Abstract
Methionine is a nutritionally essential sulfur-containing amino acid found at low levels in plant tissues. Yet, the factors that regulate its synthesis and accumulation in seeds are not fully known. Recent genetic studies demonstrate that Arabidopsis seeds are able to synthesize methionine de novo through the aspartate family pathway similarly to vegetative tissues; however, additional biochemical studies suggest that the S-methylmethionine (SMM) cycle also plays a major role in methionine synthesis in seeds. To better understand the contribution of these two pathways to methionine synthesis, we have sampled various vegetative and reproductive tissues during the Arabidopsis life cycle and determined the contents of soluble and protein-incorporated methionine, SMM, as well as the expression levels of the key genes involved in these two pathways. Our results strengthen the hypothesis that SMM that is produced in the rosette leaves from methionine contributes to methionine accumulation in seeds. However, the SMM cycle may have additional functions in plant tissues since its key genes were expressed in all of the examined tissues, although at different rates. The accumulation patterns of soluble and protein-incorporated methionine during the Arabidopsis life cycle were found to be similar to most of the other amino acids, especially to those belonging to the branched-chain and aromatic amino acids that are produced in chloroplasts together with methionine. This indicates that similar factors regulate the levels of amino acids during development.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Methionine is a nutritionally essential sulfur-containing amino acid found at low levels in plants and in their seeds. Therefore, its level limits the nutritional value of plants as a source of dietary protein for humans and animals. In plants, methionine not only serves as a building block for protein synthesis, but through its main derivative, S-adenosyl-methionine (SAM), controls the levels of several key metabolites such as ethylene, polyamines and biotin. SAM is also the primary methyl group donor that regulates various essential processes in plants [reviewed by (Amir 2010; Galili and Amir 2013; Roje 2006)]. For these reasons, studies were performed to assess the factors that regulate methionine metabolism and its accumulation in vegetative tissues and seeds, though most of them focused on factors regulating its synthesis [reviewed by Amir et al. 2012; Galili and Amir 2013; Hesse and Hoefgen 2003; Jander and Joshi 2010)]. The level of methionine in vegetative tissues is regulated mainly by the expression level of its first committed enzyme, CYSTATHIONINE γ-SYNTHASE (CGS), which combines the carbon/amino skeleton derived from aspartate with the sulfur moiety derived from cysteine (Kim et al. 2002; Ravanel et al. 1998) (Fig. 1). Recent studies performed in legumes, tobacco and Arabidopsis seeds show that seed-specific expression of feedback-insensitive mutated forms of Arabidopsis CGS (AtCGS) leads to significantly higher levels of soluble methionine (Hanafy et al. 2013; Matityahu et al. 2013; Song et al. 2013; Cohen et al. 2014), indicating that methionine can be synthesized de novo in seeds via the aspartate family pathway through CGS activity similarly to vegetative tissues.
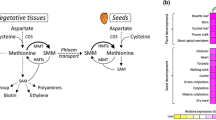
Metabolic pathways and genes associated with Met metabolism. Key enzymes in the aspartate family pathway and the SMM cycle are underlined. Methionine catabolic products are presented in italics. Dashed arrows represent several metabolic steps. OAS O-acetylserine, MTHF methyltetrahydrofolate, SMM S-methylmethionine, SAM S-adenosylmethionine, CGS cystathionine-γ-synthase, MS methionine synthase, MGL methionine-γ-lyase, HMTs homocysteine S-methyltransferases, MMT Met S-methyltransferase, SAMS SAM synthase
On the other hand, several other studies suggest that methionine is synthesized in seeds through an additional and/or alternative pathway in which methionine that is produced in vegetative tissues is converted to S-methyl-methionine (SMM) by the activity of METHIONINE S-METHYLTRANSFERASE (MMT) and then transported via the phloem into the developing seeds at the onset of the reproductive period. In the seeds, the transported SMM is then re-converted back to soluble methionine by HOMOCYSTEINE S-METHYLTRANSFERASE (HMT), thus contributing to the synthesis of methionine in seeds (Fig. 1) (Bourgis et al. 1999; Lee et al. 2008). This assumption is based on several observations. (1) Transgenic Arabidopsis plants overexpressing AtCGS and methionine over-accumulating Arabidopsis mutants exhibited relatively high levels of soluble methionine in their young rosette leaves. When these mutants began to flower, the contents of methionine decreased in the rosette leaves and significantly increased in the inflorescence apex region, immature siliques and seeds. Interestingly, the expression levels of AtCGS were found to be similar in leaves during these two developmental stages (Bartlem et al. 2000; Inaba et al. 1994; Kim et al. 2002), suggesting that methionine is transferred from the leaves toward the reproductive organs including the seeds. (2) Radioactive experiments showed that methionine is transported from leaves to seeds in the form of SMM (Bourgis et al. 1999), which was found to be the second major sulfur metabolite after glutathione in Arabidopsis and wheat phloem sap (Tan et al. 2010). (3) Flux experiments in wheat suggested that the major conversion in leaves is from methionine to SMM, and in seeds from SMM back to methionine. These researchers defined that the activity levels of HMT in Arabidopsis seeds are higher than that those of MMT, and the incorporation of 35S into the protein was the same whether 35S-methionine or 35S-SMM was supplied (Ranocha et al. 2001). (4) Arabidopsis hmt2 mutants exhibited significant accumulation of methionine in their seeds, suggesting that SMM that is transported to the seeds is re-converted to methionine by HMT isoenzymes other than HMT2. All the above lines of evidence strongly imply that soluble methionine is translocated as SMM towards the sink organs at the onset of the reproductive stage, thus exhibiting temporal and spatial accumulation patterns.
The relative contribution of the aspartate family and SMM pathways to methionine synthesis in seeds is probably more complex than assumed. Integrated comparative proteomic and transcriptomic analyses of different sections of Medicago truncatula seeds suggest that these two pathways contribute to methionine synthesis in seeds at different stages of seed development (Gallardo et al. 2007). The researchers imply that methionine is synthesized through the aspartate family pathway in endosperm and seed coat up to the mid-stages of seed filling, but in the late stages of seed filling methionine is synthesized from the SMM pathway at the seed coat (Gallardo et al. 2007). This assumption is based on the expression levels of the three genes/proteins of HMT, MS (METHIONINE SYNTHASE) and SAMS (SAM SYNTHASE) (Fig. 1). Despite the fact that these results are based on transcripts and protein levels rather than on distinct metabolic changes, they suggest roles played by these two pathways in methionine synthesis in seeds.
