Brazilian Antarctic Station Comandante Ferraz (62°5′12.869″ S; 58°23′42.312″ W). Photo L.H. Rosa
DNA extraction, data analyses and fungal and bacterial identification
Three samples of each of healthy, infected and dead mosses were processed separately to recover the total fungal and bacterial DNA. Total DNA was extracted using the QIAGEN DNeasy PowerLyzer PowerSoil Kit, following the manufacturer’s instructions. Extracted DNA was used as template for generating PCR-amplicons. For fungi, the internal transcribed spacer 2 (ITS2) of the nuclear ribosomal DNA was used as a DNA barcode for molecular species identification (Chen et al. 2010; Richardson et al. 2015). PCR-amplicons were generated using the universal primers ITS3 (5′-GCATCGATGAAGAACGCAGC-3′) and ITS4 (5′-TCCTCCGCTTATTGATATGC-3′) for fungi (White et al. 1990). For bacteria, we used the 16S rRNA gene V3-V4 region and primers341F (5′-CCTACGGGNGGCWGCAG-3′) and 805R (5′-GACTACHVGGGTATCTAATCC-3′) that produce an amplicon of ~ 460 bp (Herlemann et al. 2011; Klindworth et al. 2013). These amplicons were subjected to high-throughput sequencing at Macrogen Inc. (South Korea) on an Illumina MiSeq sequencer (3 × 300 bp), using the MiSeq Reagent Kit v3 (600-cycle) following the manufacturer’s protocol.
Raw fastq files were filtered using BBDuk version 38.34 (BBMap—Bushnell B.—sourceforge.net/projects/bbmap/) to remove Illumina adapters, known Illumina artefacts and the PhiX Control v3 Library. Quality read filtering was carried out using Sickle version 1.33-q 30-l 50 (Joshi and Fass 2011), to trim 3′ or 5′ ends with low Phred quality score, and sequences shorter than 50 bp were also discarded. The remaining sequences were imported to QIIME2 version 2019 for bioinformatics analyses (Bolyen et al. 2019). For fungi, the qiime2-dada2 plugin is a complete pipeline that was used for filtering, dereplication, turn paired-end fastq files into merged and remove chimeras (Callahan et al. 2016). Taxonomic assignments were determined for amplicon sequence variants (ASVs) using the qiime2-feature-classifier (Bokulich et al. 2018) classify-sklearn against the UNITE fungal ITS database version 8.2 (Abarenkov et al. 2020) trained with Naive Bayes classifier and a confidence threshold of 98.5%. For bacteria, sequences were quality filtered using “quality-flter q-score-joined” plugin to improve diversity (Bokulich et al. 2013). Sequences were denoised using deblur (Amir et al. 2017) with-p-trim-length parameter of 300 and were taxonomically assigned to sub-operational-taxonomic-units (sOTU) against the Silva 138 Ref NR 99 database pre-trained with Naive Bayes classifier using the “feature-classifier classify-sklearn” plugin.
Many factors, including extraction, PCR and primer bias, can affect the number of reads obtained (Medinger et al. 2010) and thus lead to misinterpretation of absolute abundance (Weber and Pawlowski 2013). However, Giner et al. (2016) concluded that such biases did not affect the proportionality between reads and cell abundance, implying that more reads are linked with higher abundance (Deiner et al. 2017). Therefore, for comparative purposes, we used the number of reads as a proxy for relative abundance. Fungal classification followed Kirk et al. (2011), Tedersoo et al. (2018), MycoBank (http://www.mycobank.org) and the Index Fungorum (http://www.indexfungorum.org).
Diversity, distribution and ecological analysis
To quantify species diversity, richness and dominance, we used the following indices: (i) Fisher’s α, (ii) Margalef’s and (iii) Simpson’s, respectively, to assess alpha diversity. In addition, the Sorensen and Bray–Curtis similarity indices were used to assess beta diversity among the fungal and bacterial assemblages present in the mosses representing the different disease stages. The relative abundance of the OTUs was used to quantify the taxa present in the different disease stages as described by Rosa et al. (2021), where OTUs with relative abundance > 10% were considered dominant, those with relative abundance of 1–10% intermediate and those with < 1% minor (rare) components of the microbial community. All of the results were obtained with 95% confidence and bootstrap values were calculated from 1,000 iterations. Taxon accumulation curves were obtained using the Mao Tao index. All diversity index calculations and t tests were performed using PAST, version 1.90 (Hammer et al. 2001). To prepare Kronar charts, QIIME2 taxonomy classifications and the table of taxa abundance were converted to tsv and biom format, respectively. The table of fungal abundance was converted to tsv by using biom convert and combined with taxonomy classification with a custom script krona_qiime.py (https://github.com/lokeshbio/Amplicon_course/blob/master/krona_qiime.py). The Krona Tools (v. 2.7.1) (Ondov et al. 2011) program, ktImportText.pl, was used to provide interactive visualization of identified fungi species. Venn analysis to compare the fungal diversity obtained from the different sampling locations was carried out using the program available at http://bioinformatics.psb.ugent.be/webtools/Venn/.
