Abstract
Feathers are the most complex skin appendages of vertebrates. Mature feathers consist of interconnected dead keratinocytes that are filled with heavily cross-linked proteins. Although the molecular architecture determines essential functions of feathers, only few feather proteins have been characterized with regard to their amino acid sequences and evolution. Here, we identify Epidermal Differentiation protein containing DPCC Motifs (EDDM) as a cysteine-rich protein that has co-evolved with other feather proteins. The EDDM gene is located within the avian epidermal differentiation complex (EDC), a cluster of genes that has originated and diversified in amniotes. EDDM shares the exon-intron organization with EDC genes of other amniotes, including humans, and a gene encoding an EDDM-like protein is present in crocodilians, suggesting that avian EDDM arose by sequence modification of an epidermal differentiation gene present in a common ancestor of archosaurs. The EDDM protein contains multiple sequence repeats and a higher number of cysteine residues than any other protein encoded in the EDC. Immunohistochemical analysis of chicken skin and skin appendages showed expression of EDDM in barb and barbules of feathers as well as in the subperiderm on embryonic scutate scales. These results suggest that the diversification and differential expression of EDDM, besides other EDC genes, was instrumental in facilitating the evolution of the most complex molecular architecture of feathers.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Feathers are the characteristic for birds and more complex than any other skin appendages of vertebrates. They are also diverse in shape in order to accomplish different functions, including thermal insulation, communication, and flight. Pennaceous feathers, i.e., the main type of feathers in adult birds, are comprised of a shaft (rachis) and two levels of branches that are called barbs and barbules. This hierarchical organization is established in a process of differential growth, cell death, and cornification of epithelial cells, as described in extensive reviews (Chuong 1993; Prum and Dyck 2003; Prum 2005; Maderson et al. 2009; Chen et al. 2015; Alibardi 2017).
The evolutionary origin of feathers has been traced back to extinct members of the clade Dinosauria. The only extant representatives of Dinosauria are the birds and their closest modern relatives are crocodilians. Various models for the modification of ancestral development and differentiation programs of skin cells have been proposed to explain the evolution of feathers (Yu et al. 2002; Ng et al. 2015; Wu et al. 2015; Mlitz et al. 2014). Importantly, feather barbs and barbules and the subperiderm, a layer of the embryonic epidermis present not only in birds but also in crocodilians, share the same differentiation markers. Together with topological considerations, this shared gene expression signature has led to the hypothesis that the evolution of feather growth and regeneration depended on the modification of an embryonic development program of archosaurian skin for new purposes (Sawyer and Knapp 2003; Sawyer et al. 2005; Strasser et al. 2015; Alibardi et al. 2016), thus representing an example of evolutionary co-option (True and Carroll 2002; Prum 2005).
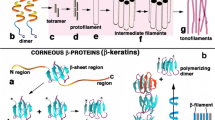
The mechanical stability and stress resistance of feathers are facilitated by their microarchitecture and biochemical composition. Besides a minor content of lipids, proteins constitute more than 90% of the feather mass (Bolliger and Varga 1961). The major protein components of feathers are Corneous Beta Proteins (CBPs), traditionally referred to as beta-keratins (Gregg and Rogers 1986; Fraser and Parry 2008; Greenwold and Sawyer 2011, 2013; Greenwold et al. 2014; Alibardi 2017; Holthaus et al. 2018a). CBPs are not related to keratin intermediate filament proteins (Eckhart and Ehrlich 2018; Holthaus et al. 2018a) but to proteins encoded in the Epidermal Differentiation Complex (EDC), a cluster of genes for protein components of cornifying keratinocytes of amniotes (Strasser et al. 2014). Many CBP genes are located in a sub-cluster of the EDC (Strasser et al. 2014). Recent reports have, however, shown that, besides CBPs, other EDC-encoded proteins and also intermediate filament keratins which are encoded by genes outside the EDC are present in different regions of feathers (Ng et al. 2014; Ng et al. 2015; Wu et al. 2015; Alibardi 2013). Most notably epidermal differentiation cysteine-rich protein (EDCRP) (Strasser et al. 2015) and epidermal differentiation proteins starting with MTF motif, Met-Thre-Phe, and rich in Histidine (EDMTFH) (Alibardi et al. 2016) are important feather components. While the mechanism by which EDMTFH may contribute to feather stabilization is unclear, EDCRP is likely to form multiple intermolecular disulfide bonds and thereby to enhance protein cross-linking in feathers (Strasser et al. 2015). The analysis of proteome data of chicken skin appendages has suggested that further proteins encoded by EDC genes are present in feathers (Rice et al. 2013; Strasser et al. 2014). Epidermal Differentiation protein containing DPCC (Asp-Pro-Cys-Cys) amino acid Motifs (EDDM) was one of these EDC proteins abundant in the proteome of chicken feathers but its expression pattern in feather cells and the possible role of EDDM in the evolution of feathers have remained unknown.