In spite of these lines of evidence, the role of SMM in methionine synthesis in seeds requires additional studies, since it was shown that the lack of SMM cycle in Arabidopsis and maize mmt mutants does not alter the levels of methionine, thiols and total sulfur compared to wild-type seeds (Kocsis et al. 2003). Additionally, transgenic tobacco plants overexpressing the AtCGS together with the bacterial feedback-insensitive form of AK(ASPARTATE KINASE), which exhibited 176- and 39-fold higher levels of methionine and SMM, respectively, showed only slightly higher methionine contents in their seeds (Hacham et al. 2008). Similarly, an examination of the uptake, transport and conversion of 13C-SMM and 13C-methionine in Arabidopsis hmt2 and mmt mutants revealed that SMM is a non-essential intermediate in the movement of methionine from vegetative tissues into the seeds (Lee et al. 2008).
In the current study, we aimed at gaining more knowledge about the relative contribution of these two pathways to methionine synthesis and accumulation in Arabidopsis seeds. To achieve this, we measured the levels of methionine and SMM, as well as the expression levels of genes encoding the key enzymes involved in these two pathways (CGS, MMT, and HMTs) during the development of Arabidopsis wild-type plants in its various organs. We also studied how soluble and protein-incorporated levels of other amino acids were changed in these organs during development, and compared them with those of methionine to reveal whether methionine possesses unique patterns of accumulation.
Materials and methods
Plant material and growth conditions
Arabidopsis thaliana (ecotype C-24) plants were used in this study. In all experiments, the plants were first grown on Petri dishes containing 1 % MS (Murashige and Skoog, Duchefa), 1 % sucrose (Baker) and 7 mg/mL plant agar (Duchefa), and then the young seedlings were planted on fertilized soil at 22 ± 3 °C under a 16/8 h light/dark cycle at 100 μmol m−2 s−1 with 50–70 % relative humidity.
All analyses performed in this study were done on the vegetative and reproductive tissues during the Arabidopsis life cycle as follows: imbibed seeds at 0, 1 and 5 days after imbibition (DAI), rosette leaves at 10, 20, 30, 40 and 44 DAI, cauline leaves at 40 and 44 DAI, stems at 40 DAI, flowers at 40 DAI, siliques at 40 and 44 DAI, and seeds at 40, 44 and 55 DAI. Seeds at 55 DAI represent mature dry seeds. All of the samples were collected at 10:00 a.m. and immediately snap-frozen in liquid nitrogen, cryo-lyophilized for 3 days and stored at −80 °C until further analysis.
RNA isolation and quantitative real-time PCR (qRT-PCR) analysis
For the expression level analysis of AtCGS, AtHMT1, AtHMT2, AtHMT3, AtMMT and AtSAMS3, total RNA was extracted from 20 mg of the different tissues. All RNA samples were extracted and analyzed as previously described (Hacham et al. 2013). To normalize the variance among samples, the PP2A-A3 transcript level was used as endogenous control (Czechowski et al. 2005) and the relative gene expression was normalized to the values determined in rosette leaves at 30 DAI. The values presented are the mean of three biological replicates, each with three technical replicates. The primers used for the expression analysis of AtCGS were 5′-GGCGGCTGCTGCTACCTCGTC-3′ (forward) and 5′-CCCTGATCCCCT CCTCGGCC-3′ (reverse); for AtHMT1 5′-CGCAAATTCGCCAAGCTGACG-3′ and 5′-GCTCCAAGGTCACG CCATTTCG-3′; for AtHMT2 5′-AGTGTACCCAAACAGCGGAGAAAGC-3′ and 5′-TGGTT GGTGTTGTGCGGCAGC-3′; for AtHMT3 5′-TTTGCGACGGAGCTACAGCG-3′ and 5′-TCC CACCGATAGACCTTTTGCCA-3′; for AtMMT 5′-TGAACCTCCAGCTGGCAGCAAA-3′ and 5′-GCCAGCTCCTCGGAAGTTGTCG-3′; and for AtSAMS3 5′-GGAGATGCAGGGCTTA CCGGC-3′ and 5′-TGCACAATGACCCGCCTCGC-3′.
Amino acids and S-methylmethionine (SMM) determination using GC–MS and HPLC
For soluble and protein-incorporated amino acid determinations, tissue samples throughout the Arabidopsis life cycle were collected as four distinct biological pools. Following extraction, amino acids were detected using the single ion method (SIM) as previously described (Matityahu et al. 2013). Protein-incorporated amino acids were analyzed following protein hydrolysis (Matityahu et al. 2013). The levels of SMM were detected using HPLC as previously described (Hacham et al. 2002).
Statistical and correlation analyses
Statistical analyses were performed using the GraphPad Prism 5.01 scientific software (http://www.graphpad.com/). In all experiments throughout the manuscript, significance was calculated using the one-way ANOVA test of p < 0.05. Principal component analysis (PCA), and correlations and patterns analyses (Spearman’s rank correlation) of GC–MS data were done using MetaboAnalyst 2.0, a comprehensive tool suite for metabolomic data analysis [http://metaboanalyst.ca/; (Xia et al. 2009, 2012)], following data log10 transformation and pareto scaling (mean centered and divided by the square root of standard deviation of each variable) manipulations.
Results
Soluble methionine contents vary between different tissues during the life cycle of Arabidopsis
To better understand the factors that regulate methionine accumulation in Arabidopsis seeds and the changes in methionine levels during plant development, we collected samples from vegetative and reproductive tissues during the different developmental stages of the Arabidopsis ecotype C-24 life cycle. These include three stages of germination, rosette leaves during five developmental stages up to their senescence, stems, two stages of cauline leaves, flowers, two stages of siliques, and developing seeds in three different stages including dry seeds (total of 17 samples). The samples were analyzed for their soluble methionine content using GC–MS (Table S1).
