Abstract
Nitrogenases catalyze the biological fixation of inert N2 into bioavailable ammonium. They are bipartite systems consisting of the catalytic dinitrogenase and a complementary reductase, the Fe protein that is also the site where ATP is hydrolyzed to drive the reaction forward. Three different subclasses of dinitrogenases are known, employing either molybdenum, vanadium or only iron at their active site cofactor. Although in all these classes the mode and mechanism of interaction with Fe protein is conserved, each one encodes its own orthologue of the reductase in the corresponding gene cluster. Here we present the 2.2 Å resolution structure of VnfH from Azotobacter vinelandii, the Fe protein of the alternative, vanadium-dependent nitrogenase system, in its ADP-bound state. VnfH adopts the same conformation that was observed for NifH, the Fe protein of molybdenum nitrogenase, in complex with ADP, representing a state of the functional cycle that is ready for reduction and subsequent nucleotide exchange. The overall similarity of NifH and VnfH confirms the experimentally determined cross-reactivity of both ATP-hydrolyzing reductases.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The chemistry of biological nitrogen fixation is challenging in that it involves the reduction of the stable triple bond of the N2 molecule, requiring it to overcome a bond enthalpy of − 946 kJ mol−1 [1]. This reaction is equivalent to the industrial Haber–Bosch process that since its development in the early twentieth century has profoundly transformed the face of the planet and has enabled an unprecedented growth of the human population by effectively removing the bioavailability of the element nitrogen as a limiting factor for crop production [2, 3]. While the industrial process is promoted by high temperature and pressure on an iron catalyst that reacts atmospheric N2 with hydrogen gas, the biological reaction operates at ambient conditions, driven solely by the hydrolysis of adenosine triphosphate (ATP) [1, 4]. The only known enzyme able to mediate this reaction is the metalloprotein nitrogenase, a dynamic complex of two components, commonly referred to as dinitrogenase and nitrogenase reductase [5]. Nitrogenase reductase—also known as the Fe protein—is a homodimer of approximately 60 kDa and a member of the family of P-loop NTPases, whose protomers are connected exclusively via a cubane-type [4Fe:4S] cluster [6]. In this system, the task of the reductase is to provide a single electron to the catalytic dinitrogenase component, concomitant with spending two ATP in an intricate mechanism that effectively leads to the transfer of a low-potential electron to the active site and according to current knowledge proceeds as follows: upon reduction by a ferredoxin or flavodoxin, Fe protein exchanges a bound ADP molecule per monomer for ATP [7]. This leads to a conformational change that allows Fe protein to dock onto the catalytic dinitrogenase component, in a position that brings its metal cluster in close proximity to the first of two metal clusters in the dinitrogenase component [8]. This is P-cluster, a unique [8Fe:7S] electron relay site that is already fully reduced, i.e., in an all-ferrous state in the resting state of the enzyme [9, 10]. Consequently, Fe protein cannot merely transfer an electron from its [4Fe:4S] center to P-cluster, but instead is thought to employ what has been very aptly described as a deficit-spending mechanism [11]. In this model, it is the docking of Fe protein and the dinitrogenase that first triggers the release of a low-potential electron from P-cluster in the latter, followed by ATP hydrolysis in the former. Only in the following step this electron is then replenished from the [4Fe:4S] cluster in Fe protein, and the oxidized, ADP-bound Fe protein then dissociates from dinitrogenase to be replenished for the next one-electron transfer. The purpose of this mechanism is in lowering the redox potential of the electron-leaving P-cluster, and we may speculate that this is achieved by a slight conformational rearrangement of the immediate surrounding of this site, caused by the interaction of both components, or the ATP-induced power stroke of Fe protein. This intricate sequence of events converts the chemical energy of an ATP phosphodiester bond via the mechanical action of a conformational change into the lowered electrochemical potential of the electron that eventually reaches the active site of dinitrogenase. In essence, this is equivalent to the elevated temperature and pressure of the Haber–Bosch process, but the entire, orchestrated interaction of the two protein components must be repeated eight times for the generation of two molecules of ammonium from a single N2 [12].
