Abstract
In this study, we have synthesized a series of dinuclear and trinuclear gold(III) complexes of the general formula [Au2(N–N)Cl6] (1–3) for dinuclear and [Au3(N–N)2Cl8]+ (4–6) for trinuclear compounds, respectively, in which N–N is a bidentate ligand (1,4-diaminobutane; 1,6-diaminohexane or 1,8-diaminooctane). These complexes were characterized by elemental analysis, molar conductivity, and spectroscopic techniques (IR, UV–Vis, 1H NMR, ESI–MS). We performed DFT calculations to get insight into the geometry of the studies complexes. DNA-binding studies were performed by UV–Vis spectrophotometry and fluorescence spectroscopy. The results of competitive reactions between gold(III) complexes and ethidium bromide (EB) towards DNA have shown that selected complexes can displace EB from DNA-EB adduct. In addition, these experiments confirm that polynuclear gold(III) complexes interact with DNA covalently or via intercalation. Furthermore, high values of binding constants of gold(III) complexes towards bovine serum albumin (BSA) protein indicate good binding affinity. In addition, redox stability of complexes in the presence of DNA/BSA was confirmed by cyclic voltammetry. Results of the interactions between gold(III) complexes with DNA/BSA were discussed in reference to molecular docking data obtain by Molegro virtual docker. The cytotoxic activity of synthesized gold(III) complexes was evaluated on human breast cancer cell line (MDA-MB-231), human colorectal cancer cell line (HCT-116), and normal human lung fibroblast cell line (MRC-5). All complexes dose-dependently reduced cancer and normal cells viabilities, with significant cytotoxic effects (IC50 < 25 μM) for trinuclear gold(III) complexes (4, 5) on HCT-116 cells.
Graphic abstract

Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The discovery of anticancer activity of platinum complexes (cisplatin, carboplatin, oxaliplatin) developed high interest towards evolution of different metal-based antitumor agents. Numerous reports have revealed that other transition metal complexes can possibly be applied in medicine and pharmacy [1,2,3]. The use of cisplatin is limited due to several significant disadvantages, such as normal tissue toxicity and resistance to treatment [4]. The poor chemotherapeutic index of this clinically used drug has led the researches to focus on the synthesis of metal complexes with better selectivity for cancerous cells, improved cytotoxic properties, and less side effects [5, 6].
The pharmacologic properties of gold compounds have been known since the end of 19th century. They have been used for different studies, even though they are usually used for the treatment of rheumatoid arthritis [7]. Therefore, in the last decade, gold complexes have received increased attention due to the variety of their applications. More recently, gold compounds have been investigated as potential anticancer, antimicrobial, or chemotherapeutic agents and the results shown that they are the most promising alternative for platinum-based metallodrugs [8,9,10,11,12,13,14,15,16,17]. Gold(III) complexes are isostructural and isoelectronic to platinum(II) compounds, and they could exhibit prospective and fascinating anticancer, cytotoxic, and antitumor properties [2]. In general, gold(III) complexes are unstable and rapidly hydrolyze and reduce to gold(I) or gold(0) under physiological conditions, which is the main problem of their development as therapeutic drug [18].
Multimetallic gold(III) complexes are important for catalysis. They have been thoroughly investigated for photophysical and photoluminescent properties due to attractive aurophilic d8–d8 interactions [19, 20]. The efficacy of metal complexes usually depends on the type of metal ion, types of ligands, and geometry of the coordination compounds. To attain gold-based drugs possessing medicinal activities, specific ligands are needed for a good stabilization of gold(III) centers. Nevertheless, gold(III) ions generally prefer ligands bearing “hard” Lewis base donor sites such as nitrogen and oxygen [2].
It is already known that DNA is the main target for platinum-based anticancer compounds, while gold(III) complexes shown a variety of interactions, including direct DNA damage, inhibition of thioredoxin reductase or proteasome, and alteration of cell cycles. Some gold(III) complexes with nitrogen-containing polydentate ligands have potent cytotoxic properties and represent efficient DNA binders [21]. Furthermore, the activity of gold(III) complexes with aliphatic diamine decreases when the chain length of diamine is extended [22, 23]. Knowing these modes of action is very important for further investigation in the field of gold(III) complexes to develop potential cytotoxic properties against cancer cells, especially multidrug-resistant cell lines [10, 24,25,26,27,28,29,30]. According to previous, we have synthesized and characterized three dinuclear (1–3) and three trinuclear (4–6) gold(III) complexes with different aliphatic nitrogen-donor ligands (1,4-diaminobutane, 1,6-diaminohexane or 1,8-diaminooctane) (Fig. 1). These new complexes were characterized by various spectroscopic techniques, such as elemental analysis, molar conductivity, 1H NMR, IR, UV–Vis, and ESI–MS. We also investigated the DNA-binding ability, bovine serum albumin (BSA)-binding ability, redox stability in the presence of DNA/BSA, and cytotoxicity on human breast cancer cell line (MDA-MB-231), human colon cancer cell line (HCT-116), and normal human lung fibroblast cell line (MRC-5) of synthesized gold(III) complexes.
Materials and methods
Potassium tetrachloridoaurate(III) (K[AuCl4]) and nitrogen-containing ligands (1,4-diaminobutane, 1,6-diaminohexane, 1,8-diaminooctane) are commercially available and they were used without purification. All the other chemicals were of analytical reagent grade and used without further purification. Ethidium bromide (EB), calf thymus DNA (CT-DNA), and bovine serum albumin (BSA) were obtained from Sigma Chemicals Co. (USA). The stock solution of CT-DNA was prepared in 0.01 M phosphate-buffered saline (PBS) (Sigma-Aldrich) at pH 7.4, which gave a ration of UV absorbance at 260 nm and 280 nm (A260/A280) of ca. 1.8–1.9, indicating that the DNA was sufficiently free of protein. The concentration was determined by UV absorbance at 260 nm after 1:20 dilution using (ɛ = 6600 M−1 cm−1) [31, 32]. Stock solution of BSA was prepared by dissolving the solid BSA in 0.01 M PBS buffer at pH 7.4 and the concentration was kept fixed at 2 μM. All stock solutions were stored at 277 K and used within 5 days. For cytotoxic assay, Dulbecco’s Modified Eagle Medium (DMEM) and phosphate-buffered saline (PBS) were obtained from GIPCO, Invitrogen, USA, while dimethyl sulfoxide (DMSO) and 3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide (MTT) were obtained from SERVA, Germany.
