Abstract
Geobacillus spp. are moderate thermophiles that have great potential for use in diverse applications. For effective utilization of the species, genetic tools have been extensively studied; however, an overexpression vector remains to be developed. Here we constructed a plasmid vector that can shuttle between Escherichia coli and Geobacillus spp., and which contained a maltose-inducible promoter from Geobacillus kaustophilus HTA426. Although the vector (termed pGKE119) was originally designed for basal gene expression, it surprisingly directed robust protein production in G. kaustophilus. Protein production essentially occurred in an auto-inducible manner without maltose; however, some proteins were produced more efficiently in the presence of maltose. Although the productivity was affected by culture conditions, three proteins were successfully produced with abundance ratios of 12–27% (on a total protein basis) and yields of 77–170 mg (per L culture). pGKE119 directed substantial protein production even in Geobacillus subterraneus, Geobacillus thermoglucosidasius, and Geobacillus thermoleovorans. This suggests that pGKE119 can use a range of Geobacillus spp. as hosts and widely expand their genetic toolbox. Because Geobacillus spp. are highly proliferative bacteria that are distinct from organisms used as protein production hosts, pGKE119 may also provide a novel platform for hyperproduction of recombinant proteins.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The genus Geobacillus comprises aerobic or facultative anaerobic Gram-positive bacteria that grow preferentially at 55–65 °C. Members of the genus are of historical importance as producers of thermostable proteins that serve as robust enzymatic catalysts or biomimetic structures (Hussein et al. 2015). In addition, Geobacillus spp. have attracted interest as hosts for whole-cell applications because the species often exhibit unique properties that are useful in bioproduction, biodegradation, biomineralization, and bioremediation (Hussein et al. 2015), and high-temperature processes using thermophiles have several advantages over moderate processes using mesophiles (Wiegel and Ljungdahl 1986).
Genetic modification of microorganisms facilitates their use in whole-cell applications; thus, genetic tools available for Geobacillus spp. have been extensively studied. Numerous genetic tools are summarized in previous reviews (Hussein et al. 2015; Kananavičiūtė and Čitavičius 2015). Other genetic tools recently reported include plasmids (Reeve et al. 2016), reporter proteins (Frenzel et al. 2018), mutant sequences for promoters and ribosome-binding sites (Pogrebnyakov et al. 2017; Reeve et al. 2016), and gene knockout systems via allele-coupled exchange (Bacon et al. 2017; Sheng et al. 2017). However, a convenient vector that directs substantial gene expression remains to be established in Geobacillus spp.
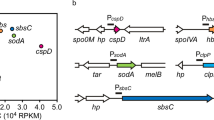
Geobacillus kaustophilus HTA426 has served as a pilot organism for biotechnological studies on Geobacillus spp. (Suzuki 2017). The strain originated from a mud sample from the bottom of the Challenger Deep in the Mariana Trench (Takami et al. 1997), and the complete genome sequence of this strain has been published (Takami et al. 2004). In a previous report (Suzuki et al. 2013b), we reported that the genome contained a possible operon for starch utilization (Fig. 1a) and that the promoter (termed the gk704 promoter) was induced by maltose. The present study was originally designed to construct a plasmid that contained the gk704 promoter for basal gene expression in Geobacillus spp.; however, contrary to our expectations, the plasmid directed substantial production of diverse heterologous proteins. We report here on the development of an overexpression vector for Geobacillus spp., which not only expands the genetic tools available for Geobacillus spp. but also suggests a novel platform for recombinant protein production.
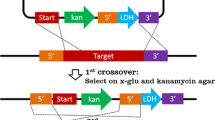
Construction of the pGKE119 vector. a Gene organization surrounding the gk704 promoter in Geobacillus kaustophilus HTA426. GK0704–GK0708 potentially constitute an operon for starch utilization where GK0704–GK0706 encode sugar ABC transporter whereas GK0707 and GK0708 encode α-amylase and LacI family regulator, respectively. The gk704 promoter (Pgk704) is located upstream of the operon. b Schematic representation of pGKE119 construction. Abbreviations used are the following: bla, ampicillin resistance gene; cat, chloramphenicol resistance gene; lacZα-T7term, lacZα gene followed by the T7 terminator; oriT, conjugative transfer origin; pBST, pBST1 replicon for autonomous replication in Geobacillus spp.; pUC and p15A, replicons for autonomous replication in E. coli; and TK101, kanamycin resistance gene functional at elevated temperatures. Relevant sites digested by restriction endonucleases are also indicated
Materials and methods
Bacterial strains
Geobacillus kaustophilus MK244 (JCM 31,151) was previously constructed from G. kaustophilus HTA426 (Suzuki et al. 2013a). G. kaustophilus MK244 has the genotype: ΔpyrF, ΔpyrR, ΔhsdM1S1R1, and Δ(mcrB1–mcrB2–hsdM2S2R2–mrr). Geobacillus subterraneus DSM 13,552, Geobacillus (Parageobacillus) thermoglucosidasius DSM 2542, and Geobacillus thermoleovorans DSM 5366 were purchased from the Bacillus Genetic Stock Center (Columbus, OH, USA). Escherichia coli strains DH5α and BL21(DE3) were purchased from Takara Bio (Otsu, Japan) and Merck Millipore (Darmstadt, Germany), respectively. E. coli DH5α, carrying either pUB307 or pRK2013, was used as a helper strain for ternary conjugation (Suzuki et al. 2013a).