The aims of the present study were to determine the expression pattern of EDDM in chicken tissues, to identify avian and non-avian orthologs of EDDM and to develop a model for the evolutionary history of the EDDM gene.
Materials and methods
Comparative genomics and sequence analysis
Using the amino acid sequence of chicken (Gallus gallus) EDDM protein as query in tBLASTn searches (Altschul et al. 1990) at the NCBI GenBank website (http://www.ncbi.nlm.nih.gov/), EDDM orthologs were identified in the genome sequences of duck (Anas platyrhynchos), pigeon (Columba livia), falcon (Falco cherrug), Adélie penguin (Pygoscelis adeliae), emperor penguin (Aptenodytes forsteri), loon (Gavia stellata), flycatcher (Ficedula albicollis), canary (Serinus canaria), cuckoo roller (Leptosomus discolor), ostrich (Struthio camelus), greater rhea (Pterocnemia pennata), and great spotted kiwi (Apteryx haastii) (Jarvis et al. 2014); sequences of crocodilian EDDML were identified in the genome sequences of the American alligator (Alligator mississippiensis), the gharial (Gavialis gangeticus), and the saltwater crocodile (Crocodylus porosus) (Green et al. 2014; Holthaus et al. 2018b) (Suppl. Table S1). Amino acid sequences were obtained by translation of the coding region present in exon 2 of these genes. The amino acid sequences were aligned using Multalin (http://multalin.toulouse.inra.fr/multalin/) (Corpet 1988) with manual adjustment. Sequence logos were generated online with the Weblogo software (Crooks et al. 2004). Orthologs of the non-coding exon 1 of chicken EDDM were identified by BLASTn search of nucleotide sequences on the 5′-side of EDDM and EDDML genes.
Preparation of chicken tissues
Chickens were maintained and eggs were incubated according to published protocols (Eresheim et al. 2014). At Hamburger-Hamilton (HH) stages 35, 39, and 44 (Hamburger and Hamilton 1992), tissues were prepared from chicken embryos that were euthanized by decapitation. The tissue samples were fixed with 7.5% formaldehyde and embedded in paraffin as described previously (Mlitz et al. 2014). RNA was prepared with the Trifast reagent (VWR) according to a published protocol (Mlitz et al. 2014).
Quantitative reverse-transcription polymerase chain reaction
RNA was reverse-transcribed to cDNA with iScript cDNA Synthesis Kit (Bio-Rad, Hercules, CA) and the quantitative PCR was performed using the Lightcycler 480 DNA SYBR Green I master kit on a Roche LightCycler® (LC480) according to the manufacturer’s protocol. EDDM mRNA was amplified with the intron-spanning primer pair EDDM-f (5′-CGGCATTACTCCATCAGCTG-3′) and EDDM-r (5′-AACATCGGAGGGCTCAAGAA-3′). As a control transcript, Casp3 mRNA was amplified with primers reported previously (Strasser et al. 2015).
Generation of an antibody against EDDM
The peptide CYYARVPQGTTTYLKL, corresponding to amino acid residues 49–64 of chicken EDDM (Suppl. Fig. S1) was synthesized and coupled to keyhole limpet hemocyanin (KLH) by GeneCust, Ellange, Luxembourg. Six injections of 100 μg KLH-coupled peptide were performed to generate an antiserum in mice according to a published protocol (Eckhart et al. 2008).
