Abstract
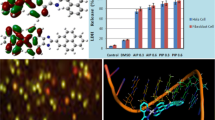
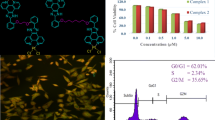
Three new ruthenium(II)-arene complexes with pyrido[2′,3′:5,6]pyrazino[2,3-f][1, 10]phenanthroline (ppf) of general formula: C1 ([(ƞ6-benzene)Ru(ppf)Cl]PF6, C2 ([(ƞ6-toluene)Ru(ppf)Cl]PF6) and C3 ([(ƞ6-p-cymene)Ru(ppf)Cl]PF6) have been synthesized. The structures of complexes were determined by elemental analysis, IR, ESI–MS, as well as with 1H and 13C NMR spectroscopy. Cytotoxic activity has been evaluated in three different human neoplastic cell lines (A549, A375, LS 174T) and in one human non-tumor cell line (MRC-5), by the MTT assay. Complexes C1–C3 showed IC50 values in the micromolar range below 100 µM. Complex C3, carrying ƞ6-p-cymene as the arene ligand, exhibited cytoselective activity toward human malignant melanoma A375 cells (IC50 = 15.8 ± 2.7 µM), and has been selected for further analyses of its biological effects. Drug-accumulation study performed in the A375 cells disclosed that C3 possess lower ability of entering the cells compared to cisplatin and distributes approximately equally in the cytosol and membrane/organelle fraction of cells. Investigations in the 3D model of A375 cells, disclosed different effects of the complex C3 and cisplatin on growth of multicellular tumor spheroids (MCTSs). While the size of cisplatin-treated MCTSs decreased with time, MCTSs treated with C3 continued to growth. Differences in structural organization and biological activity of this type of ruthenium(II)-arene complexes versus cisplatin in A375 malignant melanoma cells pointed out their different modes of action, and necessity for further biological studies and optimizations for potential applications.
Graphical abstract

Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
There is a necessity for the development and screening of potential anticancer agents, because of rapid increase in cancer cases worldwide. More than 50% of the treatments are based on cisplatin (cis-diamminedichloroplatinum(II)) and its derivatives [1]. Beside their effectiveness, they also produce severe adverse effects [2, 3]. Due to the resistance of some types of tumors to platinum, their effectiveness decreases or even makes them ineffective, causing treatment failure [4, 5]. Numerous other metal-based drugs have been tested to find an inexpensive metal complex with less harmful and improved curative effects. Ruthenium complexes, with their ability to mimic iron by binding to some biological molecules, reduced general toxicity, slow ligand exchange kinetics and higher affinity to cancer tissues compared to normal tissues, have desirable properties for suitable replacement of commercially available anticancer drugs [6]. Plenty of ruthenium complexes have been described in the literature, but only a few of them show remarkable anticancer activity. NAMI-A [ImH][trans-RuCl4(DMSO)(Im)] (Im = imidazole, DMSO = dimethylsulfoxide), the first ruthenium complex in clinical trials, has low direct cytotoxicity towards cancer cells in vitro; but, in vivo, it inhibits tumor metastasis. Due to its low therapeutic efficiency, progression of the disease in the clinical studies (phase I) and partial response (phase I/II) limited further clinical use of NAMI-A and resulted in the failure of the clinical development [7]. In addition, other ruthenium complex KP1019, [transtetrachlorobis-(1H-indazole)ruthenate(III)], designed by the Keppler’s group entered cancer clinical trials [8]. However, its low solubility limited its further examination and KP1019 is replaced with better soluble sodium salt, KP1339 [9].
Another type of complexes that attracted attention in recent years are arene ruthenium(II) complexes. General formula of this type of complexes is [(η6-arene)Ru(X)(Y)(Z)], and they are also called piano-stool complexes, because arene moiety looks like a seat, and ligands X, Y and Z are standing legs of chair. Widely known arene complexes were designed by Sadler and Dyson groups [10, 11]. Arene moiety (benzene and its derivatives) enables lipophilicity of complexes; type of substituents at the ring determine the electron distribution at the ruthenium(II) complex molecule, which affects the stability and facilitates the entry of compound into the cells [12]. Another feature of these complexes is possibility of use N,N-chelating ligands as X and Y ligand. This type of ligands can be aliphatic diamine, aromatic diamine and pyridine derivatives. Arene ruthenium complexes with ethylenediamine (en) chelating ligand have been studied by Sadler and coworkers. They reported remarkable effects in chemical and biological activity [13].
Many researches proved that ruthenium(II) polypyridyl complexes have significant biological properties [14, 15]. Most complexes are very reactive, imaging capable, with binding ability, and redox chemistry, which rank them as potential diagnostic and therapeutic drugs for cancer. Most of these ruthenium complexes are kinetically inert, because they have N,N-chelating ligands and octahedral structures. Complexes interact with biological molecules including DNA, proteins and RNA, both as probes and inhibitors. Interaction with these biological molecules often induces damage or toxicity. Many Ru(II) polypyridyl complexes are photosensitizers suitable for cancer photodynamic therapy (PDT), because of their photophysical and photothermal properties (long luminescence lifetime, significant two-photon absorption and photostability) [16,17,18]. These properties make them convenient in PDT, because of its requirement for both a non-toxic photosensitizer and a harmless light source. In the best case, PDT affects directly cancer cells, inducing their cell death by distinct mechanisms.
Barton et al. [19] first reported that [Ru(bpy)2dppz]2+ interacts with both mismatched and well-matched targets in the oligonucleotide. These, saturated tris(bidentate)ruthenium(II) complexes are lipophilic and cationic, and their unique octahedral structure contributes to a diversity of biological applications [20]. Biological activity of ruthenium(II) polypyridyl complexes was first reported more than 65 years ago. Dwyer and coworkers demonstrated that different types of enantiomeric [Ru(bpy)3]2+ and [Ru(phen)3]2+ have different biological activities [21]. The intercalating action of the Ru(II) polypyridyl complexes with DNA is a common and frequent mechanism of their anticancer activity [22]. In recent years, many organoruthenium(II) complexes coordinated to the arene moiety were synthesized, characterized and evaluated for in vitro biological activity in cancer and non-cancer cell lines, with results pointing out their interesting and quite distinct mode of action vastly different from the widely used chemotherapeutic platinum-based drugs [23, 24].
For these reasons, we investigated ruthenium(II) complexes containing different arene moieties (benzene, toluene and cymene) with pyrido[2′,3′:5,6]pyrazino[2,3-f][1, 10]phenanthroline (ppf) with the aim to study influence of arene moiety on the cytotoxicity of compounds, and to have a closer look into the mechanism of action.
