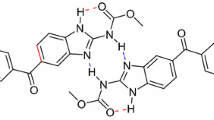
Abstract
Organometallic half-sandwich IrIII complexes of the type [(η5-Cpx)Ir(N^N)Cl]PF6 1–6, where Cpx = C5Me5 (Cp*), C5Me4C6H5 (Cpxph), C5Me4C6H4C6H5 (Cpxbiph), N^N is imionopyridine chelating ligand, were prepared and characterized. The X-ray crystal structure of complex 1 has been determined. Four compounds displayed higher anticancer potency than clinically used anticancer drug cisplatin against A549 cancer cells, especially complex 3 which is 8 times more active than cisplatin. No hydrolysis was observed by NMR and UV–Vis for complexes 3 and 6; however, these complexes show big differences in nucleobase binding, mainly decided by the imionopyridine chelating ligand. Complex 3 is stable in the presence of glutathione, but 6 reacted rapidly with glutathione. The octanol/water partition coefficients (log P) of 3 and 6 have been determined. In addition, these complexes display effective catalytic activity in converting coenzyme NADH to NAD+ by accepting hydride to form an Ir hydride adduct. The mechanism of actions of these complexes involves apoptosis induction, cell cycles arrest, and significant increase of reactive oxygen species levels in A549 cancer cells.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Since the successful discovery of cisplatin in the treatment of types of cancers, platinum-based anticancer drugs have attracted great attention in the chemotherapeutic studies [1, 2]. Simultaneously, however, serious side effects have been limiting the further uses of the classic platinum-based drugs. To overcome the drawbacks, various non-platinum transition metals, such as ruthenium, osmium, gold, palladium, and rhodium-based drugs have been investigated as potential anticancer therapeutics [3,4,5,6,7,8,9,10,11,12,13,14,15,16,17,18,19,20]. Very recently, possible biological applications of iridium complexes have received much interests [21,22,23,24,25,26,27,28,29,30], in which organoiridium anticancer agents have shown very promising antiproliferative activity toward cancer cells [31,32,33,34]. In addition, iridium complexes were found displaying novel anticancer mechanism of actions (MoAs) different from platinum drugs, and showing high antiproliferative activity against cisplatin-resistant cancer cells [35, 36], thus could be a potential candidates for treatment of platinum resistant cancers.
One of the most important reasons people select metal complexes as potential anticancer drugs is their structural diversity. A wide variety of metals and ligands with steric and electronic features can be selected to make a huge amount of combinations and incorporations. Moreover, any subtle changes in the composition or in the structure of complex may cause totally different effects on chemical and biological activities. For example, antiproliferative activity of organoiridium complexes against cancer cells can be switched on readily by replacing one nitrogen atom in 2,2′-bipyridine with one carbon atom to form ligand phenyl pyridine [37], where the new complex displayed a much stronger binding to nucleobases and higher hydrophobicity.
In the potential ligand candidates, N^N-donor chelating ligands present one of the main subjects which have been widely used in chemical and biological evaluation [38,39,40]. Iminopyridine type of ligands has been attracting much attention in catalysis [41] and may play an important role in redox procedure of transition metal complexes [42, 43]. Interestingly, anticancer agents and catalysts are sometimes inseparable [44]. Some ruthenium transfer hydrogenation catalysts with poor catalytic activity showed high anticancer activity [45]. In addition, investigation on the catalytic behavior of iridium complexes helped to understand their MoAs as anticancer agents [46]. Very recently, iminopyridine type of ligands started to be used in the design of metal-based anticancer agents. Sadler group and our group have reported a panel of osmium(II) [47] and iridium(III) [48] phenyliminopyridine complexes, respectively, which have shown very promising anticancer activities against various cancer cell lines.
On the basis of our previous work, herein we newly explored adamantyl iminopyridine and oxime pyridyl as novel N^N-chelating iminopyridine ligands, and synthesized eight iridium half-sandwich complexes containing Cp* and its phenyl substituents. Their hydrolysis chemistry, nucleobase binding, catalytic transfer hydride reactions, in vitro cancer cell toxicity, and basic mechanism of action were studied.
Experimental section
Materials
IrCl3·nH2O (99%), 1, 2, 3, 4, 5-pentamethyl-cyclopentadiene (95%), 2, 3, 4, 5-tetramethyl-2-cyclopentanone (95%), 4-bromobiphenyl (98%), butyllithium solution (1.6 M in hexane), ammonium hexafluorophosphate (NH4PF6), 9-ethylguanine, and 9-methyladenine were purchased from Sigma-Aldrich. Solvents of laboratory grade were used for syntheses without further purification. The starting Ir(III) dimers [(η5-Cp*)IrCl2]2, [(η5-Cpxph)IrCl2]2 and [(η5-Cpxbiph)IrCl2]2, and ligand L1 were prepared according to reported methods [49]. Ligand L2 (methyl 2-pyridyl ketoxime) were purchased from Alfa Aesar. For the biological experiments, phosphate-buffered saline (PBS), fetal bovine serum, DMEM medium, trypsin/EDTA, and penicillin/streptomycin mixture were purchased from Sangon Biotech. Testing compounds were dissolved in DMSO and diluted with the tissue culture medium before use. All reactions were performed under an N2 atmosphere.
Synthesis of the [(η 5-Cpx)Ir(N^N)Cl]PF6
General method
A solution of chloride bridged dimer [(η5-Cpx)IrCl2]2 (0.05 mmol) and iminopyridine ligand (0.12 mmol) in MeOH (20 ml) was heated under reflux in an N2 atmosphere for 10 h and filtered. The volume was slowly reduced to half on a rotary evaporator and NH4PF6 (0.42 mmol) was added. After standing at 298 K, a microcrystalline product formed. This was collected by filtration, washed with diethyl ether, and recrystallized from CH2Cl2/diethyl ether. The 1H NMR (500.13 MHz, CDCl3 or DMSO-d6) peak integrals of complexes 1–6 are shown in Figs. S7–S12. The high-resolution mass spectrometry data of complexes 1–6 are shown in Fig. S13.