Culture conditions
Escherichia coli was cultured at 37 °C in Luria–Bertani (LB) medium (Nacalai Tesque, Kyoto, Japan). E. coli, carrying the TK101 marker, was cultured in the presence of 20 mg/L kanamycin. Geobacillus spp. were cultured at 60 °C in LB or semisynthetic (MC and MY) liquid media that contained inorganic salts (0.3 g/L K2SO4, 2.5 g/L Na2HPO4·12H2O, 1 g/L NH4Cl, 0.4 g/L MgSO4, 3 mg/L MnCl2·4H2O, 5 mg/L CaCl2·2H2O, and 7 mg/L FeCl3·6H2O), 0.1% trace element solution (Amartey et al. 1991), 10 mM Tris–HCl (pH 7.5), and 10 mg/L uracil. MC medium also contained 10 g/L casamino acids (Becton, Dickinson and Company, Franklin Lakes, NJ, USA), whereas MY medium contained 10 g/L yeast extract (Becton, Dickinson and Company). Geobacillus transformants carrying the TK101 marker were cultured in the presence of 5 mg/L kanamycin. Protein production was examined under constitutive, inductive, and non-inductive expression of the gk704 promoter. Non-inductive expression was examined via consecutive incubation in LB and semisynthetic media that contained 1% D-glucose. Constitutive expression was performed via consecutive incubation in media supplemented with 1% maltose. Inductive expression was performed via preincubation for 6 h (until the culture reached the stationary phase) in media without maltose and then incubation in the presence of 1% maltose.
Genetic materials
Table 1 summarizes the plasmids used in this study. pUC19ΔNdeI was constructed from pUC19 (Takara Bio) via a silent mutation in the lacZα gene to abolish the NdeI site (5′-CATATG-3′ to 5′-TATATG-3′). The mutagenesis was performed using a QuikChange Site-directed Mutagenesis Kit (Agilent Technologies, Santa Clara, CA, USA). venusGK was artificially synthesized along with codon optimization for G. kaustophilus HTA426. The sequence was deposited in DDBJ/EMBL/GenBank databases with accession number LC488130.
pGKE119 construction
The p15A replicon was amplified from pIR200 using the primers 5′-CCCAGATCTGGATCCGCGTAACGGCAAAAGCACC-3′ and 5′-CCCAGATCTCCCTTGAGAGCCTTCAACCC-3′ (BglII sites underlined and BamHI site in italics) and trimmed with BglII. The pBST1 replicon and TK101 gene were together amplified from pGKE75 using the primers 5′-GCGGGATCCACTAGTTTCCTTAAGGAACGTACAGACGG-3′ and 5′-GCGGGATCCTCAAAATGGTATGCGTTTTGACAC-3′ (BamHI sites underlined and SpeI site in italics) and trimmed with BamHI. The two fragments were ligated to give pGKE98 (Fig. 1b), which was then digested with MunI and ligated with an oriT fragment that was excised from pGKE75 with EcoRI. Subsequently, the oligonucleotides 5′-GATCAAGCTTGCATGCATATGCTGCAGCTCGAGTCGACGGATCCGAATTC-3′ and 5′-AAGCTTGCATGCATATGCTGCAGCTCGAGTCGACGGATCCGAATTCGATC-3′ were hybridized and cloned at the BamHI site to give pGKE106. pGKE119 was constructed from pGKE106 via ligation with the gk704 promoter, T7 terminator, and lacZα fragments. The gk704 promoter was excised from pGKE75 and cloned between HindIII and SphI sites. The lacZα fragment, devoid of the NdeI site, was amplified from pUC19ΔNdeI using the primers 5′-CAGGAAACAGCTATGAC-3′ and 5′-GGGACTAGTATCGATCAATTGCTATGCGGCATCAGAGCAG-3′ (SpeI site underlined) and cloned between EcoRI and SpeI sites. The T7 terminator was generated via hybridization of the oligonucleotides 5′-AATTGCTAGCATAACCCCTTGGGGCCTCTAAACGGGTCTTGAGGGGTTTTTTGA-3′ and 5′-CTAGTCAAAAAACCCCTCAAGACCCGTTTAGAGGCCCCAAGGGGTTATGCTAGC-3′ and cloned between MunI and SpeI sites.
pGKE119 construction of pGKE119-bgaB, pGKE119-catA138T, and pGKE119-venusGK
The venusGK fragment was cloned between SphI and BamHI sites of pGKE119 to give pGKE119-venusGK. The bgaB and catA138T fragments were excised from pGAM46-bgaB and pGKE75-catA138T, respectively, and subcloned between SphI and BamHI sites of pGKE119 to give pGKE119-bgaB and pGKE119-catA138T.