Experimental section
Materials and measurements
All chemicals were of reagent-grade quality or higher, obtained from commercial suppliers and were used without further purification. Solvents were used as received. RuCl3·3H2O was purchased from Johnson Matthey (London, United Kingdom). [(η6-p-cymene)-RuCl2]2, [(η6-toluene)-RuCl2]2 and [(η6-benzene)-RuCl2]2 were prepared according to a published procedure [25].
1H and 13C NMR spectra were recorded in deuterated solvents on a Bruker Ultrashield Advance III spectrometer 500 (1H: 500 MHz) or VarianGemini-200 (1H: 200 MHz) MHz spectrometers at room temperature. The chemical shifts, δ, are reported in ppm (parts per million), and coupling constants (J) in Hertz. The residual solvent peaks have been used as an internal reference. The abbreviations for the peak multiplicities are as follows: s (singlet), d (doublet), dd (doublet of doublets), t (triplet), q (quartet), m (multiplet), and br (broad). Elemental analysis was carried out using an Elemental Vario EL III microanalyzer. Infrared spectra were recorded on a Nicolet 6700 FT-IR spectrometer using the ATR technique. ESI mass spectra measurements of complexes were carried out on a MS system LTQ Orbitrap XL with heated ESI ionization in methanol solutions. Measurement of ruthenium(II) content in cells was analyzed using ICP-MS and a Thermo Scientific iCAP Qc ICPMS (Thermo Scientific, Bremen, Germany) spectrometer with Qtegra operational software.
Synthesis of ligand and complexes
Synthesis of pyrido[2′,3′:5,6]pyrazino[2,3-f][1, 10]phenanthroline (ppf)
Pyridine-2,3-diamine (104 mg, 0.951 mmol, 1 equiv) was added to the boiling solution of 1,10-phenanthroline-5,6-dione [26] (200 mg, 0.951 mmol, 1 equiv) in methanol (16 ml) and mixture was refluxed for 1.5 h. Formed yellow precipitate was filtered and washed with cold MeOH, Et2O and dried in vacuo. Yield: 253 mg (94%). IR (ATR): ṽ 3052, 2977, 2925, 1580, 1480, 1412, 1356, 1274, 1075, 810, 737 cm−1. 1HNMR (200 MHz, CDCl3) δH 9.81 (H2, dd, J = 8.2, 1.8 Hz, 1H), 9.64 (H19, dd, J = 8.2, 1.8 Hz, 1H), 9.40 (H10, dd, J = 4.0, 1.9 Hz, 1H), 9.33 (H4 iH17, dd, J = 4.5, 1.6 Hz, 2H), 8.76 (H12, dd, J = 8.5, 1.9 Hz, 1H), 7.96–7.80 (H3, H18 iH11, m, 3H).
Synthesis of [(η 6-benzene)Ru(ppf)Cl]PF6 (C1)
The suspension of [(η6-benzene)RuCl2]2 (40 mg, 0.08 mmol, 1 equiv) in isopropanol (4 ml) was added to a solution of ppf (45 mg, 0.16 mmol, 2 equiv) in isopropanol/CH2Cl2 mixture (10 ml) 1/1 (V/V). The mixture was stirred at room temperature for 5 h and then filtered. NH4PF6 (39 mg, 0.24 mmol, 3 equiv) was then added to the solution and left to stir over night at room temperature. Formed yellow precipitate was filtered and washed with cold Et2O, MeOH and dried in vacuo. Yield: 82 mg (80%). Elemental analysis: Calcd. For C23H15ClF6N5PRu C, 42.97; H, 2.35; N, 10.89. Found: C, 43.16; H, 2.44; N, 10.93. 1H NMR (500 MHz, DMSO-d6) δ 10.17 (H2 i H19, dd, J = 11.5, 5.0 Hz, 2H), 9.70 (H4, d, J = 7.9 Hz, 1H), 9.64 (H17, d, J = 8.0 Hz, 1H), 9.50 (H10, dd, J = 3.8, 1.7 Hz, 1H), 8.89 (H12, dd, J = 8.5, 1.6 Hz, 1H), 8.31 (H3 i H18, ddd, J = 18.2, 8.1, 5.4 Hz, 2H), 8.15 (H11, dd, J = 8.5, 3.9 Hz, 1H), 6.43 (H23-PhH, s, 6H). 13C NMR (125 MHz, DMSO) δ 158.72, 158.56, 158.23, 158.23, 156.83, 149.80, 148.97, 141.91, 140.54, 140.54, 139.06, 138.81, 136.55, 136.22, 129.85, 129.51, 128.29, 128.24, 125.39, 87.62. (+)ESI–MS(m/z): ([M-PF6]+) calculated 498.00, found 498.00.
Synthesis of [(η 6-toluene)Ru(ppf)Cl]PF6 (C2)
The suspension of [(η6-toluene)RuCl2]2 (28 mg, 0.053 mmol, 1 equiv) in methanol (5 ml) was added to a solution of ppf (30 mg, 0.106 mmol, 2 equiv) in methanol/CH2Cl2 mixture (10 ml) 1/1 (V/V). The mixture was stirred at room temperature for 5 h and then filtered. NH4PF6 (26 mg, 0.159 mmol, 3 equiv) was then added to the solution and left to stir overnight. Formed yellow precipitate was filtered and washed with cold Et2O, MeOH and dried in vacuo. Yield: 54 mg (78%). Elemental analysis: Calcd. For C24H19ClF6N5PRu∙H2O C, 42.71; H, 2.84; N, 10.38. Found: C, 42.60; H, 2.67; N, 10.09. 1H NMR (500 MHz, DMSO-d6) δ H 10.07 (H2 i H19, d, J = 4.2 Hz, 2H), 9.74 (H4 i H17, dd, J = 9.7, 8.4 Hz, 2H), 9.52 (H10, dd, J = 4.0, 1.9 Hz, 1H), 8.94 (H12, dd, J = 8.5, 1.8 Hz, 1H), 8.34 (H3 i H18, ddd, J = 8.3, 5.4, 3.0 Hz, 2H), 8.18 (H11, dd, J = 8.6, 4.1 Hz, 1H), 6.52 (H26 i H26′, t, J = 6.0 Hz, 2H), 6.14 (H25 i H25′, d, J = 6.1 Hz, 2H), 5.94 (H27, t, J = 5.7 Hz, 1H), 2.32 (H24, s, 3H). 13C NMR (125 MHz, DMSO) δ 158.60, 158.44, 158.25, 149.86, 149.15, 148.95, 142.07, 140.71, 139.14, 138.84, 136.51, 136.18, 129.87, 129.53, 128.37, 128.31, 106.50, 90.82, 83.68, 81.03, 19.55. (+)ESI–MS(m/z): ([M-PF6]+) calculated 512.02, found 512.02.