Elemental analysis was performed on a Carlo Erba Elemental Analyser 1106. The molar conductivities of freshly prepared 1.0 × 10−3 M solutions of all complexes in dimethylformamide (DMF) solutions at 25 °C were measured by Crison EC-Meter Basic 30 +. Using the relation Λm = K/C, the molar conductance of complexes was calculated, where C is the molar concentration of metal complex solutions. The 1H NMR spectra were acquired on a Varian Gemini 2000, 200 MHz NMR spectrometer at 295 K. The measurements were performed with a commercial 5 mm Bruker broadband probe. All chemical shifts are referenced to TSP (trimethylsilylpropanoic acid). IR spectra were recorded as KBr pallets on a Perkin-Elmer Spectrum One spectrometer over the range 450–4000 cm−1. The UV–Vis spectra were obtained on a Perkin-Elmer Lambda 35 double beam spectrophotometer, using 1.0 cm path-length quartz cuvettes (3.0 mL). Fluorescence measurements were carried out on an RF-1501 PC spectrofluorometer (Shimadzu, Japan). The excitation and emission bandwidths were both 10 nm. Mass spectrometry was performed at Amazon SL mass spectrometer Bruker Daltonics (Bremen, Germany). Cyclovoltammetric (CV) measurements were performed in a one-compartment three-electrode cell using a glassy carbon (GC) electrode as working electrode, an Ag/AgCl as reference electrode and a platinum wire as a counter electrode. Measurements were recorded with an Autolab PGSTAT 302N at room temperature. The working electrode surface was polished with alumina on a microcloth with water as the lubricant. All CVs were recorded with a scan rate of 0.1 V/s at room temperature, while the potential of the electrode was cycled between + 1.5 V, 0 V and + 1.5 V. The supporting electrolyte was a 0.1 M NaCl. Solution of NaClO4 and PBS buffer were not employed because of possible redox reactions that can interfere. The certain amount (100 µL—concentration 2 × 10−5 M) of BSA or (100 µL—concentration 1.24 × 10−3 M) DNA and (10 mL—concentration 10−3 M) Au(III) complexes (1–6) were mixed. The experiments were repeated at least three times at room temperature. The obtained data were collected and analyzed using the OriginPro8 and Microsoft Office Excel 2007 programs. The working electrode was polished every day or even more often if that was necessary.
Synthesis and characterization of complexes
For complexes 1–3, the solution of 0.065 mmol of corresponding nitrogen-containing ligand [6.5 μL (ρ = 0.877 g/mL) of 1,4-diaminobutane; 9 μL (ρ = 0.84 g/mL) of 1,6-diaminohexane, and 9.4 mg of 1,8-diaminooctane] in 2.0 mL of ethanol was added slowly under the stirring to the solution containing 0.13 mmol of K[AuCl4] (50 mg) dissolved in 5.0 mL of ethanol. The yellow precipitate was formed immediately after addition of ligand. The reaction mixture was stirred in the dark at room temperature for 5 h. Precipitate was filtered off (complex 1) and dried in the dark at room temperature or left to evaporate at room temperature (complexes 2 and 3). For complexes 4–6, to the solution of 0.17 mmol of the corresponding nitrogen-containing ligand [17.1 μL (ρ = 0.877 g/mL) of 1,4-diaminobutane; 23.5 μL (ρ = 0.84 g/mL) of 1,6-diaminohexane and 24.5 mg of 1,8-diaminooctane] in 2.0 mL of ethanol was dropped under the stirring the solution that contains 0.26 mmol of K[AuCl4] (100 mg) dissolved in 5.0 mL of ethanol. The reaction mixture was stirred in the dark at room temperature for 3 h, and then left to evaporate at room temperature. After evaporation, yellow precipitate was collected.
[Au2(NH2(CH2)4NH2)Cl6] (1) Yield: 61.40%. Anal. Calc. (%): C, 6.91; H, 1.74; N, 4.03. Found: C, 6.86; H, 1.81; N, 3.99. 1H NMR (D2O, 200 MHz): δ 1.75 (m, 4H, C2–H, C3–H), 3.04 (t, 4H, C1–H, C4–H), IR (KBr, cm−1): 3526 s, 3479 s (N–H stretch), 2918 s (C–H stretch), 1575 s (N–H bend), 1499 s (C–H bend), 1142 m (C–N stretch). UV–Vis (H2O, nm): 285. m/z 692.87.
[Au2(NH2(CH2)6NH2)Cl6] (2) Yield: 41.90%. Anal. Calc. (%): C, 9.97; H, 2.23; N, 3.88. Found: C, 9.91; H, 2.31; N, 3.84. 1H NMR (D2O, 200 MHz): δ 1.41 (m, 4H, C3–H, C4–H), 1.69 (m, 4H, C2–H, C5–H), 2.99 (t, 4H, C1–H, C6–H). IR (KBr, cm−1): 3500–3400 s (N–H stretch), 2935 s (C–H stretch), 1560 s (N–H bend), 1469 s (C–H bend), 1175 m (C–N stretch). UV–Vis (H2O, nm): 286. m/z 720.94.
[Au2(NH2(CH2)8NH2)Cl6] (3) Yield: 63.63%. Anal. Calc. (%): C, 12.80; H, 2.68; N, 3.73. Found: C, 12.67; H, 2.75; N, 3.68. 1H NMR (D2O, 200 MHz): δ 1.35 (m, 8H, C3–H, C4–H, C5–H, C6–H), 1.64 (m, 4H, C2–H, C7–H), 2.98 (t, 4H, C1–H, C8–H). IR (KBr, cm−1): 3460–3420 s (N–H stretch), 2925 s (C–H stretch), 1648 s (N–H bend), 1460 s (C–H bend), 1253 m (C–N stretch). UV–Vis (H2O, nm): 293. m/z 748.97.
[Au3(NH2(CH2)4NH2)2Cl8]Cl (4) Yield: 58.42%. Anal. Calc. (%): C, 8.85; H, 2.23; N, 5.16. Found: C, 8.78; H, 2.27; N, 5.04. 1H NMR (D2O, 200 MHz): δ 1.75 (m, 4H, C2–H, C3–H), 3.04 (t, 4H, C1–H, C4–H). IR (KBr, cm−1): 3538 s, 3486 s (N–H stretch), 2933 s (C–H stretch), 1599 s (N–H bend), 1499 s (C–H bend), 1109 m (C–N stretch). UV–Vis (H2O, nm): 293. m/z 1050.04.
