Abstract
Thermostable lipases offer major biotechnological advantages over mesophilic lipases. In this study, an intracellular thermostable and organic solvent-tolerant lipase-producing strain YB103 was isolated from soil samples and identified taxonomically as Xanthomonas oryzae pv. oryzae. The lipase from X. oryzae pv. oryzae YB103 (LipXO) was purified 101.1-fold to homogeneity with a specific activity of 373.9 U/mg. The purified lipase showed excellent thermostability, exhibiting 51.1 % of its residual activity after incubation for 3 days at 70 °C. The enzyme showed optimal activity at 70 °C, suggesting it is a thermostable lipase. LipXO retained 75.1–154.1 % of its original activity after incubation in 20 % (v/v) hydrophobic organic solvents at 70 °C for 24 h. Furthermore, LipXO displayed excellent stereoselectivity (e.e.p >99 %) toward (S)-1-phenethyl alcohol in n-hexane. These unique properties of LipXO make it promising as a biocatalyst for industrial processes.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Lipase is a class of versatile enzyme capable of showing high enantioselectivity and broad substrate specificity simultaneously. Lipases (triacylglycerol hydrolases, EC 3.1.1.3) catalyze both the hydrolysis and the synthesis of acylglycerides and other fatty acid esters at high activity, regioselectivity, and stereoselectivity in aqueous and nonaqueous media (Gotor-Fernández et al. 2006; Tran et al. 2012). Although lipases are produced by animals, plants, and microorganisms (Franken et al. 2011; Cardenas et al. 2001), microbial lipases have an immense prospective for commercial applications (Haki and Rakshit 2003). They are widely utilized in a number of applications in various industries, including the biodiesel, pharmaceutical, detergent, cosmetic, and food industries (Dheeman et al. 2011; Dandavate et al. 2009).
Thermostable lipases usually function effectively at high temperatures, in comparison to mesophilic or cold-active lipases which show little activity at high temperature. In this respect, the thermostability of lipases is of paramount importance for any bioprocess in industry due to most of other enzymes being inherently unstable (Ebrahimpour et al. 2011; Faoro et al. 2012; Mahadevan and Neelagund 2014). Utilization of thermostable lipases in elevated temperatures can take several advantages, including increasing the reaction rates due to a decrease in viscosity and an increase in diffusion coefficient of substrates, reducing the risk of contamination by common mesophiles, and improving the process yield due to increased solubility of substrates (Haki and Rakshit 2003; Ebrahimpour et al. 2011; Vieille and Zeikus 2001). Therefore, discovery of novel thermostable lipases is prerequisite for industrial application.
Organic solvent-tolerant lipases offer new possibilities such as enabling the use of hydrophobic substrates and shifting of the thermodynamic equilibria in favor of synthesis (Ahmed et al. 2010; Doukyu and Ogino 2010; Lima et al. 2004). Furthermore, of particular importance has been the discovery that enzymatic stereoselectivity (substrate, stereo-, regio- and chemoselectivity) can be markedly affected by the solvent (Klibanov, 2001; Jaeger and Eggert 2002). If the lipases are intrinsically stable and active in solvents, it should make them more useful for enzymatic reaction in chemical industry.
In this work, we describe the identification, purification, and characterization of a novel thermostable and organic solvent-tolerant lipase (LipXO) from strain Xanthomonas oryzae pv. oryzae YB103. This lipase has been found stable not only at high temperature, but also in the presence of organic solvents. Furthermore, the lipase showed excellent stereoselectivity toward (S)-1-phenethyl alcohol in n-hexane. The characteristics of LipXO make it a potential biocatalyst for industrial biochemical processes.
Materials and methods
Materials
p-Nitrophenol (pNP), p-nitrophenyl-acetate, p-nitrophenyl-butyrate, p-nitrophenyl-caproate, p-nitrophenyl-octanoate (pNPO), p-nitrophenyl-decanoate, p-nitrophenyl-laurate (pNPL), p-nitrophenyl-myristate, p-nitrophenyl-palmitate, p-nitrophenyl-stearate, triacylglycerols (glyceryl triacetate, glyceryl tributyrate, glyceryl trihexanoate, glyceryl trioctanoate, glyceryl tridecanoate, glyceryl tridodecanoate, glyceryl trimyristate, glyceryl tripalmitate and glyceryl tristearate), (R, S)-1-phenethyl alcohol and (R)-1-phenethyl acetate were acquired from Sigma-Aldrich Co., Ltd. (Shanghai, China). All other chemicals used were of analytical grade from Guoyao Co. Ltd. (Shanghai, China).