[(η 5-Cp*)Ir(L1)Cl]PF6 (1)
Yield: 46.4 mg, 62.0%. 1H NMR (500.13 MHz, CDCl3) δ 8.98 (s, 1H), 8.72 (d, J = 5.4 Hz, 1H), 8.23 (d, J = 7.6 Hz, 1H), 8.08 (t, J = 7.2 Hz, 1H), 7.69 (m, 1H), 2.30 (s, 3H), 2.20 (d, 3H), 2.05 (d, 3H), 1.77 (s, 6H), 1.69 (s, 15H). Anal. calcd for [(η5-Cp*)Ir(L1)Cl]PF6 (748.18): C, 41.74; H, 4.72; N, 3.74. Found: C, 41.69; H, 4.76; N, 3.77. MS: m/z 603.42 [(η5-C5Me5)Ir(L1)Cl]+. Crystals suitable for X-ray diffraction were obtained by slow evaporation of a CH2Cl2/diethyl ether solution at ambient temperature.
[(η 5-Cpxph)Ir(L1)Cl]PF6 (2)
Yield: 34.5 mg, 42.6%. 1H NMR (500.13 MHz, CDCl3) δ 9.00 (s, 1H), 8.24 (dd, J = 14.5, 6.8 Hz, 2H), 8.01 (t, J = 7.7 Hz, 1H), 7.51 (m, J = 6.1 Hz, 5H), 7.36 (m, 1H), 2.26 (s, 3H), 2.20 (d, J = 11.9 Hz, 4H), 2.02 (m, 5H), 1.90 (d, J = 4.2 Hz, 6H), 1.74 (d, J = 3.0 Hz, 6H), 1.59 (s, 3H). Anal. calcd for [(η5-Cpxph)Ir(L1)Cl]PF6 (810.19): C, 45.95; H, 4.60; N, 3.46. Found: C, 45.87; H, 4.65; N, 3.51. MS: m/z 655.50 [(η5-C5Me4C6H5)Ir(L1)Cl]+.
[(η 5-Cpxbiph)Ir(L1)Cl]PF6 (3)
Yield: 62.4 mg, 70.4%. 1H NMR (500.13 MHz, CDCl3) δ 9.02 (s, 1H), 8.28 (s, 2H), 8.01 (s, 1H), 7.72 (d, 1H), 7.65 (d, 1H), 7.61 (d, 1H), 7.42–7.49 (m, 5H), 7.27 (d, 2H), 2.21 (t, 4H), 2.05 (d, 3H), 1.92 (d, J = 4.8 Hz, 3H), 1.77 (d, J = 13.8 Hz, 4H), 1.62 (s, 2H), 1.50 (s, 11H). Anal. calcd for [(η5-Cpxph)Ir(L1)Cl]PF6 (886.22): C, 50.14; H, 4.66; N, 3.16. Found: C, 50.22; H, 4.58; N, 3.24. MS: m/z 741.50 [(η5-C5Me4C6H4 C6H5)Ir(L1)Cl]+.
[(η 5-Cp*)Ir(L2)Cl]PF6 (4)
Yield: 18.2 mg, 28.3%. 1H NMR (500.13 MHz, CD3CN) δ 8.65 (d, J = 5.6 Hz, 1H), 8.02 (t, J = 7.9 Hz, 1H), 7.81 (d, J = 8.1 Hz, 1H), 7.53 (dd, J = 7.3, 5.9 Hz, 1H), 3.27 (s, 1H), 2.45 (d, J = 1.3 Hz, 3H), 1.68 (s, 15H). Anal. calcd for [(η5-Cp*)Ir(L2)Cl]PF6 (644.08): C, 31.70; H, 3.60; N, 4.35. Found: C, 31.65; H, 3.67; N, 4.42. MS: m/z 463.33 [(η5-C5Me5)Ir(L2)]2+.
[(η 5-Cpxph)Ir(L2)Cl]PF6 (5)
Yield: 19.6 mg, 27.8%. 1H NMR (500.13 MHz, CDCl3) δ 8.30 (s, 1H), 7.89 (t, 1H), 7.71 (d, 1H), 7.56 (s, 2H), 7.49 (s, 3H), 7.27 (s, 1H), 2.52 (s, 3H), 1.86 (s, 9H), 1.71 (s, 3H). Anal. calcd for [(η5-Cpxph)Ir(L2)Cl]PF6 (706.09): C, 37.42; H, 3.57; N, 3.97. Found: C, 37.45; H, 3.51; N, 3.89. MS: m/z 525.33 [(η5-C5Me4C6H5)Ir(L2)]2+.
[(η 5-Cpxbiph)Ir(L2)Cl]PF6 (6)
Yield: 20.18 mg, 25.8%. 1H NMR (500.13 MHz, CDCl3) δ 8.47 (d, J = 5.5 Hz, 1H), 8.08 (t, J = 7.9, 1.2 Hz, 1H), 7.98 (d, J = 7.6 Hz, 1H), 7.75 (d, J = 8.3 Hz, 2H), 7.70 (d, J = 8.3 Hz, 2H), 7.65 (m, 2H), 7.59 (t, 1H), 7.50 (t, J = 7.6 Hz, 2H), 7.42 (t, J = 7.4 Hz, 1H), 2.71 (s, 3H), 1.93 (s, 3H), 1.90 (s, 3H), 1.85 (s, 3H), 1.79 (s, 3H). Anal. calcd for [(η5-Cpxph)Ir(L2)Cl]PF6 (782.18): C, 43.00; H, 3.74; N, 3.58. Found: C, 43.08; H, 3.71; N, 3.62. MS: m/z 601.42 [(η5-C5Me4C6H4 C6H5)Ir(L2)]2+.
X-ray crystallography
Diffraction data of complex 1 were obtained on a Bruker Smart Apex CCD diffractometer equipped with graphite-monochromated Mo Kα radiation. SADABS program was used to correct the date for absorption effects. The crystal with the Oxford Cryosystems Cobra was kept at 100 K throughout the data collection process. The structure was solved by direct methods [SHELXS (TREF)] with additional light atoms found by Fourier methods and refined against F2 using SHELXL.
The crystal data for complex [(η5-Cp*)Ir(L1)Cl]PF6 1 are available as Fig. 1, and Tables 1 and 2, and have been deposited in the Cambridge Crystallographic Data Centre with CCDC reference number 1574955. X-ray crystallographic data in CIF format can be obtained free of charge from The Cambridge Crystallographic Data Centre via http://www.ccdc.cam.ac.uk/.
NMR spectroscopy
1H NMR spectroscopy was obtained in 5 mm NMR tubes at 298 K on Bruker DPX-500 spectrometers. 1H NMR chemical shifts were internally referenced to (CHD2)(CD3)SO (2.50 ppm) for DMSO-d6 and CHCl3 (7.26 ppm) for chloroform-d1 solutions. The data were processed using XWIN-NMR version 3.6 (Bruker UK Ltd).