Results
Taxonomy and diversity
We obtained 55,312 fungal DNA reads representing 127 OTUs and 69,755 bacterial DNA reads representing 706 OTUs (Suppl. Table 1). The Mao Tao rarefaction curves did not reach a plateau in all samples, indicating that the total number of fungal and bacterial taxa may be greater than that detected (Suppl. Figure 1). The Krona charts (Fig. 3) illustrate the changes in the fungal and bacterial assemblages with progression through the three disease stages. The fungal phylum Ascomycota dominated the assemblages in all samples, followed by Basidiomycota, Rozellomycota, Chytridiomycota, Mortierellomycota and Monoblepharomycota. Chalara sp. 1, Alpinaria sp., Helotiaceae sp. 2, Chaetothyriales sp. 1, Ascomycota sp. 1 (all Ascomycota), Rozellomycota sp. and Fungi sp. generally displayed relative abundance > 10% and were the dominant taxa, followed by 15 taxa characterized as intermediate and 109 as minor (rare) components of the total fungal community. The dominant bacterial phylum was Actinobacteriota, followed by Proteobacteria, Bacteroidota and Cyanobacteria, in rank. The class level taxon Cyanophyceae sp. (Cyanobacteria) represented the dominant prokaryotic taxa. Seventy taxa were characterized as intermediate and 643 as minor components of the bacterial community.
Fungal and bacterial alpha diversity indices across the samples are given in Table 1. The fungal communities displayed high diversity (Fisher α), richness (Margalef) and dominance (Simpson) indices, which increased progressively from the healthy to the dead moss carpet samples. For the bacterial communities the Fisher (diversity) and Margalef (richness) indices were greatest in the the healthy moss, decreasing in the fairy ring infected moss and partially recovering in the dead moss sample. The Simpson (dominance) index did not differ across all samples.
The beta diversity of the fungal and bacterial assemblages varied across the different samples (Suppl. Figure 2). Both presence-absence-based Sorensen and the abundance-related Bray–Curtis similarity indices showed that the fungal assemblages detected in the healthy and fairy ring samples were the most similar, while those of the dead moss were the most different, mainly due the dominance of Basidiomycota taxa. In contrast, the bacterial assemblages detected in the healthy and dead moss carpet samples were the most similar and those present in the fairy ring formed a separate group.
The fungal and bacterial community composition varied through the different stages of the disease (Suppl. Figure 3). For fungi (Suppl. Figure 3a), 42 taxa occurred in all samples, but each disease stage displayed different composition and 20 taxa occurred only in fairy ring and dead samples. For the bacterial communities (Suppl. Figure 3b) the largest proportion of the taxa (254) were present in all samples. Few taxa were in common between healthy/fairy ring, fairy ring/dead and healthy/dead samples.
Comparison of the dominant and intermediate fungal and bacterial communities identified different patterns of composition at the different stages of the infection (Fig. 4; Suppl. Table 2). Amongst the fungi, Chalara sp. 1, Ascomycota sp. 1, Tetracladium sp. 2 and Fungi sp. dominated the healthy moss carpet and decreased in abundance in the fairy ring infected and dead moss. In contrast, Alpinaria sp., Helotiaceae sp. 2, Helotiales sp. 1 and Rozellomycota sp. increased considerably in relative abundance between the healthy and the infected moss samples, but decreased in the dead moss. Finally, Chaetothyriales sp. 1, Serendipita sp., Agaricomycetes sp., Sebacinales sp., Knufia peltigerae, Ascomycota sp. 2, Mortierella fimbricystis, Lamprospora sp., Melanommataceae sp., Pseudogymnoascus sp. 1 and Platygloeaceae sp. were the most abundant fungi in the dead moss but had low relative abundance in the healthy and infected moss.
The relative abundance of the four prokaryote phyla varied across the three infection stages. Actinobacteriota displayed moderate abundance in healthy moss carpet, increasing to dominate the bacterial community in the infected moss and decreasing in the dead moss. In contrast, Proteobacteria, Bacteroidota and Cyanobacteria displayed moderate abundance in healthy moss, which decreased in the infected moss and increased again in dead moss. Among these phyla, Cyanophyceae sp. (Cyanobacteria) was the dominant taxon, although only reaching intermediate abundance in the infected moss (3.74%) and dominance in healthy (12.39%) and dead moss (13.05%) samples. Microbacteriaceae sp. (Actinobacteriota) and Chloroflexi sp. (Chloroflexi) occurred at intermediate abundance in all moss samples and were the most abundant taxa in the infected moss samples and least adundant in dead moss. Cyanophyceae sp. and Haliangium sp. (Myxococcota) were the most abundant taxa detected in dead moss.
Discussion
Fungal diversity
Mosses are the dominant flora in Antartica, providing habitat for multiple microbial and invertebrate taxa and communities (Ochyra et al. 2008; de Carvalho et al. 2019). Endophytic and epiphytic fungi and bacteria are considered to be the dominant microorganisms present in these habitats, known as the bryosphere (Möller and Dreyfuss 1996; Tosi et al. 2002; de Carvalho et al. 2020).