Selection and identification of thermostable lipase-producing strain
One hundred and twenty-seven soil samples were collected from different location of Hubei province, China. Each soil sample (0.1 g) was subjected to cultivation in a 50-mL olive oil medium (olive oil 1.0 %, (NH4)2SO4 1.0 %, Na2HPO4 0.3 %, MgSO4, 0.01 %, pH 7.2) at 37 °C for 48 h with a shaking rate of 200 rpm. The samples of culture were streaked on triolein agar plates. The bacterial colonies forming transparent zones after incubation for 48 h at 37 °C were further purified by repeated streaking. After incubation in the olive oil medium, the clear supernatant, as crude extracellular lipase, was obtained by centrifugation at 8000 × g and 4 °C for 10 min. The crude intracellular lipase was obtained as follows. The harvested cells were resuspended in Tris–HCl buffer (50 mM, pH 7.2) and disrupted by sonication (240 W). The resultant cell debris was removed by centrifugation at 12,000 × g and 4 °C for 20 min, and the clear supernatant was used as intracellular lipase. The crude (intracellular and extracellular) lipase was incubated at 70 °C for 24 h, and then the remaining activity was determined at 50 °C by the colorimetric method using pNPO as substrate. The strain was identified based on morphological and 16S rDNA sequence by the method described previously (Hold et al. 2002).
Purification of intracellular LipXO
A 24-h growth of the isolated bacterial strain YB103 was inoculated into 100 mL medium (olive oil 1.0 %, yeast extract 1.0 %, tryptone 1.0 %, NaCl 0.5 %, (NH4)2SO4 1.0 %; pH 7.2) contained in 500 mL flask. The flask was incubated at 37 °C with shaking (200 rpm) for 48 h. Lipase was purified as we described previously with a little modification (Li et al. 2013). The crude intracellular lipase was ultrafiltrated using a 40 kDa cutoff centrifugal membrane filter (Millipore, USA). The resultant concentrated solution was precipitated by adding solid ammonium sulfate at saturation of 65 %, and maintained at 4 °C for 18 h. Then the solution was centrifuged at 8000 × g for 10 min. Pellet was collected and suspended in minimum volume of Tris–HCl buffer (25 mM, pH 7.2). The highly active fractions were pooled and applied on phenyl Sepharose column (75 cm × 1.25 cm, GE, USA) previously equilibrated with Tris–HCl buffer (25 mM, pH 7.2). The unbound proteins were eluted by same buffer while the bound proteins were eluted with (NH4)2SO4 gradient from 1.4 to 0.0 M. The column was run at flow rate of 0.5 mL/min. The resultant fractions were detected for lipase hydrolytic activity by the colorimetric method using pNPL as the substrate, and stored at 4 °C for further study. The lipase preparations (crude and purified lipase) were electrophoresed on a SDS-PAGE gel (10 % polyacrylamide).
Biochemical characterization
Effect of temperature on lipase activity and stability
Optimum temperature of LipXO was determined by assessing the hydrolytic activity at various temperatures (4–90 °C, at the interval of 10 °C) under standard conditions (50 mM CHES buffer, pH 9.0). To study thermostability, purified lipase was incubated at various temperatures (4, 50 and 70 °C) for up to 7 days. After the lipase solution was heated to appointed temperature, the pH value was adjusted to the desired value at this temperature on a Mettler Toledo S470-USP/EP Seven Excellence pH/Conductivity Meter (Schwerzenbach, Switzerland). Before the assay of activity, the temperature and pH of solution were adjusted to 70 °C and 9.0, respectively. Aliquots of enzyme solution were withdrawn at various time intervals, and the residual hydrolytic activity was measured by the colorimetric method using pNPL as the substrate at 70 °C.
Circular dichroism (CD) spectra
Circular dichroism spectroscopy was performed on a Jasco 1500 spectropolarimeter equipped with peltier-type temperature controller (Jasco, Inc., Easton, MD). Purified LipXO protein samples were prepared as described above and then diluted to 5.0 μmol/L in 50 mM CHES buffer (pH 9.0) for CD analysis in a 1.0-mm path length cell. Variable wavelength measurements of protein solutions were scanned at 25 °C from 180 to 260 nm with a scan rate 50 nm/min and data points were collected every 0.1 nm. The signal was averaged over six scans. Each spectra was acquired independently three times. Thermal denaturation experiments were conducted using 5 μmol/L protein in a 1.0 mm quartz cuvette. The protein samples were heated from 5 to 90 °C at a rate of 1.0 °C/min. Thermal denaturation profiles were displayed in molar ellipticity at 222 nm. For all experiments, 222 nm voltages were within accepted limits (<700 V at 216 nm and significantly lower for 222 nm), which allowed for monitoring of thermally induced loss of secondary structure by plotting ellipticity at 222 nm versus temperature. The results were analyzed with Standard Analysis software (JACSO) and expressed as mean residue molar ellipticity [θ].