All known nitrogenases form a monophyletic group [13] and use Fe protein for electron transfer. For the catalytic dinitrogenase, three different classes are known that differ primarily in their active site cofactor, the second metal site beside P-cluster. In the best-studied variant, molybdenum nitrogenase, this cofactor is FeMoco, a [Mo:7Fe:9S:C]:homocitrate cluster [14,15,16], while the alternative vanadium nitrogenase employs FeVco, which is of composition [V:7Fe:8S:C]:CO3:homocitrate [17]. It is less active in N2 reduction, but instead reduces CO, an inhibitor of FeMoco, to hydrocarbons [18], and a ligand-bound structure most recently reported for this enzyme has provided essential hints for an atomic-level understanding of its mechanism [19]. The third type of nitrogenases solely relies on iron, with a FeFeco whose precise architecture remains to be elucidated. Its N2-reducing activity again is lower than the one of FeVco, but it also has a special power in that it reduces CO2 that remains untouched by the other two nitrogenase types [20]. Some diazotrophs, including the paradigmatic soil bacterium Azotobacter vinelandii, encode all three types of nitrogenase in their genome [21], preferably expressing the most potent, Mo-dependent system. They switch to producing vanadium nitrogenase under Mo-depleted growth conditions and will eventually resort to the iron-only enzyme if both heterometals become scarce. Although we assume that all dinitrogenase components feature highly similar P-clusters, the distinct gene clusters for Mo (nif), V (vnf) and Fe (anf) nitrogenase each contain their dedicated Fe protein ortholog, encoded by the nifH, vnfH and anfH genes, respectively [22]. To date, high-resolution crystal structures are available for molybdenum-dependent dinitrogenases (MoFe proteins) of A. vinelandii, Klebsiella pneumoniae, Clostridium pasteurianum and Gluconacetobacter diazotrophicus [23], as well as for the analogous VFe protein of A. vinelandii [17], but the only Fe protein studied structurally is NifH of A. vinelandii [6], although the isolation of VnfH from a ΔnifHDK was already reported in 1986 [24]. This may be due to the high sensitivity of the protein towards dioxygen, leading to a half-decay time of less than 10 s in the presence of 2 vol% of O2 [25]. For the Fe proteins of molybdenum and vanadium nitrogenases, a cross-reactivity has been reported, and the polypeptide chains of the two A. vinelandii proteins exhibit a sequence identity of 90.7%. Here we report on the isolation, crystallization and structural analysis of A. vinelandii VnfH in the ADP-bound state.
Experimental procedures
Cultivation of A. vinelandii under Mo-depleted conditions
Azotobacter vinelandii strain Lipman 1903 [26] was grown in nitrogen-free, modified Burk medium at pH 7.5 [27]. The medium contained 58 mM of sucrose, 0.9 mM CaCl2, 1.67 mM MgSO4, 0.04 mM FeSO4, 0.2 mM of citric acid, 2.5 mM KH2PO4, and 10 mM K2HPO4. Initial cultures were supplemented with 10 mM NH4Cl and 0.01 mM Na3MoO4 and grown at 30 °C and 180 rpm agitation under aerobic conditions in baffled Erlenmeyer flasks. When the culture reached OD600nm = 3.0, 1 mL of medium was plated on Mo-containing Burk agar plates (Burk medium supplemented with 5% (w/v) of Agar–Agar) and incubated at 30 °C over night. In the resulting colonies, active nitrogen fixation led to a visible, brown color of the cells [28]. Single colonies were then used to inoculate Burk medium supplemented with 10 mM NH4Cl and 0.01 mM Na3VO4 and grown at 30 °C and 180 rpm agitation until an OD600nm of 3.0 was reached and the cells were then singled out on Burk agar plates (5% (w/v) agar–agar) without vanadium and nitrogen and incubated at 30 °C to obtain single colonies. This cycle of growth and singling out was repeated for five times and the resulting cells did no longer produce any detectable Mo nitrogenase and were used at larger scale (10 L) for producing VnfH.