Immunohistochemistry
Immunohistochemical stainings were performed according to published protocols (Mlitz et al. 2014; Alibardi et al. 2016). In brief, tissues were sectioned at 4-μm thickness and antigens were demasked with citrate buffer, pH 6 (Dako). Endogenous peroxidase was blocked with hydrogen peroxide. Mouse anti-EDDM antiserum at a dilution of 1:200 was used as primary antibody. Biotinylated sheep anti-mouse immunoglobulin (RPN1001V, lot 9793564, GE Healthcare, Chalfont, UK) at a dilution of 1:200 was used as secondary antibody, and sheep serum (10%) was added to prevent unspecific binding. In control experiments, the primary antibody was replaced with preimmune serum. The samples were incubated with streptavidin-biotin-horseradish peroxidase (HRP) complex and 3-amino-9-ethylcarbazole (DakoCytomation, Glostrup, Denmark) to develop red color. Nuclei were counterstained with hematoxylin.
Results
The EDDM gene encodes the protein with the highest number of cysteine residues among chicken EDC proteins
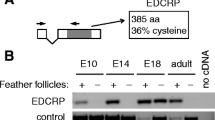
The EDDM gene is located in the EDC and has two exons. As the entire coding sequence is located in exon 2, it belongs to the Single-coding exon EDC (SEDC) genes (Strasser et al. 2014), which are located on the 5′-side of the evolutionarily conserved Cornulin (Crnn) gene in both chicken and human (Fig. 1). EDDM does not have an ortholog in humans. Cysteine residues that allow cross-linking via disulfide bonds are present in different numbers in SEDC proteins both in chicken and humans. EDCRP has 140 cysteine residues in a total of 385 amino acid residues which represents the highest relative content of cysteine (36%) among chicken EDC proteins (Strasser et al. 2015). However, the highest absolute number cysteine residues (n = 152) is present in EDDM (total number amino acid residues, n = 657) (Fig. 1; Suppl. Table S1), suggesting that this protein is capable of serving a role as a cross-linkable structural protein.
EDDM is located in the EDC and encodes a cysteine-rich protein. The EDC gene locus of humans (chromosome 1q21) and chicken (chromosome 25) is schematically depicted. The orientation of the genes is indicated by arrows; orthologs are connected by vertical lines. Colors distinguish genes within different domains of the EDC. The position of the corneous beta protein (CBP)/beta-keratin gene cluster is indicated by a striped box in the chicken EDC. The number of cysteine residues in proteins encoded by Single-coding exon EDC (SEDC) genes is shown in diagrams above the chicken and below the human EDC schematics. Values above bars indicate the number of cysteine residues in the proteins. The bars above the CBP cluster indicate the minimum (min), mean, and maximum (max) number of cysteine residues in CBPs
EDDM is expressed in feather barbs and in the subperiderm
To determine the distribution of EDDM in chicken tissues, we generated a mouse antiserum against a unique internal peptide of EDDM (Suppl. Fig. S1), and used this antibody for immunohistochemical studies. The studies of gene expression were carried out on embryonic tissues to compare different growth stages of skin appendages and to investigate embryo-specific epithelial cells of the periderm and subperiderm. Feather buds at development stages HH35 (Suppl. Fig. S2a) and HH39 (Suppl. Fig. S2b) and the pulp and sheath of feathers at HH44 (Fig. 2a) were immunonegative. By contrast, EDDM was detected at highest signal intensity in barbs and barbules of feathers (Fig. 2a; Suppl. Fig. S2c). EDDM was also detected in the subperiderm layer on scutate scales at stage HH44 (Fig. 2c). The staining intensities in the epidermis, the periderm (Fig. 2c), and the epithelium of the tongue (Suppl. Fig. S2e) were low or absent. Negative control stainings in which the EDDM antiserum was replaced by preimmune serum (Fig. 2b, d; Suppl. Fig. S2d) showed no signals in feathers and subperiderm. The intensity of immunostaining of EDDM protein correlated with EDDM mRNA abundance in feathers during embryonic development (Suppl. Fig. S2f). In summary, immunohistochemical and RT-PCR analysis demonstrated that EDDM is abundantly expressed in feathers and, at lower levels, in the subperiderm of embryonic scutate scales.