Measurements of seeds following 48 h of imbibition [0 days after imbibition (DAI)] showed relatively high levels of soluble methionine, suggesting the degradation of seed storage proteins (Fig. 2a). The levels of soluble methionine continued to increase up to 5 DAI, when the cotyledons appeared in the young seedlings. The relatively high level of methionine at this stage is apparently the result of continuous degradation of the seed proteins and/or de novo synthesis of soluble methionine (Fig. 2a). The levels of soluble methionine in the rosette leaves at 10 DAI showed significantly lower levels compared to imbibed seeds. The level of methionine continued to decrease up to 40 DAI, when the reproductive tissues emerged (Fig. 2a). At this stage, stems carrying the cauline leaves were already developed, including flowers with green siliques and immature seeds. While the levels of soluble methionine decreased in the rosette leaves at 40 DAI, the contents of methionine in the reproductive tissues of flowers and developing seeds significantly increased up to 10- and 21-fold, respectively, compared to rosette leaves (Fig. 2a). We also measured the levels of soluble methionine in seeds at three different maturation stages: 40 DAI (immature green seeds), 44 DAI (seeds at their full size at the beginning of the desiccation processes) and 55 DAI (mature dry seeds). The results strongly indicate a significant reduction (up to 66-fold) in soluble methionine contents as the seeds desiccate and mature (Fig. 2a).
The levels of methionine and S-methylmethionine (SMM) in various tissues during the life cycle of Arabidopsis. a Soluble methionine contents. b Protein-incorporated methionine contents. c SMM contents. Soluble and protein-incorporated methionine contents were determined via GC–MS analysis, while SMM contents were determined via HPLC analysis. Data shown are mean ± SE of four independent biological replicates. Significance was calculated using the two-way ANOVA test of p < 0.05 and identified by different letters
Total methionine levels in different tissues during Arabidopsis development
The levels of soluble methionine and of other soluble amino acids are relatively low compared to their total levels, which include protein-incorporated amino acids (Hacham et al. 2008). Thus, to have a broader view of the level of methionine, we also analyzed total methionine levels following protein hydrolysis in all tissues examined (Table S2).
Total methionine contents were about 10- to 100-fold higher than the soluble amino acid (Fig. 2a, b). Generally, total methionine showed different patterns of change in the various tissues during the Arabidopsis life cycle compared to those observed for soluble methionine. The highest levels were found in dry seeds at the beginning of imbibition and in mature dry seeds (Fig. 2b), most likely due to the accumulation of high levels of total methionine within the seed storage proteins. Following imbibition, a consistent moderate reduction in total methionine contents was observed up to 10 DAI, indicating that storage proteins were being degraded, thus explaining the higher levels of soluble methionine found at these stages (Fig. 2a). In the rosette leaves, there was a slight reduction in total methionine content, from 10 to 40 DAI. Between 40 and 44 DAI, when the reproductive tissues are fully developed, the total methionine content was significantly reduced in the rosette leaves. At that stage, the levels of methionine in the developing seeds significantly increased (Fig. 2a).
SMM contents exhibit inverted accumulation patterns from soluble methionine
To assess the role of SMM in methionine synthesis in seeds, the accumulation patterns of SMM were determined during the Arabidopsis life cycle using HPLC (Table S3). The results show a significant increase in SMM levels in the rosette leaves from 10 DAI, when the leaves were formed, up to 44 DAI, when the reproductive organs were fully mature and contained the developing seeds (Fig. 2c). The largest increase occurred in the rosette leaves between 30 and 40 DAI, indicating that SMM was accumulated massively in these tissues at this stage, apparently before being transferred to the reproductive organs (Fig. 2c). Supporting this transfer are the relatively high levels of SMM found in the stems and flowers (Fig. 2c). Higher levels of SMM (up to fourfold than in leaves 10 DAI) were also detected in the cauline leaves. A significant reduction was observed in the siliques and the developing seeds compared to the rosette leaves during the reproductive stage, implying that in these reproductive tissues SMM were converted back to soluble methionine to supply methionine toward its incorporation into the seed storage proteins (Fig. 2c). Indeed, mature dry seeds (55 DAI) had the lowest SMM contents during the Arabidopsis life cycle (Fig. 2c). It seems that the conversion from SMM to methionine occurred less in the flowers, as the SMM levels were found to be relatively high in these tissues.
Transcript levels of key genes involved in methionine synthesis through the aspartate family pathway and the SMM cycle
To gain a broader picture of the synthesis and accumulation patterns of methionine and SMM, we conducted a quantitative real-time PCR (qRT-PCR) analysis to measure the transcript levels of AtCGS that operate in the aspartate family pathway, and of genes encoding enzymes that operate in the SMM cycle. There is only one gene in Arabidopsis for CGS and MMT and three genes encoding HMTs.
CGS (At3g01120) was expressed in all tissues examined. The highest levels were measured in the rosette leaves, from 5 DAI up to 40 DAI (Fig. 3a). However, the expression level decreased in these tissues at 44 DAI, displaying similar expression levels to those detected in the reproductive tissues. The elevation in levels of CGS at the early stages of seed germination from 1 DAI up to 5 DAI suggests that methionine is synthesized de novo in addition to methionine that is released from the degradation of seed storage proteins. The newly synthesized methionine is required to support what is needed for soluble methionine and its associated derivatives (such as ethylene, polyamines, glucosinolates) that are apparently required less in seeds, where most of the soluble methionine is incorporated into the seed storage proteins. Notably, the observed expression patterns of CGS are not compatible with the levels of methionine (Fig. 2a, b), suggesting that additional regulatory metabolic factors are involved in methionine accumulation. These factors may include methionine catabolic enzymes, which lead to the synthesis of SAM and its associated metabolites, incorporation into proteins and/or conversion into SMM (Amir 2010; Roje 2006).