Plasmid transformation
Competent E. coli cells were prepared using a standard method (Inoue et al. 1990). pGKE119 derivatives were transferred from E. coli DH5α (donor) to Geobacillus spp. (recipient) via ternary conjugation with E. coli DH5α carrying pUB307 or pRK2013 (as helper). Conjugation was performed following the previously published procedure (Suzuki et al. 2013a). Briefly, the recipient, donor, and helper strains were cultured to the proliferative phase and mixed at a ratio of 10:1:1. Cells were concentrated on a nitrocellulose membrane (0.22 µm) via suction filtration and incubated on an LB plate at 37 °C for 16 h to achieve conjugation. Cells were then recovered from the membrane and incubated at 60 °C on LB plates supplemented with or without kanamycin (5 mg/L) to determine the concentration of transformant or recipient cells, respectively. The conjugation efficiency was expressed as the number of transformants per total number of recipients. Data are presented as the mean ± standard error (n = 3–4).
Flow cytometric analysis
Geobacillus kaustophilus MK244 [pGKE119-venusGK], where square brackets indicate a carrier state of the plasmid, was precultured in LB medium at 60 °C. An aliquot of the culture (50 µL) was inoculated in LB medium (5 mL) in a test tube and aerobically incubated at 60 °C for 24 h with rotary shaking at 180 rpm. Cell fluorescence was first detected with light-emitting diodes that irradiated green light at a wavelength of about 500 nm and was then analyzed by flow cytometry using FACSAria (Becton, Dickinson and Company) with excitation at 488 nm and detection at 519 nm. The analysis was performed for 104 cells. G. kaustophilus MK244 [pGKE119] was used as the negative control.
Quantitative assay of cell fluorescence
Geobacillus kaustophilus MK244 [pGKE119-venusGK] was precultured in LB medium at 60 °C. Aliquots of culture (200 µL) were inoculated in different media (20 mL) in Erlenmeyer flasks (100 mL). The medium was aerobically incubated for 3 d, and aliquots (1 mL) were collected every 12 h. The culture was quickly frozen using liquid nitrogen and stored at − 80 °C until use. The frozen sample was spontaneously thawed at room temperature and appropriately diluted in 50 mM Tris–HCl (pH 8.0). Culture fluorescence was determined using a FP-8300 DS fluorescence spectrophotometer (JASCO Corp., Tokyo, Japan), with excitation at 515 nm and detection at 528 nm. In parallel, the optical density at 600 nm (OD600) of the culture was determined using a UV-1700 spectrophotometer (Shimadzu, Kyoto, Japan). The intensity of cell fluorescence was defined as culture fluorescence per OD600, where the mean value after incubation at 60 °C for 24 h in LB medium was taken to be one unit. G. kaustophilus MK244 [pGKE119] was used as the negative control.
Estimation of protein productivity
Geobacillus spp. were precultured in LB medium at 60 °C. Aliquots of culture (200 µL) were inoculated into LB medium (20 mL) in an Erlenmeyer flask (100 mL). After the medium was aerobically incubated for 24 h or 48 h at 50–65 °C, cells were harvested by centrifugation (5000×g for 10 min) and stored at − 80 °C. Frozen cells were suspended in 50 mM sodium phosphate at pH 7.0 (1 mL) and homogenized by sonication. Homogenates were centrifuged to obtain clear lysates, in which protein concentrations were determined using the Bradford assay with bovine serum albumin as the standard. The lysate was further analyzed using sodium dodecyl sulfate–polyacrylamide gel electrophoresis (SDS–PAGE). Proteins in gels were stained with Coomassie Brilliant Blue. The intensity of the protein bands was quantified using the ImageJ program (https://rsb.info.nih.gov/ij) to calculate the intensity ratio of the recombinant protein to total proteins, which was defined as the abundance ratio. The yield of recombinant proteins (mg protein per L culture) was calculated based on the abundance ratio and protein concentration.
Results
Genetic characteristics of the pGKE119 vector
Escherichia coli transformants with pGKE119 formed substantial colonies (>1 mm in diameter) on LB plates supplemented with 20 µg/mL kanamycin following incubation at 37 °C for 24 h. False positives appeared on plates with 10 µg/mL kanamycin, whereas no colonies appeared in the presence of 50 µg/mL kanamycin possibly because the TK101 marker was inefficient at moderate temperatures. The transformation efficiency (n = 3) was determined as (4.8 ± 1.1) × 105 cfu/µg when examined using competent cells whose efficiency was 107 cfu/µg of pUC19. The lower efficiency is attributable to the differences in plasmid size (pGKE119, 5.8 kb; pUC19, 2.7 kb) and/or the replicon (pGKE119, p15A replicon; pUC19, pUC replicon). It is unlikely that the lower efficiency arose from the selection marker (pGKE119, TK101; pUC19, bla) because a plasmid that carried these markers produced comparable frequencies of transformants upon kanamycin and ampicillin selections (Reeve et al. 2016). In the presence of 5-bromo-4-chloro-3-indolyl-β-d-galactopyranoside, colonies exhibited pale blue coloration because of the lacZα gene in pGKE119. Although coloration required incubation for > 24 h, E. coli transformants that carried pGKE119 derivatives (e.g., pGKE119-bgaB, pGKE119-catA138T, and pGKE119-venusGK) were strictly selected on the basis of colony coloration. G. kaustophilus accepted pGKE119 transferred from E. coli via ternary conjugation. The efficiencies (n = 4) were (1.1 ± 0.8) × 10−4 and (1.2 ± 0.7) × 10−5 when pUB307 and pRK2013, respectively, were used as conjugation helper plasmids. The efficiency was comparable to that observed for a plasmid previously constructed (Suzuki et al. 2013a). When pGKE119 was purified from G. kaustophilus [pGKE119], it produced only a slight band on agarose gel electrophoresis analysis. The observation implied that pGKE119 replicated at a low copy number in G. kaustophilus.