Mass spectrometry
Electrospray ionization mass spectra (ESI–MS) were obtained on LCQ FLEET. Samples were prepared in 60% MeOH/40% H2O (v/v). The mass spectra were recorded with a scan range of m/z 200–1500 for positive ions.
UV–Vis spectroscopy
A TU-1901 UV–Vis recording spectrophotometer with 1 cm path-length quartz cuvettes (3 ml) was used. All spectra were processed with UVWinlab software. All experiments were carried out at 298 K unless otherwise stated.
Hydrolysis studies
The final concentrations of compounds 3 and 6 are 1 mM in 30% MeOD-d4/70% D2O (v/v), obtained by dissolving compounds 3 and 6 in MeOD-d4 followed by rapid dilution with D2O. All 1H NMR spectra of different time intervals were recorded at 298 K. The final concentrations of compounds 3 and 6 are 50 µM in 20% MeOH/80% H2O (v/v), obtained by dissolving compounds 3 and 6 in MeOH followed by rapid dilution with H2O. All 1H NMR spectra of different time intervals were recorded at 298 K.
Catalytic oxidation of NADH to NAD+
Complexes 3–6 were evaluated for the catalytic oxidation of NADH to NAD+ by UV–Vis spectroscopy after various time intervals in 20% MeOH/80% H2O at 298 K. The concentration of complexes that remained fixed at 1 μM and NADH concentrations is around 75 μM. The conversion of NADH to NAD+ was followed by absorption at 339 nm (εNADH = 6220 cm−1 M−1) to allow evaluation of kinetic data. As NADH is converted to NAD+ by O2 overtime, a NADH control under the same condition was used for all the UV–Vis experiments.
The reaction between complex 6 (0.25 mM) and NADH (3.5 mol equiv) in 30% MeOD/70% D2O at 298 K was monitored by 1H NMR spectroscopy.
Interaction with GSH
GSH (5 mol equiv) was added to an NMR tube containing a 2 mM solution of complex 3 and 6 in 50% MeOD-d4/50% D2O (v/v) at 298 K under N2 atmosphere. 1H NMR spectra of the resulting solution were recorded at 298 K after various time intervals.
Log P determination
Water saturated octanol (WSO) and octanol-saturated water (OSW) were prepared using 0.05 M aqueous NaCl solution (to suppress hydrolysis of the chlorido complexes) and analytical grade octanol. Mix aliquots of stock solutions of iridium complexes 3 and 6 in OSW and WSO in equal volume and shaken in an IKA Vibrax VXC basic shaker for 4 h at 500 g/min, 298 K, to allow partition at room temperature. The aqueous layer was attentively separated from the octanol layer for iridium analysis. Partition coefficients of metal complexes were calculated using the equation log P = log ([M]WSO/[M]OSW), where [M]WSO was obtained by subtraction of the metal content of the aqueous layer after partition from the metal content of the aqueous layer before partition.
Reactions with nucleobases
Interactions of complexes 3 and 6 (ca. 1 mM) with 9-ethylguanine (9-EtG) and 9-methyladenine (9-MeA) (ca. 1 mM) were studied by 1H NMR spectroscopy after various time intervals after addition of the nucleobase (s) in 30% MeOD/70% D2O (v/v) at 298 K using a Bruker DPX500 NMR spectrometer.
Cell culture
A549 human lung cancer cells were purchased from Shanghai Institute of Biochemistry and Cell Biology (SIBCB). A549 cells were cultured in Dulbecco’s modified Eagle medium (DMEM). All medium were replenished with 10% fetal bovine serum and 1% penicillin–streptomycin solution. All cells were cultured at 37 °C in a humidified incubator under a 5% CO2 atmosphere.
Cell viability assay (MTT assay)
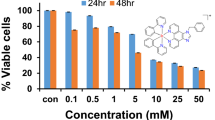
A549 cells were seeded in 96-well plates at a density of 5 × 103 cells per well. After the cells were preincubated in drug-free medium at 37 °C for 24 h, different concentrations of the compounds tested were added. The complexes were dissolved in dimethyl sulfoxide (DMSO) to obtain the stock solution of the drug. This stock was further diluted using cell culture medium to get working concentrations. MTT solution (15 µl, 5 mg/ml) was added to form a purple formazan after 24 h of drug exposure. Then 100 μl of DMSO was added to each well to dissolve the purple formazan. Results were measured by a microplate reader (DNM-9606, Perlong Medical, Beijing, China) at an absorbance of 570 nm. All processing were performed in triplicate. IC50 values quoted are mean ± SEM.
Crystal violet assay
A549 cells were seeded in 96-well plates at a density of 5 × 103 cells per well. After the cells were preincubated in drug-free medium at 37 °C for 24 h, different concentrations of the compounds tested were added. The complexes were dissolved in DMSO to obtain the stock solution of the drug. This stock was further diluted using cell culture medium to get working concentrations. Monolayers were washed twice with PBS and fixed and stained 0.5% crystal violet for 30 min after 24 h of drug exposure. Afterward, plates were washed with hyperpure water and dried and dye was eluted with 0.1 M sodium citrate (pH 4.2) in ethanol (1:1). Results were measured by a microplate reader (DNM-9606, Perlong Medical, Beijing, China) at an absorbance of 570 nm. All processing were performed in triplicate. IC50 values quoted are mean ± SEM.
Cell cycle analysis
A549 cells were seeded in 6-well plates at a density of 1.0 × 106 cells per well. After the cells were preincubated in drug-free medium at 37 °C for 24 h, drugs were added at concentrations of 0.25 × IC50 and 0.5 × IC50. MTT solution (15 µl, 5 mg/ml) was added to form a purple formazan after 24 h of drug exposure. The supernatants were removed after 24 h of drug exposure, and the cells were washed twice with PBS. Afterward, cells were harvested using trypsin–EDTA and fixed in ice-cold 70% ethanol for 24 h. Then, cells rinsed twice with PBS and DNA were stained with propidium iodide (PI) and RNase at 37 °C for 30 min. Cell pellets were washed and resuspended in 200 µl PBS before being analyzed using a flow cytometer (ACEA NovoCyte, Hangzhou, China). Data were analyzed using NovoExpress™ software.
