Abstract
Attention-deficit/hyperactivity disorder (ADHD) is a common neurodevelopmental disorder, but the underlying pathophysiological mechanisms of ADHD remain unclear. Gut microbiota has been recognized to influence brain function and behaviors. Therefore, this study aimed to determine whether imbalanced gut microbiomes identified by a 16S rRNA sequencing approach are involved in the pathophysiology of ADHD. We recruited a total of 30 children with ADHD (mean age: 8.4 years) and a total of 30 healthy controls (mean age: 9.3 years) for this study. The dietary patterns of all participants were assessed with the food frequency questionnaire. The microbiota of fecal samples were investigated using 16S rRNA V3V4 amplicon sequencing, followed by bioinformatics and statistical analyses. We found that the gut microbiota communities in ADHD patients showed a significantly higher Shannon index and Chao index than the control subjects. Furthermore, the linear discriminant analysis effect size (LEfSe) analysis was used to identify differentially enriched bacteria between ADHD patients and healthy controls. The relative abundance of Bacteroides coprocola (B. coprocola) was decreased, while the relative abundance of Bacteroides uniformis (B. uniformis), Bacteroides ovatus (B. ovatus), and Sutterella stercoricanis (S. stercoricanis) were increased in the ADHD group. Of all participants, S. stercoricanis demonstrated a significant association with the intake of dairy, nuts/seeds/legumes, ferritin and magnesium. B. ovatus and S. stercoricanis were positively correlated to ADHD symptoms. In conclusion, we suggest that the gut microbiome community is associated with dietary patterns, and linked to the susceptibility to ADHD.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Attention-deficit/hyperactivity disorder (ADHD) is a common neurodevelopmental disorder that occurs in childhood and can persist into adulthood. This disorder affects 3–10% of school-age children throughout the world [1] and may result in failures related to academic achievement, interpersonal relationships, and mental health [2, 3]. The underlying pathophysiological mechanisms of ADHD are multidimensional. In recent years, compelling evidence has revealed multicomponent bidirectional signaling pathways between the gut and the brain [4]. The concept of a “gut–brain axis”, which refers to a link between gut microbiota and brain function, has been applied to several neuropsychiatric disorders [5, 6]. Therefore, investigating the profiles of gut microbiota in patients with ADHD may provide us a new insight of the pathophysiology to this neurodevelopmental disorder [7].
The gut microbiota of humans is determined by genetic, epigenetic, and dietary factors [8]. Previous evidence has indicated that host–microbe interactions play a key role in brain development [9,10,11,12,13]. For example, both probiotics and prebiotics can have different impacts on the central nervous system (CNS) according to the administered microbial species or oligosaccharides [12]. A 3-year follow-up study revealed that children that received antibiotics in the first year of life subsequently had more behavioral difficulties and symptoms of depression during the follow-up [14]. Another study revealed that probiotic supplementation during the first 6 months of life may reduce the risk of neuropsychiatric disorder development (i.e., ADHD and autistic spectrum disorder) later in childhood [15]. Moreover, dietary habits and balanced nutrients may affect children’s behavior and learning [16]. Since ADHD is a neurodevelopmental disorder characterized by behavioral problems and learning disabilities, some researchers have proposed that ADHD may be associated with “unhealthy” diets, such as increased intake of sweetened desserts or fried foods [17,18,19]. Dietary habits play a key role in the modulation of gut microbiota composition [20]. However, the profile of gut microbiota and the potential relationship between diet and gut microbiota in children with ADHD continue to be poorly understood.
The development of high-throughput sequencing technology, such as advances in next-generation sequencing (NGS), has facilitated significant breakthroughs in microbial ecology studies [21]. NGS has also led to the establishment and rapid expansion of research in the field of “metagenomics”, which is often defined as the analysis of DNA from microbial communities in environmental samples without the need for culturing [22]. 16S ribosomal RNA (16S rRNA) is a component of the 30S small sub-unit of prokaryotic ribosomes [23]. Studies of 16S rRNA are useful phylogenetic markers for identifying bacterial taxa in a given sample [24, 25]. Many 16S rRNA sequencing statistical/computational tools and databases have since been developed to enable the utilization of the huge influx of data [26]. As a result, combining 16S rRNA sequencing data with a case–control format can provide candidate pathogens that may be associated with complex diseases and serve as a reference for further studies aimed at clarifying the pathogenesis of a disease [27]. Aarts et al. [28] investigated the profile of gut microbiota in patients with ADHD using 16S rRNA marker gene sequencing (16S) to identify bacterial taxa and their predicted gene functions. The researchers found no difference in alpha diversity between patients with ADHD and controls, but ADHD patients had a 12.7% to 20.5% increased Bifidobacterium genus. Moreover, the increased proportion of Bifidobacterium could predict the function of dopamine precursor synthesis and were associated with decreased neural responses to reward anticipation identified using functional magnetic resonance imaging (fMRI). To date, few studies have investigated the role of gut microbiota in ADHD using a 16S rRNA sequencing approach [28,29,30].
A comprehensive analysis of human gut microbiota would be helpful for revealing the mechanisms of these host–microbe interactions. We hypothesize that gut microbiota profiles may differ considerably between ADHD patients and healthy control subjects. Furthermore, since dietary intake plays a vital role in modulating microbiota composition, we propose that dietary habits are involved in gut microbiome and are thus further connected to the pathogenesis of ADHD. This study aimed to determine whether an imbalance of gut microbiomes was involved in the pathophysiology of ADHD using the 16S rRNA sequencing approach.