venusGK expression from pGKE119
Venus is a yellow fluorescent protein that is monomeric, tolerant to acidic conditions, and excellent in terms of protein folding and brightness (Nagai et al. 2002). We synthesized the gene (venusGK) and used it as a reporter gene to examine gene expression from pGKE119 in G. kaustophilus. Biochemical analysis confirmed that mature VenusGK was sufficiently thermostable in vitro to retain > 90% fluorescence at 70 °C for 12 h. G. kaustophilus [pGKE119-venusGK] exhibited obvious yellow fluorescence when irradiated with green light of the excitation wavelength (Fig. 2a). The cells retained fluorescence at 70 °C with a half-life of > 12 d in the presence of antibiotics (streptomycin and rifampicin) to prevent further protein synthesis, suggesting that mature VenusGK is extremely thermostable in vivo. Flow cytometric analysis clearly differentiated cell fluorescence between G. kaustophilus carrying pGKE119-venusGK and pGKE119 (Fig. 2b). Non-fluorescent cells were negligibly detected in G. kaustophilus [pGKE119-venusGK]; therefore, it is likely that pGKE119-venusGK was stably maintained during cell division.
Cell fluorescence of Geobacillus kaustophilus [pGKE119-venusGK]. a Visual cell fluorescence of G. kaustophilus that carried pGKE119-venusGK (+) or pGKE119 (−). Cells were cultured at 60 °C for 24 h and observed with (lower panel) or without (upper panel) green light irradiation. b Cell fluorescence distribution of G. kaustophilus that carried pGKE119-venusGK (yellow) or pGKE119 (gray). The analysis was performed for 104 cells using flow cytometric analysis
Effects of culture conditions
Geobacillus kaustophilus [pGKE119-venusGK] was cultured in LB medium at 60 °C while cell fluorescence was analyzed (Fig. 3a). Cell growth reached the stationary phase at 12 h, and yellow fluorescence associated with VenusGK increased markedly at 36 h. This suggests that venusGK expression was spontaneously induced during the stationary phase even in the absence of maltose. Cells were also cultured under constitutive and inductive conditions to determine whether maltose increased venusGK expression; however, neither of the conditions had any marked effects on cell fluorescence (Fig. 3b). Similar autoinduction was observed in MC medium but not in MY medium. In MC medium, constitutive conditions caused higher cell fluorescence at the middle stationary phase than at the late stationary phase; therefore, it is likely that maltose served as an inducer in MC medium, a finding in agreement with the previous observation (Suzuki et al. 2013b). In MY medium, inductive conditions slowly increased cell fluorescence, but constitutive conditions did not. Among the culture conditions examined, VenusGK was most efficiently produced in LB medium at the late stationary phase under non-inductive conditions.
VenusGK production is dependent on culture conditions. aGeobacillus kaustophilus that carried pGKE119-venusGK (solid) or pGKE119 (hollow) was cultured in LB medium at 60 °C and analyzed for cell fluorescence. bG. kaustophilus [pGKE119-venusGK] was analyzed for cell fluorescence following culture at 60 °C for 12 h (solid) and 48 h (hollow) under non-inductive (No), constitutive (Cs), or inductive conditions (In). The analysis was performed for three independent clones. Data are presented as the mean ± standard error. Single and double asterisks indicate that P values (one-way ANOVA) are < 0.1 and < 0.01, respectively
pGKE119 directs robust protein production
Geobacillus kaustophilus [pGKE119-venusGK] was cultured in LB medium at different temperatures, after which intracellular soluble proteins were analyzed using SDS–PAGE (Fig. 4). The protein band of VenusGK (27 kDa) was observed at 48 h but not at 24 h when cells were cultured at 60 °C. This observation was consistent with the time course of cell fluorescence, which was much higher at 48 h than at 24 h (Fig. 3a). The VenusGK band was undetectable at 65 °C but became more prominent at < 55 °C. On the basis of band intensity, VenusGK was produced at 48 h with an abundance ratio of 27% and a yield of 170 mg/L. The higher productivity at 50 °C than at 60 °C could be because VenusGK folding was more efficient at lower temperatures and misfolded proteins produced at higher temperatures were degraded by intrinsic biological processes. This idea is consistent with the observation that BgaB, which is a thermostable β-galactosidase from G. kaustophilus ATCC 8005, was produced over a broad temperature range of 50–65 °C with abundance ratios of 5–14% and yields of 11–72 mg/L (Fig. 4). catA138T encodes a thermostable variant of chloramphenicol acetyltransferase, which was produced exclusively at 50 °C with an abundance ratio of 6–7% and a yield of 32–34 mg/L. Because the protein originates from a mesophilic bacterium (Staphylococcus aureus), its folding may also be inefficient at elevated temperatures, although the mature form was thermostable even at > 60 °C (Kobayashi et al. 2015a). We further examined BgaB and CatA138T production in MY medium under inductive conditions. Both proteins were produced more efficiently at 50 °C and at 24 h, where the abundance ratio and yield were 24% and 140 mg/L for BgaB and 12% and 77 mg/L for CatA138T, respectively. The results confirmed that culture conditions affected protein production from pGKE119 and that inductive conditions might increase the productivity of certain proteins.