Induction of apoptosis
Apoptotic populations of A549 cells caused by exposure to iridium complexes were carried out using the Annexin V-FITC Apoptosis Detection Kit (Beyotime Institute of Biotechnology, China) by flow cytometry analysis. Momently, A549 cells (1.5 × 106/2 ml per well) were seeded in a 6-well plate. After the cells were preincubated in drug-free medium at 298 K for 24 h, drugs were added at concentrations of 1 × IC50, 2 × IC50, and 3 × IC50. After 24 h of complexes treatment, cells were harvested, washed twice with PBS, and resuspended in 195 μl of binding buffer which was then added to 5 μl of annexin V-FITC and 10 μl of PI, and then incubated at ambient temperature under dark conditions for 15 min. Afterward, the samples were placed in an ice bath in the dark and were analyzed by a flow cytometer (ACEA NovoCyte, Hangzhou, China).
ROS determination
ROS generation in A549 cells caused by exposure to IrIII complexes was carried out using the ROS Assay Kit (Beyotime Institute of Biotechnology, Shanghai, China) by flow cytometry analysis. Momently, 1.0 × 106 A549 cells per well were seeded in a 6-well plate. The cells were preincubated in drug-free medium at 37 °C for 24 h in a 5% CO2 humidified atmosphere, and then drugs were added at concentrations of 0.25 × IC50, 0.5 × IC50. After 24 h of drug exposure, cells were washed twice with PBS and then incubated with the DCFH-DA probe (10 µM) at 37 °C for 30 min, and then washed it three times immediately with PBS. The fluorescence intensity was analyzed by flow cytometry (ACEA NovoCyte, Hangzhou, China). Data were analyzed using NovoExpress™ software. Samples were kept in the darks all the time to avoid light-induced ROS production.
Results and discussion
Three iridium cyclopentadienyl dimers [(η5-Cp*)IrCl2]2 (dimer 1), [(η5-Cpxph)IrCl2]2 (dimer 2) and [(η5-Cpxbiph)IrCl2]2 (dimer 3), where Cp* = C5Me5, Cpxph = C5Me4C6H5, Cpxbiph = C5Me4C6H4C6H5, were synthesized according to a reported method [50]. Six IrIII half-sandwich iminopyridine complexes 1–6 of the type [(η5-Cpx)Ir(N^N)Cl]PF6 were prepared by reaction of corresponding dimers 1–3 with iminopyridine ligand L1–L2 in 1:2.4 ratio in methanol at ambient temperature followed by ion exchange from Cl− to PF6− (Fig. 2; Scheme 1). All the six complexes are newly prepared and fully characterized by mass spectroscopy, 1H NMR spectroscopy and elemental analysis.
X-ray crystal structure
Crystal of [(η5-Cp*)Ir(L1)Cl]PF6 (1) was grown from CH2Cl2/diethyl ether by vapor diffusion. The molecular structure of 1 was determined (Fig. 2). Crystallographic data and selected bond lengths and angles are listed in Tables 1 and 2. Complex 1 adopts the half-sandwich piano-stool geometry. The distance between iridium to the centroid of bound η5-cyclopentadienyl ligand is 1.8153 Å. The bond length of Ir–Cl is 2.3986(12) Å for 1, which is almost identical to the reported [38] cationic polypyridyl iridium complexes [(η5-C5Me4C6H4C6H5)Ir(bpy)Cl]PF6 (2.3840 (14) Å) and [(η5-C5Me4C6H5)Ir(bpy)Cl]PF6 (2.3859 (5) Å). The Ir–N1 (chelating ligand) bond length (2.087(4) Å) is dramatically shorter than the bond length of Ir–N2 (2.157 (4) Å). The single crystal structure of the cationic part of complex 4 has been reported [51].
Cell viability assay
The effect of cell viability of complexes 1–6 on human lung cancer A549 cells was evaluated by 3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide (MTT) assay, and the IC50 values (concentration at which 50% of the cell growth is inhibited) after 24 h drug exposure are listed in Table 3. The Cp* complex 1 deemed as inactive as its IC50 was > 100 µM. However, the rest five complexes were all active against A549 cell line, displaying IC50 values range from 75.9 to 2.7 μM. Basically, for those complexes containing the same chelating ligands, their cell viability decreased with the number of the phenyl ring on the Cp ring in the trend of Cpxbiph < Cpxph < Cp*, which is consistent with our previous results [50]. Introduction of one phenyl ring switched on anticancer activity from complex 1 (IC50 > 100 μM) to complex 2 (IC50 16.9 μM), and the cell viability decreased significantly from complex 2 to 3 (IC50 2.7 μM) arising from introduction of the second phenyl ring on Cp. The additional phenyl ring increases the lipophilicity of the complexes, which in turn enhances the cell membrane penetration [38]. Varying chelating ligands also caused significant differences on the cell viability. The complex 3 containing adamantyl iminopyridine ligand displayed the most potent anticancer activity with IC50 value of 2.7 μM, ca. 4 and 8 ×more active than its oxime pyridyl analog and cisplatin, respectively. Given the best anticancer activity displayed by complexes 3 and 6, the two complexes were chosen to perform more experiments to investigate their MoAs. The cell viability assay was also determined by crystal violet method (Table S1). There is a significant difference from the results of MTT assay.
Hydrolysis studies
In comparison with metal chloride complexes, metal aqua complexes often display more reactive activity, and therefore the hydrolysis of the M–Cl bonds (M = metal) can represent an activation step for transition metal-based anticancer complexes, for example cisplatin [38]. The hydrolysis of complexes 3 and 6 was studied by 1H NMR spectroscopy in 30% MeOD-d4/70% D2O at 298 K, where methanol was used to avoid precipitation of the complexes in aqueous solution. However, no hydrolysis was observed by NMR for complexes 3 and 6 in 30% MeOD-d4/70% D2O at 298 K (Figure S1). The UV–Vis spectra also could not detect obvious hydrolysis for complexes 3 and 6 (50 µM) in 20% MeOH/80% H2O (v/v) (Figure S2), indicating that these complexes remained stable under the above conditions.
Interaction with nucleobases
Like cisplatin, transition metal anticancer complexes often bind to DNA and thus cause cell death. The binding of complexes 3 and 6 to nucleobase models 9-EtG (9-ethylguanine) and 9-MeA (9-methyladenine) was investigated. Nucleobases 9-EtG or 9-MeA was added to an solution containing iridium complex (1 mM) in 30% MeOD-d4/70% D2O (v/v), and the solution was recorded by 1H NMR spectra at 298 K after various time intervals. Based on 1H NMR peak integrals, the percentages of iridium nucleobase adduct formed after 24 h are shown in Table 4. Although no hydrolysis for complex [(η5-Cpxbiph)Ir(L2)Cl]PF6 (6), the addition of 9-EtG to an solution of 6 led to the appearance of one set of new peaks assignable to the 9-EtG adduct, showing that 86% of 6 reacted after 24 h (Fig. 3a). Complex 6 also showed the high affinity for 9-MeA with 40% nucleobase adducts formation after 24 h (Fig. 3b). However, no any nucleobase adduct formation was observed for complex 3 with 9-EtG and 9-MeA (Figure S3), most likely due to the steric hindrance of the adamantane. These results indicate that the chelating ligand plays a crucial role in reaction with nucleobases.