Methods
Study participants
The Institutional Review Board (IRB) at Chang Gung Memorial Hospital in Taiwan approved our research protocol. This study consisted of eligible patients with ADHD treated in the outpatient Department of Child Psychiatry at Chang Gung Children’s Hospital in Taiwan and healthy control children. We obtained the written informed consent from the parents or guardians of the participants as required by the IRB prior to the start of this study.
The criteria for ADHD patients consisted of the following: (1) a clinical diagnosis of ADHD by a senior child psychiatrist based on the criteria provided in the Diagnostic and Statistical Manual of Mental Disorders, Fourth Edition, Text Revision (DSM-IV-TR) through structured interviews conducted based on the Chinese version of the schedule for affective disorders and schizophrenia for school-age children, epidemiologic version (K-SADS-E) [31]; (2) aged between 6 and 16 years; and (3) must have never taken any medications to treat their ADHD. We excluded the following from our study: (1) patients with a history of neuropsychiatric diseases or major physical illnesses (such as intellectual disabilities, autism spectrum disorder, bipolar disorders, major depressive disorders, psychotic disorders, substance dependence, epilepsy, severe head trauma, or gastrointestinal disorders); and (2) patients who are vegetarians or were currently taking probiotics or antibiotics.
The healthy control subjects were children without ADHD between the ages of 6 and 16 years within the same catchment area. They had no known major physical illnesses or any of the aforementioned major neuropsychiatric diseases. We also excluded those currently taking probiotics or antibiotics from the healthy control group.
Sample collection
We collected fecal samples from both ADHD patients and healthy controls using the standard method of scooping a pea-sized piece of feces, placing it in a 50-ml Falcon tube, and storing it at 4 °C immediately after collection and then at − 80 °C within 24 h. Total DNA extraction of the fecal samples was carried out using a QIAamp® DNA Stool Mini Kit (QIAGEN, Tokyo, Japan) following the manufacturer’s instructions. To increase the recovery of bacterial DNA, particularly from Gram-positive bacteria, we pretreated the samples with lytic enzymes using the stool kit prior to extraction. Briefly, 100 mg of fecal sample was suspended in 10 mL of Tris–EDTA buffer (pH 7.5), and 50 μL of 100 mg/mL lysozyme type VI purified from chicken egg white (MPBIO, Derby, UK) and 50 μL of 1 mg/mL purified achromopeptidase (Wako, Osaka, Japan) were added. The solution was incubated at 37 °C for 1 h with mixing, 0.12 g of sodium dodecyl sulfate (final conc. 1%) was added, and the suspension was mixed until clear. Next, 100 μL of 20 mg/mL proteinase K (Wako) was added, followed by incubation at 55 °C for 1 h with mixing. The cell lysate was then subjected to ethanol precipitation. The precipitant was dissolved in 1.6 mL of ASL buffer from the stool kit and subsequently purified using the QIAamp® DNA Stool Mini Kit (QIAGEN).
16S rRNA amplicon sequencing
The DNA samples were subjected to the first-run PCR reaction, where the specified forward primer (TCGTCGGCAGCGTCAGATGTGTATAAGAGACAGCCTACGGGNGGCWGCAG) and reverse primer (GTCTCGTGGGCTCGGAGATGTGTATAAGAGACAGGACTACHVGGGTATCTAATCC) were designed to amplify the V3–V4 genomics region of bacterial 16S rRNA genes. Using approximately 550 bp PCR products confirmed with gel electrophoresis, the products were then subjected to library preparation for 16S rRNA sequencing. We prepared a DNA library according to the 16S rRNA Sequencing Library Preparation instructions (Illumina, California, USA). The prepared amplicons were sequenced on the MiSeq sequencer (Illumina, California, USA) using the 600-cycle sequencing reagent and specifying the pair-end mode.
Clinical measurements
A senior psychiatrist used the K-SADS-E diagnostic tool [31] to conduct interviews with all the participants in both the ADHD patient group and the control group. The K-SADS-E is a semistructured diagnostic interview that is designed to assess current and past episodes of psychopathology in children and adolescents according to DSM-IV criteria [31]. The K-SADS-E is administered by interviewing the parent(s) and the child and finally acquiring summary ratings that include all sources of information. The validity and reliability of the Chinese version of K-SADS-E has been established in Taiwan [32].
Furthermore, an experienced child psychologist conducted the Wechsler Intelligence Scale for Children—Fourth Edition (WISC-IV) [33] with individual patients in a room designed to reduce variability in testing conditions. The Swanson, Nolan, and Pelham Version IV Scale (SNAP-IV) parent form and SNAP-IV teacher form were completed by the patients’ parents and teacher, respectively [34,35,36]. Patients were also interviewed by a clinician using the ADHD rating scale (ADHD-RS) [37].
Dietary pattern assessment
We adopted a validated food frequency questionnaire (FFQ) to evaluate participants’ dietary intake in the previous year [38]. This questionnaire included the frequencies and amounts of 49 food items consumed from eight food groups. The FFQ had a fixed format and open-ended questions with regard to major staple foods, oils/fats, sugars, and supplements and also included their frequency of consumption. Standard portion size and frequency were recorded to estimate the daily intake of foods and nutrients. We calculated daily dietary patterns and nutrients intake levels by multiplying the amount of food eaten daily, the level estimated from frequency, and nutrient concentrations. We determined calorie intake level by summing the calories from the food and beverages consumed. All nutrients were calorie-adjusted using the residue method. The validity of FFQ has been verified in our previous study [39].