A range of Antarctic moss species have been documented to be vulnerable to infection by ‘fairy ring disease’. In the current study comparing the microbial communities present in healthy, visibly infected (within the ring) and dead moss from the same moss carpet, we detected complex and diverse fungal and bacterial communities using a metabarcoding approach. Overall, the fungal community was richer (3.2 times greater) than that reported recently (Rosa et al. 2020a) in a study using culture methods which detected 40 taxa in eight moss species sampled from different locations in the north-west Antarctic Peninsula region. Among the taxa reported by Rosa et al. (2020a), only representatives of the genera Alpinaria, Helotiales, Cladosporium, Cadophora, Pseudogymnoascus, Glarea, Chalara, Ophiocordycipitaceae, Juncaceicola and the species Mortierella fimbricystis and Gyoerffyella entomobryoides were shared with the current study.
In the current study, relative abundance of the fungal taxa Alpinaria sp., Helotiaceae sp. 2, Coleophoma sp., Helotiales sp. 1, Chytridiomycota sp. 2, Rozellomycota sp. and Fungi sp. increased between healthy and infected moss samples, but was lower in dead moss. The genus Alpinaria (Melanommataceae) includes only a single described species (A. rhododendri) and seems to be common in the subalpine to alpine zone worldwide on twigs or buds of Rhododendron spp. (Ericaceae) (Hashimoto et al. 2017). It has recently been reported on mosses affected by fairy ring disease in maritime Antarctica (Rosa et al. 2020a). The family Melanommataceae includes plant pathogenic species such as Gemmamyces piceae (Jaklitsch and Voglmayr 2017) and, according to the FunGuild database (Nguyen et al. 2016), A. rhododendri is considered a probable plant pathogenic and/or wood saprotrophic species.
The genus Coleophoma includes species reported as plant pathogenic, saprophytic or endophytic on different plant species (Crous and Groenewald 2016). Plant pathogens in the genus include C. fusiformis on leaves of Rhododendron (Sutton 1980), C. eucalypti and C. eucalyptorum on Eucalyptus (Yuan 1996), C. gevuinae on Gevuina (Bianchinotti and Rajchenberg 2004), C. empetri on Vaccinium (Polashock et al. 2009) and C. proteae on Protea caffra (Crous et al. 2012). Rozellomycota species are common in temperate, sub-Arctic and Antarctic environments (Rosa et al. 2020b). According to Grossart et al. (2016), all known Rozellomycota taxa are obligate pathogens of eukaryotes, including amoebae, fungi and algae. However, there are no reports of Rozellomycota acting as plant pathogens. Chytridiomycota, known as chytrids, primarily includes free-living saprophytic taxa present in aquatic and terrestrial environments. However, some species are reconized as plant pathogens, such as Synchytrium endobioticum that causes potato wart disease (van de Vossenberg et al. 2019).
The taxa Chaetothyriales sp. 1, Serendipita sp., Agaricomycetes sp., Sebacinales sp. and Knufia peltigerae are notable here due to their increase in abundance in dead relative to healthy moss carpet. They may, therefore, represent major decomposing taxa in the ecological succession following the death of the moss. The order Chaetothyriales (Ascomycota) includes species with multiple ecological roles, including soil saprophytes, human and animal opportunistic pathogens and plant epi- and/or endophytes (Madrid et al. 2016). In addition, some representatives of Chaetothyriales are known to colonize extreme environments characterized by drought, oligotrophic conditions, extreme temperatures and high UV-radiation exposure (Tsuneda et al. 2011). Some species are known phytopathogens (Gueidan et al. 2014).
Serendipita is a genus with eight known species (Kirk et al. 2011), including S. indica, formerly known as Piriformospora indica (Weiß et al. 2016), an endophytic fungus detected in low-nutrient desert soil in Rajasthan, India (Verma et al. 1998), which acts to increase nutrient uptake and utilization in its host (Yadav et al. 2010; Ngwene et al. 2016). Serendipita has been reported as an endophyte of bryophytes (Varma et al. 2012). Agaricomycetes is a class of Basidiomycota that includes almost 21,000 described species (Kirk et al. 2011) whose members play different ecologicals role such as decomposers, pathogens and mutualists in different environments (Hibbett et al. 2014).
The order Sebacinales (Agaricomycetes, Basidiomycota) includes species recognized to show diverse interactions with plants, which range from mutualistic root endophytes (obligate biotrophs, mycorrhizae) to saprophytes (Weiß et al. 2016). Within the order, members of the family Serendipitaceae have been reported from the Antarctic Peninsula associated with the liverwort Barbilophozia hatcheri and the mosses Chorisodontium aciphyllum and Sanionia uncinata (Zhang et al. 2013).