[Au3(NH2(CH2)6NH2)2Cl8]Cl (5) Yield: 63.21%. Anal. Calc. (%): C, 12.69; H, 2.82; N, 4.90. Found: C, 12.57; H, 2.93; N, 4.79. 1H NMR (D2O, 200 MHz): δ 1.41 (m, 4H, C3–H, C4–H), 1.67 (m, 4H, C2–H, C5–H), 2.99 (t, 4H, C1–H, C6–H). IR (KBr, cm−1): 3591 s, 3518 s (N–H stretch), 2933 s (C–H stretch), 1560 s (N–H bend), 1468 s (C–H bend), 1174 m (C–N stretch). UV–Vis (H2O, nm): 287. m/z 1106.91.
[Au3(NH2(CH2)8NH2)2Cl8]Cl (6) Yield: 46.24%. Anal. Calc. (%): C, 16.03; H, 3.36; N, 4.67. Found: C, 15.97; H, 3.45; N, 4.57. 1H NMR (D2O, 200 MHz): δ 1.35 (m, 8H, C3–H, C4–H, C5–H, C6–H), 1.65 (m, 4H, C2–H, C7–H), 2.98 (t, 4H, C1–H, C8–H). IR (KBr, cm−1): 3450–3400 s (N–H stretch), 2925 s (C–H stretch), 1596 s (N–H bend), 1498 s (C–H bend), 1132 m (C–N stretch). UV–Vis (H2O, nm): 287. m/z 1163.24.
Quantum chemical calculations
All calculations were done applying B3LYP functional [33,34,35] in combination with the basis set cc-pVTZ [36]. For the gold atoms, the relativistic effective core potential LanL2TZ(f) [37] was utilized. The performance of this level has been documented by others to be very effective in DFT calculation of similar systems [38]. The characterization as minima was done by computation of vibrational frequencies at the same level. Relative energies were corrected for zero-point vibrational energies (ZPE). The influence of bulk solvent was evaluated via single point calculations using the CPCM formalism [39, 40] on the same level and water as a solvent. All calculations were performed using the Gaussian 09 program package [41].
DNA-binding studies
Absorption spectroscopic studies
The binding of complexes 1–6 towards CT-DNA was studies by UV–Vis spectroscopy to investigate the possible DNA-binding modes as well as to calculate the binding constants (Kb). The series of complex-DNA solutions were prepared by mixing the fixed concentration of complex solution (8 µM) with increments of DNA stock solution (2.63 mM).
Ethidium bromide (EB) displacement studies
The EB-competitive studies of gold(III) complexes were carried out by fluorescence emission spectroscopy to examine whether the complexes 1–6 can displace EB from DNA–EB complex. DNA-EB was initially prepared by mixing of 21 µM EB and 21 µM CT-DNA in 0.01 M PBS buffer solution (pH 7.4). The possible binding effect of complexes was investigated step by step after the addition of certain amount of complex solution into the solution of DNA-EB. Fluorescence intensities were measured with the excitation wavelength set at 527 nm and fluorescence emission at 612 nm. Before measurements, each system was shaken and incubated at room temperature for 5 min. The emission was recorded in the range 550–750 nm.
Albumin-binding studies
The protein-binding study was performed by tryptophan fluorescence quenching experiments using bovine serum albumin (BSA, 2 µM) in 0.01 M PBS buffer solution (pH 7.4). The quenching of the emission intensity of tryptophan residues of BSA at 365 nm was monitored using complexes 1–6 as quenchers with increasing concentration (up to 40 µM). Fluorescence spectra were recorded in the range 300–500 nm at an excitation wavelength of 295 nm. The fluorescence spectra of compounds in buffered solutions were recorded under the same experimental conditions and no fluorescence emission was detected.
Viscosity measurements
The viscosity of a DNA solution was measured in the presence of increasing amounts of complexes 1–6. The flow time was measured with a digital stopwatch. Each sample was measured six times and the average flow time was calculated. The data were presented as (η/η0)1/3 against r, where η is the viscosity of DNA in the presence of complex and η0 is the viscosity of DNA alone in the buffered solution. The viscosity values were calculated from the observed flow time of DNA-containing solutions (t) corrected for the flow time of buffer alone (t0), η = (t–t0)/t0.
Molecular docking
Optimized structures of investigated complexes were done as previously described in experimental section. In the rigid DNA structure, flexible compounds, such as gold(III) complexes, were docked using Molegro Virtual Docker (MVD, version 2013.6.0.1) [42]. For DNA model, B-DNA dodecamer (CGCGAATTCGCG)2 (PDB code 1BNA) [43] was used to study the interaction between metal complex and DNA. The 3D-crystal structure of bovine serum albumin with PDB code 4F5S [44] was downloaded from Brookhaven Protein Data and loaded to Molegro Virtual Docker (MVD). All water molecules were removed. Grid resolution of the binding side was 0.3 Å. The parameters of docking procedure were: maximum number of iterations 1500, population size 50, energy threshold 100.00, and maximum number of steps 300. The MolDock SE as a search algorithm was used with the number of runs set to 100 and the number of generated poses was 5. The estimation of gold(III) complexes and DNA/BSA interactions was described by the MVD-related scoring functions: MolDock, Docking, Rerank, and Hbond. A maximum population of 100 and maximum number of iterations of 10 000 were used for each run. The five best poses were retained. Docked poses were visualized using CHIMERA [45] molecular graphics program. Molegro scores were evaluated in a relative fashion.
Cytotoxic activity and cell viability assay (MTT assay)
The gold(III) complexes (1–6), as well as K[AuCl4] (7), were dissolved in DMEM medium for cell treatment (final concentration of DMSO < 0.5%). The MDA-MB-231, HCT-116, and MRC-5 cell lines were obtained from the American Tissue Culture Collection (Manassas, VA, USA). Standard maintaining conditions were adjusted according to our previous publication [46]. All experiments were done with cells at 70–80% confluence.