Transcript levels of key genes involved in methionine synthesis through the aspartate family pathway and the SMM cycle. Transcript levels of: a AtCGS; b AtMMT; c AtHMT1; d AtHMT2; e AtHMT3 and f AtSAMS3. The expression levels of all investigated genes were determined following qRT-PCR analysis and normalized to the PP2A gene and the relative gene expression in rosette leaves 30 DAI. Data shown are mean ± SE of three independent biological replicates. Significance was calculated using the two-way ANOVA test of p < 0.05 and identified by different letters
Next, we examined the expression levels of the key genes participating in the SMM cycle. First, we measured the expression patterns of MMT (At5g49810) that catalyzes the formation of SMM from methionine and SAM. The results demonstrate that this gene is expressed in all tissues examined at relatively stable expression levels except in seeds at 0 DAI, which showed fourfold lower expression levels than those detected in the rosette leaves at 40 DAI (Fig. 3b). This implies that there is lower MMT activity when the seed storage proteins are degraded, supporting the requirements of methionine at this stage. The highest expression levels were detected in the rosette leaves at 40 and 44 DAI (Fig. 3b), when the highest SMM contents were observed (Fig. 2c).
MMT is also expressed in seeds (Fig. 3b). To determine if the SMM in seeds is synthesized de novo from MMT activity or transported mainly from other tissues, we examined the expression level of SAMS3 (At3g17390), which represents the expression of SAMSs in seeds (according to http://bar.utoronto.ca). This enzyme produces SAM, the second substrate of MMT. The analysis showed that the expression levels were significantly reduced during the late two stages of seed development (Fig. 3f), suggesting a low level of SAM production and thus lower activity of MMT in seeds at these stages.
Expression analysis of the three HMT genes revealed that their expression levels were relatively high in the late stage of developing seeds, especially during the desiccation and maturation stage, compared to the first two stages of seed development (Fig. 3c–e). Among these three isoforms, the expression level of HMT3 (At3g22740) decreased significantly from 0 DAI up to 20 DAI and then increased by about 7.5-fold in leaves at 44 DAI (Fig. 3e). Its expression level increased greatly in the maturing seeds (about 3.3- and 26-fold compared to rosette leaves at 44 and 20 DAI, respectively) (Fig. 3e). These findings imply that this isoform may play a more functional role in the conversion of SMM into methionine at the late stages of seed development. Expression analysis of HMT1 (At3g25900) revealed that its levels decreased in the rosette leaves from5 DAI up to 44 DAI, reaching similar levels to those detected in the cauline leaves and the stems. However, its expression level increased greatly in the flowers, siliques and developing seeds (up to fourfold) compared to the levels observed in the rosette leaves at 40 DAI (Fig. 3c). Its expression levels are quite similar to the levels of soluble methionine (compare Fig. 3c to Fig. 2a). Finally, the expression level of HMT2 (At3g63250) showed only minor fluctuations during the whole life cycle compared to HMT1 and HMT3 (Fig. 3d) and were not consistent with those observed for soluble methionine.
The accumulation patterns of other amino acids during the Arabidopsis life cycle
Soluble and protein-incorporated amino acids levels were measured to assess the similarity of their accumulation patterns with those of methionine and to reveal whether methionine exhibits unique independent patterns (Tables S1 and S2). When measuring total soluble amino acid contents, we could clearly observe higher contents in the rosette leaves at 5 DAI, flowers at 40 DAI, siliques at 44 DAI and developing seeds at 40 DAI (Fig. 4a), which also showed distinct separation compared to all other tissues examined when plotted on principal component analysis (PCA) (Fig. S1).
GC–MS analysis of amino acids in various tissues during the life cycle of Arabidopsis. a Total soluble amino acids; b total protein-incorporated amino acids. All soluble and protein-incorporated amino acid contents were determined via GC–MS analysis. Data shown are mean ± SE of five independent biological replicates
Next, we were interested in investigating the individual patterns of the amino acids (Fig. 5). The results point to two global trends. (1) The levels of all soluble amino acids were dramatically reduced during seed development from 40 DAI until full maturation and desiccation at 55 DAI, most likely due to the massive incorporation of soluble amino acids into seed storage proteins that occurs during seed development and maturation. (2) High levels of total soluble amino acids were detected in rosette leaves at 5 DAI and flowers at 40 DAI (Figs. 4a, 5). Notably, only threonine and serine showed different profiles, since their levels remained relatively stable in the vegetative tissues even when the reproductive stage had already begun, and flowers, siliques and developing seeds operate as strong sink organs (Fig. 5).
Next, we performed the same manipulations on the data obtained from the GC–MS analysis of the protein-incorporated amino acids, as we did for the soluble amino acid data, allowing us to compare the changes during the Arabidopsis life cycle between soluble and protein-incorporated amino acids. Analysis of the total protein-incorporated amino acids showed a relatively consistent high level in imbibed seeds at 0 DAI up to 20 DAI for rosette leaves. From this point up to 44 DAI, a gradual decline was observed (Fig. 4b). The cauline leaves and flowers exhibited relatively high levels, while the lowest levels were observed in the stems and siliques (Fig. 4b). Finally, a consistent increase in total protein-incorporated amino acids content was observed during seed development up to full maturation at 55 DAI (Fig. 4b), implying that amino acids were incorporated into the seed storage proteins. All these tissue samples also showed clear separation according to PCA (Fig. S2).
The accumulation patterns of individual protein-incorporated amino acids are shown in Fig. 6. The results reveal several trends. (1) Most of the amino acids accumulate highly in the rosette leaves and flowers. (2) The levels of most amino acids are reduced in the rosette leaves when the reproductive organs appear at 40 and 44 DAI. (3) The contents of all amino acids increase during seed development as expected due to massive synthesis of seed storage proteins at these stages.
Do other amino acids exhibit an accumulation patterns similar to methionine?
The data described above suggest that methionine is transferred from the mature leaves toward the reproductive tissues in the form of SMM. To assess whether other amino acids display similar patterns, or whether this behavior is unique to methionine during development, we performed correlation tests. These tests were performed between methionine levels and the levels of the other soluble and protein-incorporated amino acids during the entire developmental stage using the statistical tools embedded in the MetaboAnalyst 2.0 metabolomic data analysis software (see “Materials and methods”). The seven highest correlation values were found to be between soluble methionine and the levels of the branched-chain amino acids, leucine, valine and isoleucine, the aromatic amino acids phenylalanine, tryptophan and tyrosine, and asparagine (Fig. 7a). Apart from serine, threonine and homoserine that did not exhibit significant correlation with methionine, all the other amino acids showed significant correlation, although their values of significance were relatively low during the developmental processes.