Statistical and bioinformatics analysis
We analyzed data with the statistical software package SPSS, version 16.0 (SPSS Inc., Chicago, IL, USA). Variables were presented as either the mean (standard deviation) or frequency. Two-tailed p values < 0.05 were considered statistically significant. We applied the Chi-square test or Fisher’s exact test to compare gender distribution between the ADHD patients and the controls. Furthermore, we used an independent t test or Mann–Whitney U test to determine the potential differences in age and clinical assessments between the ADHD patients and healthy controls. Multiple linear regression was performed to analyze the relationships between the relative abundance of bacteria, dietary patterns, and ADHD clinical symptoms, respectively (controlling for age, sex, body mass index and FSIQ).
16S rRNA gene amplicon sequence results were analyzed using Mothur v1.39.5 [40] in accordance with the MiSeq SOP [41]. In short, the 16S rRNA V3–V4 sequencing reads were initially demultiplexed using MiSeq Reporter v2.6. The demultiplexed paired reads were then assembled into a single contig with the following parameters: minimum length of 405 bp, maximum length of 428 bp, and no ambiguity. The single contig consisted of the effective reads from all samples clustered into OTUs based on a 97% sequence similarity according to Illumina MiSeq SOP [41] and following the steps described here: (1) sequencing filtering and trimming to de-replicate the working sequence set and align it with the SILVA bacteria reference 16S alignment (release 132) distributed with Mothur (v1.38.1) [40]. (2) Sequencing error reduction. (3) PCR chimera removal after screening with UCHIME (v4.2) [42]. (4) Taxonomy assignment of sequences. We ran the Mothur implementation of the Naïve Bayesian Classifier [43] against the homemade RDP rRNA training set (v9) to create a taxonomic assignment for every sequence with a minimum bootstrap confidence score of 80%. (5) Clustering into OTU. The clustering of sequences was carried out at a threshold identity of 0.03% using the average neighbor algorithm.
We performed alpha rarefaction analysis, including observed OTUs, Chao1, Shannon, Simpson, and ACE index, using custom R scripts and data acquired from Mothur output files or calculated by QIIME [44]. Beta diversity analysis, which included weighted and unweighted UniFrac distance metrics calculated using QIIME and PCoA plots were generated using R scripts. Finally, we used linear discriminant analysis effect size (LEfSe) [45] to determine the taxa that most likely explains the differences between the experimental samples and the control samples.
Results
Demographic data
We collected fecal samples from 30 children with ADHD (mean age: 8.4 years, 76.7% male) and 30 healthy controls (mean age: 9.3 years, 60% male). The characteristics of the ADHD patients and healthy controls are summarized in Table 1. Patients with ADHD were significantly shorter, had lower scores in the full scale intelligence quotient (FSIQ), perceptual reasoning index (PRI), working memory index (WMI), and processing speed index (PSI), and displayed a greater severity of inattention and hyperactivity/impulsivity symptoms as rated by parents (SNAP-IV parent form), teachers (SNAP-IV teacher form), and clinicians (ADHD-RS).
We extracted total DNA samples from the fecal samples and analyzed them for 16S rRNA V3V4 amplicon sequencing and bacterium category identification. The average number of raw paired reads per sample was 266,022 ± 21,720 reads for ADHD patients and 373,471 ± 26,496 reads for healthy controls (Supplementary Table 1). After selecting the qualified reads, the average quality-filtered reads per sample were 54,670 ± 5296 for ADHD and 70,099 ± 4544 for healthy controls.
Alpha diversity and beta diversity
Microbial diversity was assessed either within a community (alpha diversity) or between the collection of samples (beta diversity). We calculated three different values to assess the alpha diversity: “Chao1” estimates species abundance; observed OTU estimates the amount of unique OTUs found in each sample, and “Shannon index” accounts for both richness and evenness. Gut microbiota communities from the ADHD patients demonstrated Shannon index (p = 0.0378) and Chao index (p = 0.0351) that were significantly increased compared to the healthy controls (Fig. 1a, b and Supplementary Table 2). The Simpson index was significantly lower in ADHD patients compared with healthy controls (Fig. 1c). The ACE was similar between the ADHD and control groups (Supplementary Table 2).
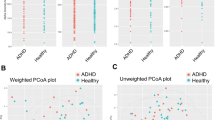
The beta diversity was assessed using a principal coordinate analysis (PCoA) plot for healthy controls and ADHD patients. The unweighted unifrac and weighted unifrac PCoA plots indicated that the gut microbiomes were similar between the healthy controls and ADHD patients (Fig. 1d). Overall, these results indicated that the gut microbiota communities were similar between ADHD patients and healthy controls.
Gut microbiome profiling
The gut microbiome dataset from healthy controls and ADHD patients revealed a total of nine phyla, five of which accounted for 99% of all the bacteria (Table 2). The healthy controls demonstrated a dominance of Bacteroidetes (73.68%), followed by Firmicutes (11.71%), Proteobacteria (8.21%), Fusobacteria (0.02%), and Actinobacteria (0.33%). The bacteria composition at the phylum level of the ADHD group closely resembled that of the healthy controls. The abundance of Fusobacteria was higher in the ADHD group (median = 0.28%) than in the healthy control group (mean = 0.02%) (p = 0.041).