The cell viability was determined by MTT assay [47] after treatments with all gold(III) complexes. This test is based on the color reaction of mitochondrial dehydrogenase from living cells with MTT (3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide). MDA-231, HCT-116, and MRC-5 cells were seeded in a 96-well plate (104 cells per well). After 24 h of pre-incubation, cells were treated with 100 µL of each concentration of investigated gold(III) complexes (0.1, 1, 10, 50, 100, and 200 μM) for 24 and 72 h. Untreated cells served as a control. At the end of the treatment period, MTT (final concentration 5 mg/mL in PBS) was added to each well and then incubated at 37 °C in 5% CO2 for 2 h. The colored crystals of formazan were dissolved in DMSO and the absorbance was measured at 550 nm. The effects on cell viability were calculated as a ratio of the absorbance of the treated group divided by the absorbance of control group, multiplied by 100 to give percentage of viable cells.
The data are expressed as the means of two independent experiments, performed in triplicate for each dose. The magnitude of correlation between variables was done using a SPSS statistical software package (SPSS for Windows, ver. 17, 2008). The cytotoxic effect of each tested complex was expressed by IC50 (inhibitory dose which inhibit 50% growth cells). The IC50 values were calculated from the dose curves by a computer program CalcuSyn v. 2.1 [48].
Results and discussion
Dinuclear gold(III) complexes 1–3 with general formula [Au2(N–N)Cl6] and trinuclear gold(III) complexes 4–6 with general formula [Au3(N–N)2Cl8]+, where N–N is a bidentate ligand (1,4-diaminobutane; 1,6-diaminohexane, or 1,8-diaminooctane), have been synthesized according to the procedure presented in Scheme 1. Their structures were confirmed by elemental analysis, molar conductivity, and different spectroscopic techniques (IR, UV–Vis, 1H NMR, and ESI–MS).
The molar conductivity values, given in Table 1, confirmed the non-electrolyte nature of complexes 1–3 as well as ionic 1:1 nature of complexes 4–6, respectively. These results are supported by the results of chemical analysis.
The 1H NMR, IR, UV–Vis, and ESI–MS spectroscopic data for dinuclear and trinuclear gold(III) complexes (Figures S1–S7, ESI) are listed in the Experimental section. They are consistent with the structures presented in Fig. 1 and Scheme 1.
In the 1H NMR spectrum of ligand 1,4-diaminobutane, the signal of 4H (C1–C4) appeared at 2.775 ppm (triplets) and the signal of 4H (C2–C3) appeared at 1.614 ppm (multiplet). Both signals in the 1H NMR spectrum of complexes 1 and 4 are moved to higher values. In the 1H NMR spectrum of complexes 2 and 5, all signals are also moved to higher chemical shifts relative to signals of uncoordinated ligand 1,6-diaminohexane, in which the signal of 4H (C1–C6) appeared at 2.829 ppm (triplets), the signal of 4H (C2–C5) appeared at 1.672 ppm (multiplet), and the signal of 4H (C3–C4) appeared at 1.580 ppm (multiplet). In the 1H NMR spectrum of 1,8-diaminooctane, all tree signals of free ligand [4H (C1–C8) at 2.606 ppm (triplets), 4H (C2–C7) at 1.436 ppm (triplets), and 8H (C3, C4, C5, C6) at 1.314 ppm (singlet)] are moved downfield after coordination (Table 2). All spectra are given in ESI (Figs. S1 and S2).
The IR spectrum of gold(III) complexes 1–6 (ESI, Fig. S3) confirms the presence of characteristic bands belonging to NH2 group (3600–3200 cm−1) and bands specific for the vibration for aliphatic C–H bond (2920–2940 cm−1 for all complexes).
The UV–Vis spectra of all complexes were recorded in water and in PBS buffer (ESI, Fig. S4). The shape of the spectra (ESI, Fig. S4) and the values of λmax are almost identical for all complexes (λmax = 285 nm for 1, λmax = 286 nm for 2, λmax = 293 nm for 3, λmax = 293 nm for 4, λmax = 287 nm for 5, and λmax = 287 nm for 6) indicating the same coordination mode of ligands, respectively. Considering that all cytotoxicity measurements were performed in medium that consist small amount of DMSO (less than 0.5%), the presence of possible interactions was followed by UV–Vis as well. The results have shown negligible changes in the absorption (Fig. S5, ESI).
ESI–MS spectra of complexes 1–6 are also given in ESI (Figs. S6 and S7). In all spectra are detected signals that correspond to the complexes 1–6. Regarding the conditions during the analysis, different fragmentations were observed without higher mass fragments.
DFT calculations
To get structural insight of investigated complexes 1–6, quantum chemical calculations were performed. The geometry-optimized structures and extracted data of dinuclear gold(III) complexes are shown in Fig. 2 and Table 3, and for trinuclear gold(III) complexes in Fig. 3 and Table 4. The observed results shown that d8 gold(III) centers adopt distorted square-planar coordination geometry, all Au–Cl bonds are in the range of 2.27–2.32 Å and all Au–N bonds are in the range between 2.10 and 2.13 Å which is in a good alignment with the previously published results of X-ray analysis [49].
With fully awareness of limitations, calculating HOMO and LUMO frontier orbital energies by DFT methods [50] from Fig. 4 can be seen that HOMO frontier orbitals of complexes 1–3 are concentrated on one gold(III) center, but when it goes to the LUMO frontier orbitals, they are mainly shifted towards another gold(III) center, with the exception of the complex 2 in which the LUMO orbitals are evenly distributed on both metal centers. In the case of complexes 4–6 (Fig. 5), LUMO orbitals are shifted to the gold(III) center standing in the middle of trinuclear complex.
DNA-binding studies
Absorption spectroscopic studies
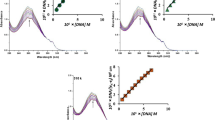
DNA is an important potential biological target for many metal-based anticancer agents. Because of that, it is very important to understand DNA-binding properties of potential anticancer agents. UV–Vis spectroscopy is one of the most universally employed methods for determination of binding modes and binding extent of metal complexes with DNA. Already is known that metal complexes can bind to double-stranded DNA via covalent (replacement of the labile ligand of complex by a nitrogen base of DNA, e.g., guanine N7) or noncovalent (intercalation, electrostatic, or groove binding) interactions [51]. The absorption intensity of complexes in the presence of CT-DNA may decrease (hypochromism) or increase (hyperchromism) with slightly increase in the absorption wavelength (bathochromism). The absorption spectra of complexes 3 and 4 in the absence and presence of DNA (at constant concentration of the complex) are given in Fig. 6, while for other studied complexes, they are presented in ESI (Fig. S8).