Correlation analyses between amino acids and methionine. a Soluble amino acids; b protein-incorporated amino acids. The analyses were performed according to the Spearman’s rank correlation method (see “Materials and methods”). Calculated correlation values appear within bars; asterisks represent statistical significance of P value <0.001
The levels of soluble amino acids are approximately 25-fold lower than those incorporated in the proteins. Thus, the correlation values between protein-incorporated methionine and other incorporated amino acids were measured as well. The tests revealed that phenylalanine and tyrosine showed the highest correlation values to methionine (Fig. 7b), as observed in their soluble forms (Fig. 7a). Interestingly, serine also showed high correlation values to methionine, even though its soluble form exhibited the lowest correlation values (Fig. 7a). Lysine, glutamate, proline and threonine also showed relatively high correlation values to methionine (Fig. 7b), while the branched-chain amino acids of leucine, isoleucine and valine showed lower levels (Fig. 7b), even though they were highly correlated to methionine in their soluble form (Fig. 7a).
Discussion
Accumulation patterns of methionine and SMM during the Arabidopsis life cycle
The current study supports the assumption that in addition to the de novo synthesis of methionine in seeds (Cohen et al. 2014; Hanafy et al. 2013; Matityahu et al. 2013; Song et al. 2013), methionine is also transferred from vegetative tissues in the form of SMM and contributes to methionine accumulation in seeds. The assumption is based on several observations. (1) The levels of soluble and protein-incorporated methionine decreased in the rosette leaves at 40 and 44 DAI when the reproductive tissues of flowers and developing seeds were developed, whereas the contents of methionine increased significantly by 10- and 21-fold, respectively, in these organs compared to the rosette leaves (Fig. 2a, b). (2) The level of SMM is significantly higher in the rosette leaves (from 30 DAI up to 44 DAI) and stems and flowers, indicating that SMM is formed massively in the rosette leaves at these stages and then transferred into the reproductive organs through the stems (Fig. 2c). (3) The lower levels of SMM in the siliques and developing seeds compared to the levels detected in flowers (about 2.75-fold lower level; Fig. 2c) imply that SMM is converted back to soluble methionine in these tissues and is available for the synthesis of the seed storage proteins. Accordingly, mature dry seeds (55 DAI) exhibited the lowest SMM contents during the Arabidopsis life cycle (Fig. 2c). (4) The fact that SMM showed inverted accumulation patterns than those of methionine strengthens the notion that methionine synthesized in the leaves is converted to SMM mainly during the senescence of these leaves. (5) The expression levels of MMT are slightly, but significantly higher in leaves at 40 and 44 DAI, indicating that the production of SMM is elevated at these developmental stages. This observation, together with the reduction in the soluble and total methionine contents at these stages, strengthens the assumption that SMM is formed in the mature rosette leaves before being transferred into the developing reproductive tissues and that MMT plays a prominent role in converting methionine to SMM. (6) The transcript pattern of HMT1, which is similar to that of soluble methionine and the higher expression level of HMT3 in the late stages of seed development, supports the assumption that SMM is converted to methionine in the developing seeds through the activity of these isoforms. (7) The lower expression level of CGS in the reproductive tissues compared to the rosette leaves and the higher levels of methionine found in the reproductive tissues suggest that methionine is transferred to these tissues and synthesized less in situ. (8) The expression levels of the three isoforms of HMT are significantly increased at the last stage of seed development, suggesting that re-conversion of SMM back to soluble methionine occurs at these stages. Higher levels of HMT were also observed in the last stages of seed development of Medicago truncatula, suggesting that it contributes to methionine accumulation at this stage of development (Gallardo et al. 2007).
Our observations stand in agreement with previous studies (Bourgis et al. 1999; Kocsis et al. 2003; Lee et al. 2008; Ranocha et al. 2001), suggesting that the SMM cycle is an important contributor to methionine synthesis in seeds, even though the tissues contributing to SMM production have not yet been discovered. Due to the lower levels of soluble and protein-incorporated methionine at 40 and 44 DAI and the higher levels of methionine in the reproductive tissues, we assume that the rosette leaves play an important role in transporting SMM toward the seeds, as suggested previously by Bourgis et al. (1999). However, since relatively high levels of SMM were also detected in the flowers and stems, we cannot exclude the possibility that these tissues are also significant contributors to SMM production. This proposal was raised previously by Lee et al. (2008), suggesting that SMM was produced mainly at the top of the flower stalk (cauline leaves, stems, flowers, silique hulls and seeds) and less in the rosette leaves. However, the higher SMM contents in the flowers and stems might also result from SMM transfer through these tissues rather than de novo synthesis of SMM. Thus, the precise functional role and contribution of the SMM cycle to methionine synthesis in seeds requires additional studies (Kocsis et al. 2003; Hacham et al. 2008).
Roles of the SMM cycle
Our gene expression analyses indicated that the enzymes constituting the SMM cycle (HMTs and MMT) were expressed in all of the tissues examined. These findings are in agreement with a previous study, which showed that Arabidopsis leaves, roots and developing seeds can metabolize 35S-methionine to 35S-SMM and vice versa (Ranocha et al. 2001). Principally, the tandem action of MMT and HMTs, together with those of SAMS and S-ADENOSYLHOMOCYSTEINE HYDROLASE (ADOHCY HYDROLASE), set up a futile cycle in which methionine is converted to SMM and vice versa (Mudd and Datko 1990). In each turn of the cycle, a molecule of ATP is hydrolyzed into adenosine, pyrophosphate and phosphate (Mudd and Datko 1990). Thus, it might be that methionine formed from SMM will be re-converted back to SMM by the activity of MMT in these tissues, leading to a loss of energy. The observation that MMT and HMTs are detected in these tissues suggests that the contribution of the SMM cycle to methionine synthesis in seeds is only one of this cycle’s roles in the plant’s metabolism. Indeed, several previous studies have proposed other roles for the SMM cycle. Among them are a short-term control of SAM levels (Ranocha et al. 2001), stabilization of the free methionine pool (Mudd and Datko 1990), donation of methyl groups to other plant metabolites by SMM (Giovanelli 1987) and repair of SAM suffering from chemical damage reactions under physiological conditions (Bradbury et al. 2014).