Immunohistochemial analysis of EDDM in the chicken. EDDM was detected by immunohistochemistry (red) in chicken feathers (a) and scutate scales (c) at stage HH44. In control experiments, the primary antibody was replaced by the preimmune serum (b, d). Nuclei were counterstained with hematoxylin (blue). Bars: 50 μm (a, b), 100 μm (c, d)
Avian EDDM and crocodilian EDDM-like proteins contain multiple cysteine-rich sequence repeats
To determine conserved and variable parts of the EDDM protein, we identified EDDM orthologs in genome sequences of vertebrates and compared nucleotide sequences of the genes and amino acid sequences of the encoded proteins. EDDM is conserved among birds and an EDDM-like (EDDML) gene is present in crocodilians but not in any other species of vertebrates investigated (Fig. 3a). In addition to the EDDML genes of the American alligator and the saltwater crocodile reported previously (Holthaus et al. 2018b), we could also identify EDDML of the gharial (Suppl. Table S1; Fig. 3a). The non-coding exon 1 and the proximal promoter was identified in EDDM genes of birds and in EDDML genes of crocodilians (Fig. 3b; Suppl. Fig. S3). Interestingly, the promoters of avian EDDM genes contained a canonical TATA box, whereas this important element of transcriptional control was modified in sequence in the promoters of crocodilian EDDML genes (Suppl. Fig. S3).
An EDDM-like (EDDML) gene is present in crocodilians. (a) Schematic depiction of the EDDM gene locus in birds and homologous loci in crocodilians and turtle. Species: chicken (Gallus gallus), pigeon (Columba livia), ostrich (Struthio camelus), American alligator (Alligator mississippiensis), saltwater crocodile (Crocodylus porosus), gharial (Gavialis gangeticus), turtle (Chrysemys picta). (b) Schematic organization of EDDM and EDDML genes. cds., coding sequence
Amino acid sequences, obtained by in silico translation of EDDM and EDDML coding sequences, were aligned to define common and divergent sequence features. EDDM and EDDML proteins contain 3 domains, all of which are characterized by high cysteine content. The amino- and carboxy-terminal domains are only partially conserved between birds and crocodilians whereas the central domain consists of at least 18 repeats of a sequence motif in both clades (Fig. 4a–c). Crocodilian EDDML proteins have 19–21 imperfect repeats of a 16-residue sequence and avian EDDM proteins have 18–54 imperfect repeats of a 15-residue sequence (Fig. 4b). In birds, the repeat sequence includes the DPCC motif, that is referred to in the protein name “EDDM.” The carboxy-terminal cysteine of this motif is not present in EDDML proteins of crocodilians, but cysteine is conserved at two other positions in birds and crocodilians (Fig. 4c). Another cysteine residue is present in the repeat of crocodilians, so that the average number of cysteine residues per repeat is 4 in all archosaurs. The sequence comparisons led to the evolutionary model depicted in Fig. 4d, which suggests that an EDDM-like gene originated in a common ancestor of archosaurs after the divergence from the turtle lineage, a central sequence motif underwent amplification in stem archosaurs, and further sequence changes both within and outside of the repeats led to divergent features of EDDM and EDDML in modern archosaurs. The number of central sequence repeats varied between the subclades of birds without an obvious correlation with an integumentary feature or lifestyle trait (Suppl. Fig. S4). Importantly, both the high cysteine content and the repetitive central domain are suggested to have emerged prior to the split of the avian and crocodilian lineages. Thus, our data point to an evolutionary origin of an epidermal differentiation protein with EDDM-like features in a common ancestor of birds and crocodilians, and this protein was subsequently co-opted for a new role as a component of feathers evolving in birds.
Amino acid sequence features and evolution of EDDM. The amino acid sequences of the amino-terminus (a), the central repeats (b), and the carboxy-terminus (c) of avian EDDM and crocodilian EDDML proteins were aligned. Colored fonts indicate the amino acid residues C, K, P, Q, and R. The symbol “:” below the sequence alignments in a and c mark positions of amino acid residues that are conserved in representatives of both EDDM and EDDML. (b) The conservation of residues in the sequence repeats in the central region of EDDM and EDDML is indicated by sequence logos. Species: chicken (Gallus gallus), duck (Anas platyrhynchos), pigeon (Columba livia), falcon (Falco cherrug), penguin (Pygoscelis adeliae), canary (Serinus canaria), ostrich (Struthio camelus), American alligator (Alligator mississippiensis), saltwater crocodile (Crocodylus porosus), gharial (Gavialis gangeticus). (d) Schematic model of the evolution of EDDM and EDDML. The boxes represent the organization of EDDM and EDDML proteins whereby repeat units in the central domain are indicated by red boxes. n, number of central sequence repeats
Discussion
The present study extends previous investigations on avian epidermal differentiation by determining the expression pattern and evolution of a gene that encodes the cysteine-rich protein EDDM in the chicken. Our immunolocalization of EDDM in feather barbs and barbules contributes to ongoing characterization of the complex molecular architecture that makes feathers mechanically resistant, yet elastic skin appendages, and the results of comparative genomics provide new insights into the molecular evolution of feathers.