Isolation of VnfH
A buffer containing 50 mM Tris/HCl at pH 7.4, 100 mM NaCl and 10 mM Na2S2O4 was used to thoroughly resuspend the cell pellet after harvesting by centrifugation. Cells were disrupted in a homogenizer (Emulsiflex-C5, Avestin) at 15,000 psi under an N2 atmosphere. The lysate was centrifuged at 100,000×g for 30 min and the supernatant was subsequently loaded onto a HiTrap Q HP anion exchange column (GE Healthcare) equilibrated with 50 mM Tris/HCl at pH 7.4, 100 mM NaCl and 2.5 mM Na2S2O4. To separate the vanadium nitrogenase components, the column was developed with a linear NaCl gradient, with VnfH eluting at 360 mM NaCl. After concentration (Vivaspin 20, 30 kDa MWCO, Sartorius) under 4 bar N2 pressure, VnfH was loaded onto a size exclusion column (Superdex 200 pg, 26/600, GE Healthcare) equilibrated with 20 mM Tris/HCl buffer at pH 7.4, 100 mM NaCl and 2.5 mM Na2S2O4. Fractions containing pure VnfH were identified by SDS-PAGE, pooled, concentrated and stored in liquid nitrogen until further use.
Crystallization and data collection
Azotobacter vinelandii VnfH was crystallized by sitting drop vapor diffusion, in an anoxic chamber containing 95% N2 and 5% H2. A protein solution containing 4–7 mg mL−1 VnfH, 10 mM MgCl2 and 2.5 mM ADP was used for screening with reservoir solutions containing polyethylene glycol of various molecular weights in buffers ranging from pH 4.5 to 9.5. The final condition contained 100 mM Tris/HCl buffer at pH 8.5 and 25% (w/v) PEG 3350 in the reservoir solution. 1 µL of protein solution was added to 1 µL of reservoir solution on a 96-well crystallization plate (MRC 2, SwissSci) and subsequently sealed. After 6 weeks, crystals were mounted in a nylon loop after addition of 10% (v/v) 2R,3R-butane diol as a cryoprotectant and flash frozen in liquid N2. Diffraction data were collected at the Swiss Light Source (Paul Scherrer Institute, Villigen, CH) on beamline X06SA, equipped with an Eiger 16 M detector (Dectris) at a wavelength of 1 Å. An oscillation angle of 0.1° per diffraction image was used, covering a total range of 360° of crystal rotation per dataset. The X-ray beam was readjusted to a different spot on the crystal for each dataset.
Structure solution and refinement
Data were indexed and integrated with XDS [29] and subsequently merged and scaled with AIMLESS from the CCP4 suite [30]. Two datasets from the same crystal were merged during data processing. The structure was solved by molecular replacement using Phaser-MR [31]. ADP-bound NifH served as a model structure (PDB-entry: 1FP6), yielding a clear solution with four VnfH dimers in the asymmetric unit of the C 2221 unit cell. The structure of VnfH was then built in COOT [32] and refined in iterative cycles with further model improvement using Buster/TNT 2.10 [33]. Data collection and refinement statistics are summarized in Table 1. All figures were generated with PyMOL [34], electrostatic potential maps were calculated with APBS [35].