Synthesis of [(η 6-p-cymene)Ru(ppf)Cl]PF6 (C3)
The solution of [RuCl2(η6-p-cymene)]2 (50 mg, 0.082 mmol, 1 equiv) in CH2Cl2 (4 ml) was added dropwise to a solution of ppf (46 mg, 0.164 mmol, 2 equiv) in methanol/CH2Cl2 mixture (10 ml) 1/1 (V/V). The mixture was stirred at room temperature for 5 h and then filtered. NH4PF6 (40 mg, 0.246 mmol, 3 equiv) was then added to the solution and left to stir overnight. Formed yellow precipitate was filtered and washed with cold Et2O, MeOH and dried in vacuo. Yield: 91 mg (83%). Elemental analysis: Calcd. For C27H23ClF6N5PRu C, 46.39; H, 3.32; N, 10.02. Found: C, 47.10; H, 2.77; N, 10.38. 1H NMR (500 MHz, DMSO-d6) δ H 10.05 (H2 i H19, m, 2H), 9.75 (H4, dd, J = 8.2, 1.0 Hz, 1H), 9.70 (H17, dd, J = 8.1, 1.0 Hz, 1H), 9.49 (H10, dd, J = 3.9, 1.9 Hz, 1H), 8.90 (H12, dd, J = 8.5, 1.8 Hz, 1H), 8.34 (H3 i H18, dt, J = 8.2, 5.6 Hz, 2H), 8.15 (H11, dd, J = 8.5, 4.0 Hz, 1H), 6.41 (H25 i H25′, d, J = 6.5 Hz, 2H), 6.18 (H26 i H26′d, J = 6.4 Hz, 2H), 2.71 (H28, m, J = 13.7, 6.8 Hz, 1H), 2.23 (H24, s, 3H), 1.01 (H29 i H29′, d, J = 6.9 Hz, 6H). 13C NMR (126 MHz, DMSO) δ 158.41, 158.25, 158.20, 149.86, 149.06, 148.86, 142.26, 140.89, 139.10, 138.78, 136.57, 136.24, 130.05, 129.72, 128.46, 128.29, 105.56, 103.50, 86.61, 84.89, 31.12, 22.43, 18.91. (+)ESI–MS(m/z): ([M-PF6]+) calculated 554.06, found 554.06.
Cell lines and culture conditions
Three human cancer cell lines: human lung adenocarcinoma (A549), human malignant melanoma (A375), human colorectal adenocarcinoma (LS 174T) and one human non-cancer lung fibroblast cell line (MRC-5) were maintained in nutrient medium (Roswell Park Memorial Institute) RPMI 1640 (Sigma-Aldrich). RPMI 1640 medium was supplemented with 10% fetal calf serum (FCS) (pH 7.2) (Sigma-Aldrich), 4-(2-hydroxyethyl)piperazine-1-ethanesulfonic acid (HEPES) (25 mM), l-glutamine (3 mM), penicillin (100 U/mL) and streptomycin (200 µg/mL). Cells were grown as flat monolayer culture in tissue culture flasks (Thermo Scientific Nunc™), at 37 °C in a humidified incubator with 5% CO2.
MTT assay
Cells were seeded into 96-well culture plates (Thermo Scientific Nunc™) at cell densities of 6000 c/w (A375, LS 174T) and 5000 c/w (A549, MRC-5) in 100 µL of cell culture medium. After 24 h, the cells were treated with serial dilutions of the investigated ruthenium(II)-arene complexes C1–C3 and ligand ppf. Tested compounds were dissolved in DMSO at a concentration of 10 mM as stock solution (the final concentration of DMSO did not exceed 1% (v/v) per well) immediately prior to use. Cisplatin was used in this study as a reference compound. After 72 h of continuous drug incubation, MTT solution (3-(4.5-dimethylthiazol-2-yl)-2.5-diphenyltetrazolium bromide, Sigma-Aldrich), was added to each well (5 mg/mL) [27]. The culture plates were incubated for the next 4 h at 37 °C, and finally 10% sodium dodecyl sulfate (SDS) was added to dissolve formed formazan crystals. Absorbances were read on a microplate reader (ThermoLabsystemsMultiscan EX 200–240 V) after 24 h at a wavelength of 570 nm. Results are eventually expressed as IC50 values (concentration of investigated compound that cause 50% decrease in the number of viable cells in a treated cell population compared to a non-treated control).
Trypan blue exclusion assay
Trypan blue exclusion assay was applied to investigate whether the result of metabolic MTT assay represents a real viability status of the C3-treated A375 cells. Basic principle of the test is that viable cells possess intact cell membranes which exclude trypan blue dye, whereas non-viable cells do not. A375 cells were treated with IC50 concentration of C3 for 24 h, 48 h and 72 h. After the incubation period, cells were harvested and cell suspension stained by trypan blue solution (0.4%), according to the previously described protocol [28]. Cells were visually examined by light microscopy and cell number determined using a hemocytometer chamber. Viable cells appear with clear cytoplasm, while non-viable appear with blue cytoplasm.
Flow cytometric analysis of cell cycle phase distribution (PI staining)
Flow-cytometric analysis of the cell cycle phase distribution was performed in fixed A375 cells after staining with PI [29]. 2 × 105 A375 cells were seeded into 6-well plates (Thermo Scientific Nunc™) in the nutrient medium. After 24 h of growth, cells were continually exposed to C3 or cisplatin at concentrations corresponding to 0.5 × IC50, IC50 and 2 × IC50 for additional 24 h, 48 h, and 72 h. The cells were collected and fixed overnight in 1 mL of 70% ice-cold ethanol. After fixation, cells were washed with cold phosphate-buffered saline (PBS), and incubated with 100 µg/mL of RNaseA (1 mg/mL in PBS) for 30 min, at 37 °C. Immediately before flow-cytometric analysis, cells were stained in the dark with 50 µg/mL of PI (400 µg/mL in PBS). 10.000 cells were analyzed for each sample by a fluorescence-activated cell sorting (FACS) BD Calibur flow cytometer (Becton–Dickinson, Heidelberg Germany), at 488 nm excitation line. Collected data were analyzed by Cell Quest computer software.
Morphological analysis of cell death by fluorescent microscopy
A375 cells (5000 c/w) were seeded into 96-well plate (Thermo Scientific Nunc™) in 100 µL of cell culture medium. After 24 h of growth, cells were treated with IC50 and 2 × IC50 concentrations of C3 or cisplatin and left for another 72 h to incubate with the investigated drugs. Further, cells were stained with the acridine orange (AO, 5 µg/mL) and ethidium bromide (EtBr, 3 µg/mL) [30], and immediately after observed under the Bio-Rad™ ZOE™ Fluorescent Cell Imager, using 20x/0.4 objective.