The top ten most abundant genera are shown in Table 3. At the genus level, the top five genera were Bacteroides, Prevotella, Parabacteroides, Phascolarctobacterium, and Escherichia Shigella. The abundance of Fusobacterium was elevated in the ADHD group (0.28%) compared to the healthy control group (0.02%) (p = 0.041). Overall, the bacteria profiles of the healthy controls and ADHD patients were similar.
LEfSe analysis
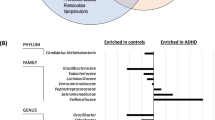
We used the LEfSe analysis to identify the specific bacteria phylotypes that were differentially altered between the healthy controls and ADHD patients. The LEfSe plot as a result of the ADHD and control groups is shown in Fig. 2a. The relative abundance of Bacteroides coprocola (B. coprocola) in the ADHD group was significantly lower than in the control group, while the relative abundance of Bacteroides uniformis (B. uniformis), Bacteroides ovatus (B. ovatus), and Sutterella stercoricanis (S. stercoricanis) in the ADHD group were significantly higher than in the control group (Fig. 2b). The distribution of enriched bacteria was also identified at genus level using LEfSe analysis (Supplementary Fig. 1). The relative abundance of Fusobacterium was enriched in ADHD group; otherwise, the relative abundance of Lactobacillus was enriched in healthy controls.
Supplementary Table 3 lists the dietary and nutrient status between the ADHD group and control group. Compared to the control children, children with ADHD demonstrated a higher intake proportion of refined grains (p = 0.027) and a lower proportion of dairy (p = 0.020) and vitamin B2 (p = 0.033). Table 4 shows the correlations between the relative abundance of bacteria, dietary patterns, and ADHD symptoms among all participants (ADHD patients and controls). We found that the amounts of S. stercoricanis had a significant correlation to the intake of dairy, nuts/seeds/legumes, ferritin and magnesium. B. uniformis was correlated to fat and carbohydrate intake. However, B. ovatus and B. coprocola was not associated with any of the dietary patterns identified by the FFQ. The relative abundance of B. ovatus, and S. stercoricanis had a positive correlation with ADHD symptoms, while B. uniformis and B. coprocola levels were not significantly correlated with ADHD symptoms.
Discussion
We used a 16S rRNA sequencing platform to screen the bacteria communities in the fecal samples of both ADHD children and healthy subjects. The gut microbiota communities in ADHD showed a significantly higher alpha (Shannon index and Chao index) than the control subjects. The Simpson index was significantly reduced in ADHD patients. The relative abundance of Fusobacterium and Lactobacillus was enriched in ADHD group and controls, respectively. We found significant differences in four species (B. uniformis, B. ovatus, B. coprocola, and S. stercoricanis) between the ADHD group and the control group using LEfSe analysis. Furthermore, the relative abundances of B. uniformis, B. ovatus, and S. stercoricanis were correlated with ADHD symptoms and dietary indices. We therefore suggest that the four previously mentioned bacterial species may serve as potential microbiota markers for ADHD.
According to a previous study [24], 2000 Illumina sequence reads for each sample are sufficient to estimate unbiased relative abundances of bacterial species. In this study, our samples averagely reached over 50,000 sequence reads, much more than the suggested 2000. Therefore, our samples should have reached unbiased estimates of relative abundances of bacterial species in both control and case sets. We found that the Shannon index, Chao index and Simpson index were significantly different between ADHD patients and healthy controls. However, the ACE index, a measure of richness which is similar with Chao1, did not show group differences. The inconsistent trends of Chao1 and ACE index may be explained by discrepancy in the calculation equations of these two indexes [46], or the marginal difference of the diversity of gut microbiome between patients with ADHD and healthy controls. A previous case–control study [28] found no significant difference in alpha diversity (using the same metrics as in the current paper) nor in relative abundance of Fusobacterium between ADHD patients and healthy controls. In addition, Aarts et al. [28] discovered that young adults with ADHD showed an increased amount in the Bifidobacterium genus. However, we observed no difference in the Bifidobacterium genus between the ADHD group and the control group. One possible explanation for these discrepancies is that the population recruited in the aforementioned study was adults with ADHD, while our study population was children with ADHD. In a hypothesis-driven approach, Aarts et al. [28] targeted candidate microbial taxa associated with dopamine neurotransmission based on pathway analysis and predicted bacterial gene function. Clinical and preclinical data have shown that gut microbiota can regulate complex physiological reactions, such as the expression of BDNF [47], HPA axis responsiveness [48], and the immune system [49]. The microbiome poses peripheral immune homeostasis and predisposes host susceptibility to neuropsychiatric disorders [50, 51]. Therefore, continuous research is necessary to clarify whether the role of gut microbiota in the pathogenesis of ADHD is moderated by immunological function.