Results
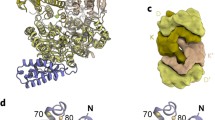
Azotobacter vinelandii VnfH crystallized in space group C 2221 with four homodimers in the asymmetric unit that are highly similar, with a positional root-mean-squared deviation of 0.18 ± 0.09 Å for all atoms. As expected based on its high sequence homology to the NifH protein from the same organism, the VnfH dimer is linked by a canonical [4Fe:4S] cluster at the dimer interface, coordinated by two cysteine residues, C98 and C133 from each monomer. The 290 amino acid VnfH attains the Rossman-type βαβ-fold with a parallel, 7-stranded β-sheet that is typical for P-loop NTPases (Fig. 1). In each monomer, a bound ADP molecule was clearly defined in the electron density map, residing in a specific binding pocket on the surface of each monomer (Fig. 1b). In the VnfH dimer, these binding pockets are buried deep within a central cleft on the surface of the protein that is opposite from the bound [4Fe:4S] cluster (Fig. 1c). Nucleotide binding and hydrolysis occurs at a conserved structural feature, the P-loop (or Walker A) motif GX4GKS located at residues 11–17 in loop 1 that connects strand β1 with helix α1 of the protein, employing the helix dipole and residue K16 to create a positively charged pocket for binding the pyrophosphate moiety of the ADP ligand (Fig. 2). Residue S17 is a key part of the Walker A motif, as it coordinates a Mg2+ cation, whose octahedral ligand environment is completed by an oxygen atom of the β-phosphate of ADP, as well as four well-defined water molecules. S71 and one of these water ligands also form short hydrogen bonds (2.6 and 2.7 Å, respectively) to residue D126, the signature residue of the switch II (or Walker B) motif that changes its conformation according to whether a nucleoside di- or triphosphate is bound to the P-loop (Fig. 2). D126 is placed at the end of strand β4, from where the protein chain continues past D130, the second key residue in the switch II region, to C133, one of the two ligands to the [4S:4S] cluster. A second part of the protein that undergoes ligand-dependent conformational changes is the switch I region, which in VnfH encompasses residues 39–45, immediately following sheet β2. Adjacent to the nucleotide binding site, four residues in this loop region are in contact with the water ligands of the Mg2+ cation and can thus react to its different positioning in the ATP- and ADP-bound states (Fig. 2).
Three-dimensional structure of A. vinelandii VnfH. a Front view of the VnfH dimer in the ADP-bound state. The two monomers are exclusively connected through the coordination to the [4Fe:4S] cluster located on the twofold symmetry axis. b The bound nucleoside diphosphate was well defined in its binding pocket on a VnfH monomer. The electron density map shown is a 2Fo − Fc omit map contoured at the 1.5 σ level. c Surface representation of a bottom view of the VnfH dimer. The deep cleft between the monomers provides access for MgADP to dissociate from the enzyme when exchanged for ATP. d VnfH monomer in cartoon representation, colored from blue at the N terminus of the 291 aa peptide chain to red at the C terminus. The [4Fe:4S] cluster, as well as the bound ADP molecule and the Mg2+ ion are shown as spheres or sticks, respectively
Nucleotide binding site of VnfH. The stereo figure shows MgADP binding to a monomer of A. vinelandii VnfH. As typical for NTPases of this family, the phosphoanhydride chain of the ligand is bound to a P-loop region (11–17), centered on a 15GKS17 motif, in which the lysine interacts with the β-phosphate of ADP, while the serine coordinates the Mg2+ ion. Typical for P-loop NTPases, the binding of ATP vs. ADP leads to conformational changes in the switch I and switch II regions, with immediate consequences for the positioning of the [4Fe:4S] cluster bound at the end of the switch II motif (C133). This change brings the two halves of the dimer close together, allowing for K11 and D130 to interact with the nucleotide bound in the other monomer of the VnfH dimer, thereby promoting ATP hydrolysis