The addition of CT-DNA to the solution of complexes 3 and 4 resulted in the appearance of a new band centered at 258 nm (see Fig. 6). The behavior of other studied complexes was quite similar upon addition of increasing amount of CT-DNA (ESI, Fig. S8). The appearance of a new signal in UV–Vis spectra indicates the presence of interaction between gold(III) complexes and CT-DNA.
The intrinsic binding constants Kb (M−1) for all studied complexes calculated by the equation (Eq. S1, ESI) are given in Table 5. The high values for Kb indicate the binding of all complexes to CT-DNA. Complex 2 exhibits the highest Kb value compared to other, while in comparison with Kb value for classical intercalator ethidium bromide (EB), Kb = (1.23 ± 0.07) × 105 M−1 [52], studied complexes exhibit lower values of binding constants.
Fluorescence quenching measurements
Ethidium bromide (EB = 3,8-diamino-5-ethyl-6-phenylphenanthridinium bromide) is an intercalator that gives significant fluorescence emission intensity when it intercalates to CT-DNA through the planar EB phenanthridine ring in between adjacent DNA base pairs. DNA-induced EB fluorescence emission could be quenched after addition of complex that is able to replace EB and form strong interactions with DNA [53]. The fluorescence quenching curves of EB bound to DNA in the absence and the presence of complexes 1 and 6 are shown in Fig. 7 and curves for other complexes are presented in ESI Fig. S9.
Emission spectra of EB bound to CT-DNA in the presence of complexes 1 and 6. CEB = 21 μM, CDNK = 21 μM, Ccomplex = 0–29.4 μM for complex 1 and CEB = 18 μM, CDNK = 18 μM, Ccomplex = 0–25.7 μM for complex 6; λex = 527 nm. Arrows show the intensity changes upon increasing the concentration of complex. Insert: Stern–Volmer quenching plot of DNA-EB for complexes 1 and 6
The increase of complex concentration in all studied systems resulted in significant decrease in intensity of the emission band at 612 nm. This indicates the competition of complexes with EB in binding to DNA (Fig. 7 and Fig. S9, ESI). The observed quenching of DNA-EB fluorescence suggests that all studied complexes displace EB from the DNA-EB complex and interact with DNA [31, 32].
The Stern–Volmer plots of EB-DNA (insert Fig. 7 and Fig. S9) confirm the fluorescence emission quenching. Stern–Volmer quenching constants (Ksv) are calculated from the slopes of the plots of I0/I vs. [Q] (Eq. S2, ESI, insert Fig. 7 and Fig. S9) and reported in Table 5. All studied gold(III) complexes showed high values of quenching constants, indicating their great efficiency to replace EB and bind to DNA. Complex 4 exhibits the highest Ksv values in comparison with other gold(III) complexes.
Albumin-binding studies
Serum albumin (SA) is the most abundant plasma protein in humans (and other mammals) and plays an important role in the transport of ions and drugs through the bloodstream to cells and tissues [54]. Bovine serum albumin (BSA) is the most extensively studied serum albumin because of its high structural homology with human serum albumin (HSA). The interactions of complexes 1–6 with BSA were studied by fluorescence spectroscopy, as this method allows a quantitative assessment of the binding strength. Observed quenching may be attributed to the changes in protein conformation, subunit association, substrate binding, or denaturation.
The values of Stern–Volmer quenching constant (Ksv) and the quenching constant (kq) for the interaction of complexes with BSA are calculated by Stern–Volmer quenching equation (Eq. S3, ESI, insert Fig. 8 and Fig. S10) and are given in Table 6. The values of BSA-binding constant (K) and the number of binding sites per albumin (n) are calculated from the Scatchard equation (Eq. S4 and Fig. S11, ESI). A good quenching ability of BSA fluorescence was found for each complex. Furthermore, the maximum of the bands was slightly shifted from 361 to 365 nm for all complexes (Fig. 8; Fig. S10). The red shift implies the formation of gold(III)-BSA adducts, which altered the polarity of microenvironment in the vicinity of tryptophan.
All gold(III) complexes show good quenching ability, while complex 4 exhibits the strongest, kq = (1.320 ± 0.004) × 1013 M−1 s−1. The kq values for all complexes (1012 M−1 s−1 or 1013 M−1 s−1) are higher than diverse kinds of quenching for biopolymer fluorescence (1010 M−1 s−1) indicating the existence of a static quenching mechanism [55].
Viscosity measurements
Viscosity measurements of DNA solutions were performed in the absence and presence of complexes to confirm the modes of binding of complexes 1–6 to CT-DNA. The viscosity of DNA is sensitive to length changes and regarded as the least ambiguous and the most critical clues of the DNA-binding mode in solution [56, 57]. The addition of increasing amount (up to r = 1.0) of complexes 1–6 (0–12 μM) to a DNA solution (12 μM) generally resulted in an increase of relative viscosity of DNA (Fig. 9), which was more pronounced upon addition of complex 3. In the case of classic intercalation DNA, base pairs are separated to host the bound compound resulting in increased DNA viscosity. The magnitude of interaction is usually in accordance with the strength of the interaction because of the lengthening of the DNA helix. Therefore, the observed viscosity increase may be explained by an increase in the overall DNA length provoked by the insertion of complex in between the DNA base pairs due to interaction via intercalation.
Cyclic voltammetry
The redox stability of complexes in the presence of DNK/BSA was studied by cyclic voltammetry. Reaction between 100 µL BSA (concentration 2 × 10−5 M) and 10−3 M gold(III) complexes (1–6) in 0.1 M NaCl aqueous solution was followed. The results displayed a shift of the cathodic peak (Fig. 10; Fig. S12, ESI): for complex 1 from 0.43 to 0.57 V; for complex 2 from 0.56 to 0.58 V; for complex 3 from 0.39 to 0.46 V; for complex 4 from 0.61 to 0.57 V; for complex 5 from 0.45 to 0.36 V; and for complex 6 from 0.51 to 0.50 V. Change to a higher value of potential is evidence for the reduction of Au(III) to Au(I) and formation of Au(I) complex [58]. For the reverse sweep, clear oxidation waves were observed and shift of the anodic peak from 1.02 to 1.16 V (complex 1); from 1.01 to 1.05 V (complex 2); from 1.01 to 0.99 V (complex 3); from 1.08 to 1.19 V (complex 4); from 0.95 to 0.98 V (complex 5); and from 1.11 to 1.20 V (complex 6). However, the possible numerous oxidation steps can be occurred further [59,60,61].