Bacteroides are commonly found in the human intestine where they participate in a symbiotic host–bacterial relationship with humans. They help break down food and produce valuable nutrients and energy that the body needs [52]. B. uniformis is a putative bacterial species associated with the degradation of the isoflavone genistein in human feces. In high degraders, B. uniformis may be a candidate for genistein degradation based on fecal isoflavone degradation in the presence of these species [53]. Although part of the normal human gut flora, B. ovatus is the least common of the Bacteroides intestinal isolates. In one comprehensive 16S rRNA sequence-based enumeration of the colonic microbiota of three healthy adult humans, it represents, on average, 0.034% of all 16S rRNA sequences and 0.041% of the sequences in its division. This organism is occasionally isolated from clinical specimens and can also be isolated from poultry [54,55,56]. B. coprocola produces many extracellular enzymes that assist in the breakdown of such complex plant polysaccharides as cellulose and hemicellulose, as well as host-derived polysaccharides like mucopolysaccharides [57]. Our findings agree with that by Aarts et al. [28] which observed overrepresentation of B. uniformis and B. ovatus in ADHD patients. Bacteroides were found to be associated with development of the frontal lobe, cerebellum, and hippocampus region in healthy women [58]. We propose that the aforementioned three bacterial species may potentially alter development and function of the brain and play a role in the pathophysiology of ADHD.
Sutterella stercoricanis is a Gram-negative, oxidase- and catalase-negative, anaerobic and microaerophilic, non-spore-forming, rod-shaped bacterium from the genus Sutterella in the Sutterellaceae family and has been isolated from canine feces [59]. One previous study demonstrated that Sutterella is a major component of the microbiota in more than half of autistic children [60]. Our results revealed that S. stercoricanis was strongly correlated with diet (dairy, nuts/seeds/legumes, ferritin and magnesium). Our previous study has found that multidimensional allergic and nutritional factors may play a role in the pathophysiology of ADHD [61, 62]. The current study result suggests that S. stercoricanis may be associated with dietary patterns and linked to the susceptibility to ADHD. Although the causal inference of gut microbiota, diet and ADHD characteristics needs further clarification, S. stercoricanis may serve as a preferred target of dietary modification for treatment of ADHD.
We found that ADHD children and healthy controls have different dietary and nutrient patterns. Compared to the control group, the ADHD group demonstrated a higher proportion of refined grains intake, a lower proportion of dairy and vitamin B2 intake. These findings were mostly comparable with our previous work [39]. Taken together, the results herein support the association between ADHD and unhealthy diets or nutrient deficiencies. Unhealthy diet such as high fat and sugar consumption may change the healthy microbiota composition which leads to an imbalanced microbial population in the gut, a phenomenon known as “gut dysbiosis” [63]. It is possible that the gut microbiota species we identified are related to the alterations in dietary intake in patients with ADHD. It warrants further investigation whether the four species we have found in this study are reliable biomarkers of ADHD of their own right, or just an outcome fluctuated with diet/nutrient patterns. Our study showed that the OTUs that correlated with dietary pattern in the regression models did significantly differentiate cases from controls. The difference in the abundance of bacterial species between ADHD and controls may be related to both the pathophysiology and the dietary pattern of patients with ADHD. Future studies are warranted to unravel whether specific ADHD-symptom-related bacteria may be changed in abundance by changes in diet pattern or supplements including vitamins/pro-/pre-/syn-biotics.
This study has several limitations that should be noted. First, this study was conducted using a cross-sectional method. Even though we identified four microbiota markers between ADHD patients and healthy control subjects, early life events during initial colonization and microbiota development (e.g., birth delivery mode, breastfeeding or not, previous antibiotic use, and dietary patterns) can influence general and mental health later in life [64,65,66]. This study was unable to depict the longitudinal change and associated factors of gut microbiota. Second, while we observed that four bacterial species were correlated with ADHD clinical symptoms, the mechanisms underpinning the association between gut microbiota and ADHD have yet to be clearly explained. Third, vegetarians were excluded in the current study. Our study result may not be applicable in those with different diet patterns such as gluten free, vegan or specific diets like paleo, low carb. Future studies may be needed to explore patterns of microbiome in these specific dietary groups. Moreover, the patient and control groups were not perfectly age- and sex-matched. We excluded patients with major neurodevelopmental comorbidities to eliminate the potential confounding effect of comorbidities. According to the normative data of the Chinese version of the SNAP-IV [67], the symptom severity of our ADHD samples were relatively low compared to typical ADHD patients. The study sample may not represent overall ADHD population. Finally, we found that the most abundant phylum is Bacteroidetes followed by Firmicutes both in the ADHD and in the control group in Taiwan. However, a previous study indicated that the most abundant phylum was Firmicutes among the Singapore Chinese adult population [68]. Therefore, the gut microbiota community may be varied across different age groups or socio-environmental circumstance. The difference in ethnic populations may also contribute to the different results in the Aarts et al. [28] study. It warrants further verification whether our finding can be generalized to ADHD children in other ethnic populations.
In conclusion, we found different variability of gut microbiome in alpha diversity and our findings suggest that increased proportion of B. uniformis, B. ovatus, and S. stercoricanis and decreased proportion of B. coprocola may be associated with susceptibility to ADHD. This indicates that gut microbiome dysbiosis may be associated with dietary habit, and potentially alter the pathophysiology of ADHD. However, the mechanisms supporting the association between gut microbiota and ADHD still require further investigation.