The high cysteine content of EDDM is comparable to that of avian EDCRP, another cysteine-rich protein component of feathers (Strasser et al. 2015), and mammalian cysteine-rich keratin-associated proteins (KRTAPs), which are components of hair and nails (Rogers et al. 2001; Deb-Choudhury 2018; Plowman 2018; Wu and Irwin 2018). While KRTAPs are encoded by genes outside of the EDC, the EDCRP gene is located in the EDC but within a different region than EDDM (Fig. 1). While the avian EDC segment localized between S100A9/S100A12 and Loricrin lacks a counterpart in the mammalian EDC (Henry et al. 2012; Poterlowicz et al. 2017), the EDC segment containing EDDM in birds is syntenic with the human cluster of Late cornified envelope (LCE), Cysteine-rich C-terminal 1 (CRCT1), and Keratinocyte proline–rich protein (KPRP) genes (Fig. 1). Among human EDC proteins, KPRP has the highest number of cysteine residues. KPRP was detected by proteomics, in human and mouse nails (Rice et al. 2010; Jaeger et al. 2019) and, by immunohistochemistry, also in the granular layer of human epidermis (Lee et al. 2005). Thus, proteins competent in the formation of disulfide bonds via multiple cysteine residues likely contribute to the mechanical and chemical resistance of cornified skin derivatives in diverse amniotes.
Our immunostainings detected EDDM in the cornifying cells of barb and barbules as well as in the embryonic subperiderm. This pattern is similar to that of feather CBPs (Sawyer et al. 2003), EDCRP (Strasser et al. 2015), and EDMTFH (Alibardi et al. 2016), and suggests common mechanisms of gene regulation for the concerted synthesis of these proteins. The most mature portions of feathers were immunonegative for EDDM and despite testing several conditions of protein extraction under reducing conditions, we could not detect EDDM by western blot analysis. Most likely heavy intermolecular cross-linking of EDDM to other structural proteins in differentiated cells of feathers prevents access to antibodies and extraction of EDDM for detection as a soluble protein. As EDDM was previously detected in a proteomic analysis of feathers, which involved proteolytic digestion and mass spectrometry of peptides (Rice et al. 2013; Strasser et al. 2014), the immunohistochemical detection of EDDM in feathers is supported by a mechanistically independent method.
The identification of EDDM orthologs in crocodilians, which represent the closest phylogenetic relatives of birds, and the detection of EDDM in the subperiderm of scutate scales of chicken embryos suggest that EDDM has not specifically evolved as a feather protein. The crocodilian orthologs of EDDM share many sequence features, including the repeat-rich central domain and the high cysteine content, with avian EDDM. Therefore, these features have most likely been inherited from the last common ancestor of birds and crocodilians which lived around 240 million years ago (Kumar et al. 2017) and according to current knowledge, did not have feathers. Interestingly, the evolutionary origin of another feather protein, EDCRP, could also be traced back to the last common ancestor of extant archosaurs (Holthaus et al. 2018b). Therefore, the co-option of epidermal differentiation for new roles as components of feathers appears to be an important theme, comparable to the co-option of claw keratins as structural proteins of hair in mammals (Eckhart et al. 2008).
Taken together, our data add EDDM to the catalog of feather proteins, also including feather CBPs (feather beta-keratins), EDCRP, and EDMTFH, which are encoded by single-coding exon genes within the EDC, also known as SEDC genes (Strasser et al. 2014). It is thus remarkable that the diversification of SEDC proteins within the EDC of archosaurs provided the molecular substrates for the evolution of feathers.