Heterologous protein production from pGKE119 derivatives. Geobacillus kaustophilus that carried pGKE119-venusGK, pGKE119-bgaB, pGKE119-catA138T, or pGKE119 (empty) was cultured in LB medium for 24 h (a) and 48 h (b) at temperatures indicated on top of gels. Subsequently, intracellular soluble proteins (40 µg) were analyzed using SDS–PAGE. Recombinant proteins are indicated by arrows. The abundance ratios and yields of recombinant proteins were calculated based on band intensities. Minus indicates that the recombinant protein band was unclear
VenusGK production at moderate temperatures
VenusGK expression was examined at lower temperatures (Fig. 5). Because G. kaustophilus barely grows at < 45 °C, cells were first precultured at 60 °C until the stationary phase was reached and then incubated at 40 °C. Cell fluorescence was slowly increased and reached a plateau at 60 h. According to SDS–PAGE analysis, VenusGK was produced at 60 h with an abundance ratio of 15% and a yield of 73 mg/L. Cell fluorescence also increased during incubation at 30 °C and 35 °C, albeit with slower increase rates. The results show that pGKE119 can direct protein production even at moderate temperatures, although longer incubation times may be required.
VenusGK production at moderate temperatures. Geobacillus kaustophilus [pGKE119-venusGK] was precultured in LB medium at 60 °C for 6 h and then incubated at 40 °C (solid squares), 35 °C (hollow squares), or 30 °C (solid circles). Subsequently, cell fluorescence was analyzed. The analysis was performed for three independent clones. Data are presented as the mean ± standard error
pGKE119 functions widely in Geobacillus spp.
VenusGK expression was examined using G. subterraneus, G. thermoglucosidasius, and G. thermoleovorans as host cells. Transformants carrying pGKE119-venusGK were cultured in LB medium at different temperatures, following which intracellular soluble proteins were analyzed by SDS-PAGE (Fig. 6). G. subterraneus [pGKE119-venusGK] showed lower productivity for unknown reasons; however, G. thermoglucosidasius [pGKE119-venusGK] and G. thermoleovorans [pGKE119-venusGK] produced substantial amounts of VenusGK with abundance ratios of 17–25% and yields of 49–170 mg/L at < 55 °C.
VenusGK production in other Geobacillus spp. Geobacillus subterraneus (a), Geobacillus thermoglucosidasius (b), and Geobacillus thermoleovorans (c) were transformed with pGKE119-venusGK and pGKE119 (empty). Cells were cultured in LB medium for 24 h (upper panels) and 48 h (lower panels) at temperatures indicated on top of gels. Subsequently, intracellular soluble proteins (40 µg) were analyzed using SDS–PAGE. Gels focus on protein bands around VenusGK. The abundance ratios and yields of VenusGK were calculated based on band intensities. Minus indicates that the VenusGK band was unclear
Discussion
pGKE119 was constructed to facilitate gene expression in Geobacillus spp. To this end, the plasmid contained multiple cloning sites in lacZα under the control of the gk704 promoter. The lacZα gene was functionally expressed in E. coli and allowed us to screen for pGKE119 derivatives based on colony coloration. TK101 was contained as a sole selectable marker. This contributed to a reduction in the plasmid size, whereas E. coli transformants were readily selected based on kanamycin resistance without false positives. pGKE119 contains p15A and pBST1 replicons for autonomous replications in E. coli and Geobacillus spp., respectively. The p15A replicon replicates at a moderate copy number (~20 copies) in contrast to the pUC replicon (>500 copies), which was commonly used in previously studied plasmids that shuttle between E. coli and Geobacillus spp. (Hussein et al. 2015; Kananavičiūtė and Čitavičius 2015; Reeve et al. 2016). pGKE119 employed the p15A replicon to decrease involuntary gene expression from the gk704 promoter in E. coli. The T7 terminator was arranged downstream of lacZα to depress read-through transcription from the gk704 promoter. We noted that venusGK was not expressed from a pGKE119 derivative that lacked the T7 terminator in G. kaustophilus (data not shown), indicating that the terminator contributed to the stable gene expression from pGKE119. Since the pBST1 replicon is located downstream of the gk704 promoter, the terminator may prevent read-through transcription from impairing plasmid replication in G. kaustophilus.