References
Polanczyk GV, Willcutt EG, Salum GA, Kieling C, Rohde LA (2014) ADHD prevalence estimates across three decades: an updated systematic review and meta-regression analysis. Int J Epidemiol 43(2):434–442
American Psychiatric Association (2000) Diagnostic and statistical manual of mental disorders (DSM-IV-TR). American Psychiatric Association, Washington
Spencer TJ, Biederman J, Mick E (2007) Attention-deficit/hyperactivity disorder: diagnosis, lifespan, comorbidities, and neurobiology. J Pediatr Psychol 32(6):631–642
Stilling RM, Dinan TG, Cryan JF (2014) Microbial genes, brain and behaviour—epigenetic regulation of the gut–brain axis. Genes Brain Behav 13(1):69–86
Montiel-Castro AJ, Gonzalez-Cervantes RM, Bravo-Ruiseco G, Pacheco-Lopez G (2013) The microbiota-gut–brain axis: neurobehavioral correlates, health and sociality. Front Integr Neurosci 7:70
Lima-Ojeda JM, Rupprecht R, Baghai TC (2017) “I Am I and My Bacterial Circumstances”: linking gut microbiome, neurodevelopment, and depression. Front Psychiatry 8:153
Sandgren AM, Brummer RJM (2018) ADHD-originating in the gut? The emergence of a new explanatory model. Med Hypotheses 120:135–145
Yadav M, Verma MK, Chauhan NS (2018) A review of metabolic potential of human gut microbiome in human nutrition. Arch Microbiol 200(2):203–217
Borre YE, O’Keeffe GW, Clarke G, Stanton C, Dinan TG, Cryan JF (2014) Microbiota and neurodevelopmental windows: implications for brain disorders. Trends Mol Med 20(9):509–518
Cenit MC, Nuevo IC, Codoner-Franch P, Dinan TG, Sanz Y (2017) Gut microbiota and attention deficit hyperactivity disorder: new perspectives for a challenging condition. Eur Child Adolesc Psychiatry 26(9):1081–1092
Dore J, Blottiere H (2015) The influence of diet on the gut microbiota and its consequences for health. Curr Opin Biotechnol 32:195–199
Principi N, Esposito S (2016) Gut microbiota and central nervous system development. J Infect 73(6):536–546
Felice VD, O’Mahony SM (2017) The microbiome and disorders of the central nervous system. Pharmacol Biochem Behav 160:1–13
Slykerman RF, Thompson J, Waldie KE, Murphy R, Wall C, Mitchell EA (2017) Antibiotics in the first year of life and subsequent neurocognitive outcomes. Acta Paediatr 106(1):87–94
Partty A, Kalliomaki M, Wacklin P, Salminen S, Isolauri E (2015) A possible link between early probiotic intervention and the risk of neuropsychiatric disorders later in childhood: a randomized trial. Pediatr Res 77(6):823–828
Park S, Cho SC, Hong YC, Oh SY, Kim JW, Shin MS, Kim BN, Yoo HJ, Cho IH, Bhang SY (2012) Association between dietary behaviors and attention-deficit/hyperactivity disorder and learning disabilities in school-aged children. Psychiatry Res 198(3):468–476
Stevenson J (2006) Dietary influences on cognitive development and behaviour in children. Proc Nutr Soc 65(4):361–365
Sinn N (2008) Nutritional and dietary influences on attention deficit hyperactivity disorder. Nutr Rev 66(10):558–568
Millichap JG, Yee MM (2012) The diet factor in attention-deficit/hyperactivity disorder. Pediatrics 129(2):330–337
Bibbo S, Ianiro G, Giorgio V, Scaldaferri F, Masucci L, Gasbarrini A, Cammarota G (2016) The role of diet on gut microbiota composition. Eur Rev Med Pharmacol Sci 20(22):4742–4749
Wang WL, Xu SY, Ren ZG, Tao L, Jiang JW, Zheng SS (2015) Application of metagenomics in the human gut microbiome. World J Gastroenterol 21(3):803–814
Bragg L, Tyson GW (2014) Metagenomics using next-generation sequencing. Methods Mol Biol 1096:183–201
Janda JM, Abbott SL (2007) 16S rRNA gene sequencing for bacterial identification in the diagnostic laboratory: pluses, perils, and pitfalls. J Clin Microbiol 45(9):2761–2764
Caporaso JG, Lauber CL, Walters WA, Berg-Lyons D, Lozupone CA, Turnbaugh PJ, Fierer N, Knight R (2011) Global patterns of 16S rRNA diversity at a depth of millions of sequences per sample. Proc Natl Acad Sci USA 108(Suppl 1):4516–4522
Culligan EP, Marchesi JR, Hill C, Sleator RD (2014) Combined metagenomic and phenomic approaches identify a novel salt tolerance gene from the human gut microbiome. Front Microbiol 5:189
Gilbert JA, Dupont CL (2011) Microbial metagenomics: beyond the genome. Ann Rev Mar Sci 3:347–371
Oulas A, Pavloudi C, Polymenakou P, Pavlopoulos GA, Papanikolaou N, Kotoulas G, Arvanitidis C, Iliopoulos I (2015) Metagenomics: tools and insights for analyzing next-generation sequencing data derived from biodiversity studies. Bioinform Biol Insights 9:75–88
Aarts E, Ederveen THA, Naaijen J, Zwiers MP, Boekhorst J, Timmerman HM, Smeekens SP, Netea MG, Buitelaar JK, Franke B, van Hijum S, Arias Vasquez A (2017) Gut microbiome in ADHD and its relation to neural reward anticipation. PLoS One 12(9):e0183509