Catalysis of NADH oxidation
Coenzyme NADH and NAD+ is crucial in numerous biocatalyzed processes, such as electron transfer in the mitochondrial electron chain. Previously, we have shown that cyclopentadienyl IrIII aqua complexes can catalytically convert NADH to its oxidized form NAD+ by robbing hydride from NADH, and pass the hydride to oxygen to finally generate reactive oxygen species (ROS H2O2), thus provide a pathway to an oxidant MoA [46, 52]. First, we investigated the reaction between complex 6 (0.25 mM) and NADH (3.5 mol equiv) in 30% MeOD-d4/70% D2O (v/v), which was recorded by 1H NMR at 298 K. When NADH was added to a solution of complex 6, NADH was gradually converted to NAD+ and a sharp singlet correspond to Ir–H at − 11.4 ppm was detected.
To evaluate the catalytic abilities of the iridium complexes, the reaction between NADH (around 75 μM) and complexes 3–6 (fixed concentration of complex at 1.0 μM) in 20% MeOH/80% H2O (v/v) was monitored by UV–Vis spectroscopy over 8 h at 298 K (Fig. 4b; Figure S4). As NADH has an obvious absorbance at 339 nm while NAD+ does not, the decrease of absorbance intensity at 339 nm was monitored and the data were used for calculation of the turnover number (TON) for each complex (Fig. 4c). Basically, the complexes displayed good catalytic activity in conversion of NADH to NAD+, with TONs range from 6.8 of complex 5 to 19.2 of complex 6. Complexes 3 and 6 showed higher catalytic ability than did 4 and 5, which is consistent with their anticancer activity. The TONs of these complexes are similar with the half-sandwich IrIII N-heterocyclic carbene complexes [50]. The Sadler group has reported that the iridium complex could modify the NAD+/NADH ratio in A2780 cancer cells [52]. Therefore, the potent catalytic activity of these complexes in converting NADH to NAD+ may provide a basis to modulate the redox balance and generate ROS in cells [46, 53]. In a solution of 6 (0.5 mM) with NADH (3 mol equiv) in 50% MeOH/50% H2O (v/v) at 298 K, the ROS hydrogen peroxide was detected by appearing blue on hydrogen peroxide test paper (Fig. 4d). It is found that the concentration of H2O2 is about 0.18 mM, and its level may be limited by the oxygen solubility (ca. 0.23 mM at 288 K) [46].
a 1H NMR spectra recorded the reaction between complex [(η5-Cpxbiph)Ir(L2)Cl]PF6 (6) (0.25 mM) and NADH (3.5 mol equiv) in 30% MeOD/70% D2O (v/v) at 298 K. b UV–Vis spectra recorded the reaction of NADH (about 75 µM) with complex 6 (1 µM) in 20% MeOH/80% H2O (v/v) at 298 K for 8 h. c Turnover numbers of complexes 3–6. d Detection of hydrogen peroxide (H2O2) in a solution of complex 6 (0.5 mM) with NADH (3 mol equiv) in 50% MeOH/50% H2O (v/v) at 298 K. After 24 h, H2O2 (ca. 0.18 mM) was detected by peroxide-test paper
Interaction with GSH
GSH can coordinate to the metals in anticancer drugs to form less toxic GSH conjugates, thus detoxifying anticancer drugs [46]. Therefore, the stability of Cpxbiph complexes 3 and 6 to GSH was explored. GSH (5 mol equiv) was added to an 50% MeOD-d4/50% D2O (v/v) solution containing complex 3 or 6 (2 mM) under N2 atmosphere, and the solution was recorded by 1H NMR spectra at 298 K after various time intervals. No new peaks were observed by NMR for complex 3 (Figure S5). The result shows that adamantyl iminopyridine complex 3 is stable in the solution of GSH, which may avoid inactivating before reaching its biological targets. However, 1H NMR spectra showed that 6 had reacted with GSH after 10 min (Figure S6). The less stability of complex 6 in the presence of GSH may lead to less potency than complex 3 against cancer cells.
Hydrophobicity (log P)
The octanol/water partition coefficients (log P) are usually a factor relevant for cellular uptake and anticancer activity [38]. The log P values for the complexes 3 and 6 were determined (Table 5). To make sure that the determined log P values are for the chlorido complexes, NaCl (0.05 M) was added to the solution of the compounds to inhibit hydrolysis. The log P values for complexes 3 and 6 are 1.16 and 0.70, respectively, suggesting that complex 3 possesses higher hydrophobicity than 6, and therefore contributed to the more potent anticancer activity of complex 3 than 6. This result showed that the hydrophobicity of the complexes is related to their anticancer activity.
Apoptosis assay
Cpxbiph complexes 3 and 6 were chose to explore more details on their MoAs using flow cytometry. Apoptosis is the process where cells die in a programmed fashion under particular stimuli. A large number of publications have reported that metal-based anticancer agents could inhibit cell growth by causing apoptosis [54,55,56,57]. To investigate whether the cell viability of the complexes is a result of apoptosis, A549 cells were exposed to complexes 3 and 6 at IC50, 2 × IC50 and 3 × IC50 concentrations for 24 h, followed by annexin V/propidium iodide staining, and then determined by flow cytometry. Cell populations can be distinguished as viable (annexin V−/PI−), nonviable (annexin V−/PI+), early apoptosis (annexin V+/PI−), and late apoptosis (annexin V+/PI+). Compared to the high viability of untreated cells (93.1%), obvious induction of apoptosis was observed in those cells treated with complexes 3 and 6 (Fig. 5; Table S2). The plots showed that around 4.3%, A549 cells were in early apoptotic phase and 23.2% cells were in the late apoptotic phase after 24 h of exposure to complex 3 at a concentration of IC50. The proportion of cells in late apoptosis increased significantly with higher concentration of complex 3, where 94.3% of cells were observed in this stage upon exposure to 3 at 3 × IC50. Interestingly, complex 6 remained most cells in early apoptosis with 84.2% in early apoptosis and only 8.9% in late apoptosis at 3 × IC50.