Flow cytometric analysis of apoptotic potential (Annexin V-FITC/PI staining)
A375 cells were seeded into 6-well plates (2 × 10 [5] c/w) and after 24 h of growth treated with 0.5 × IC50, IC50 and 2 × IC50 concentrations of C3 or cisplatin. Following the 24-h and 48-h incubation time, cells were harvested and resuspended in 200 µL 1 × Binding Buffer (10 mM HEPES/NaOH pH 7.4, 140 mM NaCl, 2.5 mM CaCl2). 100 µL of cell suspension (~ 1 × 105 cells) was transferred to a 5 mL round-bottom polystyrene tube (Falcon, Corning) and mixed with 5 µL of both FITC Annexin V (BD Pharmingen) and PI (50 µg/mL in PBS) [31]. After incubation of 15 min in the dark, at 37 °C, 400 µL of 1 × Binding Buffer was added to each tube and samples were analyzed using a FACS BD Calibur flow cytometer and Cell Quest computer software.
Cellular uptake and subcellular distribution studies
For cellular uptake and subcellular distribution studies, 2 × 106 A375 cells were grown in 75 cm2 culture flasks (Thermo Scientific Nunc™). After 24 h, cells were treated with equimolar concentrations (10 µM) of C3 or cisplatin, and incubated for 24 h. Cells were harvested by trypsinization, and cell pellet was collected by centrifugation at 1500 rpm for 10 min. Cell viability was determined by the Trypan blue exclusion assay. For subcellular distribution studies, cell pellet was further lysed with Subcellular Protein Fractionation Kit for Cultured Cells, according to the manufacturer’s instructions (Thermo Scientific, cat.no. 78840). Total intracellular accumulation and accumulation in subcellular compartments of C3 and cisplatin in A375 cells were analyzed using the inductively coupled plasma mass spectrometry (ICP-MS) and a Thermo Scientific iCAP Qc ICP-MS (Thermo Scientific, Bremen, Germany) spectrometer with operational software Qtegra [32]. The levels of C3 and cisplatin, found in cells after the treatment, were expressed as the amount of metal (ruthenium or platinum) (ng) taken up per 106 cells.
Generation and analysis of MCTSs
A375 cells were seeded at cell density of 1000 c/w in 200 µL of RPMI containing 10% FCS in a low attachment NunclonSphera 96-well ultra-low attachment plate (Thermo Scientific Nunc™) to form multicellular tumor spheroids (MCTSs) [33]. MCTS aggregates of approximately 400 µm in diameter were formed after one day incubation with 5% CO2 and 20% O2 at 37 °C. The formation and growth of MCTSs were examined and imaged with a Bio-Rad™ ZOE™ Fluorescent Cell Imager, using 20x objective.
A375 MCTSs (diameters approximately 500 µm) were treated by carefully replacing 50% of the medium with fresh nutrient medium for control MCTSs or with C3-supplemented medium for treated MCTSs. MCTSs were incubated with freshly made serial dilution of C3 (up to 120 µM) for another 72 h. The cytotoxicity of C3 toward the A375 tumor spheroids was investigated by MTT assay, after transferring MCTSs into flat-bottomed 96-well plate (Thermo Scientific Nunc™).
For the live/dead analysis of A375 MCTSs, dual fluorescent staining with Calcein-AM (Sigma-Aldrich) and propidium iodide (PI, Sigma-Aldrich) was performed. The intracellular esterases convert non-fluorescent cell permeable Calcein dye to intensely fluorescent Calcein in live cells, allowing the assessment of cell viability (green fluorescence). Propidium iodide is able to enter dead cells with a porous membrane and to intercalate with nucleic acid to give a fluorescence signal (red fluorescence), allowing the assessment of cell death [34]. After the formation, A375 spheroids were treated with C3 or cisplatin (concentrations corresponding to 3 × IC50). Briefly, after the 96 h treatment, the MCTSs were incubated with Calcein-AM (3 µM) and PI (5 µM) solutions for 30 min, in the dark, at 37 °C, and imaged directly using a Bio-Rad™ ZOE™ Fluorescent Cell Imager (20x/0.4 objective).
Results and discussion
Synthesis and characterization of the complexes
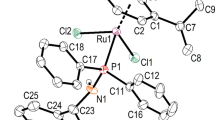
All complexes were synthesized following the synthetic routes described in Scheme 1. The resulting complexes were found to be soluble in DMSO, methanol and acetonitrile.
All complexes were characterized by 1H NMR, IR, elemental analysis and mass spectrometry. In the 1H NMR spectra of the complexes, all protons and carbons appear at chemical shifts expected for such types of compounds. Importantly, as determined by the integrals of the 1H NMR spectra, the arene to ppf ratio was found to be 1:1.
In complexes spectra there is a deviation of chemical shifts in relation to the free ligand. Chemical shifts of ppf in C1 are closest to ones of free ligand, due to presence of benzene arene moiety. Changes of ligand proton shifts are more noticeable in complexes with toluene (C2) and cymene (C3) moiety. Peak positions are at the chemical shifts expected for such type of complexes. This suggested a piano-stool geometry for the complexes with a coordinated chloride ligand. The composition of the inner sphere is also confirmed by analysis of the ESI–MS spectra, where a single peak was found at m/z 498.00, 512.02, and 554.06 for the complexes C1, C2 and C3, respectively, without a counter ion PF6−.
Since all complexes were found to be well soluble in DMSO, stock solutions for IC50 were prepared in this solvent. In these experimental conditions, the arene ligand and chlorido anion from the complex can undergo a ligand exchange with the DMSO molecule, leading potentially to problems during the IC50 value examination [35]. For this reason, we first investigated the stability of all complexes in DMSO-d6 solutions using 1H NMR spectroscopy. We followed the changes in the spectra of the complexes for 24 h. The signals of the ligands did not change over time in all complexes, indicating that the complexes are stable in DMSO solution.