References
Alibardi L (2013) Immunolocalization of alpha-keratins and feather beta-proteins in feather cells and comparison with the general process of cornification in the skin of mammals. Ann Anat 195:189–198
Alibardi L (2017) Review: cornification, morphogenesis and evolution of feathers. Protoplasma 254:1259–1281
Alibardi L, Holthaus KB, Sukseree S, Hermann M, Tschachler E, Eckhart L (2016) Immunolocalization of a histidine-rich epidermal differentiation protein in the chicken supports the hypothesis of an evolutionary developmental link between the embryonic subperiderm and feather barbs and barbules. PLoS One 11:e0167789
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ (1990) Basic local alignment search tool. J Mol Biol 215:403–410
Bolliger A, Varga D (1961) Feather lipids. Nature 190:1125
Chen CF, Foley J, Tang PC, Li A, Jiang TX, Wu P, Widelitz RB, Chuong CM (2015) Development, regeneration, and evolution of feathers. Annu Rev Anim Biosci 3:169–195
Chuong CM (1993) The making of a feather: homeoproteins, retinoids and adhesion molecules. Bioessays 15:513–521
Corpet F (1988) Multiple sequence alignment with hierarchical clustering. Nucleic Acids Res 16:10881–10890
Crooks GE, Hon G, Chandonia JM, Brenner SE (2004) WebLogo: a sequence logo generator. Genome Res 14:1188–1190
Deb-Choudhury S (2018) Crosslinking between trichocyte keratins and keratin associated proteins. Adv Exp Med Biol 1054:173–183
Eckhart L, Ehrlich F (2018) Evolution of trichocyte keratins. Adv Exp Med Biol 1054:33–45
Eckhart L, Dalla Valle L, Jaeger K, Ballaun C, Szabo S, Nardi A, Buchberger M, Hermann M, Alibardi L, Tschachler E (2008) Identification of reptilian genes encoding hair keratin-like proteins suggests a new scenario for the evolutionary origin of hair. Proc Natl Acad Sci U S A 105:18419-18423
Eckhart L, Lippens S, Tschachler E, Declercq W (2013) Cell death by cornification. Biochim Biophys Acta 1833:3471–3480
Eresheim C, Plieschnig J, Ivessa NE, Schneider WJ, Hermann M (2014) Expression of microsomal triglyceride transfer protein in lipoprotein-synthesizing tissues of the developing chicken embryo. Biochimie 101:67–74
Fraser RD, Parry DA (2008) Molecular packing in the feather keratin filament. J Struct Biol 162:1–13
Green RE, Braun EL, Armstrong J, Earl D, Nguyen N, Hickey G, Vandewege MW, St John JA, Capella-Gutiérrez S, Castoe TA, Kern C, Fujita MK, Opazo JC, Jurka J, Kojima KK, Caballero J, Hubley RM, Smit AF, Platt RN, Lavoie CA, Ramakodi MP, Finger JW Jr, Suh A, Isberg SR, Miles L, Chong AY, Jaratlerdsiri W, Gongora J, Moran C, Iriarte A, McCormack J, Burgess SC, Edwards SV, Lyons E, Williams C, Breen M, Howard JT, Gresham CR, Peterson DG, Schmitz J, Pollock DD, Haussler D, Triplett EW, Zhang G, Irie N, Jarvis ED, Brochu CA, Schmidt CJ, McCarthy FM, Faircloth BC, Hoffmann FG, Glenn TC, Gabaldón T, Paten B, Ray DA (2014) Three crocodilian genomes reveal ancestral patterns of evolution among archosaurs. Science 346:1254449
Greenwold MJ, Sawyer RH (2011) Linking the molecular evolution of avian beta keratins to the evolution of feathers. J Exp Zool 316:609–616
Greenwold MJ, Sawyer RH (2013) Expression of archosaurian beta-keratins: diversification and expansion of archosaurian beta-keratins and the origin of feather beta-keratins. J Exp Zool B Mol Dev Evol 320:393–405
Greenwold MJ, Bao W, Jarvis ED, Hu H, Li C, Gilbert MTP, Zhang G, Sawyer RH (2014) Dynamic evolution of the alpha (α) and beta (β) keratins has accompanied integument diversification and the adaptation of birds into novel lifestyles. BMC Evol Biol 14:249–265
Gregg K, Rogers GE (1986) Feather keratins: composition, structure and biogenesis. In: Bereither-Hahn J, Matoltsy GA, Sylvia-Richards K (eds) Biology of the integument, vertebrates 2. Springer-Verlag, Berlin, pp 666–694