In nitrogenase Fe proteins, the conformational switch of the conserved P-loop NTPase scaffold is thus recruited to mediate a movement of the immediate vicinity of the metal cluster, with obvious functional consequences for electron transfer to the dinitrogenase component. The ADP-bound state in the present structure only represents a single structural snapshot in the dynamic interaction of Fe protein with the catalytic component of dinitrogenase, and currently a complex structure of VnfH with VnfDKG is not available. However, the high similarity between VnfH and NifH allows to conclude that the most prominent functional features are retained. VnfH and the ADP-bound form of NifH (PDB 1FP6) align with a root-mean-squared deviation of 0.3 Å for all atoms (Fig. 3a, b), and only one of the 27 amino acid exchanges separating the proteins, H174 that in NifH corresponds to N173, is located within the surface region of the Fe protein that interacts with the catalytic component of nitrogenase (Fig. 3c). In the ADP·AlF4−-stabilized complex of NifH and NifDK (PDB 1N2C) [36], N173 relocates slightly and does not interact directly with NifDK, and the same is likely the case for VnfH interacting with VnfDGK. Furthermore, the conformation of NifH with bound ADP·AlF4−, considered to mimic the transition state of ATP hydrolysis, corresponds well to the one obtained with AMPPCP, a non-hydrolysable analog of ATP (PDB 2AFK) [37]. In both cases, the different positioning of the switch I and II regions causes the two protomers of the NifH dimer to approach each other more closely. This is supported by the binding to NifDK, which is a prerequisite for ATP hydrolysis, so that in this case the enzyme acts as the activator, in analogy to what is found in many G protein-based signaling systems. Two amino acid residues in NifH are important for this activation, and both are conserved in VnfH: K11 forms part of the Walker A motif, and D130 is the second signature residue of the Walker B motif (Fig. 2). In the ADP-bound state of VnfH reported here, both residues are oriented towards the gap in the dimer interface that is closed in the NifHDK complex structures (Fig. 1c), and this rearrangement then brings both into close proximity of the MgATP bound to the other monomer of the dimer. The closing of the dimer thus is required for efficient ATP hydrolysis, and the conservation of K11 and D130 in VnfH strongly indicates that an analogous mechanism will be in place when binding to VnfDKG.
Structural comparison of VnfH with NifH. a The homologous Fe proteins of molybdenum (black) and vanadium (red) nitrogenase in their respective, ADP-bound states in ribbon representation. The [4Fe:4S] clusters and Mg2+ ions are shown as spheres, the ADP ligands in stick representation. The near identical quaternary structures of the two proteins emphasize that the ADP-bound state, in which nucleotides are exchanged upon reduction, are structurally well defined rather than intrinsically flexible. b Top view of the dimer, represented as in a, highlighting the location of the metal clusters on the twofold dimer axis. c Stereo representation of the VnfH monomer (cartoon representation) and NifH (black). Residues differing in VnfH are shown as red sticks. While located mainly on the protein surface, they are distributed across the entire protein, omitting the interaction surface with the dinitrogenase
For molybdenum nitrogenase, the interaction of the two component proteins was suggested to be initially guided primarily through electrostatic interactions that help to orient and pre-align both NifH and NifDK, followed by a probing of the surface of the latter by the smaller Fe protein [38]. In the productive complex of the two components, i.e., the state where ATP hydrolysis and electron transfer take place, the twofold symmetry axis of the Fe protein dimer aligns fully with the pseudo-twofold axis relating the D- and K-subunits of the reductase. In the Fe protein, the [4Fe:4S] cluster bridges the monomers and is therefore of course located directly on the symmetry axis, and in the catalytic subunit the same is true for P-cluster [36, 37]. Their juxtaposition in the complex leads to the shortest possible distance, which in turn benefits electron transfer efficiency. Given the overall similarity of the vanadium nitrogenase system, an analogous mode of interaction of both proteins seems to be a reasonable assumption. The electrostatic surface properties of VnfH and VnfDKG provide strong support for this hypothesis. While VnfH shows an overall negative electrostatic surface potential, a distinct, positive patch is located in the immediate surroundings of the iron–sulfur cluster (Fig. 4a). Recently, the three-dimensional structure of VnfDKG has revealed that the D–K interface of this component retains the characteristic ridge that serves as a docking point for Fe protein in NifDK [17]. The electrostatic surface potential of VnfDKG around this presumed docking site is highly complementary to the charge distribution on VnfH, with an overall negative potential on the docking ridge, surrounded by patches of positive surface potential (Fig. 4b). As elaborated elsewhere, VnfH docking on this ridge would not interfere with the position of VnfG, the additional subunit of vanadium nitrogenase that is not present in the molybdenum-based enzyme [17].