The genus Knufa comprises black fungi and has six known species (He et al. 2013). Knufia peltigerae is a lichenicolous fungus (Gueidan et al. 2014) which, according to Lawrey and Diederich (2003), represents an important ecological group that forms obligate associations with lichens. The ascomata of K. peltigerae (originally reported as Capronia peltigerae) was first described on thalli of the lichen Peltigera rufescens (Fuckel 1874; Untereiner et al. 2011). Peltigera rufescens is a cosmopolitan lichen that occurs on sub-Antarctic South Georgia, the South Orkney Islands and in various locations along the Antarctic Pensinula (both east and west coasts, including James Ross and Alexander Islands) (Øvstedal and Smith 2001). Possibly analogous to the bleaching effect of fairy rings on mosses, Untereiner et al. (2011) reported the presence of K. peltigerae ascomata on decolourized or moribund P. rufescens thalli. However, it is unclear if K. peltigerae was responsible for the discolouration or represents an opportunistic fungus occurring on aging parts of the lichen thalli. The species has been rarely recorded taxa in Antarctica using culture approaches. de Souza et al. (2021) detected the DNA of K. peltigerae in cotton baits deposited in a lake at Hennequin point, King George Island, close to a moss carpet that was under attack from fairy ring disease.
The taxa Serendipita sp. and Agaricomycetes sp. occurred exclusively and were dominant in the dead moss carpet. Serendipita species include root fungal endophytes and arbuscular mycorrhizal fungi (AMF) known as plant growth promoters (Verma et al. 1998). Bridge and Newsham (2009) reported Serendipita-like Sebacinale fungi in soil at Mars Oasis, Alexander Island, in the southern maritime Antarctic. The class Agaricomycetes includes about 21,000 described mushroom-forming species with ecological roles such as decomposers, pathogens and mutualists in different terrestrial and aquatic environments (Hibbett et al. 2014).
Chalara sp. displayed high dominance in the healthy moss carpet, decreasing in dominance in infected moss. The genus includes 103 widespread species with multiple ecological functions (Kirk et al. 2011). Among Chalara species, C. fraxinea (teleomorph: Hymenoscyphus pseudoalbidus) has been reported as an emerging epidemic plant pathogen that has severely affected ash tree stands in Europe since 1990 (Kowalski 2006; Husson et al. 2011).
Previous studies have concluded that the causative agent of the fairy ring disease in Antarctica is Psychronectria hyperantarctica, identified using classical morphological techniques from its fruiting body (Hawksworth 1973; Pawłowska et al. 2017). However, despite the potentially high taxonomic resolution of the metabarcoding approach, we did not detect sequences of P. hyperantarctica in any samples. Rather, our data indicated the presence of several other recognized plant pathogenic fungi, supporting the suggestion of Rosa et al. (2020a) that the disease may be caused by multiple fungal infections in parallel. The fungal taxa Alpinaria sp., Helotiaceae sp. 2, Coleophoma sp., Helotiales sp., Rozellomycota sp. and Chytridiomycota sp. 2 showed high levels of dominance in infected moss showing fairy ring symtoms, which deserve further detailed taxonomic characterization and assays in vivo using plant models to confirm whether they are able to cause plant disease symptoms. Robinson et al. (2018) demonstrated that moss vegetation in the Windmill Islands, East Antarctica is changing rapidly in response to a drying climate causing declining viability in some species. It is possible that the incidence of fungal attack, evidenced by the fairy ring disease, might be connected to a decrease in moss health resulting from climatic changes in the Antarctic Peninsula region in recent decades, although no studies have specifically addressed this or directly quantified disease incidence.
Bacterial diversity
Few studies have addressed the bacterial communities associated with Antarctic mosses. Park et al. (2013) studied endophytic bacteria associated with healthy material of the moss Sanionia uncinata. To our knowledge, no studies have focused on the bacterial diversity present specifically in mosses affected by the fairy ring disease in Antartica. However, the overall dominance of the phyla Actinobacteriota, Proteobacteria, Bacteroidota and Cyanobacteria documented here is consistent with studies such as those of Holland-Moritz et al. (2018) and Wang et al. (2018), which reported that moss-associated bacterial communities were commonly dominated by Proteobacteria and Bacteroidetes. Using molecular phylogenetic techniques to analyse the bacterial diversity associated with aquatic moss pillars in continental Antarctic lakes, Nakai et al. (2012) reported Proteobacteria, Cyanobacteria and Firmicutes as dominant groups. Park et al. (2013) and Câmara et al. (2021) reported highest relative abundances of sequences representing the phylum Actinobacteria in a transplanted S. uncinata carpet moss in a study also carried out on the Keller Peninsula. Raymond (2016) reported Actinobacteria (genera Conexibacter, Rhodococcus, Marmoricola, Micromonospora and Streptomyces) and Bacteroidetes (genera Flavobacterium, Segetibacterium, Epilithonimonas and Pedobacter) from Bryum argenteum leaves.
The dominance of Microbacteriaceae sp. (Actinobacteriota) in the bacterial assemblage of fairy ring affected moss may be notable. Representatives of Actinobacteria are among the most common prokaryotic organisms in Antarctic terrestrial environments (Pearce et al. 2012). They are also known as prolific producers of bioactive natural products (Liu et al. 2012), including some able to suppress plant diseases (Palaniyandi et al. 2013). Gu et al. (2020) analysed the diversity and composition of fungal and bacterial communities in continuous cropping soil from Chinese chive cultivation, reporting dominance of Actinobacteria in the same samples where potential phytopathogenic fungi were detected. We found a similar high Actinobacteria abundance pattern to that reported by Gu et al. (2020) in fairy ring infected moss.