The observation that a higher expression level of HMT3 was found in the rosette leaves 40 and 44 DAI simultaneously with an elevated expression of MMT suggests other roles of this cycle at this stage. Also, the expression levels of HMT2 that remain relatively stable through all of the developmental stages imply that this isoform may have additional yet unknown roles other than the contribution to methionine synthesis in seeds.
However, most probably other factors are involved in the control of this cycle in addition to the expression levels of the genes encoding to enzymes within the cycle. One of these factors could be the availability and the contents of SAM, the second substrate for MMT. An indication of this possibility comes from the observation that the expression level of SAMS3 was significantly reduced in the late stages of seed development. This strongly implies that the level of SAM decreased at this stage and probably also decreased the activity of MMT at these stages in seeds. Low expression levels of this gene were also detected in the endosperm and seed coat of M. truncatula seeds during seed filling (Gallardo et al. 2007), suggesting that the SMM pool within seeds is derived mainly from the phloem rather than inter-production in seeds. Additional studies are required to reveal the different roles of this cycle in metabolism and its important functions in plants.
Accumulation patterns of other amino acids during plant development
Analyses of the soluble and protein-incorporated amino acids gave us the opportunity to define whether methionine has unique accumulation patterns during the development, or whether its patterns are like most other amino acids. Our correlation analyses revealed that patterns of soluble methionine were similar to those exhibited by most other soluble amino acids, with the exception of threonine and serine. These two amino acids are associated with the metabolism of methionine via the aspartate family pathway (Fig. 1). Threonine synthesis competes with that of methionine (Ravanel et al. 1998; Curien et al. 2003), and serine is the donor for cysteine synthesis, which is the thiol precursor required for methionine synthesis (Nikiforova et al. 2005). Their accumulation pattern suggests that these amino acids have their own patterns that are not related to methionine and to the other amino acids.
Notably, the two groups of amino acids showing the highest correlation values to soluble methionine are the branched-chain and aromatic amino acids, which are synthesized within the chloroplast together with methionine [reviewed by (Maeda and Dudareva 2012)]. A recent study had also showed high correlations between methionine and the levels of these amino acids in transgenic Arabidopsis seeds expressing a feedback-insensitive form of AtCGS (Cohen et al. 2014). The branched-chain amino acids are synthesized primarily through threonine, but it was demonstrated previously that they could also be synthesized from methionine through the activity of METHIONINE γ-LYASE (MGL), particularly during times of stress (Joshi and Jander 2009). Since, in the current study, the samples were taken from non-stressed plants, it would be interesting in the future to determine if a metabolic link exists between their accumulation patterns to that of methionine, as well as with the aromatic amino acids.
Previous genetic studies showed a relationship between methionine metabolism and other soluble amino acids. Arabidopsis thaliana antisense lines with repressed CGS activity exhibited an overall increase in their total soluble amino acids content, although little change in the content of soluble methionine and SMM were reported (Kim et al. 2002; Kim and Leustek 2000). Contrary to these results, several recent studies strongly imply a positive correlation between AtCGS and the accumulation of methionine and other amino acids since transgenic soybean, tobacco and Arabidopsis expressing the AtCGS showed significant elevations in soluble methionine and significantly higher contents of other soluble amino acids (Matityahu et al. 2013; Song et al. 2013; Hacham et al. 2008; Cohen et al. 2014). In addition, Arabidopsis plants that suppress the expression of SAMS exhibited much higher levels of total soluble amino acids, and methionine, arginine, proline and lysine were the most increased amino acids (Kim et al. 2002). These results suggest that changes in the expression of genes that are associated with methionine metabolism might affect the levels of other amino acids. Generally, little is known about the global regulation of amino acids in plants, and it appears that the disruption of a single amino acid such as methionine can lead to pleiotropic effects on the homeostasis of all amino acids (Guyer et al. 1995). Hence, additional studies are required to determine if the accumulation patterns of soluble amino acids during plant development are related to methionine or one of the other amino acids. Alternatively, it is possible that other transcriptional and/or developmental factors play prominent regulatory roles in determining the levels of amino acids, since most exhibited similar accumulation patterns during the Arabidopsis life cycle. Still, a master regulatory factor involved in these processes, such as the yeast GENERAL CONTROL PROTEIN 4 (GCN4), which is responsible for the activation of more than 30 genes required for amino acids biosynthesis, has not yet been detected in plants (Guyer et al. 1995), thereby addressing the necessity for additional research to define the factors regulating the levels of amino acids during plant development.
The accumulation patterns of protein-incorporated methionine in the various tissues are also significantly correlated with other protein-incorporated amino acids. The only amino acids showing low correlation values with methionine are glycine and aspartate. In general, an examination of the patterns of protein-incorporated amino acids indicated that most of the amino acids produced in the rosette leaves were incorporated into the vegetative storage proteins and other proteins from 5 to 30 DAI. However, at later stages from 40 DAI when the reproductive tissues start their development, the vegetative proteins are degraded to release amino acids for the synthesis of these tissues. These findings are in agreement with other studies suggesting that vegetative protein degradation is the major process responsible for the dramatic increases in the remobilization of nitrogen into the sink organs during leaf senescence (Soudry et al. 2005). It was previously demonstrated that these senescence-associated processes are accompanied by a transition from anabolic to catabolic processes (Soudry et al. 2005), and that the SENESCENCE-ASSOCIATED GENES (SAGs) are the key proteins involved in the regulation of these processes (Gepstein et al. 2003). SAGs encode hydrolytic enzymes responsible for the intensive breakdown of the various macromolecules such as proteins, nucleic acids, polysaccharides and lipids (Gepstein et al. 2003), as well as for nitrogen recycling, which is the most recycled nutrient during senescence (Tegeder and Rentsch 2010). It was shown that in the leaves of Arabidopsis at these late stages, glutamate, glutamine, aspartate and asparagine constitute 60–64 % of the total free amino acids that are transported in the vascular tissues, and that glutamine and asparagine are among the most modified amino acids being transported (Lam et al. 1996). Based on the results of our study, we suggest that methionine is also subjected to such modifications, being converted to SMM before being translocated into the reproductive tissues. Since methionine is a hydrophobic amino acid and SMM is a polar compound, we assume that this modification enables methionine to be transported more effectively in the phloem flux. The transportation of amino acids from leaves to seeds is prominent not only for the synthesis of seed storage proteins, but also for many other metabolic processes. Amino acids play critical metabolic roles due to their conversion to many other essential metabolites (e.g., Tzin and Galili 2010). Some of these metabolites are being used by the tricarboxylic acid (TCA) cycle as an energy source and/or as the precursors for the synthesis of many secondary metabolites (e.g., phenylpropanoids, alkaloids, glucosinolates), act as signaling molecules, (e.g., hormones), structural components (e.g., lignin) and defense substances (e.g., glucosinolates, anthocyanins, nicotine) (Maeda and Dudareva 2012; Tzin and Galili 2010). Therefore, the transportation of amino acids from leaves to seeds might play additional important roles in seed biology.