MTT assay
The cytotoxicity of three new ruthenium(II)-arene complexes C1–C3, as well as their corresponding ligand, ppf, were investigated by the colorimetric MTT assay in a panel of three human neoplastic cell lines (A549, A375, LS 174T) and one human non-tumor cell line (MRC-5), in comparison to cisplatin as a referent compound. Results obtained after 72 h of continuous drug treatment, are presented as IC50 values (µM) (Table 1), provided from cell survival diagrams (Fig. 1). Complexes C1–C3 showed IC50 values in the micromolar range of 15.8–95.9 µM, except of C1 which was found to be non-active in LS 174T cells, in the tested range of concentrations up to 100 µM. Investigated complexes C1–C3 displayed moderate cytotoxic activity in a panel of tested cell lines, but within every of these cell lines their activity remained consistent, with C3 being the most potent. C3 exhibited the highest activity and cytoselectivity toward malignant melanoma A375 cells. With IC50 value being 15.8 µM, C3 was approximately six times more active than C1 and C2. Also C3 was capable of reducing viability of A375 cells four times more efficiently than of the MRC-5 cells, as seen from the SI values (Table 1). Obtained results showed that cytotoxic activity of these ruthenium(II)-arene complexes obviously depends on the coordinated ƞ6-arene ligand [36]. This observation is in agreement with a previous report describing the more potent activity of ruthenium complexes containing arene ligands such as p-cymene (as in C3) [37]. Ligand exhibited consistent activity in all tested cell lines, with IC50 value of approximately 6 µM, which is in line with cisplatin. Results show that ligand was at least 2.4 times more active than its corresponding complexes C1–C3. It is previously demonstrated that some intercalative ligands can display higher activity toward cancer cell lines in comparison to their corresponding ruthenium(II)-arene complexes [38]. According to the results of MTT assay, C3 was chosen for further biological studies of its effects in A375 malignant melanoma cells.
IC50 values (µM) are presented as an average (± SEM) of three independent experiments. *SI-selectivity index for tested complexes, ligand and cisplatin, in tumor cell lines (A549, A375 and LS 174T), related to non-tumor MRC-5 cell line: SIA549 (IC50 MRC-5/IC50 A549), SIA375 (IC50 MRC-5/IC50 A375), SILS 174T (IC50 MRC-5/IC50 LS 174T).
Trypan blue exclusion assay
The cell death assays can be divided in two major groups: assays that measure bona fide cell death (direct measuring of cell death), and assays that quantify biochemical processes that are viewed as viability markers (indirect measuring of cell death) [39]. The MTT assay determines cell viability by measuring mitochondrial dehydrogenase activity. Therefore, as metabolic type of assays it cannot distinguish between cytotoxic and antiproliferative effects. The investigation of morphological changes induced by investigated complexes (see Fig. 4, bright field) showed that A375 cells exposed for 72 h to C3, at concentrations corresponding to IC50 and 2 × IC50, still have preserved morphology and number of cells is not dramatically changed as expected based on the IC50 values obtained from MTT assay. To investigate viability status of the A375 cells, considering the cellular membrane integrity, under the C3-treatment and to compare it to the result of metabolic MTT assay, the cells were submitted to the Trypan blue exclusion assay after the 24 h, 48 h and 72 h of continuous drug incubation. Number of live and dead cells in treated cell population in comparison to non-treated cells is evaluated visually by light microscopy, and cell growth curves were plotted according to the viable cell numbers (Fig. 2). Results showed that A375 cells treated with IC50 concentration (15.8 µM, see Table 1) of C3 (obtained from the MTT assay) were not only capable of dividing and growing during the 3 days continuous drug incubation time, but also 98% of the cells appeared as viable after the 72 h treatment, since they excluded Trypan blue dye. C3-treated A375 cells showed doubling time of 34 h, comparing to 22 h doubling time in control cell population. These results differ from the result of MTT assay [40], and indicate that C3 displays specific effects on the proliferation and metabolism of A375 cells.
Flow cytometric analysis of cell cycle phase distribution (PI staining)
To investigate whether the C3 interferes with cell cycle progression, flow cytometry analysis of the DNA content was performed in A375 cells using PI staining. A375 cells were treated with 0.5 × IC50, IC50 and 2 × IC50 concentrations of C3 or cisplatin for 24 h, 48 h and 72 h. As shown in Fig. 3, upon exposure of the A375 cells to C3, cells distribution over the cell cycle phases has not changed in considerable amount through 3 days continuous C3-treatment, when compared to the non-treated cell population (Fig. 3a). After 72 h incubation with C3, approximately 6% of the cells were detected in hypodiploid sub-G1 region, representing a cell population undergoing programmed cell death or necrosis (Fig. 3c). It seems from these results, that neither increased C3 concentration nor prolonged drug incubation time, triggered substantial alterations in the cell cycle phase distribution of cells, and C3 does not seems to interfere with A375 cells replication. On the contrary, cisplatin showed prominent effects on A375 cell cycle phase distributions, comparing to control cells. After the 24-h treatment, cisplatin-induced strong S phase arrest, up to 48.02% (0.5 × IC50), 61.18% (IC50) and 59.38% (2 × IC50), versus 20.97% in control cells (Fig. 3a). Cisplatin-induced S phase arrest in the A375 cells, suggest block of the DNA replication and is in agreement with the literature regarding the effect of cisplatin on cell cycle progression [41]. Prolonged treatment (48 h) with cisplatin, induced further changes characterized by decrease in percentage of cells in S phase, compensated with the accompanying increase in the percentage of the cells arrested in the G2-M phase of the cell cycle (Fig. 3b). The 72 h incubation of A375 cells with cisplatin triggered cell death, since cells rapidly and dosage-dependently accumulated in the sub-G1 fraction.
Fluorescence microscopy analysis of cell death (AO/EtBr staining)
Acridine orange/ethidium bromide (AO/EtBr) dual staining of A375 cells treated with IC50 and 2 × IC50 concentrations of C3 or cisplatin was performed to analyze morphological characteristics of cell death by fluorescence microscopy. Morphological appearance of cells is often used as criteria for distinguishing among the different types of cell death, without clear remarks about underlying biochemical mechanisms [42]. Photomicrographs of A375 cells stained by AO/EtBr dyes, following 72 h treatment were captured under the bright field and green channel, and presented in Fig. 4. Control cells are dense, polygonal in shape and light green colored. The A375 cells treated with lower concentration (IC50) of cisplatin, started to lose their normal morphology, reduced in number and majority of cells became rounded, with appearance of enlarged individual cells with long pseudopods, implying cell senescence [43]. Mixed cell population was present, with predominantly early apoptotic cells, with intact membranes and condensed green fluorescent chromatin, and individual cells resembling late apoptotic morphology (orange to red fluorescing nuclei with fragmented chromatin). With the cisplatin dosage increase, necrotic, swollen cells with red fluorescing nucleus are present in noticeable extent. Under the 72 h continual drug treatment, C3 caused less obvious cytological changes in A375 cells, when compared to cisplatin. Concentration-dependent reduction in A375 cell number is observed; cells became more rounded with green fluorescent nuclei. Disruption of intercellular connections and increase in percentage of cells with orange/red fluorescent nucleus, with or without low chromatin condensation, is noticeable. However, other characteristic hallmarks of apoptosis, such as membrane blebbing and apoptotic bodies, have not been noticed.