The pBST1 replicon has been widely employed in E. coli–Geobacillus shuttle plasmids including pUCG18T (Suzuki and Yoshida 2012), pGKE75 (Kobayashi et al. 2015a), and pG1AK (Reeve et al. 2016). In previous observations, pUCG18T was undetectable following agarose gel electrophoresis when recovered from transformant cells of Geobacillus spp., whereas pSTE33T, which has another type of replicon and replicates at a moderate copy number (~16 copies), was clearly detected (Suzuki and Yoshida 2012; Tominaga et al. 2016). On the basis of the observation, we assumed that the pBST1 replicon replicated at low copy numbers in Geobacillus spp. and designed pGKE119 as a plasmid that could direct basal gene expression in Geobacillus spp. However, although pGKE119 identity was still unclear following agarose gel electrophoresis, it directed the hyperproduction of heterologous proteins in G. kaustophilus with abundance rates of 12–27% and yields of 77–170 mg/L. The reason for such hyperproduction is unclear, but it is possible that pGKE119 actually replicates at a high copy number. This idea is supported by copy number analysis using PCR-based methods, which suggest that pGKE75 derivatives replicate at > 17 copies in G. kaustophilus (Kobayashi et al. 2015b) and that pG1AK replicates at > 80 copies in G. thermoglucosidasius (Reeve et al. 2016). The discrepancy in terms of plasmid copy numbers determined by PCR-based methods and agarose gel electrophoresis may be explained by the large plasmid concatemers that are generated during plasmid replication via a rolling circle mechanism (Khan 1997). The concatemers could express multiple copies of a gene even though their identity on agarose gel electrophoresis is unclear because of the wide distribution of sizes. Although the pBST1 replicon exhibits a weak similarity to theta-type replicons (Taylor et al. 2008), it is still possible that the replication proceeds via a rolling circle mechanism because this mechanism is common in plasmids of Gram-positive bacteria (Khan 1997).
Numerous promoter sequences have been identified from Geobacillus spp. The ldh promoter from Geobacillus stearothermophilus NCA1503 expresses under conditions of oxygen limitation and is employed to modify Geobacillus spp. for fermentative biofuel production (Taylor et al. 2008). The 2n38 (Bartosiak-Jentys et al. 2013), pfl (Pogrebnyakov et al. 2017), pta (Frenzel et al. 2018), RHIII (Blanchard et al. 2014), and uppT12 (Daas et al. 2016) promoters are all constitutively expressed, whereas the βglu (Bartosiak-Jentys et al. 2013), surP (Blanchard et al. 2014), and xylA (Pogrebnyakov et al. 2017) promoters are induced by xylose, cellobiose, and sucrose, respectively. In addition, the rplS and groES promoters have been used to generate mutant sequences that exhibit expression levels spanning ~ 100-fold ranges (Pogrebnyakov et al. 2017; Reeve et al. 2016). Among these promoters, only the gk704 promoter in pSTE33T has been examined with respect to the amount of proteins that it can produce in Geobacillus spp.; however, this plasmid had not been optimized as an expression vector and showed lower productivity than pGKE119 (Suzuki et al. 2013b). Importantly, gene clusters homologous to GK0704–GK0708 have been widely identified in whole genome sequences of Geobacillus spp., including G. stearothermophilus, G. subterraneus, Geobacillus thermocatenulatus, Geobacillus thermodenitrificans, G. thermoglucosidasius, and G. thermoleovorans. This observation suggests that pGKE119 should function in these species and widely expand the genetic toolboxes available for Geobacillus spp. In fact, pGKE119 has been shown to function in G. subterraneus, G. thermoglucosidasius, and G. thermoleovorans (Fig. 6).
The development of pGKE119 not only expands the genetic tools available for Geobacillus spp. but also suggests a novel host–vector system for recombinant protein production. E. coli is commonly used as a host for recombinant protein production since the species can rapidly grow and numerous genetic tools are available to facilitate its modification (Kaur et al. 2018). Yeasts and fungi are also used as protein production hosts, which are advantageous for producing eukaryotic proteins that undergo post-translational modifications unique to eukaryotes (Baghban et al. 2019). In theory, any protein can be produced in such microorganisms; however, it is known that protein productivity varies to a large degree in terms of the combination of target protein and host species. This is attributable to biological and/or physicochemical factors such as codon usage biases and intracellular conditions that affect protein folding (Kaur et al. 2018; Suzuki et al. 2013b). Efficient production of recombinant proteins requires the consolidation of overexpression vectors for diverse microorganisms.