Effect of pH on lipase activity and stability
The optimum pH of the purified lipase was investigated by measuring its hydrolytic activity at 50 °C at pH ranging from 7.0 to 12.0. The following buffers were used at 50 mM concentration; MOPS buffer for pH 6.5–7.5, Tricine buffer for pH 7.5–8.5, CHES buffer for pH 8.5–10.0, and CAPS buffer for pH 10.0–11.0. The pH stability was determined by incubating 1 mL purified lipase in the aforementioned buffers (20 mM) for 60 min at 70 °C, and its hydrolytic activity was detected in the presence of pNPL in 50 mM CHES buffer (pH 9.0) at 70 °C. The stability was determined as the relative activity to the control.
Effect of organic solvents on lipase stability
One milliliter of lipase solution (CHES buffer, 50 mM, pH 9.0) was mixed with 0.25 organic solvents at a final concentration of 20 % (v/v). The mixture was incubated at 70 °C and 200 rpm for 24 h. The residual activity was determined by the colorimetric method using pNPL as the substrate at 70 °C. The organic solvents used included dimethyl sulfoxide (DMSO), methanol, ethanol, acetonitrile, toluene, chloroform, n-hexane and n-heptane.
Substrate specificity
Various p-nitrophenyl esters containing different chain length of acyl group (C 2–C 18) were tested to determine the substrate specificity of the enzyme. The hydrolytic activity was assayed at 50 °C and pH 9.0 by the colorimetric method. Moreover, triacylglycerols were also selected as substrates for determination of substrate specificity. The lipase activity was measured at 70 °C and pH 9.0 by the titrimetric method the colorimetric method.
Transesterification of (R, S)-1-phenethyl alcohol
The lipase (5.0 mg/mL), (R,S)-1-phenethyl alcohol (100 mM) and vinyl acetate (100 mM) were mixed with anhydrous n-hexane (10 mL) in a 20-ml screw-capped vial, and the reaction mixture was shaken at 200 rpm and 70 °C for 12 h. The resulting mixture was sampled several times to determine enantiomeric excess (e.e.p) and conversion using GC analysis.
Analysis method
Lipase hydrolysis activity assay
In general, the reaction mixture contained 950 μL of CHES buffer (50 mM, pH 9.0), 45 μL of pNP-ester (40 mM) solution. After 5 min of incubation at 70 °C, the reaction was initiated by the addition of 5 μL of lipase solution. The enzyme reaction mixture was centrifuged at 12,000 × g for 30 s at room temperature, and the activity was measured at 410 nm on a spectrophotometer (Shimadzu, Japan) (Hess et al. 2008). The pNP-esters were beforehand dissolved in isopropanol to 40 mM. One enzyme unit (U) is defined as 1.0 μmol of pNP enzymatically released from the pNP-esters per minute at 70 °C.
Lipase activity of the samples was also determined with the pH–stat titrimetry (Glogauer et al. 2011). The reaction was carried out in a glass vessel thermostated at 70 °C containing 9.8 mL of substrate emulsion and 0.2 mL of lipase. Lipase activity was detected by the production of free fatty acids which were titrated automatically in a pH–stat titrator (Metrohm, Switzerland) with 0.05 M NaOH for 5 min. The triacylglycerol emulsion was prepared by mixing 75 mM triacylglycerol, 3 % (w/v) gum arabic, and 2.5 mM CHES buffer (pH 9.0). The solution was emulsified for 10 min and then for an additional 2 min immediately before use. One unit (U) of lipase activity was defined as the amount of lipase liberating 1 μmol equivalent of fatty acid from triacylglycerol per min at 70 °C.
GC analysis
The optical purity and conversion were determined by GC-7890 gas chromatography (Agilent Technologies, USA) equipped with FID detector and CP7501 Chiral column (50 m × 0.39 mm, 0.25 mm) from Varian Co. Ltd. (USA). n-Dodecane was used as an internal standard, and N 2 was used as carrier gas. The injector, detector and column temperatures were set at 260, 250 and 110 °C, respectively. Enantiomeric excess (e.e.p) and conversion were calculated as described by Chen et al. (1982).