Photomicrographs of A375 control cells and cells exposed for 72 h to C3 or cisplatin, at concentrations corresponding to IC50 and 2 × IC50. Left—A375 cells investigated under the bright field, and right—acridine orange/ethidium bromide-stained A375 cells observed under the Bio-Rad™ ZOE™ Fluorescent Cell Imager, using 20×/0.4 objective. Arrows indicate characteristic changes in cell morphology after the treatment with the tested compounds. Scale bar: 100 µm
Flow cytometric analysis of apoptotic potential (Annexin V-FITC/PI staining)
Investigation of apoptosis/necrosis induction potential of complex C3 and cisplatin in A375 cells is assessed by flow cytometry following Annexin V-FITC/PI staining of treated A375 cells. The obtained experimental data revealed that C3 after 24 h and 48 h of A375 cells treatment does not seems to initiate neither apoptotic nor necrotic cell death, since approximately same percent of C3-treated and non-treated, control A375 cells stained as FITC(+)/PI(−) (early apoptotic cells), FITC(+)/PI(+) (late apoptotic or necrotic cells) and FITC(−)/PI(+) (dead cells) (Figure S10). Additionally, 24-h treatment with cisplatin did not induce apoptosis, only percent of cells already dead increased to 13.11% (with IC50 concentration treatment) compared to 8.95% in control, which is in accordance with the literature data [44]. Further incubation with cisplatin (48 h) promoted very small increase in percent of early apoptotic cells (up to 5.21% (2 × IC50), versus 0.45% in control cells). Approximately similar percent of cells stained as double positive for both FITC and PI markers, and appeared as late apoptotic. The 48-h treatment with cisplatin introduced 22.06% (IC50) and 31.61% (2 × IC50) of A375 cells in cell death (stained as FITC(-)/PI(+)), supporting high cytotoxic activity of cisplatin toward A375 cells, without clear externalization of phosphatidylserine (Annexin V-FITC-staining) as one of the markers of apoptosis. Gained results are in accordance with the findings from AO/EtBr analyses. It seems that C3 exerts its activity in A375 cells in a programmed cell death independent way, and precise mechanism remains to be further investigated.
Cellular uptake and localization
The total cellular uptake and subcellular distribution of Ru (from C3) and Pt (from cisplatin) in the A375 cells was determined by ICP-MS, to reveal a possible relationship between the cellular uptake and in vitro cytotoxicity. After 24 h of treatment with equimolar concentrations (10 µM), both C3 and cisplatin entered the cells, but in a different amounts. Complex C3 entered the A375 cells less efficiently compared to cisplatin, as there were 1.55 ± 0.04 ng of Ru in 106 treated cells and 28.58 ± 0.8 ng of Pt in 106 treated cells (Fig. 5a). Lower C3-intracellular content may be consequence of continued proliferation of A375 cells under the C3-treatment, having as a result reduced uptake and enhanced efflux of the drug. The ICP-MS analysis of subcellular localization of the tested compounds revealed different distribution of C3 (39.1% localized in the cytosol, 44.6% membrane/organelle, 8.2% nucleus, 3.9% chromatin proteins, 4.2% cytoskeletal fraction) and cisplatin (18.2% in the cytosol, 67.5% membrane/organelle, 7.7% nucleus, 3.6% chromatin proteins, 3.0% cytoskeletal fraction) (Fig. 5b). Cisplatin accumulated in the highest amount in the membrane fraction of cells, while C3 accumulated in approximately equal amount in the membrane and cytoplasmic fraction of cells. This could be related to the different reactivity of C3 and cisplatin with proteins and other subcellular components. Accumulation of cisplatin in membrane/organelle fraction of A375 cells is expected [45], but susceptibility to cisplatin damage depends on specific cellular/tissue characteristics and cisplatin interactions with macromolecules, while trafficking through diverse subcellular structures. Mitochondria are the key intracellular organelles responsible for the generation of energy and regulation of cellular metabolism. Therefore, tumor cells need to reprogram mitochondrial metabolism to meet increased demands for energy and building blocks required for uncontrolled proliferation. Growing evidences indicate the significance of mitochondrial reprogramming for proliferation of BRAF-mutated melanomas (such as A375 cells) [46], rationalizing mitochondria as a key therapeutic target for these tumors [47]. This may explain remarkably accumulation of cisplatin and also C3 in mitochondrial fraction of A375 cells.
Analyses of the efficacy of C3 in a 3D MCTS cancer model
Primary investigations of the efficacy of potential anticancer drugs are usually performed in two-dimensional (2D) monolayer cell cultures, which fail to truly represent the in vivo environment and complex interactions between cells and extracellular matrix components. Cells grown as 2D culture differ morphologically, physiologically and metabolically from the cells grown in 3D culture environment [48]. While cells in 2D model systems are uniformly exposed to oxygen, nutrients and other biochemical signals, cells in 3D models are exposed to gradients of these conditions and subsequent cellular layers of different proliferative capability appoint different drug response. Multicellular tumor spheroids (MCTSs) are heterogeneous cellular aggregates that can be introduced as 3D models that resemble properties of solid tumors in vitro. 3D model of A375 cells was introduced to investigate the efficacy of C3 on the viability and growth kinetics of MCTSs over a period of time, in comparison to cisplatin. 72 h of continuous drug incubation of MCTSs with C3 revealed sevenfold lower sensitivity of A375 MCTSs to C3 (IC50 = 110 µM) than in 2D cell culture (see Table 1), which is in accordance with the stated observation of different organization and physiology of tumor spheroids (Fig. 6).
Investigations in the 3D model of A375 cells further disclosed different effects of 4-day treatment with C3 (3 × IC50) and cisplatin, on growth of MCTSs. The sizes of the MCTSs that were untreated (control) and treated with tested C3, continued to growth over time (Fig. 7a), indicating that C3 was not able to stop the proliferation of A375 cells in tumor spheroid with investigated concentrations. This result can be compared to the results of Trypan blue exclusion assay that showed sustained proliferation of A375 cells under the C3-treatment, with prolonged doubling time, compared to control A375 cell population. On the contrary, cisplatin-treated MCTSs decreased in size over time. Their growth was stopped already 24 h after the treatment. Bright field images show distinct cell shrinkage and time-dependent increase of dark dead cells in the center of cisplatin-treated MCTSs. These observations are far less present in C3-treated MCTSs.