Cyanophyceae sp. and Haliangium sp. (Myxococcota) were present in all moss samples, but were the most abundant taxa detected in dead moss. Cyanobacteria are the dominant phototrophs in Antarctic terrestrial and freshwater ecosystems (Taton et al. 2003) and represent the greatest accumulation of biomass the benthic habitats of lakes and ponds (Vincent 2000). Pandey et al. (2004) reported several cyanobacterial taxa in association with mosses sampled in the Schirmacher Oasis, continental Antarctica. The primary habitats of myxobacteria such as Haliangium are rich in organic matter (Fudou et al. 2002). These bacteria are strictly aerobic and usually live in the surface layers of the soil, but can also be found in decaying plant material (Reichenbach 1999). The dominance of these two taxa in dead moss may be due the high concentration of minerals released during the organic decomposition of the moss.
Conclusions
Previous culture-based and morphological studies have proposed fungi to be the causative agent of the ‘fairy ring’ disease in different Antarctic mosses species. The use of a metabarcoding approach to assess the diversity of microbial communities associated with different stages of the disease in a carpet of the moss S. antarctica revealed, based on sequence assignment, a greater diversity of associated mutualisticic, phytopathogenic and decomposer fungi than previously recognised, with clear community differences as the disease progressed. In contrast with the fungal community, bacterial diversity decreased in infected relative to healthy moss carpet. We recognise that the metabarcoding approach identifies sequences presence and does not confirm the viability or functional activity of the taxa detected. For these reasons, further traditional isolation studies to recover phytopathogenic fungi are necessary to understand if and how fungi may contribute to the disease and, consequently, moss death. In addition, future long-term monitoring of microbial community dynamics associated with moss carpets may provide a novel bioindicator of moss health in Antarctica.
Data availability statement
All raw sequences have been deposited in the NCBI database under the codes SAMN17612011, SAMN17612012, SAMN17612013, SAMN17612014, SAMN17612015, SAMN17612016 and SAMN17612017.
References
Abarenkov K, Allan Z, Timo P, Raivo P, Filipp I, Nilsson, Henrik R, Urmas K (2020) UNITE QIIME release for eukaryotes. Version 04.02.2020. UNITE Community. doi: https://doi.org/10.15156/BIO/786386
Amir A, McDonald D, Navas-Molina JA, et al. et al (2017) Deblur rapidly resolves single-nucleotide community sequence patterns. mSystems. https://doi.org/10.1128/mSystems.00191-16
Bianchinotti MV, Rajchenberg M (2004) Coleophoma gevuinae comb. nov., a foliar pathogen on Gevuina avellana (Proteaceae). Sydowia –Horn 56:1–5
Bokulich N, Subramanian S, Faith J et al (2013) Quality-filtering vastly improves diversity estimates from Illumina amplicon sequencing. Nat Methods 10:57–59
Bokulich NA, Kaehler BD, Rideout JR, Dillon M, Bolyen E, Knight R, Huttley GA, Gregory Caporaso J (2018) Optimizing taxonomic classification of marker-gene amplicon sequences with QIIME 2’s q2-feature-classifier plugin. Microbiome 6:90
Bolyen E, Rideout JR, Dillon MR et al (2019) Reproducible, interactive, scalable and extensible microbiome data science using QIIME 2. Nat Biotechnol 37:852–857
Bridge PD, Newsham KK (2009) Soil fungal community composition at Mars Oasis, a southern maritime Antarctic site, assessed by PCR amplification and cloning. Fungal Ecol 2:66–74
Callahan BJ, McMurdie PJ, Rosen MJ, Han AW, Johnson AJ, Holmes SP (2016) DADA2: high-resolution sample inference from Illumina amplicon data. Nat Methods 13:581–583
Câmara PEAS, Soares A, Henriques D, Peralta D, Bordin J, Carvalho-Silva M, Stech M (2019) New insights into the species diversity of Bartramia Hedw. (Bryophyta) in Antarctica from a morpho-molecular approach. Antarct Sci 31:208–215
Câmara PEAS, Convey P, Rangel SB et al (2021) The largest moss carpet transplant in Antarctica and its bryosphere cryptic biodiversity. Extremophiles 25:369–384
de Carvalho CR, Santiago IF, Coelho LC, Câmara PEAS, Carvalho-Silva M, Stech M, Rosa CA, Rosa LH et al (2019) Fungi associated with plants and lichens of antarctica. In: Rosa LH (ed) Fungi of Antarctica: diversity, ecology and biotechnological applications. Springer, Berlin, pp 165–199