Therefore, both complexes 3 and 6 dramatically induced apoptosis of A549 cells in a dose-dependent manner. However, there were big differences in their mode of action. In comparison to the complex 3 which caused most cells in late apoptosis, complex 6 was mainly concentrated in early apoptosis. The stronger ability to cause apoptosis of complex 3 is consistent with its lower the cell viability compared to complex 6.
Cell cycle arrest
Arrest of cell cycle of cancer cells has been recognized as an effective strategy in chemotherapy [57]. In this work, compounds 3 and 6 were investigated their effects on the cell cycle of A549 cells. Treating of A549 cells with compound 3 at 0.5 ×IC50 led to obvious G1 phase arrest, where the percentages of cells increased 12.6% compared to untreated cells (Fig. 6; Table S3). In addition, a sub-diploid peak corresponds to sub-G1 was detected, usually indicating cells are undergoing apoptosis. This result is consistent with the apoptosis assay. Upon exposure of the cells to complex 6 at a concentration of 0.5 × IC50, there were significant increases in the proportions of cells at S phase and G2/M phase, about 6.6% and 23.2%, respectively. Compared to the untreated control, complex 3 and 6 disturbed the cell cycle at G1 phase, and S and G2/M phase, respectively.
ROS induction
Reactive oxygen species (ROS) play important roles as regulating cell proliferation, death and signal transduction [58, 59]. The ROS level in A549 cells induced by complexes 3 and 6 (concentrations used: 0.25 × IC50) was monitored by flow cytometry (Fig. 7; Table S4). Compared to untreated cells, a dramatic increase in the ROS levels was observed for both complexes even at low concentration (0.25 × IC50) after 24 h of drug exposure. Complex 3 caused even higher ROS levels in cells than complex 6, causing 93.4% of A549 cells in high ROS levels at concentration of 0.25 ×IC50. The higher ability to generate ROS in cells of complex 3 is correlated to its more potent anticancer activity in comparison with complex 6. The generation of ROS by the complexes may provide a basis for antiproliferative activity.
Conclusion
In this work, six IrIII half-sandwich complexes [(η5-Cpx)Ir(N^N)Cl]PF6 containing two types of iminopyridine chelating ligand were designed and prepared. Four complexes exhibited superior anticancer potency toward A549 cancer cells than the clinical used cisplatin. The most potent complex 3 is 8 × more active than cisplatin against A549 cells. The anticancer activity can be tuned by varying both Cp*-based ligands and iminopyridine ligands, in the order of Cpxbiph > Cpxph > Cp*.
Basically, both complexes 3 and 6 containing different iminopyridine ligands did not show hydrolysis in aqueous condition; however, the chelating ligand played a very important role in nucleobase binding selectivity and in the reaction with glutathione. Complex 6 showed strong binding to the nucleobases, preferring 9-EtG than 9-MeA. These results suggest that DNA could be a potential target, although other targets appear to be also important because complex 3 containing adamantyl iminopyridine is a potent anticancer agent even no nucleobase binding. Complex 3 is much more stable than complex 6 in the presence of GSH, and is more hydrophobic than complex 6, which appears contribute to its higher anticancer activity. This type of complexes are effective catalysts in converting coenzyme NADH to NAD+ with formation of an Ir hydride adduct, thus possible affect the redox environment in cells. Further mechanistic studies showed that complexes 3 and 6 induced obvious apoptosis, arrested cell cycles at G1, S, and G2/M phase, respectively, and significantly increased the ROS levels in A549 cancer cells. Our study demonstrates that these iridium iminopyridine complexes could be potential anticancer agents.
References
Lozano-Perez AA, Gil AL, Perez SA, Cutillas N, Meyer H, Pedreno M, Aznar-Cervantes SD, Janiak C, Cenis JL, Ruiz J (2015) Antitumor properties of platinum(IV) prodrug-loaded silk fibroin nanoparticles. Dalton Trans 44(30):13513–13521
Pizarro AM, McQuitty RJ, Mackay FS, Zhao Y, Woods JA, Sadler PJ (2014) Cellular accumulation, lipophilicity and photocytotoxicity of diazido platinum(IV) anticancer complexes. ChemMedChem 9(6):1169–1175
Muhammad N, Sadia N, Zhu C, Luo C, Guo Z, Wang X (2017) Biotin-tagged platinum(IV) complexes as targeted cytostatic agents against breast cancer cells. Chem Commun 53(72):9971–9974
Chen Y, Guan R, Zhang C, Huang J, Ji L, Chao H (2016) Two-photon luminescent metal complexes for bioimaging and cancer phototherapy. Coord Chem Rev 310:16–40
Zeng L, Kuang S, Li G, Jin C, Ji L, Chao H (2017) A GSH-activatable ruthenium(II)-azo photosensitizer for two-photon photodynamic therapy. Chem Commun 53(12):1977–1980
Shen J, Kim H-C, Wolfram J, Mu C, Zhang W, Liu H, Xie Y, Mai J, Zhang H, Li Z, Guevara M, Mao Z-W, Shen H (2017) A liposome encapsulated ruthenium polypyridine complex as a theranostic platform for triple-negative breast cancer. Nano Lett 17(5):2913–2920
Zhang D-Y, Zheng Y, Tan C-P, Sun J-H, Zhang W, Ji L-N, Mao Z-W (2017) Graphene oxide decorated with Ru(II)–polyethylene glycol complex for lysosome-targeted imaging and photodynamic/photothermal therapy. ACS Appl Mater Interfaces 9(8):6761–6771
Wang M, Mao Z, Kang T-S, Wong C-Y, Mergny J-L, Leung C-H, Ma D-L (2016) Conjugating a groove-binding motif to an Ir(III) complex for the enhancement of G-quadruplex probe behavior. Chem Sci 7(4):2516–2523