Growth inhibition of A375 multicellular tumor spheroids treated with 3 × IC50 concentrations of C3 or cisplatin. a A375 multicellular tumor spheroids observed under the bright field, and b Calcein-AM/PI dual stained A375 multicellular tumor spheroids after 96-h treatment with C3 or cisplatin (3 × IC50). Red signal indicates dead cells; green signal, viable cells. Images obtained using a Bio-Rad™ ZOE™ Fluorescent Cell Imager (20x/0.4 objective). Scale bar: 100 µM
The live/dead analysis of A375 MCTSs is performed after the 96 h treatment with 3 × IC50 concentrations of C3 or cisplatin, by dual fluorescence staining using the Calcein-AM and PI dyes. Calcein-AM/PI dual staining displayed sustained compactness of control spheroids with predominantly live, green fluorescent cells, and appearance of small proportion of red fluorescing cells in the sphere’s core, presumably due to the overgrowing of sphere and deprivation of cells in oxygen and other nutrients, which consequently led to cell death (Fig. 7b). The compactness of cisplatin-treated MCTSs was disturbed and spheroids dispersed, mostly from the sphere’s periphery, to individual cells. Fluorescent staining confirmed that in the disintegrated sphere, majority of disseminated cells were dead, with small population of live core cells. It seems that the structure of cisplatin-treated spheroids was highly sensitive to the slightest mechanical disturbances caused by addition of dyes, and hence resulted in decomposed spheroid (Fig. 7b), which suggests that cisplatin possess capability to induce disruption of intercellular connections in spheroids. On the other hand, compactness of C3-treated MCTSs after the Calcein-AM/PI staining was preserved, with appearance of red fluorescent necrotic core with predominantly live peripheral edge of cells. These results, once again suggest that investigated compound C3 has a different mode of action in comparison to cisplatin in A375 cells.
Conclusions
In this work, three new half-sandwich Ru(II)–arene complexes: C1 ([(ƞ6-benzene)Ru(ppf)Cl]PF6, C2 ([(ƞ6-toluene)Ru(ppf)Cl]PF6) and C3 ([(ƞ6-p-cymene)Ru(ppf)Cl]PF6); ppf = pyrido[2′,3′:5,6]pyrazino[2,3-f][1, 10]phenanthroline were synthesized and characterized. According to spectral and elemental analysis, all complexes possess “piano-stool” geometry with arene ligand as stool seat. The continuous drug treatment for 72 h showed cytotoxicity in micromolar range in three human neoplastic cell lines (A549, A375, LS 174T). In addition, the demonstrated activity of the investigated ruthenium(II)-arenes clearly depended on the structure of coordinated ƞ6-arene ligand, as complex containing p-cymene moiety, C3, showed the highest activity and cytoselectivity toward malignant melanoma A375 cells, with IC50 value being 15.8 µM. Biological studies to unveil the mechanism of action of C3 in A375 cells, revealed that C3 displayed specific effect on the proliferation and metabolism of A375 cells, although did not notably interfere with cell cycle progression. The investigation of the total cellular uptake and subcellular distribution of Ru and Pt in the A375 cells, determined by ICP-MS, showed that C3 entered cells less efficiently comparing to cisplatin. Detailed analysis of subcellular localization revealed different distribution of C3 and cisplatin. Cisplatin accumulated in the highest amount in the membrane (organelle) fraction of cells, while C3 accumulated in approximately equal amount in the membrane and cytoplasmic fraction. Tested ruthenium(II)-arene complexes possess positive charge, but carry ppf ligands with intercalating and lipophilic characteristics, which may explain the intracellular distribution of C3. Negative charges of double-stranded DNA and single-stranded RNA (which is present both in nucleus and cytoplasm), as well as lipid membrane bilayers of organelles (such as mitochondria), allow C3 to target these biomolecules or organelles. Still in the 3D model of melanoma cells, C3 was not able to stop the growth of spheroids, neither to compromise their compactness, while cisplatin evidently did so. Based on the results presented in this work, we can conclude that investigated ruthenium complex C3 exhibited lower toxicity in vitro and displayed different mechanism of action comparing to cisplatin. Therefore, these types of complexes could give an appropriate guideline for further structural modifications of ruthenium-based compounds that could combat cisplatin resistance in aggressive and metastatic tumors, such as melanoma.
Abbreviations
- NAMI-A:
-
[ImH][trans-RuCl4(DMSO)(Im)]
- KP1019:
-
[transtetrachlorobis-(1H-indazole)ruthenate(III)]
- KP1339:
-
Sodium [transtetrachlorobis-(1H-indazole)ruthenate(III)]
- DMSO:
-
Dimethyl sulfoxide
- DNA:
-
Deoxyribonucleic acid
- RNA:
-
Ribonucleic acid
- PDT:
-
Photodynamic therapy
- A549:
-
Human lung adenocarcinoma cells
- A375:
-
Human malignant melanoma cells
- LS 174T:
-
Human colorectal adenocarcinoma cells
- MRC-5:
-
Non-tumor human lung fibroblast cells
- ICP-MS:
-
Inductively coupled plasma mass spectrometry
- RPMI 1640:
-
Roswell Park Memorial Institute nutrient medium (1640)
- FCS:
-
Fetal calf serum
- HEPES:
-
4-(2-hydroxyethyl)piperazine-1-ethanesulfonic acid
- MTT:
-
3-(4.5-dimethylthiazol-2-yl)-2.5-diphenyltetrazolium bromide dye
- SDS:
-
Sodium dodecyl sulfate
- PI:
-
Propidium iodide
- PBS:
-
Phosphate-buffered saline
- RNaseA:
-