Jiang HY, Zhou YY, Zhou GL, Li YC, Yuan J, Li XH, Ruan B (2018) Gut microbiota profiles in treatment-naive children with attention deficit hyperactivity disorder. Behav Brain Res 347:408–413
Prehn-Kristensen A, Zimmermann A, Tittmann L, Lieb W, Schreiber S, Baving L, Fischer A (2018) Reduced microbiome alpha diversity in young patients with ADHD. PLoS One 13(7):e0200728
Kaufman J, Birmaher B, Brent D, Rao U, Flynn C, Moreci P, Williamson D, Ryan N (1997) Schedule for affective disorders and schizophrenia for school-age children-present and lifetime version (K-SADS-PL): initial reliability and validity data. J Am Acad Child Adolesc Psychiatry 36(7):980–988
Gau SF, Soong WT (1999) Psychiatric comorbidity of adolescents with sleep terrors or sleepwalking: a case-control study. Aust N Z J Psychiatry 33(5):734–739
Baron IS (2005) Test review: wechsler Intelligence Scale for Children-Fourth Edition (WISC-IV). Child Neuropsychol 11(5):471–475
Bussing R, Fernandez M, Harwood M, Wei H, Garvan CW, Eyberg SM, Swanson JM (2008) Parent and teacher SNAP-IV ratings of attention deficit hyperactivity disorder symptoms: psychometric properties and normative ratings from a school district sample. Assessment 15(3):317–328
Gau SS, Lin CH, Hu FC, Shang CY, Swanson JM, Liu YC, Liu SK (2009) Psychometric properties of the Chinese version of the Swanson, Nolan, and Pelham, Version IV Scale-Teacher Form. J Pediatr Psychol 34(8):850–861
Gau SS, Shang CY, Liu SK, Lin CH, Swanson JM, Liu YC, Tu CL (2008) Psychometric properties of the Chinese version of the Swanson, Nolan, and Pelham, version IV scale—parent form. Int J Methods Psychiatr Res 17(1):35–44
Zhang S, Faries DE, Vowles M, Michelson D (2005) ADHD Rating scale IV: psychometric properties from a multinational study as a clinician-administered instrument. Int J Methods Psychiatr Res 14(4):186–201
Lee MS, Pan WH, Liu KL, Yu MS (2006) Reproducibility and validity of a Chinese food frequency questionnaire used in Taiwan. Asia Pac J Clin Nutr 15(2):161–169
Chou WJ, Lee MF, Hou ML, Hsiao LS, Lee MJ, Chou MC, Wang LJ (2018) Dietary and nutrient status of children with attention-deficit/hyperactivity disorder: a case-control study. Asia Pac J Clin Nutr 27(6):1325–1331
Schloss PD, Westcott SL, Ryabin T, Hall JR, Hartmann M, Hollister EB, Lesniewski RA, Oakley BB, Parks DH, Robinson CJ, Sahl JW, Stres B, Thallinger GG, Van Horn DJ, Weber CF (2009) Introducing mothur: open-source, platform-independent, community-supported software for describing and comparing microbial communities. Appl Environ Microbiol 75(23):7537–7541
Kozich JJ, Westcott SL, Baxter NT, Highlander SK, Schloss PD (2013) Development of a dual-index sequencing strategy and curation pipeline for analyzing amplicon sequence data on the MiSeq Illumina sequencing platform. Appl Environ Microbiol 79(17):5112–5120
Edgar RC, Haas BJ, Clemente JC, Quince C, Knight R (2011) UCHIME improves sensitivity and speed of chimera detection. Bioinformatics 27(16):2194–2200
Wang Q, Garrity GM, Tiedje JM, Cole JR (2007) Naive Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl Environ Microbiol 73(16):5261–5267
Caporaso JG, Kuczynski J, Stombaugh J, Bittinger K, Bushman FD, Costello EK, Fierer N, Pena AG, Goodrich JK, Gordon JI, Huttley GA, Kelley ST, Knights D, Koenig JE, Ley RE, Lozupone CA, McDonald D, Muegge BD, Pirrung M, Reeder J, Sevinsky JR, Turnbaugh PJ, Walters WA, Widmann J, Yatsunenko T, Zaneveld J, Knight R (2010) QIIME allows analysis of high-throughput community sequencing data. Nat Methods 7(5):335–336
Segata N, Izard J, Waldron L, Gevers D, Miropolsky L, Garrett WS, Huttenhower C (2011) Metagenomic biomarker discovery and explanation. Genome Biol 12(6):R60
Morgan XC, Huttenhower C (2012) Chapter 12: human microbiome analysis. PLoS Comput Biol 8(12):e1002808
Bercik P, Denou E, Collins J, Jackson W, Lu J, Jury J, Deng Y, Blennerhassett P, Macri J, McCoy KD, Verdu EF, Collins SM (2011) The intestinal microbiota affect central levels of brain-derived neurotropic factor and behavior in mice. Gastroenterology 141(2):599–609 (609, e591–593)
Clarke G, Grenham S, Scully P, Fitzgerald P, Moloney RD, Shanahan F, Dinan TG, Cryan JF (2013) The microbiome-gut–brain axis during early life regulates the hippocampal serotonergic system in a sex-dependent manner. Mol Psychiatry 18(6):666–673
Sherman MP, Zaghouani H, Niklas V (2015) Gut microbiota, the immune system, and diet influence the neonatal gut–brain axis. Pediatr Res 77(1–2):127–135
Wang Y, Kasper LH (2014) The role of microbiome in central nervous system disorders. Brain Behav Immun 38:1–12
Mayer EA, Tillisch K, Gupta A (2015) Gut–brain axis and the microbiota. J Clin Invest 125(3):926–938