Cyclic voltammograms recorder after dissolution of complex 2 (black line) and after addition of 100 µL of BSA (dashed line, concentration 1.98 × 10−4 mM—left) and after addition of 100 µL of DNA (dashed line, concentration 0.012 mM—right), GC electrode, scan rate 0.1 V s−1, Estep = 0.003 V, 0.1 M NaCl as the background electrolyte
The CV of a 10−3 M solution of Au(III) complexes (1–6) in the presence of 0.1 M NaCl as a background electrolyte recorded for a GC electrode upon reaction with 100 µL DNA (concentration 1.24 × 10−3 M) displayed a shift of the cathodic peak. If the compound interacts with DNA, the peak potential and peak current of the compound show different values (Fig. 10; Fig. S13, ESI). For complex 1 from 0.44 to 0.55 V; for complex 2 from 0.56 to 0.58 V; for complex 3 from 0.39 to 0.41 V; for complex 4 from 0.61 to 0.35 V; for complex 5 from 0.59 to 0.55 V; for complex 6 from 0.5 to 0.48 V. For the reverse sweep, clear oxidation waves were observed and shift of the anodic peak from 1.02 to 1.22 V (complex 1); from 0.93 to 1.03 V (complex 2); from 0.90 to 0.97 V (complex 3); from 1.09 to 1.1 V (complex 4); from 1.02 to 1.06 V (complex 5); and from 1.12 to 1.18 V (complex 6). The directions of the movement of the peaks in some cases could predict the mode of interactions [62].
DNA and BSA docking
Molecular simulation is a useful tool that together with experimental results can develop our understanding of the drug-DNA/BSA interaction, identification of binding place, and prediction of complex binding affinities [63]. Molecular docking studies were carried out to gain information about in silico DNA/BSA-binding affinity for studied compounds 1–6. The predicted top-ranking pose with complex with the lowest energy was applied for suggesting the best possible geometry of compounds inside the DNA double helix as well as the binding inside for bovine serum albumin cavity. MolDock, Docking, Rerank, and Hbond scoring functions were used for the assessment of complex-DNA/BSA-binding affinity [42].
The best docked poses of complexes with DNA dodecamer are displayed in Fig. 11, and top-ranked poses according to used scoring functions are presented in Table 7. According to MolDock, Docking, and Rerank score values, the highest binding affinity has complex 6, while the lowest has complex 1. Obviously, the length of diamine linker plays a major role in stabilizing the DNA-Au product by fitting in the minor groove of DNA and allowing the terminal square-planar gold(III) centers of the investigated complexes better to interact.
The docking results from MVD program revealed that complexes 1–6 bind to subdomain IIA (site I) of BSA which is consistent with the experimental data by which with the increasing amount of complexes, a fluorescence quenching was observed due to the interaction between the complexes and Trp-213 residue. The results are shown in Fig. 12 for dinuclear and in Fig. 13 for trinuclear gold(III) complexes. Top-ranked poses according to the used scoring functions are presented in Table 8. According to the MolDock, Docking, Rerank, and Hbond score values, the highest binding affinity has complex 6 which is consistent with the experimental findings. Therefore, from the molecular docking studies, it can be inferred that hydrophobic and electrostatic interactions as well as hydrogen bonding play a vital role in binding gold(III) complexes binding to site I of BSA [64].
Best poses with BSA for complexes 1–3 according to Hbond values: a molecular docking results of dinuclear gold(III)–BSA complex; b complexes embedded inside the active site of BSA in the electrostatic view; c binding site of 1–3 complexes on BSA and selected amino acid residues are represented by stick models. Hydrogen bonds shown in blue dotted lines
Best poses with BSA for complexes 3–6 according to Hbond values: a molecular docking results of trinuclear gold(III)–BSA complex; b complexes embedded inside the active site of BSA in the electrostatic view; c binding site of 4–6 complexes on BSA and the selected amino acid residues are represented by stick models. Hydrogen bonds shown in blue dotted lines
Cytotoxic effects
In vitro cytotoxic activity of gold(III) complexes 1–6 and K[AuCl3] (7) against human breast (MDA-MB-231) and colorectal cancer (HCT-116) cell lines, as well as normal human lung fibroblast (MRC-5) cell lines was evaluated on the basis of cell viabilities after 24 and 72 h of treatments, given by MTT test. Gold(III) complexes 1–6 significantly reduced MDA-MB-231 cell viability (Fig. S14, ESI) on time- and dose-dependent way with coefficient correlation R > − 0.9 (ESI, Table S1). In addition, tested gold(III) complexes significantly and dose-dependently (ESI, Table S1) reduced HCT-116 cancer and MRC-5 normal cell viabilities (Figs. S14, S15, and S16, ESI), without time dependency.
Cytotoxicity of gold(III) complexes expressed as doses which killed 50% of total cells (IC50) is shown in Table 9. All complexes show cytotoxic effects on tested cells (IC50 < 100 μM). These effects are significant after 24 h of treatments when compared with cytotoxicity of K[AuCl4] (Table 9; Fig. S17, ESI), or cisplatin as positive control [65]. In general, trinuclear gold(III) complexes had better cytotoxicity. Among them, complexes 4 and 5 expressed the highest and significant cytotoxic effects on HCT-116 colorectal cancer cells (IC50 < 25 μM, Table 9). Dinuclear gold(III) complexes also showed significant cytotoxic activity on MDA-MB-231 breast cancer cells, after both time periods, compared with cisplatin activity [65]. On the basis of given data, apparently the increase of number and chain length of diamine linker in selected gold(III) complexes increase their hydrophobic and flexible characteristics enabling better entrance in the cells, respectively. However, detailed analysis of the biotransformation of gold(III) complexes in normal and cancer cells, as well as their proapoptotic or lytic/necrotic effects will be the aim of our further investigations.