Fong TT-H, Lok C-N, Chung CY-S, Fung Y-ME, Chow P-K, Wan P-K, Che C-M (2016) Cyclometalated palladium(II) N-heterocyclic carbene complexes: anticancer agents for potent in vitro cytotoxicity and in vivo tumor growth suppression. Angew Chem Int Ed 55(39):11935–11939
Chan KT, Tong GSM, To W-P, Yang C, Du L, Phillips DL, Che C-M (2017) The interplay between fluorescence and phosphorescence with luminescent gold(I) and gold(III) complexes bearing heterocyclic arylacetylide ligands. Chem Sci 8(3):2352–2364
Gasser G, Ott I, Metzler-Nolte N (2011) Organometallic anticancer compounds. J Med Chem 54(1):3–25
Graf M, Gothe Y, Metzler-Nolte N, Czerwieniec R, Sünkel K (2015) Cyclometalated iridium(III) complexes containing semicarbazone ligands: synthesis, characterization, photophysical and biological studies. Z Anorg Allg Chem 641(10):1798–1802
Raszeja LJ, Siegmund D, Cordes AL, Güldenhaupt J, Gerwert K, Hahn S, Metzler-Nolte N (2017) Asymmetric rhenium tricarbonyl complexes show superior luminescence properties in live cell imaging. Chem Commun 53(5):905–908
Zhu Z, Wang Z, Hao Y, Zhu C, Jiao Y, Chen H, Wang Y-M, Yan J, Guo Z, Wang X (2016) Glutathione boosting the cytotoxicity of a magnetic platinum(IV) nano-prodrug in tumor cells. Chem Sci 7(4):2864–2869
Albada B, Metzler-Nolte N (2016) Organometallic–peptide bioconjugates: synthetic strategies and medicinal applications. Chem Rev 116(19):11797–11839
Tan C-P, Lu Y-Y, Ji L-N, Mao Z-W (2014) Metallomics insights into the programmed cell death induced by metal-based anticancer compounds. Metallomics. 6(5):978–995
Tian M, Li J, Zhang S, Guo L, He X, Kong D, Zhang H, Liu Z (2017) Half-sandwich ruthenium(II) complexes containing N^N chelated Imino-pyridyl ligands that are selectively toxic to cancer cells. Chem Commun 53:12810–12813
Wedlock LE, Barnard PJ, Filipovska A, Skelton BW, Berners-Price SJ, Baker MV (2016) Dinuclear Au(I) N-heterocyclic carbene complexes derived from unsymmetrical azolium cyclophane salts: potential probes for live cell imaging applications. Dalton Trans 45(30):12221–12236
Peacock AFA, Melchart M, Deeth RJ, Habtemariam A, Parsons S, Sadler PJ (2007) Osmium(II) and ruthenium(II) arene maltolato complexes: rapid hydrolysis and nucleobase binding. Chem Eur J 13(9):2601–2613
Chen T, Liu Y, Zheng W-J, Liu J, Wong Y-S (2010) Ruthenium polypyridyl complexes that induce mitochondria-mediated apoptosis in cancer cells. Inorg Chem 49(14):6366–6368
Xiaomin L, Meng T, Zhenzhen T, Lai-Jin T, Mengqi L, Jing H, Zhe L (2017) Potent cyclopentadienyl iridium anticancer complexes containing CN-chelating ligands. Chin J Inorg Chem 33(7):1119–1131
Dörr M, Meggers E (2014) Metal complexes as structural templates for targeting proteins. Curr Opin Chem Biol 19:76–81
Liu Z, Lebrun V, Kitanosono T, Mallin H, Köhler V, Häussinger D, Hilvert D, Kobayashi S, Ward TR (2016) Upregulation of an artificial zymogen by proteolysis. Angew Chem Int Ed 55(38):11587–11590
Li SP-Y, Yip AM-H, Liu H-W, Lo KK-W (2016) Installing an additional emission quenching pathway in the design of iridium(III)-based phosphorogenic biomaterials for bioorthogonal labelling and imaging. Biomaterials 103:305–313
Petrini A, Pettinari R, Marchetti F, Pettinari C, Therrien B, Galindo AN, Scopelliti R, Riedel T, Dyson PJ (2017) Cytotoxic half-sandwich Rh(III) and Ir(III) β-diketonates. Inorg Chem 56(21):13600–13612
Lucas SJ, Lord RM, Wilson RL, Phillips RM, Sridharan V, McGowan PC (2012) Synthesis of iridium and ruthenium complexes with (N, N), (N, O) and (O, O) coordinating bidentate ligands as potential anti-cancer agents. Dalton Trans 41(45):13800–13802
Liu J, Jin C, Yuan B, Chen Y, Liu X, Ji L, Chao H (2017) Enhanced cancer therapy by the marriage of metabolic alteration and mitochondrial-targeted photodynamic therapy using cyclometalated Ir(III) complexes. Chem Commun 53(71):9878–9881
Ye R-R, Cao J-J, Tan C-P, Ji L-N, Mao Z-W (2017) Valproic acid-functionalized cyclometalated iridium(III) complexes as mitochondria-targeting anticancer agents. Chem Eur J 23(60):15166–15176
Wenger OS (2015) Proton-coupled electron transfer with photoexcited ruthenium(II), rhenium(I), and iridium(III) complexes. Coord Chem Rev 282–283:150–158
Lo KK-W (2015) Luminescent rhenium(I) and iridium(III) polypyridine complexes as biological probes, imaging reagents, and photocytotoxic agents. Acc Chem Res 48(12):2985–2995
Chen Y, Qiao L, Ji L, Chao H (2014) Phosphorescent iridium(III) complexes as multicolor probes for specific mitochondrial imaging and tracking. Biomaterials 35(1):2–13
Ye R-R, Tan C-P, Ji L-N, Mao Z-W (2016) Coumarin-appended phosphorescent cyclometalated iridium(III) complexes as mitochondria-targeted theranostic anticancer agents. Dalton Trans 45(33):13042–13051
Almodares Z, Lucas SJ, Crossley BD, Basri AM, Pask CM, Hebden AJ, Phillips RM, McGowan PC (2014) Rhodium, iridium, and ruthenium half-sandwich picolinamide complexes as anticancer agents. Inorg Chem 53(2):727–736
Payne R, Govender P, Therrien B, Clavel CM, Dyson PJ, Smith GS (2013) Neutral and cationic multinuclear half-sandwich rhodium and iridium complexes coordinated to poly(propyleneimine) dendritic scaffolds: synthesis and cytotoxicity. J Organomet Chem 729:20–27
Novohradsky V, Liu Z, Vojtiskova M, Sadler PJ, Brabec V, Kasparkova J (2014) Mechanism of cellular accumulation of an iridium(III) pentamethylcyclopentadienyl anticancer complex containing a C, N-chelating ligand. Metallomics 6(3):682–690