Protein
The protein content was determined by the method of Bradford using bovine serum albumin as the standard (Bradford 1976).
Results and discussion
Screening and identification of strain producing thermostable lipase
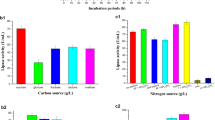
To isolating true lipase producers from microbial sources in preliminary screening plates, soil samples were incubated in the medium containing olive oil as sole carbon source (Hama et al. 2007). After cultivation at 37 °C for 48 h, the stability of intracellular and extracellular lipases from 127 soil samples was measured at 70 °C. From the strains tested, the crude intracellular lipase of strain YB103 displayed the highest stability, retaining 63 % residual activity after 24-h incubation at 70 °C for 24 h. So it was selected as the best thermostable lipase producer.
The strain YB103 was identified based on its 16S rDNA sequence (Hold et al. 2002). The BLAST search suggested a close relationship between strain YB103 (Genbank Accession No. KP313821) and the members of the X. oryzae pv. oryzae with a maximum sequence homology (99 %) and hereafter named X. oryzae pv. oryzae YB103. This strain is currently deposited in the China General Microbiological Culture Collection Center (CGMCC, Beijing, China), with the accession number CGMCC 10275.
Purification of intracellular LipXO
To exclude other intracellular proteins or enzymes which can disturb the properties of desired enzyme, the purification of enzyme was performed (Ahmed et al. 2010). In the present study, the strain X. oryzae pv. oryzae YB103 was cultivated in olive oil medium at 37 °C and 200 rpm for 48 h. The lipase hydrolytic activity (9.7 U/mL, pNPL as substrate) was observed after sonication disruption. The intracellular lipase from X. oryzae pv. oryzae YB103 (LipXO) was purified to homogeneity by a combined method of ultrafiltration concentration, ammonium sulfate precipitation and hydrophobic chromatography in sequence. The lipase was purified 101.1-fold with 15.7 % yield with a specific activity of 373.9 U/mg. A summary of the purification data is presented in Table 1. In electrophoretic analysis, LipXO migrated as a single band (about 58 kDa) corresponding to Coomassie Brilliant Blue staining of the purified LipXO fraction (Fig. 1). To the best of our knowledge, the known lipase from X. oryzae pv. oryzae has not been found to possess the same molecular mass (Aparna et al. 2007). The purified enzyme preparations were stored at 4 °C and were used to study its properties.
Thermostability and optimum temperature of LipXO
LipXO showed remarkable stability at high temperature in this study. The enzyme showed optimum temperature at 70 °C with no significant decrease at 80 °C (Fig. 2a). At 60 and 90 °C, the specific activity decreased to 329.5 U/mg (88.1 %, relative activity) and 275.1 U/mg (73.6 %, relative activity) of the maximum activity, respectively. However, less activity at lower temperatures could be due to the optimum temperature needed to trigger the lid opening of the lipase (Masomian et al. 2013). This profile is similar to those of thermostable lipases from Bacillus thermoleovorans ID-1 and Geobacillus sp. (Dong-Woo et al. 1999; Abdel-Fattah 2002). To study the effect of temperature on lipase thermostability, various incubation times and temperatures have been investigated. As a result shown in Fig. 2b, LipXO even retained 57.1 % (213.6 U/mg) of its residual activity after incubation at 90 °C for 30 min. The specific activity of a lipase from Geobacillus sp. Iso5 was 100.8 U/mg after incubation at 90 °C for 30 min (Mahadevan and Neelagund 2014).