Hamburger V, Hamilton HL (1992) A series of normal stages in the development of the chick embryo. 1951. Dev Dyn 195:231–272
Henry J, Toulza E, Hsu CY, Pellerin L, Balica S, Mazereeuw-Hautier J, Paul C, Serre G, Jonca N, Simon M (2012) Update on the epidermal differentiation complex. Front Biosci (Landmark Ed) 17:1517–1532
Holthaus KB, Eckhart L, Dalla Valle L, Alibardi L (2018a) Review: evolution and diversification of corneous beta-proteins, the characteristic epidermal proteins of reptiles and birds. J Exp Zool B Mol Dev Evol 330:438–453
Holthaus KB, Strasser B, Lachner J, Sukseree S, Sipos W, Weissenbacher A, Tschachler E, Alibardi L, Eckhart L (2018b) Comparative analysis of epidermal differentiation genes of crocodilians suggests new models for the evolutionary origin of avian feather proteins. Genome Biol Evol 10:694–704
Jaeger K, Sukseree S, Zhong S, Phinney BS, Mlitz V, Buchberger M, Narzt MS, Gruber F, Tschachler E, Rice RH, Eckhart L (2019) Cornification of nail keratinocytes requires autophagy for bulk degradation of intracellular proteins while sparing components of the cytoskeleton. Apoptosis 24:62–73
Jarvis ED, Mirarab S, Aberer AJ, Li B, Houde P, Li C, Ho SYW, Faircloth BC, Nabholz B, Howard JT, Suh A, Weber CC, da Fonseca RR, Li J, Zhang F, Li H, Zhou L, Narula N, Liu L, Ganapathy G, Boussau B, Bayzid MS, Zavidovych V, Subramanian S, Gabaldon T, Capella-Gutierrez S, Huerta-Cepas J, Rekepalli B, Munch K, Schierup M, Lindow B, Warren WC, Ray D, Green RE, Bruford MW, Zhan X, Dixon A, Li S, Li N, Huang Y, Derryberry EP, Bertelsen MF, Sheldon FH, Brumfield RT, Mello CV, Lovell PV, Wirthlin M, Schneider MPC, Prosdocimi F, Samaniego JA, Velazquez AMV, Alfaro-Nunez A, Campos PF, Petersen B, Sicheritz-Ponten T, Pas A, Bailey T, Scofield P, Bunce M, Lambert DM, Zhou Q, Perelman P, Driskell AC, Shapiro B, Xiong Z, Zeng Y, Liu S, Li Z, Liu B, Wu K, Xiao J, Yinqi X, Zheng Q, Zhang Y, Yang H, Wang J, Smeds L, Rheindt FE, Braun M, Fjeldsa J, Orlando L, Barker FK, Jonsson KA, Johnson W, Koepfli KP, O'Brien S, Haussler D, Ryder OA, Rahbek C, Willerslev E, Graves GR, Glenn TC, McCormack J, Burt D, Ellegren H, Alstrom P, Edwards SV, Stamatakis A, Mindell DP, Cracraft J, Braun EL, Warnow T, Jun W, Gilbert MTP, Zhang G (2014) Whole-genome analyses resolve early branches in the tree of life of modern birds. Science 346:1320–1331
Kumar S, Stecher G, Suleski M, Hedges SB (2017) TimeTree: a resource for timelines, timetrees, and divergence times. Mol Biol Evol 34:1812–1819
Lee WH, Jang S, Lee JS, Lee Y, Seo EY, You KH, Lee SC, Nam KI, Kim JM, Kee SH, Yang JM, Seo YJ, Park JK, Kim CD, Lee JH (2005) Molecular cloning and expression of human keratinocyte proline-rich protein (hKPRP), an epidermal marker isolated from calcium-induced differentiating keratinocytes. J Invest Dermatol 125:995–1000
Maderson PF, Hillenius WJ, Hiller U, Dove CC (2009) Towards a comprehensive model of feather regeneration. J Morphol 270:1166–11208
Mlitz V, Strasser B, Jaeger K, Hermann M, Ghannadan M, Buchberger M, Alibardi L, Tschachler E, Eckhart L (2014) Trichohyalin-like proteins have evolutionarily conserved roles in the morphogenesis of skin appendages. J Invest Dermatol 134:2682–2692
Ng CS, Wu P, Fan WL, Yan J, Chen CK, Lai YT, Wu SM, Mao CT, Chen JJ, Lu MY, Ho MR, Widelitz RB, Chen CF, Chuong CM, Li WH (2014) Genomic organization, transcriptomic analysis, and functional characterization of avian α- and β-keratins in diverse feather forms. Genome Biol Evol 6:2258–2273
Ng CS, Chen CK, Fan WL, Wu P, Wu SM, Chen JJ, Lai YT, Mao CT, Lu MY, Chen DR, Lin ZS, Yang KJ, Sha YA, Tu TC, Chen CF, Chuong CM, Li WH (2015) Transcriptomic analyses of regenerating adult feathers in chicken. BMC Genomics 16:756