Ribonuclease A
- FACS:
-
Fluorescence-activated cell sorting
- AO:
-
Acridine orange
- EtBr:
-
Ethidium bromide
- FITC:
-
Fluorescein isothiocyanate
- MCTS:
-
Multicellular tumor spheroid
References
Jakupec MA, Galanski M, Arion VB, Hartinger CG, Keppler BK (2008) Dalton Trans 2:183–194
Sledge G, Loehrer PJ, Roth BJ, Einhorn LH (1988) J Clin Oncol 6:1811–1814
Florea A-M, Büsselberg D (2011) Cancers 3:1351–1371
Liang X-J, Meng H, Wang Y, He H, Meng J, Lu J, Wang PC, Zhao Y, Gao X, Sun B (2010) Proc Natl Acad Sci USA 107:7449–7454
Fong TT-H, Lok C-N, Chung CY-S, Fung Y-ME, Chow P-K, Wan P-K, Che C-M (2016) Angew Chem Int Ed 55:11935–11939
Hu PC, Wang Y, Zhang Y, Song H, Gao FF, Lin HY, Wang ZH, Wei L, Yang F (2016) RSC Adv 6:29963–29976
Rademaker-Lakhai JM, van den Bongard D, Pluim D, Beijnen JH, Schellens JH (2004) Clin Cancer Res 10:3717–3727
Hartinger CG, Zorbas-Seifried S, Jakupec MA, Kynast B, Zorbas H, Keppler BK (2006) J Inorg Biochem 100:891–904
Bytzek AK, Koellensperger G, Keppler BK, Hartinger CG (2016) J Inorg Biochem 160:250–255
Murray BS, Babak MV, Hartinger CG, Dyson PJ (2016) Coord Chem Rev 306(part 1):86–114
Dougan SJ, Sadler PJ (2007) Chimia 61:704–715
Suss-Fink G (2010) Dalton Trans 39:1673–1688
Morris RE, Aird RE, del Socorro Murdoch P, Chen H, Cummings J, Hughes ND, Parsons S, Parkin A, Boydand G, Jodrell DI (2001) J Med Chem 44:3616–3621
Notaro A, Gasser G (2017) Chem Soc Rev 46:7317–7337
Zeng L, Gupta P, Chen Y, Wang E, Ji L, Chao H, Chen ZS (2017) Chem Soc Rev 46:5771–5804
Monro S, Colón KL, Yin H, Roque J, Konda P, Gujar S, Thummel RP, Lilge L, Cameron CG, McFarland SA (2018) Chem Rev. https://doi.org/10.1021/acs.chemrev.8b00211
Jakubaszek M, Goud B, Ferrari S, Gasser G (2018) Chem Commun 54:13040–13059
Heinemann F, Karges J, Gasser G (2017) Acc Chem Res 50:2727–2736
Song H, Kaiser JT, Barton JK (2012) Nat. Chem 4:615–620
Kilah NL, Meggers E (2012) Aust J Chem 65:1325–1332
Dwyer FP, Gyarfas EC, Rogers WP, Koch JH (1952) Nature 170:190
Tan C, Wu S, Lai S, Wang M, Chen Y, Zhou L, Zhu Y, Lian W, Peng W, Ji L (2011) Dalton Trans 40:8611–8621
Baroud AA, Mihajlović-Lalić LJE, Stanković D, Kajzerberger M, van Hecke K, Grgurić-Šipka S, Savić A (2017) J Serb Chem Soc 82(3):267–275
Jovanović K, Tanić M, Ivanović I, Gligorijević N, Dojčinović B, Radulović S (2016) J Inorg Biochem 163:362–373
Jensen SB, Rodger SJ, Spicer MD (1998) J Organomet Chem 556:151–158
Gillard RD, Hill REE, Maskill R (1970) J Chem Soc A 1447–1451
Supino R (1995) Humana Press 43:137–149
Strober W (2001) Curr Protoc Immunol Appendix 3B:1–2
Ormerod MG (1994) Oxford University Press, Oxford, 119–125
Spector DL, Goldman RD, Leinwand LA (1998) Cold Spring Harbor Laboratory Press, 1
van Engeland M, Nieland LJ, Ramaekers FC, Schutte B, Reutelingsperger CP (1998) Cytometry 31:1–9
Ott I, Biot C, Hartinger C (2014) Wiley, New York, 63–97
Selby M, Delosh R, Laudeman J, Ogle C, Reinhart R, Silvers T, Lawrence S, Kinders R, Parchment R, Teicher BA, Evans DM (2017) SLAS Discov. 22(5):473–483
Sarangapani S, Patil A, Ngeow YK, Elsa Mohan R, Asundi A, Lang MJ (2018) Integr Biol 10(5):313–324
Patra M, Joshi T, Pierroz V, Ingram K, Kaiser M, Ferrari S, Spingler B, Keiser J, Gasser G (2013) Chem Eur J 19:14768–14772
Yan YK, Melchart M, Habtemariam A, Sadler PJ (2005) Chem Commun 4764–4776
Nikolić S, Rangasamy L, Gligorijević N, Aranđelović S, Radulović S, Gasser G, Grgurić-Šipka S (2016) J Inorg Biochem 160:156–165
Valladolid J, Hortigüela C, Busto N, Espino G, Rodríguez AM, Leal JM, Jalón FA, Manzano BR, Carbayo A, García B (2014) Dalton Trans 43:2629–2645
Kepp O, Galluzzi L, Lipinski M, Yuanand J, Kroemer G (2011) Nat Rev Drug Discov 10:221–237
Berridge MV, Tan AS (1993) Arch Biochem Biophys 303(2):474–482
Wang D, Lippard S (2005) Nat Rev Drug Discov 4:307–320
Kroemer G, Galluzzi L, Vandenabeele P, Abrams J, Alnemri ES, Baehrecke EH, Blagosklonny MV, El-Deiry WS, Golstein P, Green DR, Hengartner M, Knight RA, Kumar S, Lipton SA, Malorni W, Nuñez G, Peter ME, Tschopp J, Yuan J, Piacentini M, Zhivotovsky B, Melino G (2009) Cell Death Differ 16:3–11
Sun X, Shi B, Zheng H, Min L, Yang J, Li X, Liao X, Huang W, Zhang M, Xu S, Zhu Z, Cui H, Liu X (2018) Cell Death Dis 9:260
Radisavljević S, Bratsos I, Scheurer A, Korzekwa J, Masnikosa R, Tot A, Gligorijević N, Radulović S, Rilak Simović A (2018) Dalton Trans 47(38):13696–13712
Sancho-Martínez SM, Prieto-García L, Prieto M, López-Novoa JM, López-Hernández FJ (2012) Pharmacol Therapeut 136:35–55
Corazao-Rozas P, Guerreschi P, André F, Gabert PE, Lancel S, Dekiouk S, Fontaine D, Tardivel M, Savina A, Quesnel B, Mortier L, Marchetti P, Kluza J (2016) Oncotarget 7(26):39473–39485
Hong SK, Starenki D, Wu PK, Park JI (2017) Cancer Biol Ther 18(2):106–114
Acland M, Mittal P, Lokman NA, Klingler-Hoffmann M, Oehler MK, Hoffmann P (2018) Proteomics Clin Appl 12(3):1–13
Acknowledgements
Ministry of Education, Science and Technological Development of the Republic of Serbia, Grant numbers 172035 and III 41026.
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Pavlović, M., Nikolić, S., Gligorijević, N. et al. New organoruthenium compounds with pyrido[2′,3′:5,6]pyrazino[2,3-f][1, 10]phenanthroline: synthesis, characterization, cytotoxicity, and investigation of mechanism of action. J Biol Inorg Chem 24, 297–310 (2019). https://doi.org/10.1007/s00775-019-01647-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00775-019-01647-4