Effects of temperature on the activity and stability of purified LipXO. a LipXO hydrolytic activity was determined at different temperatures (4–90 °C) in 50 mM CHES buffer (pH 9.0) using pNPL as the substrate. The lipase samples were withdrawn periodically to estimate the residual activities at 70 °C in 50 mM CHES buffer (pH 9.0) using pNPL as the substrate. b After pre-incubation of the lipase at 4–90 °C at an interval of 10 °C for 30 min, the remaining activity was determined at 70 °C in 50 mM CHES buffer (pH 9.0) using pNPL as the substrate. c Thermostability of purified LipXO for long period. The purified lipase (0.5 mg/mL) was incubated at different temperatures (filled squares 4 °C, filled downward triangles 50 °C and filled upward triangles 70 °C) for 7 days. Values represent the mean of three replicates
To comprehensively evaluate the long-term thermostability of LipXO, the lipase solution was pre-incubated at 4, 50, and 70 °C for 7 days before assay. Although good stability of bacterial and fungal lipases at high temperature for long period is rare (Haki and Rakshit 2003), LipXO retained 51.1 % (191.1 U/mg) of its original activity after incubation at 70 °C for 3 days (Fig. 2c). The hydrolytic activity of thermostable lipases from thermophilic anaerobic bacteria Thermoanaerobacter thermohydrosulfuricus and Caldanaerobacter subterraneus subsp. tengcongensis was 10.9 and 9.0 U/mg after incubated for 24 h at 70 °C (Royter et al. 2009). Thermostable lipase of Fervidobacterium changbaicum CBS-1 HSAUP0380006 showed 11.1 U/mg at 70 °C for 9 h incubation, respectively (Cai et al. 2011). Taken together, LipXO was designated a thermostable lipase. Although some work on lipases from X. oryzae pv. oryzae has been published (Aparna et al. 2007), there are no reports on a thermostable lipase from this organism.
The effect of temperature on the stability of LipXO was also determined by CD spectra. The thermal denaturation process was followed at 222 nm by CD in the range 30–95 °C (Fig. 3). Wavelength at 222 nm was set to detect α-helical to random coil transition, as they exhibited characteristic signals at this wavelength (Leow et al. 2007). The denaturation temperature (T m) is the temperature where half of the protein becomes denatured and is an estimate of protein stability (Reed et al. 2014). T m of the enzyme was 78.8 °C. This datum confirms the very high thermal stability of LipXO inferred from enzymatic activity measurements. Lipase has a hydrophobic β-sheet core surrounded by a hydrophilic α-helix surface (Carrasco-López et al. 2009). The catalytic residues of lipase were located in the hinge between β-sheet and α-helical protein domains (Nardini et al. 2000). Although CD spectra indicated that the lipase lost all its α-helical structure (Kirwan and Hodges 2014), LipXO might still have the β-sheet core structure (Khusainov et al. 2015). Thus, the results of CD spectra and thermostability suggested that the lipase could show catalytic ability in spite of losing almost all α-helical structure.
Effect of organic solvents on lipase stability
LipXO displayed high stability in all kinds of the hydrophilic and hydrophobic organic solvents tested in this study (Fig. 4). The effects of organic solvents on the activities of LipXO were determined by measuring residual activity after incubation in 20 % (v/v) solvents at 70 °C for 24 h. Although high stability of microbial lipases in hydrophilic solvents is rare (Doukyu and Ogino 2010; Zhao et al. 2008), the purified LipXO showed very stable in hydrophilic solvents with low logP values. It is noteworthy to point out that >100 % activity remained stable in presence of 20 % (v/v) methanol at 70 °C for 24 h, indicating LipXO a potential catalyst for chemical industry. Moreover, the high stability was observed in the presence of n-hexane with a 54.1 % (400.0 U/mg) increase in its activity compared to the control (259.5 U/mg, without n-hexane). Specific activity of LipXO increased to 294.6 U/mg (113.5 % of its original activity) after incubation in 20 % (v/v) n-heptane at 70 °C for 24 h. On the other hand, 194.8 U/mg and 250.1 U/mg of lipase activities were observed in the presence of 20 % (v/v) chloroform and toluene, respectively. Ogino et al. also reported very low activity of Pseudomonas aeruginosa LST-03 lipase in the presence of 25 % (v/v) solvents (Ogino et al. 2004). Thus, LipXO was an organic solvent-tolerant lipase.
Effect of pH on lipase activity and stability
LipXO turned out to be an alkaline lipase. The purified lipase exhibited high hydrolytic activity at the range of pH 6.5–11.0 with the maximum activity at pH 9.0 (Fig. 5a). At alkaline condition (pH 8.0–11.0), higher than 81.0 % of relative activity was detected compared to that at optimum pH 9.0. In pH stability experiment, the purified lipase showed good stability after 60-min incubation at pH 6.5–11.0, and the lipase even retained 74.1 % (249.0 U/mg) of residual activity at pH 11.0 (Fig. 5b). The specific activity of thermostable lipases from Staphylococcus aureus and Geobacillus sp. Iso5 was 39.2 U/mg and 37.9 U/mg at pH 11.0, respectively (Sarkar et al. 2012; Mahadevan and Neelagund 2014). The high activity and stability of LipXO over a wide alkaline pH suggests its usefulness in a range of industrial applications, such as the synthesis of biopolymers and biodiesel and the production of pharmaceuticals, agrochemicals, cosmetics, and flavor (Abdelkafi et al. 2009; Ahmed et al. 2010).