In general, the gold(III) complexes expressed cell-selective cytotoxic activities (Table 9), with the highest effects on colorectal cancer cells (HCT-116). Normal lung fibroblasts (MRC-5) are also sensitive to cytotoxic effects, indicating that synthetized gold(III) complexes had no specific antitumor activity. We focused our strategies in the development of new anticancer drugs to find new chemotherapeutics with better cytotoxic and antimetastatic activity as well as with less side physiologic effects. The most of newly synthetized metal complexes are the modification of standard cytostatic cisplatin. Considering all, investigated gold(III) complexes showed a significant cytotoxic effect compared with cisplatin [65] or with some of tested newly synthetized platinum(IV) complexes [66], selenium–hydantoin–palladium(II) complex [67], and palladium(II) complexes [46, 65]. Taking all together, novel gold(III) complexes had good predisposition for potential anticancer drugs as well as for co-therapy with natural products.
Conclusions
We described the preparation and structural characterization of six gold(III) complexes by different experimental methods and by DFT calculations. The study of the interactions of complexes 1–6 with DNA provides multiple binding mode including covalent and intercalation interaction. Complex 2 exhibits the highest Kb (intrinsic binding constant) value compared to other complexes. Furthermore, all gold(III) complexes show the high values of quenching constants indicating their great efficiency to replace EB and bind to DNA, while complex 4 exhibits the highest Ksv values in comparison with other compounds. According to viscosity measurements, complex 3 shows the best affinity for intercalation. Furthermore, relatively high binding constants towards BSA indicate a good binding affinity of all complexes. Among all, complex 4 exhibits the strongest interaction. By CV, measurements were confirmed redox stability of all complexes in the presence of DNA/BSA. Molecular docking study was carried out to gain information about in silico DNA/BSA-binding affinity for studied compounds 1–6. The highest binding affinity towards DNA and BSA has complex 6. All tested gold(III) complexes reduced cell viability with cytotoxic effect on investigated cell lines, with good predisposition for potential anticancer drugs. Among all, complexes 4 and 5 show the significant cytotoxic effects on HCT-116 colorectal cancer cells.
Abbreviations
- N–N :
-
1,4-Diaminobutane; 1,6-diaminohexane or 1,8-diaminooctane)
- CT-DNA:
-
Calf thymus DNA
- BSA:
-
Bovine serum albumine
- MDA-MB-231:
-
Human breast cancer cell line
- HTC-116:
-
Human colon cancer cell line
- MRC-5:
-
Normal human lung fibroblast cell line
- PBS:
-
Phosphate-buffered saline
- DMEM:
-
Dulbecco’s Modified Eagle Medium
- DMSO:
-
Dimethyl sulfoxide
- MTT:
-
3-(4,5-Dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide
- TSP:
-
Trimethylsilylpropanoic acid
- EB:
-
Ethidium bromide
- SPSS:
-
Statistical software package for Windows, ver. 17, 2008
- IC50 :
-
Inhibitory dose which inhibit 50% growth cells
- HOMO:
-
Highest occupied molecular orbitals
- LUMO:
-
Lowest unoccupied molecular orbitals
- DFT:
-
Density functional theory
- K b :
-
Intrinsic binding constants
- K sv :
-
Stern–Volmer quenching constant
- k q :
-
Quenching rate constant
- n :
-
Number of binding sites per albumin
- η :
-
Viscosity of DNA in the presence of complex
- η 0 :
-
Viscosity of DNA alone
- B3LYP:
-
Becke, three-parameter, Lee–Yang–Parr functional
- cc-pVTZ:
-
Correlation-consistent polarized valence-only Triple-Zeta basis set
- LanL2TZ:
-
Los Alamos effective core potential triple-zeta basis set
- ZPE:
-
Zero-point vibrational energies
- CPCM:
-
Conductor-like polarizable continuum model
- MVD:
-
Molegro Virtual Docker version 2013.6.0.1
- B-DNA:
-
Crystal structure of DNA in Protein Data Bank
- PDB:
-
Protein Data Bank
- 4F5S:
-
Crystal structure of BSA in Protein Data Bank
References
Bernard C (2017) Johns Matthey Technol Rev 61:52–59
Patanjali P, Kumar R, Sourabh, Kumar A, Chaudhary P, Singh R (2018) Main Group Chem 17:35–52
Radisavljević S, Đeković-Kesić A, Jovanović, Petrović B (2018) Trans Met Chem 43:331–338
Bugarčić ŽD, Bogojeski J, van Eldik R (2015) Coord Chem 292:91–106
Boscutti G, Nardon C, Marchio L, Crisma M, Biondi B, Dalzoppo D, Via Dalla L, Formaggion F, Casini A, Fregona D (2018) ChemMedChem 13:1131–1145
Marloye M, Berger G, Gelbcke M, Dufrasne F (2016) Future Med Chem 8:2263–2286
Bertrand B, Williams MRM, Bochmann M (2018) Chem Eur J 24:11840–11851
Baron M, Tiezza MD, Carlotto A, Tubaro C, Graiff C, Orian L (2018) J Organomet Chem 866:144–152
Glišić BĐ, Đuran MI (2014) Dalton Trans 43:5950–5969
Zou T, Lum CT, Lok CN, Zhang JJ, Che CM (2015) Chem Soc Rev 44:8786–8801
Messori L, Abbate F, Marcon G, Orioli P, Fontani M, Mini E, Mazzei T, Carotti S, O’Connell T, Zanello P (2000) J Med Soc 43:3541–3548
Marcon G, Messori L, Orioli P (2002) Expert Rev Anticancer Ther 2:337–346
Casini A, Cinellu MA, Minghetti G, Gabbiani C, Coronnello M, Mini E, Messori L (2006) J Med Chem 49:5524–5531