Electrostatic surface properties in vanadium nitrogenase. a Top view of the VnfH dimer along the twofold symmetry axis. The protein surface is colored according to its electrostatic surface potential ranging from red at − 5 kBT to blue at + 5 kBT. The icon in the lower right summarized the charge distribution. b VnfDKG, as the interaction partner of VnfH, is presumed to bind VnfH on the pseudo-twofold symmetry axis relating the D- and K-subunits. Here, the electrostatic surface potential of the enzyme (not drawn to scale with a) is ideally complementary
Discussion
Azotobacter vinelandii is a Gram-negative, free-living soil bacterium that is highly adapted to a diazotrophic lifestyle. It combines an obligate aerobic metabolism that allows for the most efficient generation of ATP from carbohydrate oxidation with a very high oxidase activity to retain its cytoplasm in a fully anoxic state. Due to the straightforward handling and the availability of a genetic system, A. vinelandii has emerged as a paradigm for studying biological nitrogen fixation. In addition to the most thoroughly characterized molybdenum nitrogenase, the organism contains two alternative enzyme systems dependent either on vanadium instead of molybdenum, or relying exclusively on iron. As clarified by the genome sequence of the organism, the three nitrogenases are organized into distinct gene clusters that provide the structural genes for the catalytic components together with a range of accessory genes required for metal uptake, cofactor biogenesis and electron transfer to sustain catalysis [21]. With characteristic differences in their active site cofactors, the evolution of separate proteins to hold these cofactors is unsurprising. On the other hand, a crucial assembly factor for nitrogenase cofactors is the radical/SAM enzyme NifB that synthesizes a [8Fe:9S:C] precursor from simple building blocks. This precursor, frequently termed the L-cluster, is common to all three nitrogenases, and to—to the best of our knowledge—they all use this enzyme, which is present with only a single copy in the genome of A. vinelandii. Nature thus does recruit compatible components for all three enzymes. For the iron protein components, however, this is not the case. The NifH protein from molybdenum nitrogenase finds its counterpart in VnfH for vanadium nitrogenase and AnfH for the iron-only enzyme. As the present structural analysis of VnfH shows—not entirely unexpectedly—the structures of both proteins are highly similar, to a far higher extent than it is the case for MoFe and VFe protein in this organism. In agreement with earlier reports, we find that both NifH and VnfH are cross-reactive and work equally well with either dinitrogenase within experimental uncertainty. The significance of having individual copies of Fe protein for each dinitrogenase variant thus remains to be elucidated. Most obviously, a coexpression with the structural genes for the enzymes might serve to ensure stoichiometric production of the components. If so, a balanced stoichiometry of these components would lead to only one Fe protein dimer produced per dinitrogenase complex (NifD2K2 or VnfD2G2K2), which is lower than commonly assumed, but might support the recent observation of half-sites reactivity in this enzyme complex [39]. It may also be connected with the additional role identified for NifH, in which it interacts with the assembly factor NifEN to promote the insertion of the molybdenum ion and the homocitrate ligand into the precursor L-cluster [40,41,42]. An analogous step exists in the biogenesis of VFe protein, occurring on VnfEN that is encoded in the vnf gene cluster. FeV cofactor, however, not only differs from its Mo-containing counterpart in the nature of the heterometal, but also in the presence of a second organic ligand, carbonate [17]. With L-cluster being the common precursor of both cofactors, this additional modification involves the removal of a µ-bridging sulfide, S5A, and its replacement by CO32−. This step is likely to occur on or near VnfEN, so that in spite of a significant sequence homology of the two EN complexes, the functional differences might be substantial, justifying a dedicated Fe protein component.
The present structure of VnfH confirms the high degree of structural conservation among nitrogenase Fe proteins and highlights that the ADP-bound form of this ATP-dependent reductase is a well-defined state, poised for the exchange of nucleotide upon subsequent reduction. Further studies to address other nucleotide-bound states, as well as a characterization of the complex of VnfH with VnfDKG, will show whether the other stages of the electron transfer choreography between nitrogenases and their corresponding reductases are as highly conserved.