Chen S, Yao H, Han J et al (2010) Validation of the ITS2 region as a novel DNA barcode for identifying medicinal plant species. PLoS ONE 5:e8613
Crous PW, Groenewald JZ (2016) They seldom occur alone. Fungal Biol 12:1392–1415
Crous PW, Summerell BA, Shivas RG et al (2012) Fungal planet description sheets: 107–127. Persoonia 28:138–182
de Carvalho CR, Ferreira MC, Gonçalves VN, Santos ARO, Carvalho-Silva M, Câmara PEAS, Rosa CA, Rosa LH (2020) Cultivable fungi associated with bryosphere of bipolar mosses Polytrichastrum alpinum and Polytrichum juniperinum in King George Island, South Shetland Islands, maritime Antarctica. Polar Biol 43:545–553
de Souza LMD, Ogaki MB, Câmara PEAS, Pinto OHB, Convey P, Carvalho-Silva M, Rosa CA, Rosa LH (2021) Assessment of fungal diversity present in lakes of Maritime Antarctica using DNA metabarcoding: a temporal microcosm experiment. Extremophiles 25:77–84
Deiner K, Bik HM, Mächler E, Seymour M, Lacoursière-Roussel A, Altermatt F, Creer S, Bista I, Lodge DM, de Vere N, Pfrender ME, Bernatchez L (2017) Environmental DNA metabarcoding: transforming how we survey animal and plant communities. Mol Ecol 26:5872–5895
Döbbeler P (1997) Biodiversity of Bryophilous Ascomycetes. Biodivers Conserv 6:721–738
Fenton JHC (1983) Concentric fungal rings in Antarctic moss communities. Trans Br Mycol Soc 80:415–420
Fuckel L (1874) Symbolae mycologicae. Zweiter Nachtrag. Jahrbu ̈cher desNassauischen Vereins fu ̈ r Naturkunde 27:1–99
Fudou R, Jojima Y, Iizuka T, Yamanaka S (2002) Haliangium ochraceumgen. nov., sp. nov. and Haliangium tepidumsp. nov.: novel moderately halophilic myxobacteria isolated from coastal saline environments. J Gen Appl Microbiol 48:109–115
Giner CR, Forn I, Romac S, Logares R, de Vargas C, Massana R (2016) Environmental sequencing provides reasonable estimates of the relative abundance of specific picoeukaryotes. Appl Environ Microbiol 82:4757–4766
Grossart HP, Wurzbacher C, James TY, Kagami M (2016) Discovery of dark matter fungi in aquatic ecosystems demands a reappraisal of the phylogeny and ecology of zoosporic fungi. Fungal Ecol 19:28–38
Gu Y, Wang Y, Wang P, Wang C, Ma J, Yang X, Ma D, Li M (2020) Study on the diversity of fungal and bacterial communities in continuous cropping fields of Chinese chives (Allium tuberosum). BioMed Res Int. https://doi.org/10.1155/2020/3589758
Gueidan C, Aptroot A, Cáceres MES, Badali H, Stenroos S (2014) A reappraisal of orders and families within the subclass Chaetothyriomycetidae (Eurotiomycetes, Ascomycota). Mycol Prog 13:1027–1039
Hammer Ø, Harper DAT, Ryan PD (2001) PAST: Paleontological statistics software package for education and data analysis. Palaeontol Electr 4:1–9
Hashimoto A, Matsumura M, Hirayama K, Fujimoto R, Tanaka K (2017) Pseudodidymellaceae fam. nov.: phylogenetic affiliations of mycopappus-like genera in Dothideomycetes. Stud Mycol 87:187–206
Hawksworth DL (1973) Thyronectria antarctica (Speg.) Seeler var. hyperantarctica D. Hawksw. var. nov. Br Antarct Sur Bull 32:51–53
He F, Lin B, Sun JZ, Liu XZ (2013) Knufia aspidiotus sp. nov., a new black yeast from scale insects. Phytotaxa 153:39–50
Herlemann D, Labrenz M, Jürgens K et al (2011) Transitions in bacterial communities along the 2000 km salinity gradient of the Baltic Sea. ISME J 5:1571–1579
Hibbett DS, Bauer R, Binder M et al (2014) 14 Agaricomycetes. Systematics and evolution. Springer, Berlin, pp 373–429
Holland-Moritz H, Stuart J, Lewis LR, Miller S, Mack MC, McDaniel SF, Fierer N (2018) Novel bacterial lineages associated with boreal moss species. Environ Microbiol 20:2625–2638
Husson C, Scala B, Caël O, Frey P, Feau N, Ioos R, Marçais B (2011) Chalara fraxinea is an invasive pathogen in France. Eur J Plant Pathol 130:311–324
Jaklitsch WM, Voglmayr H (2017) Three former taxa of Cucurbitaria and considerations on Petrakia in the Melanommataceae. Sydowia 69:81–95
Joshi NA, Fass JN (2011) Sicle: a sliding-window, adaptive, quality-based trimming tool for FastQ files (Version 1.33) [Software]. https://github.com/najoshi/sickle
Kirk PM, Cannon PF, Minter DW, Stalpers JA (2011) Dictionary of the Fungi, 10th edn. CAB International, Wallingford
Klindworth A, Pruesse E, Schweer T, Peplies J, Quast C, Horn M, Glöckner FO (2013) Evaluation of general 16S ribosomal RNA gene PCR primers for classical and next-generation sequencing-based diversity studies. Nucleic Acids Res 41:e1–e1
Kowalski T (2006) Chalara fraxinea sp. nov. associated with dieback of ash (Fraxinus excelsior) in Poland. For Pathol 36:264–270