Geobacillus kaustophilus can grow efficiently on inexpensive substrates and produce recombinant proteins as well as does E. coli, whereas codon usages are substantially different between these microorganisms (Nakamura et al. 2000). On the basis of phylogenetic relationships (Suzuki 2018), G. kaustophilus may be more effective than E. coli for the hyperproduction of numerous proteins from the family Bacillaceae. Because certain proteins from thermophiles require high temperatures for correct protein folding, G. kaustophilus is potentially superior to mesophilic hosts in terms of the production of thermophile proteins (Suzuki et al. 2013b). Intriguingly, gene expression from pGKE119 was spontaneously induced during the stationary phase. This is presumably because expression of a repressor protein to repress the operon (e.g., the LacI family protein encoded by GK0708) declined along with cell proliferation. Autoinduction is advantageous for protein production because it allows for simple and less-expensive protein production without growth inhibition. It is also noteworthy that the host–vector system produced VenusGK efficiently at 40 °C. This suggests that the system can be used to produce thermolabile proteins. Given that thermolabile proteins are accumulated as denatured forms during incubation at elevated temperatures, the system may be applicable for the production of the toxic thermolabile proteins via in vivo synthesis in the form of denatured (non-toxic) configurations, followed by refolding in vitro. Such a process could be much less expensive than in vitro protein synthesis.
Abbreviations
- OD600 :
-
Optical density at 600 nm
- LB:
-
Luria–Bertani
- SDS-PAGE:
-
Sodium dodecyl sulfate–polyacrylamide gel electrophoresis
- []:
-
A carrier state of the plasmid
References
Amartey SA, Leak DJ, Hartley BS (1991) Development and optimization of a defined medium for aerobic growth of Bacillus stearothermophilus LLD-15. Biotechnol Lett 13:621–626. https://doi.org/10.1007/BF01086315
Bacon LF, Hamley-Bennett C, Danson MJ, Leak DJ (2017) Development of an efficient technique for gene deletion and allelic exchange in Geobacillus spp. Microb Cell Fact 16:58. https://doi.org/10.1186/s12934-017-0670-4
Baghban R, Farajnia S, Rajabibazl M, Ghasemi Y, Mafi A, Hoseinpoor R, Rahbarnia L, Aria M (2019) Yeast expression systems: overview and recent advances. Mol Biotechnol 61:365–384. https://doi.org/10.1007/s12033-019-00164-8
Bartosiak-Jentys J, Hussein AH, Lewis CJ, Leak DJ (2013) Modular system for assessment of glycosyl hydrolase secretion in Geobacillus thermoglucosidasius. Microbiology 159:1267–1275. https://doi.org/10.1099/mic.0.066332-0
Bennett PM, Grinsted MJ, Richmond MH (1977) Transposition of TnA does not generate deletions. Mol Gen Genet 154:205–211. https://doi.org/10.1007/BF00330839
Blanchard K, Robic S, Matsumura I (2014) Transformable facultative thermophile Geobacillus stearothermophilus NUB3621 as a host strain for metabolic engineering. Appl Microbiol Biotechnol 98:6715–6723. https://doi.org/10.1007/s00253-014-5746-z
Daas MJA, van de Weijer AHP, de Vos WM, van der Oost J, van Kranenburg R (2016) Isolation of a genetically accessible thermophilic xylan degrading bacterium from compost. Biotechnol Biofuels 9:210. https://doi.org/10.1186/s13068-016-0618-7
Figurski DH, Helinski DR (1979) Replication of an origin-containing derivative of plasmid RK2 dependent on a plasmid function provided in trans. Proc Natl Acad Sci USA 76:1648–1652. https://doi.org/10.1073/pnas.76.4.1648
Frenzel E, Legebeke J, van Stralen A, van Kranenburg R, Kuipers OP (2018) In vivo selection of sfGFP variants with improved and reliable functionality in industrially important thermophilic bacteria. Biotechnol Biofuels 11:8. https://doi.org/10.1186/s13068-017-1008-5
Hussein AH, Lisowska BK, Leak DJ (2015) The genus Geobacillus and their biotechnological potential. In: Sariaslani S, Gadd GM (eds) Advances in applied microbiology, vol 92. Elsevier, Amsterdam, pp 1–48
Inoue H, Nojima H, Okayama H (1990) High efficiency transformation of Escherichia coli with plasmids. Gene 96:23–28. https://doi.org/10.1016/0378-1119(90)90336-P
Kananavičiūtė R, Čitavičius D (2015) Genetic engineering of Geobacillus spp. J Microbiol Methods 111:31–39. https://doi.org/10.1016/j.mimet.2015.02.002
Kaur J, Kumar A, Kaur J (2018) Strategies for optimization of heterologous protein expression in E. coli: roadblocks and reinforcements. Int J Biol Macromol 106:803–822. https://doi.org/10.1016/j.ijbiomac.2017.08.080
Khan SA (1997) Rolling-circle replication of bacterial plasmids. Microbiol Mol Biol Rev 61:442–455
Kobayashi J, Furukawa M, Ohshiro T, Suzuki H (2015a) Thermoadaptation-directed evolution of chloramphenicol acetyltransferase in an error-prone thermophile using improved procedures. Appl Microbiol Biotechnol 99:5563–5572. https://doi.org/10.1007/s00253-015-6522-4