Abbreviations
- EPR:
-
Electron paramagnetic resonance
- MR:
-
Molecular replacement
References
Rees DC (1993) Curr Opin Struct Biol 3:921–928
Leigh GJ (2004) The world’s greatest fix: A history of nitrogen in agriculture. Oxford University Press, Oxford
Smil V (2002) Ambio 31:126–131
Rees DC, Tezcan FA, Haynes CA, Walton MY, Andrade S, Einsle O, Howard JB (2005) Philos Trans R Soc Lond A 363:971–984
Howard JB, Rees DC (1996) Chem Rev 96:2965–2982
Georgiadis MM, Komiya H, Chakrabarti P, Woo D, Kornuc JJ, Rees DC (1992) Science 257:1653–1659
Burgess BK, Lowe DJ (1996) Chem Rev 96:2983–3011
Grossmann JG, Hasnain SS, Yousafzai FK, Smith BE, Eady RR, Schindelin H, Kisker C, Howard JB, Tsuruta H, Muller J, Rees DC (1999) Acta Crystallogr D 55:727–728
Peters JW, Fisher K, Newton WE, Dean DR (1995) J Biol Chem 270:27007–27013
Angove HC, Yoo SJ, Münck E, Burgess BK (1999) J Inorg Biochem 74:65
Danyal K, Dean DR, Hoffman BM, Seefeldt LC (2011) Biochemistry 50:9255–9263
Hoffman BM, Lukoyanov D, Yang ZY, Dean DR, Seefeldt LC (2014) Chem Rev 114:4041–4062
Howard JB, Kechris KJ, Rees DC, Glazer AN (2013) Multiple amino acid sequence alignment nitrogenase component 1: insights into phylogenetics and structure-function relationships. Plos One 8:e72751
Einsle O (2014) J Biol Inorg Chem 19:737–745
Spatzal T, Aksoyoğlu M, Zhang LM, Andrade SLA, Schleicher E, Weber S, Rees DC, Einsle O (2011) Science 334:940
Lancaster KM, Roemelt M, Ettenhuber P, Hu YL, Ribbe MW, Neese F, Bergmann U, DeBeer S (2011) Science 334:974–977
Sippel D, Einsle O (2017) Nat Chem Biol 13:956–960
Lee CC, Hu YL, Ribbe MW (2010) Science 329:642
Sippel D, Rohde M, Netzer J, Trncik C, Gies J, Grunau K, Djurdjevic I, Decamps L, Andrade SLA, Einsle O (2018) Science 359:1484–1489
Zheng Y, Harris DF, Yu Z, Fu Y, Poudel S, Ledbetter RN, Fixen KR, Yang ZY, Boyd ES, Lidstrom ME, Seefedt LC, Harwood CS (2018) Nat Microbiol 3:281–286
Setubal JC, dos Santos P, Goldman BS, Ertesvag H, Espin G, Rubio LM, Valla S, Almeida NF, Balasubramanian D, Cromes L, Curatti L, Du ZJ, Godsy E, Goodner B, Hellner-Burris K, Hernandez JA, Houmiel K, Imperial J, Kennedy C, Larson TJ, Latreille P, Ligon LS, Lu J, Maerk M, Miller NM, Norton S, O’Carroll IP, Paulsen I, Raulfs EC, Roemer R, Rosser J, Segura D, Slater S, Stricklin SL, Studholme DJ, Sun J, Viana CJ, Wallin E, Wang BM, Wheeler C, Zhu HJ, Dean DR, Dixon R, Wood D (2009) J Bacteriol 191:4534–4545
Dos Santos PC, Fang Z, Mason SW, Setubal JC, Dixon R (2012) Distribution of nitrogen fixation and nitrogenase-like sequences amongst microbial genomes. BMC Genom 13:162–174
Owens CP, Katz FEH, Carter CH, Oswald VF, Tezcan FA (2016) J Am Chem Soc 138:10124–10127
Hales BJ, Langosch DJ, Case EE (1986) J Biol Chem 261:5301–5306