Lawrey J, Diederich P (2003) Lichenicolous fungi: interactions, evolution, and biodiversity. Bryologist 106:80–120
Liu X, Bolla K, Ashforth EJ, Zhuo Y, Gao H, Huang P, Stanley SA, Hung DT, Zhang L (2012) Systematics-guided bioprospecting for bioactive microbial natural products. Anton Van Leeuwenhoek 101:55–66
Longton RE (1973) The occurrence of radial infection patterns in colonies of polar bryophytes. Br Antarct Sur Bull 32:41–49
Madrid H, Hernández-Restrepo M, Gené J et al (2016) New and interesting chaetothyrialean fungi from Spain. Mycol Prog 15:1179–1201
Medinger R, Nolte V, Pandey RV, Jost S, Ottenwälder B, Schlötterer C, Boenigk J (2010) Diversity in a hidden world: potential and limitation of next-generation sequencing for surveys of molecular diversity of eukaryotic microorganisms. Mol Ecol 19:32–40
Möller C, Dreyfuss MM (1996) Microfungi from Antarctic lichens, mosses and vascular plants. Mycologia 88:922–933
Nakai R, Abe T, Baba T et al (2012) Microflorae of aquatic moss pillars in a freshwater lake, East Antarctica, based on fatty acid and 16S rRNA gene analyses. Polar Biol 35:425–433
Nguyen NH, Song Z, Bates ST, Branco S, Tedersoo L, Menke J, Schilling JS, Kennedy PG (2016) FUNGuild: an open annotation tool for parsing fungal community datasets by ecological guild. Fungal Ecol 20:241–248
Ngwene B, Boukail S, Söllner L et al (2016) Phosphate utilization by the fungal root endophyte Piriformospora indica. Plant Soil 405:231–241
Ochyra R, Lewis-Smith RI, Bednarek-Ochyra H (2008) The illustrated moss flora of Antarctica. Cambridge University Press, Cambridge
Ondov BD, Bergman NH, Phillippy AM (2011) Interactive metagenomic visualization in a Web browser. BMC Bioinform 12:385
Øvstedal DO, Lewis-Smith R (2001) Lichens of Antarctica and South Georgia: a guide to their identification and ecology. Cambridge University Press, Cambridge, p 424
Palaniyandi SA, Yang SH, Zhang L, Suh JW (2013) Effects of actinobacteria on plant disease suppression and growth promotion. Appl Microbiol Biotechnol 97:9621–9636
Pandey KD, Shukla SP, Shukla PN, Giri DD (2004) Cyanobacteria in Antarctica: ecology, physiology and cold adaptation. Cell Mol Biol Paris Wegmann 50:575–584
Park M, Lee H, Hong SG, Kim OS (2013) Endophytic bacterial diversity of an Antarctic moss, Sanionia uncinata. Antarct Sci 25:51
Pawłowska J, Istel Ł, Gorczak M, Galera H, Wrzosek M, Hawksworth DL (2017) Psychronectria hyperantarctica, gen. nov., comb. nov., epitypification and phylogenetic position of an Antarctic bryophilous ascomycete. Mycologia 109:601–607
Pearce DA, Newsham KK, Thorne MA, Calvo-Bado L, Krsek M, Laskaris P, Hodson A, Wellington EM (2012) Metagenomic analysis of a southern maritime Antarctic soil. Front Microbiol 3:403
Polashock JJ, Caruso FL, Oudemans PV, McManus PS, Crouch JA (2009) The North American cranberry fruit rot fungal community: a systematic overview using morphological and phylogenetic affinities. Plant Pathol 58:1116–1127
Prather HM, Casanova-Katny A, Clements AF et al (2019) Species-specific effects of passive warming in an Antarctic moss system. R Soc Open Sci 6:190744
Putzke J, Pereira AB (2012) Fungos muscícolas na Ilha Elefante-Antártica. Cad Pesq Biol 24:155–164
Racovitza A (1959) É tude systematique et biologique des champignons bryophiles. Mém Mus NaTl Hist Nat Sér B Bot 10:1–288
Raymond JA (2016) Dependence on epiphytic bacteria for freezing protection in an Antarctic moss, Bryum argenteum. Environ Microbiol Rep 8:14–19
Reichenbach H (1999) The ecology of the myxobacteria. Environ Microbiol 1:15–21
Richardson RT, Lin CH, Sponsler DB, Quijia JO, Goodell K, Johnson RM (2015) Application of ITS2 metabarcoding to determine the provenance of pollen collected by honey bees in an agroecosystem. Appl Plant Sci 3:1400066
Robinson SA, King DH, Bramley-Alves J et al (2018) Rapid change in East Antarctic terrestrial vegetation in response to regional drying. Nat Clim Change 8:879–884
Rosa LH, de Sousa JRP, de Menezes GCA et al (2020a) Opportunistic fungal assemblages present on fairy rings spread on different moss species in the Antarctic Peninsula. Polar Biol 43:587–596
Rosa LH, da Silva TH, Ogaki MB et al (2020b) DNA metabarcoding uncovers fungal diversity in soils of protected and non-protected areas on Deception Island, Antarctica. Sci Rep 10:1–9
Rosa LH, Pinto OHB, Šantl-Temkiv T et al (2020c) DNA metabarcoding of fungal diversity in air and snow of Livingston Island, South Shetland Islands, Antarctica. Sci Rep 10:1–11