Effects of pH on the purified LipXO. a The lipase activity was detected in different buffers with varying pH values at 70 °C using pNPL as the substrate. Filled triangles 50 mM MOPS buffer (pH 6.5–7.5), filled diamonds 50 mM Tricine buffer (pH 7.5–8.5), filled circles 50 mM CHES buffer (pH 8.5–10.0) and filled squares CAPS buffer (pH 10.0–11.0). b The lipase was pre-incubated at 70 °C in 20 mM buffers with different pH values for 60 min, and the residual activity was determined at 70 °C in 50 mM CHES buffer (pH 9.0) using pNPL as the substrate
LipXO stereoselectivity
Stereoselectivity is one of the important characteristics exhibited by lipases, which is controlled by molecular properties of the enzyme (Colombo and Carrea 2002). Stereospecific lipases can catalyze the transesterification of S- or R-configuration secondary alcohol to form corresponding ester. The stereoselectivity of purified LipXO was investigated by transesterification of 1-phenethyl alcohol with vinyl acetate as acyl donor in n-hexane at 60 °C for 12 h (Scheme 1). After biocatalytic reaction, products and substrates were analyzed by chiral GC. The comparison, between the chromatograms before reaction and after 12 h transesterification, indicated that LipXO catalyzed transesterification of (R, S)-1-phenethyl alcohol to (R)-1-phenethyl acetate as main product. The reaction gave the corresponding (R)-ester product with 99 % enantiomeric excess (49 % conversion) and >200 enantioselectivity (E). However, several lipases from the Candida antarctica (Lee and Dordick 2002) and Burkholderia cepacia (Lee and Kim 2011), possessed low and moderate E (four and 101, respectively) toward (R, S)-1-phenethyl alcohol. These results indicate that LipXO was a stereoselective enzyme.
Substrate specificity of LipXO
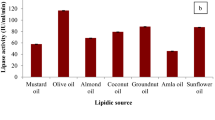
True lipases attack ester substrates that contain long-chain fatty acids (Glogauer et al. 2011; Reyes-Duarte et al. 2005). The substrate specificity of the purified LipXO was determined using p-nitrophenyl esters and triacylglycerols of varying acyl chain lengths at 70 °C and pH 9.0. The lipase demonstrated wide substrate specificity towards both classes of ester substrate (from C2 to C18) with preference to pNPL (C14, 373.9 U/mg) and glyceryl trimyristate (C14, 296.4 U/mg) (Fig. 6). The relatively high activities were observed on substrates with acyl chain length from C10 to C16, indicating LipXO showed preference to medium and long acyl chain lengths. Therefore, the lipase is a true lipase but not an esterase (Arpigny and Jaeger 1999; Reyes-Duarte et al. 2005).
References
Abdel-Fattah Y (2002) Optimization of thermostable lipase production from a thermophilic Geobacillus sp. using Box-Behnken experimental design. Biotechnol Lett 24:1217–1222
Abdelkafi S, Fouquet B, Barouh N, Durner S, Pina M, Scheirlinckx F, Villeneuve P, Carrière F (2009) In vitro comparisons between Carica papaya and pancreatic lipases during test meal lipolysis: potential use of CPL in enzyme replacement therapy. Food Chem 115:488–494
Ahmed EH, Raghavendra T, Madamwar D (2010) An alkaline lipase from organic solvent tolerant Acinetobacter sp. EH28: application for ethyl caprylate synthesis. Bioresour Technol 101:3628–3634
Aparna G, Chatterjee A, Jha G, Sonti RV, Sankaranarayanan R (2007) Crystallization and preliminary crystallographic studies of LipA, a secretory lipase/esterase from Xanthomonas oryzae pv. oryzae. Acta Crystallogr, Sect F Struct Biol Cryst Commun 63:708–710
Arpigny J, Jaeger K (1999) Bacterial lipolytic enzymes: classification and properties. Biochem J 343:177–183
Bradford MM (1976) A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem 72:248–254
Cai J, Xie Y, Song B, Wang Y, Zhang Z, Feng Y (2011) Fervidobacterium changbaicum Lip1: identification, cloning, and characterization of the thermophilic lipase as a new member of bacterial lipase family V. Appl Microbiol Biot 89:1463–1473