Hearn JM, Romero-Canelón I, Qamar B, Liu Z, Hands-Portman I, Sadler PJ (2013) Organometallic iridium(III) anticancer complexes with new mechanisms of action: nCI-60 screening, mitochondrial targeting, and apoptosis. ACS Chem Biol 8(6):1335–1343
Liu Z, Salassa L, Habtemariam A, Pizarro AM, Clarkson GJ, Sadler PJ (2011) Contrasting reactivity and cancer cell cytotoxicity of isoelectronic organometallic iridium(III) complexes. Inorg Chem 50(12):5777–5783
Liu Z, Habtemariam A, Pizarro AM, Fletcher SA, Kisova A, Vrana O, Salassa L, Bruijnincx PCA, Clarkson GJ, Brabec V, Sadler PJ (2011) Organometallic half-sandwich iridium anticancer complexes. J Med Chem 54(8):3011–3026
Nazif MA, Rubbiani R, Alborzinia H, Kitanovic I, Wolfl S, Ott I, Sheldrick WS (2012) Cytotoxicity and cellular impact of dinuclear organoiridium DNA intercalators and nucleases with long rigid bridging ligands. Dalton Trans 41(18):5587–5598
Gras M, Therrien B, Süss-Fink G, Casini A, Edafe F, Dyson PJ (2010) Anticancer activity of new organo-ruthenium, rhodium and iridium complexes containing the 2-(pyridine-2-yl)thiazole N, N-chelating ligand. J Organomet Chem 695(8):1119–1125
Carmona D, Lahoz FJ, Elipe S, Oro LA, Lamata MP, Viguri F, Mir C, Cativiela C, López-Ram de Víu MP (1998) Synthesis, characterization, properties, and asymmetric catalytic Diels–Alder reactions of chiral-at-metal imino–iridium(III) complexes. Organometallics 17(14):2986–2995
Lu CC, Bill E, Weyhermüller T, Bothe E, Wieghardt K (2008) Neutral bis(α-iminopyridine)metal complexes of the first-row transition ions (Cr, Mn, Fe Co, Ni, Zn) and their monocationic analogues: mixed valency involving a redox noninnocent ligand system. J Am Chem Soc 130(10):3181–3197
Sylvester KT, Chirik PJ (2009) Iron-catalyzed, hydrogen-mediated reductive cyclization of 1,6-enynes and diynes: evidence for bis(imino)pyridine ligand participation. J Am Chem Soc 131(25):8772–8774
Liu Z, Sadler PJ (2014) Organoiridium complexes: anticancer agents and catalysts. Acc Chem Res 47(4):1174–1185
Fu Y, Sanchez-Cano C, Soni R, Romero-Canelon I, Hearn JM, Liu Z, Wills M, Sadler PJ (2016) The contrasting catalytic efficiency and cancer cell antiproliferative activity of stereoselective organoruthenium transfer hydrogenation catalysts. Dalton Trans 45(20):8367–8378
Liu Z, Romero-Canelón I, Qamar B, Hearn JM, Habtemariam A, Barry NP, Pizarro AM, Clarkson GJ, Sadler PJ (2014) The potent oxidant anticancer activity of organoiridium catalysts. Angew Chem Int Ed 53(15):3941–3946
Fu Y, Romero MJ, Habtemariam A, Snowden ME, Song L, Clarkson GJ, Qamar B, Pizarro AM, Unwin PR, Sadler PJ (2012) The contrasting chemical reactivity of potent isoelectronic iminopyridine and azopyridine osmium(II) arene anticancer complexes. Chem Sci 3(8):2485–2494
Li J, Guo L, Tian Z, Tian M, Zhang S, Xu K, Qian Y, Liu Z (2017) Novel half-sandwich iridium(iii) imino-pyridyl complexes showing remarkable in vitro anticancer activity. Dalton Trans 46(44):15520–15534
Guo L, Jing X, Xiong S, Liu W, Liu Y, Liu Z, Chen C (2016) Influences of alkyl and aryl substituents on iminopyridine Fe(II)- and Co(II)-catalyzed isoprene polymerization. Polymers 8(11):389–400
Wang C, Liu J, Tian Z, Tian M, Tian L, Zhao W, Liu Z (2017) Half-sandwich iridium N-heterocyclic carbene anticancer complexes. Dalton Trans 46(21):6870–6883
Megumi W, Yohei K, Shigeki K, Takao I (2010) Synthesis, structure, and reversible deprotonation of a half-sandwich iridium complex bearing a chelating oxime ligand. Chem Lett 39(7):758–759
Betanzos-Lara S, Liu Z, Habtemariam A, Pizarro AM, Qamar B, Sadler PJ (2012) Organometallic ruthenium and iridium transfer-hydrogenation catalysts using coenzyme NADH as a cofactor. Angew Chem Int Ed 51(16):3897–3900
Liu Z, Deeth RJ, Butler JS, Habtemariam A, Newton ME, Sadler PJ (2013) Reduction of quinones by NADH catalyzed by organoiridium complexes. Angew Chem Int Ed 52(15):4194–4197
Zeng L, Gupta P, Chen Y, Wang E, Ji L, Chao H, Chen Z-S (2017) The development of anticancer ruthenium(II) complexes: from single molecule compounds to nanomaterials. Chem Soc Rev 46(19):5771–5804
Zhou H-Y, Li M, Qu J, Jing S, Xu H, Zhao J-Z, Zhang J, He M-F (2016) Effective antitumor candidates based upon ferrocenylseleno-dopamine derivatives: growth inhibition by induction cell apoptosis and antivascular effects. Organometallics 35(11):1866–1875
Zeng L, Chen Y, Liu J, Huang H, Guan R, Ji L, Chao H (2016) Ruthenium(II) complexes with 2-phenylimidazo[4,5-f][1,10]phenanthroline derivatives that strongly combat cisplatin-resistant tumor cells. Sci Rep 6:19449–19461
Evan GI, Vousden KH (2001) Proliferation, cell cycle and apoptosis in cancer. Nature 411:342–348
Kim H-O, Yeom M, Kim J, Kukreja A, Na W, Choi J, Kang A, Yun D, Lim J-W, Song D, Haam S (2017) Reactive oxygen species-regulating polymersome as an antiviral agent against influenza virus. Small 13(32):1700818–1700828
Chen W, Balakrishnan K, Kuang Y, Han Y, Fu M, Gandhi V, Peng X (2014) Reactive oxygen species (ROS) inducible DNA cross-linking agents and their effect on cancer cells and normal lymphocytes. J Med Chem 57(11):4498–4510
Acknowledgements
We thank the National Natural Science Foundation of China (Grant no. 21671118) and the Taishan Scholars Program for support.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Kong, D., Tian, M., Guo, L. et al. Novel iridium(III) iminopyridine complexes: synthetic, catalytic, and in vitro anticancer activity studies. J Biol Inorg Chem 23, 819–832 (2018). https://doi.org/10.1007/s00775-018-1578-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00775-018-1578-0