Rosa LH, Pinto OHB, Convey P et al (2021) DNA metabarcoding to assess the diversity of airborne fungi present over Keller Peninsula, King George Island, Antarctica. Microbial Ecol 82:165–172
Rosa LH, Zani CL, Cantrell CL, Duke SO, Van Dijck P, Desideri A, Rosa CA (2019) Fungi in Antarctica: diversity, ecology, efects of climate change, and bioprospection for bioactive compounds. In: Rosa LH (ed) Fungi of Antarctica: diversity, ecology and biotechnological applications. Springer, New York, pp 1–17
Smith RIL (1984) Terrestrial plant biology of the sub-Antarctic and Antarctic. In: Laws RM (ed) Antarctic ecology, vol 1. Academic Press, London, pp 61–162
Sutton BC (1980) The Coelomycetes. Fungi imperfecti with pycnidia, acervuli and stromata. Commonwealth Mycological Institute, Kew, Surrey, England
Taton A, Grubisic S, Brambilla E, de Wit R, Wilmotte A (2003) Cyanobacterial diversity in natural and artificial microbial mats of Lake Fryxell (McMurdo Dry Valleys, Antarctica): a morphological and molecular approach. Appl Environ Microbiol 69:5157–5169
Tedersoo L, Sánchez-Ramírez S, Kõljalg U et al (2018) High-level classification of the Fungi and a tool for evolutionary ecological analyses. Fungal Div 90:135–159
Tojo M, van West P, Hoshino T et al (2012) Pythium polare, a new heterothallic oomycete causing brown discolouration of Sanionia uncinata in the Arctic and Antarctic. Fungal Biol 116:756–768
Tosi S, Casado B, Gerdol R (2002) Fungi isolated from Antarctic mosses. Polar Biol 25:262–268
Tsuneda A, Hambleton S, Currah RS (2011) The anamorph genus Knufia and its phylogenetically allied species in Coniosporium, Sarcinomyces and Phaeococcomyces. Botany 89:523–536
Untereiner WA, Gueidan C, Orr MJ, Diederich P (2011) The phylogenetic position of the lichenicolous ascomycete Capronia peltigerae. Fungal Div 49:225–233
van de Vossenberg BTLH, Warris S, Nguyen HDT et al (2019) Comparative genomics of chytrid fungi reveal insights into the obligate biotrophic and pathogenic lifestyle of Synchytrium endobioticum. Sci Rep 9:8672
Varma A, Bakshi M, Lou B, Hartmann A, Oelmueller R (2012) Piriformospora indica: a novel plant growth-promoting mycorrhizal fungus. Agric Res 1:117–131
Verma A, Varma A, Rexer K-H et al (1998) Piriformospora indica, gen. et sp. nov., a new root colonizing fungus. Mycologia 90:896–903
Vincent WF (2000) Evolutionary origins of Antarctic microbiota: invasion, selection and endemism. Antarc Sci 12:374–385
Wang S, Tang JY, Ma J, Li XD, Li YH (2018) Moss habitats distinctly affect their associated bacterial community structures as revealed by the high-throughput sequencing method. World J Microbiol Biotechnol 34:58
Weiß M, Waller F, Zuccaro A, Selosse MA (2016) Sebacinales—one thousand and one interactions with land plants. New Phytol 211:20–40
White TJ, Bruns T, Lee S et al (1990) Amplifcation and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In: Innis MA, Gelfand DH, Sninsky JJ, White TJ (eds) PCR protocols: a guide to methods and applications. Academic Press, London, pp 315
Wilson JW (1951) Observations on concentric “fairy rings” in Arctic moss mat. J Ecol 39:407–416
Yadav V, Kumar M, Deep DK et al (2010) A phosphate transporter from the root endophytic fungus Piriformospora indica plays a role in phosphate transport to the host plant. J Biol Chem 285:26532–26544
Yuan ZQ (1996) Fungi and associated tree diseases in Melville Island, Northern Territory, Australia. Aus Syst Bot 9:337–360
Zhang T et al (2013) Molecular analysis of fungal diversity associated with three bryophyte species in the Fildes Region, King George Island, Maritime Antarctica. Extremophiles 17:757–765
Acknowledgements
We acknowledge financial support from PROANTAR CNPq (442258/2018-6), INCT Criosfera, FAPEMIG, CAPES and FNDCT. P. Convey is supported by NERC core funding to the British Antarctic Survey’s ‘Biodiversity, Evolution and Adaptation’ Team. We also thank congresswoman Jô Moraes and the Biological Sciences Institute of the University of Brasilia
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Communicated by A. Oren.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Rosa, L.H., da Costa Coelho, L., Pinto, O.H.B. et al. Ecological succession of fungal and bacterial communities in Antarctic mosses affected by a fairy ring disease. Extremophiles 25, 471–481 (2021). https://doi.org/10.1007/s00792-021-01240-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00792-021-01240-1