All in all, our findings strengthen the hypothesis that SMM is an important contributor to methionine synthesis in seeds and that it is produced mainly in the rosette leaves. Since HMT3 has a higher expression level in seeds and may be the key player in the conversion of SMM to methionine in the seeds, we assume that its overexpression in a seed-specific manner might enhance the conversion of SMM in seeds and lead to higher flux of SMM toward the seeds, resulting in higher levels of methionine in the transgenic seeds. We also found that MMT is expressed in seeds; thus we assume that even greater amounts of methionine might accumulate in the transgenic seeds if the HMT3 enzyme is expressed in the background of a seed-specific RNAi::MMT transgenic line. Future research using Arabidopsis and crop plants will determine which of these approaches is the most feasible in an effort to increase methionine contents in seeds. This is an important biotechnology goal, since the low contents of methionine limit the nutritional quality of seeds and additional approaches should be tested to increase its levels (Amir et al. 2012; Galili and Amir 2013).
References
Amir R (2010) Current understanding of the factors regulating methionine content in vegetative tissues of higher plants. Amino Acids 39(4):917–931. doi:10.1007/s00726-010-0482-x
Amir R, Han T, Ma F (2012) Bioengineering approaches to improve the nutritional values of seeds by increasing their methionine content. Mol Breed 29:915–924
Bartlem D, Lambein I, Okamoto T, Itaya A, Uda Y, Kijima F, Tamaki Y, Nambara E, Naito S (2000) Mutation in the threonine synthase gene results in an over-accumulation of soluble methionine in Arabidopsis. Plant Physiol 123(1):101–110
Bourgis F, Roje S, Nuccio ML, Fisher DB, Tarczynski MC, Li C, Herschbach C, Rennenberg H, Pimenta MJ, Shen TL, Gage DA, Hanson AD (1999) S-methylmethionine plays a major role in phloem sulfur transport and is synthesized by a novel type of methyltransferase. Plant Cell 11(8):1485–1498
Bradbury LM, Ziemak MJ, Elbadawi-Sidhu M, Fiehn O, Hanson AD (2014) Plant-driven repurposing of the ancient S-adenosylmethionine repair enzyme homocysteine S-methyltransferase. Biochem J. doi:10.1042/BJ20140753
Cohen H, Israeli H, Matityahu I, Amir R (2014) Seed-specific expression of a feedback-insensitive form of cystathionine-gamma-synthase in arabidopsis stimulates metabolic and transcriptomic responses associated with desiccation stress. Plant Physiol 163(3):1575–1592. doi:10.1104/pp.114.246058
Curien G, Ravanel S, Dumas R (2003) A kinetic model of the branch-point between the methionine and threonine biosynthesis pathways in Arabidopsis thaliana. Eur J Biochem 270(23):4615–4627
Czechowski T, Stitt M, Altmann T, Udvardi MK, Scheible WR (2005) Genome-wide identification and testing of superior reference genes for transcript normalization in Arabidopsis. Plant Physiol 139(1):5–17. doi:10.1104/pp.105.063743
Galili G, Amir R (2013) Fortifying plants with the essential amino acids lysine and methionine to improve nutritional quality. Plant Biotechnol J 11(2):211–222. doi:10.1111/pbi.12025
Gallardo K, Firnhaber C, Zuber H, Hericher D, Belghazi M, Henry C, Kuster H, Thompson R (2007) A combined proteome and transcriptome analysis of developing Medicago truncatula seeds: evidence for metabolic specialization of maternal and filial tissues. Mol Cell Proteomics 6(12):2165–2179. doi:10.1074/mcp.M700171-MCP200
Gepstein S, Sabehi G, Carp MJ, Hajouj T, Nesher MF, Yariv I, Dor C, Bassani M (2003) Large-scale identification of leaf senescence-associated genes. Plant J 36(5):629–642
Giovanelli JG (1987) Sulfur amino acids in plants: an overview. Methods Enzymol 143:419–428
Guyer D, Patton D, Ward E (1995) Evidence for cross-pathway regulation of metabolic gene expression in plants. Proc Natl Acad Sci USA 92(11):4997–5000
Hacham Y, Avraham T, Amir R (2002) The N-terminal region of Arabidopsis cystathionine gamma-synthase plays an important regulatory role in methionine metabolism. Plant Physiol 128(2):454–462. doi:10.1104/pp.010819
Hacham Y, Matityahu I, Schuster G, Amir R (2008) Overexpression of mutated forms of aspartate kinase and cystathionine gamma-synthase in tobacco leaves resulted in the high accumulation of methionine and threonine. Plant J 54(2):260–271. doi:10.1111/j.1365-313X.2008.03415.x
Hacham Y, Matityahu I, Amir R (2013) Light and sucrose up-regulate the expression level of Arabidopsis cystathionine gamma-synthase, the key enzyme of methionine biosynthesis pathway. Amino Acids 45(5):1179–1190. doi:10.1007/s00726-013-1576-z
Hanafy MS, Rahman SM, Nakamoto Y, Fujiwara T, Naito S, Wakasa K, Ishimoto M (2013) Differential response of methionine metabolism in two grain legumes, soybean and azuki bean, expressing a mutated form of Arabidopsis cystathionine gamma-synthase. J Plant Physiol 170(3):338–345. doi:10.1016/j.jplph.2012.10.018
Hesse H, Hoefgen R (2003) Molecular aspects of methionine biosynthesis. Trends Plant Sci 8(6):259–262 S1360138503001079
Inaba K, Fujiwara T, Hayashi H, Chino M, Komeda Y, Naito S (1994) Isolation of an Arabidopsis thaliana mutant, mto1, that overaccumulates soluble methionine. Plant Physiol 104(3):881–887 104/3/881 [pii]