Wexler HM (2007) Bacteroides: the good, the bad, and the nitty-gritty. Clin Microbiol Rev 20(4):593–621
Renouf M, Hendrich S (2011) Bacteroides uniformis is a putative bacterial species associated with the degradation of the isoflavone genistein in human feces. J Nutr 141(6):1120–1126
NCBI: Bacteroides ovatus. In: Normal gut bacterium. 2019. https://www.ncbi.nlm.nih.gov/genome/?term=Bacteroides%20ovatus
Wu M, McNulty NP, Rodionov DA, Khoroshkin MS, Griffin NW, Cheng J, Latreille P, Kerstetter RA, Terrapon N, Henrissat B, Osterman AL, Gordon JI (2015) Genetic determinants of in vivo fitness and diet responsiveness in multiple human gut Bacteroides. Science 350(6256):aac5992
Coyne MJ, Roelofs KG, Comstock LE (2016) Type VI secretion systems of human gut Bacteroidales segregate into three genetic architectures, two of which are contained on mobile genetic elements. BMC Genomics 17:58
Kitahara M, Sakamoto M, Ike M, Sakata S, Benno Y (2005) Bacteroides plebeius sp. nov. and Bacteroides coprocola sp. nov., isolated from human faeces. Int J Syst Evol Microbiol 55(Pt 5):2143–2147
Tillisch K, Mayer EA, Gupta A, Gill Z, Brazeilles R, Le Neve B, van Hylckama Vlieg JET, Guyonnet D, Derrien M, Labus JS (2017) Brain structure and response to emotional stimuli as related to gut microbial profiles in healthy women. Psychosom Med 79(8):905–913
Greetham HL, Collins MD, Gibson GR, Giffard C, Falsen E, Lawson PA (2004) Sutterella stercoricanis sp. nov., isolated from canine faeces. Int J Syst Evol Microbiol 54(Pt 5):1581–1584
Williams BL, Hornig M, Parekh T, Lipkin WI (2012) Application of novel PCR-based methods for detection, quantitation, and phylogenetic characterization of Sutterella species in intestinal biopsy samples from children with autism and gastrointestinal disturbances. MBio 3(1):e00261
Wang LJ, Yu YH, Fu ML, Yeh WT, Hsu JL, Yang YH, Chen WJ, Chiang BL, Pan WH (2018) Attention deficit-hyperactivity disorder is associated with allergic symptoms and low levels of hemoglobin and serotonin. Sci Rep 8(1):10229
Chou WJ, Lee MF, Hou ML, Hsiao LS, Lee MJ, Chou MC, Wang LJ (2018) Dietary and nutrient status of children with attention-deficit/hyperactivity disorder: a case-control study. Asia Pac J Clin Nutr 27(6):1325–1331
Proctor C, Thiennimitr P, Chattipakorn N, Chattipakorn SC (2017) Diet, gut microbiota and cognition. Metab Brain Dis 32(1):1–17
Diaz Heijtz R, Wang S, Anuar F, Qian Y, Bjorkholm B, Samuelsson A, Hibberd ML, Forssberg H, Pettersson S (2011) Normal gut microbiota modulates brain development and behavior. Proc Natl Acad Sci USA 108(7):3047–3052
Hopkins MJ, Sharp R, Macfarlane GT (2002) Variation in human intestinal microbiota with age. Dig Liver Dis 34(Suppl 2):S12–S18
Axelsson PB, Clausen TD, Petersen AH, Hageman I, Pinborg A, Kessing LV, Bergholt T, Rasmussen SC, Keiding N, Lokkegaard ECL (2019) Investigating the effects of cesarean delivery and antibiotic use in early childhood on risk of later attention deficit hyperactivity disorder. J Child Psychol Psychiatry 60(2):151–159
Liu YCLS, Shang CY, Lin CH, Tu CL, Gau SF (2006) Norm of the Chinese version of the Swanson, Nolan and Pelham, version IV scale for ADHD. Taiwan J Psychiatry 20:290–304
Jain A, Li XH, Chen WN (2018) Similarities and differences in gut microbiome composition correlate with dietary patterns of Indian and Chinese adults. AMB Express 8(1):104
Acknowledgements
The authors would like to thank Professor Wei-Tsun Soong for granting us the use of the Chinese version of the K-SADS, and Professor Shur-Fen Gau for granting our use of the Chinese version of the SNAP-IV.
Funding
This work was supported by grant from the Chang Gung Memorial Hospital Research Grant (CMRPG8E1441) and the Taiwan Ministry of Science and Technology (MOST 107-2628-B-182-001).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
All authors declare no biomedical financial interests or potential conflicts of interest.
Electronic supplementary material
Below is the link to the electronic supplementary material.
787_2019_1352_MOESM1_ESM.jpg
Supplementary material 1 The distribution of enriched bacteria identified at genus level in the ADHD patients and healthy controls using LEfSe analysis. Linear discriminant analysis (LDA) plots at genus levels showed the enriched bacteria in ADHD patients and healthy controls (JPEG 225 kb)
Rights and permissions
About this article
Cite this article
Wang, LJ., Yang, CY., Chou, WJ. et al. Gut microbiota and dietary patterns in children with attention-deficit/hyperactivity disorder. Eur Child Adolesc Psychiatry 29, 287–297 (2020). https://doi.org/10.1007/s00787-019-01352-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00787-019-01352-2