Cardenas F, de Castro MS, Sanchez-Montero JM, Sinisterra JV, Valmaseda M, Elson SW, Alvarez E (2001) Novel microbial lipases: catalytic activity in reactions in organic media. Enzyme Microb Technol 28:145–154
Carrasco-López C, Godoy C, de las Rivas B, Fernández-Lorente G, Palomo JM, Guisán JM, Fernández-Lafuente R, Martínez-Ripoll R, Hermoso JA (2009) Activation of bacterial thermoalkalophilic lipases is spurred by dramatic structural rearrangements. J Biol Chem 284:4365–4372
Chen CS, Fujimoto Y, Girdaukas G, Sih CJ (1982) Quantitative analyses of biochemical kinetic resolutions of enantiomers. J Am Chem Soc 104:7294–7299
Colombo G, Carrea G (2002) Modeling enzyme reactivity in organic solvents and water through computer simulations. J Biotechnol 96:23–33
Dandavate V, Jinjala J, Keharia H, Madamwar D (2009) Production, partial purification and characterization of organic solvent tolerant lipase from Burkholderia multivorans V2 and its application for ester synthesis. Bioresour Technol 100:3374–3381
Dheeman DS, Henehan G, Frías JM (2011) Purification and properties of Amycolatopsis mediterranei DSM 43304 lipase and its potential in flavour ester synthesis. Bioresour Technol 102:3373–3379
Dong-Woo L, You-Seok K, Jun KK, Byung-Chan K, HakJong C, Doo-sik K, Maggy T, Yu-ryang P (1999) Isolation and characterisation of thermophilic lipase from Bacillus thermoleovorans ID-1. FEMS Microbiol Lett 179:393–400
Doukyu N, Ogino H (2010) Organic solvent-tolerant enzymes. Biochem Eng J 48:270–282
Ebrahimpour A, Rahman RNZRA, Basri M, Salleh AB (2011) High level expression and characterization of a novel thermostable, organic solvent tolerant, 1, 3-regioselective lipase from Geobacillus sp. strain ARM. Bioresour Technol 102:6972–6981
Faoro H, Glogauer A, Couto GH (2012) Characterization of a new Acidobacteria-derived moderately thermostable lipase from a Brazilian Atlantic Forest soil metagenome. FEMS Microbiol Ecol 81:386–394
Franken B, Eggert T, Jaeger KE, Pohl M (2011) Mechanism of acetaldehyde-induced deactivation of microbial lipases. BMC Biochem 12:10–22
Glogauer A, Martini VP, Faoro H, Couto GH, Müller-Santos M, Monteiro RA, Mitchell DA, de Souza EM, Pedrosa FO, Krieger N (2011) Identification and characterization of a new true lipase isolated through metagenomic approach. Microb Cell Fact 10:54
Gotor-Fernández V, Busto E, Gotor V (2006) Candida antarctica lipase B: an ideal biocatalyst for the preparation of nitrogenated organic compounds. Adv Synth Catal 348:797–812
Haki GD, Rakshit SK (2003) Developments in industrially important thermostable enzymes: a review. Bioresour Technol 89:17–34
Hama S, Yamaji H, Fukumizu T, Numata T, Tamalampudi S, Kondo A, Noda H, Fukuda H (2007) Biodiesel-fuel production in a packed-bed reactor using lipase-producing Rhizopus oryzae cells immobilized within biomass support particles. Biochem Eng J 34:273–278
Hess M, Katzer M, Antranikian G (2008) Extremely thermostable esterases from the thermoacidophilic euryarchaeon Picrophilus torridus. Extremophiles 12:351–364
Hold GL, Pryde SE, Russell VJ, Furrie E, Flint HJ (2002) Assessment of microbial diversity in human colonic samples by 16S rDNA sequence analysis. FEMS Microbiol Ecol 39:33–39
Jaeger KE, Eggert T (2002) Lipases for biotechnology. Curr Opin. Biotech 13:390–397
Khusainov R, van Heel AJ, Lubelski J, Moll GN, Kuipers OP (2015) Identification of essential amino acid residues in the nisin dehydratase NisB. Frontiers Microbiol 6:1–8
Kirwan JP, Hodges RS (2014) Transmission of stability information through the N-domain of tropomyosin is interrupted by a stabilizing mutation (A109L) in the hydrophobic core of the stability control region (residues 97–118). J Biol Chem 289:4356–4366
Klibanov AM (2001) Improving enzymes by using them in organic solvents. Nature 409:241–246
Lee MY, Dordick JS (2002) Enzyme activation for nonaqueous media. Curr Opin Biotechnol 13:376–384