Schlesier J, Rohde M, Gerhardt S, Einsle O (2016) J Am Chem Soc 138:239–247
Lipman JG (1903) Rep N J Agric Exp Stn 24:217–285
Burk D, Lineweaver H (1930) J Bacteriol 19:389–414
Sippel D, Schlesier J, Rohde M, Trncik C, Decamps L, Djurdjevic I, Spatzal T, Andrade SLA, Einsle O (2017) J Biol Inorg Chem 22:161–168
Kabsch W (2010) Acta Crystallogr D 66:125–132
Winn MD, Ballard CC, Cowtan KD, Dodson EJ, Emsley P, Evans PR, Keegan RM, Krissinel EB, Leslie AGW, McCoy A, McNicholas SJ, Murshudov GN, Pannu NS, Potterton EA, Powell HR, Read RJ, Vagin A, Wilson KS (2011) Acta Crystallogr D 67:235–242
McCoy AJ, Grosse-Kunstleve RW, Adams PD, Winn MD, Storoni LC, Read RJ (2007) J Appl Crystallogr 40:658–674
Emsley P, Lohkamp B, Scott WG, Cowtan K (2010) Acta Crystallogr D 66:486–501
Blanc E, Roversi P, Vonrhein C, Flensburg C, Lea SM, Bricogne G (2004) Acta Crystallogr D 60:2210–2221
Schrödinger LLC (2010) The PyMOL molecular graphics system
Jurrus E, Engel D, Star K, Monson K, Brandi J, Felberg LE, Brookes DH, Wilson L, Chen JH, Liles K, Chun MJ, Li P, Gohara DW, Dolinsky T, Konecny R, Koes DR, Nielsen JE, Head-Gordon T, Geng WH, Krasny R, Wei GW, Holst MJ, McCammon JA, Baker NA (2018) Protein Sci 27:112–128
Schindelin H, Kisker C, Sehlessman JL, Howard JB, Rees DC (1997) Nature 387:370–376
Tezcan FA, Kaiser JT, Mustafi D, Walton MY, Howard JB, Rees DC (2005) Science 309:1377–1380
Owens CP, Katz FE, Carter CH, Luca MA, Tezcan FA (2015) J Am Chem Soc 137:12704–12712
Danyal K, Shaw S, Page TR, Duval S, Horitani M, Marts AR, Lukoyanov D, Dean DR, Raugei S, Hoffman BM, Seefeldt LC, Antony E (2016) Proc Natl Acad Sci USA 113:E5783–E5791
Kaiser JT, Hu YL, Wiig JA, Rees DC, Ribbe MW (2011) Science 331:91–94
Hu Y, Ribbe MW (2011) Coord Chem Rev 255:1218–1224
Hu YL, Fay AW, Lee CC, Wiig JA, Ribbe MW (2010) Dalton T 39:2964–2971
Weiss M, Hilgenfeld R (1997) J Appl Crystallogr 30:203–205
Karplus PA, Diederichs K (2012) Science 336:1030–1033
Cruickshank DWJ (1999) Acta Crystallogr D 55:583–601
Laskowski RA, MacArthur MW, Moss DS, Thornton JM (1993) J Appl Crystallogr 26:283–291
Acknowledgements
The authors thank the staff at beam line X06DA of the Swiss Light Source, Paul Scherrer Institute, Villigen, CH, for their excellent assistance with diffraction data collection. This work was supported by the Deutsche Forschungsgemeinschaft (RTG 1976 and PP 1927) and the European Research Council (Grant no. 310656).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Rohde, M., Trncik, C., Sippel, D. et al. Crystal structure of VnfH, the iron protein component of vanadium nitrogenase. J Biol Inorg Chem 23, 1049–1056 (2018). https://doi.org/10.1007/s00775-018-1602-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00775-018-1602-4