Jander G, Joshi V (2010) Recent progress in deciphering the biosynthesis of aspartate-derived amino acids in plants. Mol Plant 3(1):54–65. doi:10.1093/mp/ssp104
Joshi V, Jander G (2009) Arabidopsis methionine gamma-lyase is regulated according to isoleucine biosynthesis needs but plays a subordinate role to threonine deaminase. Plant Physiol 151(1):367–378. doi:10.1104/pp.109.138651
Kim J, Leustek T (2000) Repression of cystathionine γ-synthase in Arabidopsis thaliana produces partial methionine auxotrophy and developmental abnormalities. Plant Sci 151:9–18. doi:10.1016/S0168-9452(99)00188-0
Kim J, Lee M, Chalam R, Martin MN, Leustek T, Boerjan W (2002) Constitutive overexpression of cystathionine gamma-synthase in Arabidopsis leads to accumulation of soluble methionine and S-methylmethionine. Plant Physiol 128(1):95–107
Kocsis MG, Ranocha P, Gage DA, Simon ES, Rhodes D, Peel GJ, Mellema S, Saito K, Awazuhara M, Li C, Meeley RB, Tarczynski MC, Wagner C, Hanson AD (2003) Insertional inactivation of the methionine s-methyltransferase gene eliminates the s-methylmethionine cycle and increases the methylation ratio. Plant Physiol 131(4):1808–1815. doi:10.1104/pp.102.018846
Lam HM, Coschigano KT, Oliveira IC, Melo-Oliveira R, Coruzzi GM (1996) The molecular-genetics of nitrogen assimilation into amino acids in higher plants. Ann Rev Plant Physiol Plant Mol Biol 47:569–593. doi:10.1146/annurev.arplant.47.1.569
Lee M, Huang T, Toro-Ramos T, Fraga M, Last RL, Jander G (2008) Reduced activity of Arabidopsis thaliana HMT2, a methionine biosynthetic enzyme, increases seed methionine content. Plant J 54(2):310–320. doi:10.1111/j.1365-313X.2008.03419.x
Maeda H, Dudareva N (2012) The shikimate pathway and aromatic amino acid biosynthesis in plants. Annu Rev Plant Biol 63:73–105. doi:10.1146/annurev-arplant-042811-105439
Matityahu I, Godo I, Hacham Y, Amir R (2013) Tobacco seeds expressing feedback-insensitive cystathionine gamma-synthase exhibit elevated content of methionine and altered primary metabolic profile. BMC Plant Biol 13:206. doi:10.1186/1471-2229-13-206
Mudd H, Datko H (1990) The S-methylmethionine cycle in Lemna paucicostata. Plant Physiol 93:623–630
Nikiforova VJ, Kopka J, Tolstikov V, Fiehn O, Hopkins L, Hawkesford MJ, Hesse H, Hoefgen R (2005) Systems rebalancing of metabolism in response to sulfur deprivation, as revealed by metabolome analysis of Arabidopsis plants. Plant Physiol 138(1):304–318
Ranocha P, McNeil SD, Ziemak MJ, Li C, Tarczynski MC, Hanson AD (2001) The S-methylmethionine cycle in angiosperms: ubiquity, antiquity and activity. Plant J 25(5):575–584 tpj988 [pii]
Ravanel S, Gakiere B, Job D, Douce R (1998) The specific features of methionine biosynthesis and metabolism in plants. Proc Natl Acad Sci USA 95(13):7805–7812
Roje S (2006) S-Adenosyl-l-methionine: beyond the universal methyl group donor. Phytochemistry 67(15):1686–1698. doi:10.1016/j.phytochem.2006.04.019
Song S, Hou W, Godo I, Wu C, Yu Y, Matityahu I, Hacham Y, Sun S, Han T, Amir R (2013) Soybean seeds expressing feedback-insensitive cystathionine γ-synthase exhibit a higher content of methionine. J Exp Bot 64(7):1917–1926. doi:10.1093/jxb/ert053
Soudry E, Ulitzur S, Gepstein S (2005) Accumulation and remobilization of amino acids during senescence of detached and attached leaves: in planta analysis of tryptophan levels by recombinant luminescent bacteria. J Exp Bot 56(412):695–702. doi:10.1093/jxb/eri054
Tan Q, Zhang L, Grant J, Cooper P, Tegeder M (2010) Increased phloem transport of S-methylmethionine positively affects sulfur and nitrogen metabolism and seed development in pea plants. Plant Physiol 154(4):1886–1896. doi:10.1104/pp.110.166389
Tegeder M, Rentsch D (2010) Uptake and partitioning of amino acids and peptides. Mol Plant 3(6):997–1011. doi:10.1093/mp/ssq047
Tzin V, Galili G (2010) New insights into the shikimate and aromatic amino acids biosynthesis pathways in plants. Mol Plant 3(6):956–972. doi:10.1093/mp/ssq048
Xia J, Psychogios N, Young N, Wishart DS (2009) MetaboAnalyst: a web server for metabolomic data analysis and interpretation. Nucleic Acids Res 37(Web Server issue):W652–W660. doi:10.1093/nar/gkp356
Xia J, Mandal R, Sinelnikov IV, Broadhurst D, Wishart DS (2012) MetaboAnalyst 2.0–a comprehensive server for metabolomic data analysis. Nucleic Acids Res 40(Web Server issue):W127–W133. doi:10.1093/nar/gks374
Acknowledgments
We would like to thank Dr. Gidi Baum for his critical reading and valuable comments. This work was supported by the Israel Science Foundation (Grant 231-09).
Conflict of interest
The authors declare that they have no conflicts of interest.
Author information
Authors and Affiliations
Corresponding author
Additional information
A. Frank and H. Cohen contributed equally to this study.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Frank, A., Cohen, H., Hoffman, D. et al. Methionine and S-methylmethionine exhibit temporal and spatial accumulation patterns during the Arabidopsis life cycle. Amino Acids 47, 497–510 (2015). https://doi.org/10.1007/s00726-014-1881-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00726-014-1881-1