Kobayashi J, Tanabiki M, Doi S, Kondo A, Ohshiro T, Suzuki H (2015b) Unique plasmids generated via pUC replicon mutagenesis in an error-prone thermophile derived from Geobacillus kaustophilus HTA426. Appl Environ Microbiol 81:7625–7632. https://doi.org/10.1128/aem.01574-15
Nagai T, Ibata K, Park ES, Kubota M, Mikoshiba K, Miyawaki A (2002) A variant of yellow fluorescent protein with fast and efficient maturation for cell-biological applications. Nat Biotechnol 20:87–90. https://doi.org/10.1038/nbt0102-87
Nakamura Y, Gojobori T, Ikemura T (2000) Codon usage tabulated from international DNA sequence databases: status for the year 2000. Nucleic Acids Res 28:292–292. https://doi.org/10.1093/nar/28.1.292
Pogrebnyakov I, Jendresen CB, Nielsen AT (2017) Genetic toolbox for controlled expression of functional proteins in Geobacillus spp. PLoS ONE 12:e0171313. https://doi.org/10.1371/journal.pone.0171313
Reeve B, Martinez-Klimova E, de Jonghe J, Leak DJ, Ellis T (2016) The Geobacillus plasmid set: a modular toolkit for thermophile engineering. ACS Synth Biol 5:1342–1347. https://doi.org/10.1021/acssynbio.5b00298
Sheng LL, Kovacs K, Winzer K, Zhang Y, Minton NP (2017) Development and implementation of rapid metabolic engineering tools for chemical and fuel production in Geobacillus thermoglucosidasius NCIMB 11955. Biotechnol Biofuels 10:5. https://doi.org/10.1186/s13068-016-0692-x
Suzuki H (2017) Geobacillus kaustophilus HTA426: a model organism for moderate thermophiles. In: Berhardt LV (ed) Advances in medicine and biology, vol 114. Nova Science Publishers, New York, pp 75–108
Suzuki H (2018) Peculiarities and biotechnological potential of environmental adaptation by Geobacillus species. Appl Microbiol Biotechnol 102:10425–10437. https://doi.org/10.1007/s00253-018-9422-6
Suzuki H, Yoshida K (2012) Genetic transformation of Geobacillus kaustophilus HTA426 by conjugative transfer of host-mimicking plasmids. J Microbiol Biotechnol 22:1279–1287. https://doi.org/10.4014/jmb.1203.03023
Suzuki H, Takahashi S, Osada H, Yoshida K (2011) Improvement of transformation efficiency by strategic circumvention of restriction barriers in Streptomyces griseus. J Microbiol Biotechnol 21:675–678. https://doi.org/10.4014/jmb.1102.02038
Suzuki H, Murakami A, Yoshida K (2012) Counterselection system for Geobacillus kaustophilus HTA426 through disruption of pyrF and pyrR. Appl Environ Microbiol 78:7376–7383. https://doi.org/10.1128/aem.01669-12
Suzuki H, Wada K, Furukawa M, Doi K, Ohshima T (2013a) A ternary conjugation system for the construction of DNA libraries for Geobacillus kaustophilus HTA426. Biosci Biotechnol Biochem 77:2316–2318. https://doi.org/10.1271/bbb.1304921
Suzuki H, Yoshida K, Ohshima T (2013b) Polysaccharide-degrading thermophiles generated by heterologous gene expression in Geobacillus kaustophilus HTA426. Appl Environ Microbiol 79:5151–5158. https://doi.org/10.1128/aem.01506-13
Takami H, Inoue A, Fuji F, Horikoshi K (1997) Microbial flora in the deepest sea mud of the Mariana Trench. FEMS Microbiol Lett 152:279–285. https://doi.org/10.1016/s0378-1097(97)00211-5
Takami H, Takaki Y, Chee GJ, Nishi S, Shimamura S, Suzuki H, Matsui S, Uchiyama I (2004) Thermoadaptation trait revealed by the genome sequence of thermophilic Geobacillus kaustophilus. Nucleic Acids Res 32:6292–6303. https://doi.org/10.1093/nar/gkh970
Taylor MP, Esteban CD, Leak DJ (2008) Development of a versatile shuttle vector for gene expression in Geobacillus spp. Plasmid 60:45–52. https://doi.org/10.1016/j.plasmid.2008.04.001
Tominaga Y, Ohshiro T, Suzuki H (2016) Conjugative plasmid transfer from Escherichia coli is a versatile approach for genetic transformation of thermophilic Bacillus and Geobacillus species. Extremophiles 20:375–381. https://doi.org/10.1007/s00792-016-0819-9
Wiegel J, Ljungdahl LG (1986) The importance of thermophilic bacteria in biotechnology. CRC Rev Biotechnol 3:39–108
Acknowledgements
This work was funded by the following organizations: Japan Society for the Promotion of Science (Grant numbers: 25450105 and 17K06925), Nagase Science and Technology Foundation, and the Institute for Fermentation, Osaka, Japan.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflicts of interest. This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Communicated by Cann.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Kurashiki, R., Mizuno, T., Murata, K. et al. A plasmid vector that directs hyperproduction of recombinant proteins in the thermophiles Geobacillus species. Extremophiles 24, 147–156 (2020). https://doi.org/10.1007/s00792-019-01142-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00792-019-01142-3