Casini A, Hartinger C, Gabbiani C, Mini E, Dyson PJ, Keppler BK (2005) Messori L 102:564–575
Messori L, Marcon G (2004) Met Ions Biol 42:385–424
Messori L, Orioli P, Tempi C, Marcon G (2001) Biochem Biophys Res Commun 281:352–360
Bertrand B, Spreckelmeyer S, Bodio E, Cocco F, Picquet M, Richard P, Le Gendre P, Orvig C, Cinellu MA (2015) Casini A 44:11911–11918
Tiekink ERT (2008) Inflammopharmacology 16:138–142
Sculfort S, Braunstein P (2011) Chem Soc Rev 40:2741–2760
Bestgen S, Gamer MT, Lebedkin S, Kappes MM, Roesky PW (2014) Chem Eur J 20:1–15
Patel MN, Bhatt BS, Dosi PA (2013) Spectrochim Acta A 110:20–27
Briš A, Trošelj P, Margetić D (2017) Croat Chem Acta 90:667–677
Norrehed S, Polavarapu P, Yang W, Gogoll A, Grennberg H (2013) Tetrahedron 69:7131–7138
Jung Y, Lippard SJ (2007) Chem Rev 107:1387–1407
Ott I (2009) Chem Rev 253:1670–1681
Lum CT, Sun RWY, Zou T, Che CM (2014) Chem Sci 5:1579–1584
He L, Chen T, You Y, Hu H, Zheng W, Kwong WY, Zou T, Che CM (2014) Angew Chem Int Ed 53:12532–12536
Zhang JJ, Lu W, Sun RWY, Che CM (2012) Angew Chem Int Ed 51:4882–4886
Cinellu MA, Maiore L, Manassero M, Casini A, Arca M, Fiebig HH, Kelter G, Michelucci E, Pieraccini G, Gabbiani C, Messori L (2010) ACS Med Chem Lett 1:336–339
Chow KHM, Sun RWY, Lam JBB, Li CK, Xu A, Ma DL, Abagyon R, Wang Y, Che CM (2010) Cancer Res 70:329–337
Dimiza F, Fountoulaki S, Papadopoulos AN, Koutogiorgis CA, Tangoulis V, Raptopoulon CP, Psycharis V, Terzis A, Kessissoglou DP, Psomas G (2011) Dalton Trans 40:8555–8568
Dimiza F, Perdih F, Tangoulis V, Turel I, Kessissoglou DP, Psomas G (2011) J Inorg Biochem 105:476–489
Becke AD (1993) J Phys Chem 97:5648–5652
Lee C, Yang W, Parr RG (1988) Phys Rev B 37:785–789
Stephens PJ, Devlin FJ, Chabalowski CF, Frisch MJ (1994) J Phys Chem 98:11623
Kendal RA, Dunning TH Jr, Harrison RJ (1992) J Chem Phys 96:6796–6806
Hay PJ, Wadt WR (1985) J Chem Phys 82:270, 284, 299
Radenkovic S, Antić M, Savić ND, Glišić B (2017) New J Chem 41:12407–12415
Barone V, Cossi M (1998) J Phys Chem A 102:1995–2001
Cossi M, Rega M, Scalmani G, Barone V (2003) J Comput Chem 24:669–681
Frisch MJ et al (2009) Gaussian 09, Revision B.01. Gaussian Inc., Wallingford
Thomsen R, Christensen MH (2006) J Med Chem 49:3315–3321
Drew HR, Wing RM, Takano T, Broka C, Tanaka S, Itakura K, Dickerson RE (1981) Proc Natl Acad Sci USA 78:2179–2183
Bujacz A (2012) Acta Crystallogr Sect D Biol Crystallogr 68:1278–1289
Milutinovic M, Stankovic M, Cvetkovic D, Maksimovic V, Smit B, Pavlovic R, Markovic S (2015) J Food Biochem 39:238–250
Mosmann T (1983) J Immunol Methods 65:55–63
CalcuSyn, v 2.1, BioSoft (2009) Great Shelford, Cambridge
Warżajtis B, Glišić BĐ, Savić ND, Pavic A, Vojnovic S, Veselinović A, Nikodinovic-Runic J, Rychlewska U, Djuran MU (2017) Dalton Trans 46:2594
Dreizler RM, Gross KU (1990) Density functional theory. Springer, Berlin
Novakova O, Chen H, Vrana O, Rodger A, Sadler PJ, Brabec V (2003) Biochemistry 42:11544–11554
Recio Despaigne AA, Da Silva JG, da Costa PR, dos Santos RG, Beraldo H (2014) Molecules 19:17202–17220
Dhar S, Nethaji M, Chakravarty AR (2005) J Inorg Biochem 99:805–812
Tan C, Liu J, Li H, Zheng W, Shi S, Chen L, Ji L (2008) J Inorg Biochem 102:347–358
Wang Y, Zhang H, Zhang G, Tao W, Tang S (2007) J Lumin 126:211–218
Li DD, Tian JL, Gu W, Liu X, Yan SP (2010) J Inorg Biochem 104:171–1735
Jiang M, Li Y, Wu Z, Liu Z, Yan C (2009) J Inorg Biochem 103:833–844
Seo S, Wang X, Murray D (2009) Ionics 15:67–71
Aldous L, Silvester DS, Villagran C, Pitner WR, Compton RG, Lagunas MC, Hardacre C (2006) New J Chem 30:1576–1583
Bortoluzzi M, De Faveri E, Daniele S, Pitteri B (2006) Eur J Inorg Chem 3393-3398
Aldous L, Silvester DS, Villagran C, Pitner WR, Compton RG, Lagunas MC, Hardacre C (2007) J Phys Chem C 111:8496–8503
Carter MT, Bard AJ (1987) J Am Chem Soc 109:7528–7530
Warren GL, Andrews CW, Capelli AM, Clarke B, LaLonde J, Lambert MH, Lindvall M, Nevins N, Semus SF, Senger S, Tedesco G, Wall ID, Woolven JM, Peishoff CE, Head MS (2006) J Med Chem 49:5912–5931
Wei J, Jin F, Wu Q, Jiang Y, Gao D (2014) Talanta 126:116–121
Petrović VP, Živanović MN, Simijonović D, Djorović J, Petrović ZD, Marković SD (2015) RSC Adv 5:86274–86281
Stojković DL, Jevtić VV, Radić GP, Djacić DS, Curcić MG, Marković SD, Djinović VM, Petrović VP, Trifunović SR (2014) J Mol Struct 1062:21–28
Živanović MN, Košarić JV, Šmit B, Šeklić DS, Pavlović RZ, Marković SD (2017) Gen Physiol Biophys 36:187–196
Acknowledgements
The authors gratefully acknowledge financial support of the Ministry of Education, Science and Technological Development of the Republic of Serbia (Project Nos. 172011 and III41010)
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
There are no conflicts of interest to declare.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Radisavljević, S., Ćoćić, D., Jovanović, S. et al. Synthesis, characterization, DFT study, DNA/BSA-binding affinity, and cytotoxicity of some dinuclear and trinuclear gold(III) complexes. J Biol Inorg Chem 24, 1057–1076 (2019). https://doi.org/10.1007/s00775-019-01716-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00775-019-01716-8