Lee JK, Kim MJ (2011) Ionic liquid co-lyophilized enzyme for biocatalysis in organic solvent: remarkably enhanced activity and enantioselectivity. J Mol Catal B Enzym 68:275–278
Leow TC, Rahman RNZRA, Basri M, Salleh AB (2007) A thermoalkaliphilic lipase of Geobacillus sp. T1. Extremophiles 11:527–535
Li M, Yang LR, Xu G, Wu JP (2013) Screening, purification and characterization of a novel cold-active and organic solvent-tolerant lipase from Stenotrophomonas maltophilia CGMCC 4254. Bioresour Technol 148:114–120
Lima VMG, Krieger N, Mitchell DA, Baratti JC, de Filippis I, Fontana JD (2004) Evaluation of the potential for use in biocatalysis of a lipase from a wild strain of Bacillus megaterium. J Mol Catal B Enzym 31:53–61
Mahadevan GD, Neelagund SE (2014) Thermostable lipase from Geobacillus sp. Iso5: bioseparation, characterization and native structural studies. J Basic Microb 54:386–396
Masomian M, Rahman RNZRA, Salleh AB, Basri M (2013) A new thermostable and organic solvent-tolerant lipase from Aneurinibacillus thermoaerophilus strain HZ. Process Biochem 48:169–175
Nardini M, Lang DA, Liebeton K, Jaeger KE, Dijkstra BW (2000) Crystal structure of Pseudomonas aeruginosa lipase in the open conformation THE prototype for family I. 1 of bacterial lipases. J Biol Chem 275:31219–31225
Ogino H, Mimitsuka T, Muto T, Matsumura M, Yasuda M, Ishimi K, Ishikawa H (2004) Cloning, expression, and characterization of a lipolytic enzyme gene (lip8) from Pseudomonas aeruginosa LST-3. J Mol Microbiol Biotechnol 7:212–223
Reed CJ, Bushnell S, Evilia C (2014) Circular dichroism and fluorescence spectroscopy of cysteinyl-tRNA synthetase from Halobacterium salinarum ssp. NRC-1 demonstrates that group I cations are particularly effective in providing structure and stability to this halophilic protein. PLoS One 9(3):e89452. doi:10.1371/journal.pone.0089452
Reyes-Duarte D, Polaina J, López-Cortés N, Alcalde M, Plou FJ, Elborough K, Ballesteros A, Timmis KN, Golyshin PN, Ferrer M (2005) Conversion of a carboxylesterase into a triacylglycerol lipase by a random mutation. Angew Chem Int Edit 117:7725–7729
Royter M, Schmidt M, Elend C, Höbenreich H, Schäfer T, Bornscheuer UT, Antranikian G (2009) Thermostable lipases from the extreme thermophilic anaerobic bacteria Thermoanaerobacter thermohydrosulfuricus SOL1 and Caldanaerobacter subterraneus subsp. tengcongensis. Extremophiles 13:769–783
Sarkar P, Yamasaki S, Basak S, Bera A, Bag PK (2012) Purification and characterization of a new alkali-thermostable lipase from Staphylococcus aureus isolated from Arachis hypogaea rhizosphere. Process Biochem 47:858–866
Tran DT, Yeh KL, Chen CL, Chang JS (2012) Enzymatic transesterification of microalgal oil from Chlorella vulgaris ESP-31 for biodiesel synthesis using immobilized Burkholderia lipase. Bioresour Technol 108:119–127
Vieille C, Zeikus GJ (2001) Hyperthermophilic enzymes: sources, uses, and molecular mechanisms for thermostability. Microbiol Mol Biol Rev 65:11–43
Zhao LL, Xu JH, Zhao J, Pan J, Wang ZL (2008) Biochemical properties and potential applications of an organic solvent-tolerant lipase isolated from Serratia marcescens ECU1010. Process Biochem 43:626–633
Acknowledgments
The authors thank the National Natural Science Foundation of China (No. 31401631, No. 31171649, No. 31271834, No. 31371824, No. 20936002), Fundamental Research Funds for the Central Universities (No. 2662014BQ054) and National Undergraduate Scientific and Technological Innovation Project (No. 20150504072) for the financial supports.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by F. Robb.
Rights and permissions
About this article
Cite this article
Mo, Q., Liu, A., Guo, H. et al. A novel thermostable and organic solvent-tolerant lipase from Xanthomonas oryzae pv. oryzae YB103: screening, purification and characterization. Extremophiles 20, 157–165 (2016). https://doi.org/10.1007/s00792-016-0809-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00792-016-0809-y