Plowman JE (2018) Diversity of trichocyte keratins and keratin associated proteins. Adv Exp Med Biol 1054:21–32
Poterlowicz K, Yarker JL, Malashchuk I, Lajoie BR, Mardaryev AN, Gdula MR, Sharov AA, Kohwi-Shigematsu T, Botchkarev VA, Fessing MY (2017) 5C analysis of the epidermal differentiation complex locus reveals distinct chromatin interaction networks between gene-rich and gene-poor TADs in skin epithelial cells. PLoS Genet 13:e1006966
Prum RO (2005) Evolution of the morphological innovations of feathers. J Exp Zool B Mol Dev Evol 304:570–579
Prum RO, Dyck J (2003) A hierarchical model of plumage: morphology, development, and evolution. J Exp Zool B Mol Dev Evol 298:73–90
Rice RH, Xia Y, Alvarado RJ, Phinney BS (2010) Proteomic analysis of human nail plate. J Proteome Res 9:6752–6758
Rice RH, Winters BR, Durbin-Johnson BP, Rocke DM (2013) Chicken corneocyte cross-linked proteome. J Proteome Res 12:771–776
Rogers MA, Langbein L, Winter H, Ehmann C, Praetzel S, Korn B, Schweizer J (2001) Characterization of a cluster of human high/ultrahigh sulfur keratin-associated protein genes embedded in the type I keratin gene domain on chromosome 17q12-21. J Biol Chem 276:19440–19451
Sawyer RH, Knapp LW (2003) Avian skin development and the evolutionary origin of feathers. J Exp Zool B Mol Dev Evol 298:57–72
Sawyer RH, Salvatore BA, Potylicki TT, French JO, Knapp LW (2003) Origin of feathers: feather beta-keratins are expressed in discrete epidermal cell populations of embryonic scutate scales. J Exp Zool B Mol Dev Evol 295:12–24
Sawyer RH, Rogers L, Washington L, Glenn TC, Knapp LW (2005) Evolutionary origin of the feather epidermis. Dev Dyn 232:256–267
Strasser B, Mlitz V, Hermann M, Rice RH, Eigenheer RA, Alibardi L, Tschachler E, Eckhart L (2014) Evolutionary origin and diversification of epidermal barrier proteins in amniotes. Mol Biol Evol 31:3194–3205
Strasser B, Mlitz V, Hermann M, Tschachler E, Eckhart L (2015) Convergent evolution of cysteine-rich proteins in feathers and hair. BMC Evol Biol 15:1–11
True JR, Carroll SB (2002) Gene co-option in physiological and morphological evolution. Annu Rev Cell Dev Biol 18:53–80
Wu DD, Irwin DM (2018) Evolution of trichocyte keratin associated proteins. Adv Exp Med Biol 1054:47–56
Wu P, Ng CS, Yan J, Lai YC, Chen CK, Lai YT, Wu SM, Chen JJ, Luo W, Widelitz RB, Li WH, Chuong CM (2015) Topographical mapping of alpha-and beta-keratins on developing chicken skin integument: functional interaction and evolutionary perspectives. Proc Natl Acad Sci U S A 112:E6770–E6779
Yu M, Wu P, Widelitz RB, Chuong CM (2002) The morphogenesis of feathers. Nature 420:308–312
Acknowledgments
Open access funding provided by Austrian Science Fund (FWF). The authors thank Maria Buchberger and Bahar Golabi for excellent technical support and Bettina Ebner and Karin Brigit Holthaus for helpful discussions.
Funding
The study was supported by the Austrian Science Fund (FWF): P23801 and P28004.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
All animal procedures were approved by the Animal Care and Use Committee of the Medical University of Vienna and by the Federal Ministry for Science and Research (BMWF) (66.016/0014-II/3b/2011), and all these procedures were conducted according to the guidelines established by the committee and the ministry.
Additional information
Handling Editor: Georg Krohne
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
ESM 1
(PDF 2169 kb)
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Lachner, J., Ehrlich, F., Mlitz, V. et al. Immunolocalization and phylogenetic profiling of the feather protein with the highest cysteine content. Protoplasma 256, 1257–1265 (2019). https://doi.org/10.1007/s00709-019-01381-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00709-019-01381-3