Abstract
Plant growth-promoting rhizobacteria stimulate plant growth and development via different mechanisms. In this study, we characterized the effect of volatiles from Bacillus methylotrophicus M4-96 isolated from the maize rhizosphere on root and shoot development, and auxin homeostasis in Arabidopsis thaliana. Phytostimulation occurred after 4 days of interaction between M4-96 and Arabidopsis grown on opposite sides of divided Petri plates, as revealed by enhanced primary root growth, root branching, leaf formation, and shoot biomass accumulation. Analysis of indole-3-acetic acid content revealed two- and threefold higher accumulation in the shoot and root of bacterized seedlings, respectively, compared to uninoculated plants, which was correlated with increased expression of the auxin response marker DR5::GUS. The auxin transport inhibitor 1-naphthylphthalamic acid inhibited primary root growth and lateral root formation in axenically grown seedlings and antagonized the plant growth-promoting effects of M4-96. Analysis of bacterial volatile compounds revealed the presence of four classes of compounds, including ten ketones, eight alcohols, one aldehyde, and two hydrocarbons. However, the abundance of ketones and alcohols represented 88.73 and 8.05%, respectively, of all airborne signals detected, with acetoin being the main compound produced. Application of acetoin had a different effect from application of volatiles, suggesting that either the entire pool or acetoin acting in concert with another unidentified compound underlies the strong phytostimulatory response. Taken together, our results show that B. methylotrophicus M4-96 generates bioactive volatiles that increase the active auxin pool of plants, stimulate the growth and formation of new organs, and reprogram root morphogenesis.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Plants and their associated bacteria are no longer considered separated entities. The rhizosphere provides conditions that strengthen these mutual benefits through complex networks of molecular interactions. The so-called plant growth-promoting rhizobacteria (PGPR) improve the growth and productivity of crops, such as wheat, soybean, lettuce, bean, maize, and barley through the production of plant hormones, eliciting defense responses and enhancing nutrient acquisition (Brink 2016; Pieterse et al. 2016).
The potential of single PGPR isolates to modulate the hormonal content of plants represents a major challenge towards understanding cross-kingdom impacts on complex plant-bacteria communities. Indole-3-acetic acid (IAA) is the main naturally occurring auxin in plants. IAA coordinates growth and behavioral processes during a plant’s life cycle, and affects the division, elongation, and differentiation of cells (Woodward and Bartel 2005). Bacteria can synthesize IAA and use this phytohormone to interact with plants during colonization, which involves phytostimulation and circumvention of basal plant defense mechanisms (Lambrecht et al. 2000; Vogel 2006; Spaepen et al. 2007).
In the absence of physical contact with plant roots, certain microorganisms use other molecular mechanisms to trigger plant growth. This includes the release of volatile organic compounds (VOCs), which are perceived by the host and lead to changes in root system architecture (RSA) and improvements in shoot growth (Ryu et al. 2003; Zhang et al. 2007; Gutiérrez-Luna et al. 2010). The best studied rhizobacterium that uses volatiles to signal to plants is Bacillus subtilis GB03, which generates 3-hydroxy-2-butanone (acetoin) and 2,3-butanediol. Application of the purified compounds results in a dose-dependent stimulation of plant growth, which is likely to occur via auxin-dependent mechanisms (Zhang et al. 2007).
In this study, we characterized the rhizobacterium B. methylotrophicus M4-96 isolated from the maize rhizoplane as a new PGPR. Bacterial co-cultivation promoted the growth of Arabidopsis via the emission of VOCs, stimulating primary root growth, lateral root formation, and biomass accumulation. We monitored auxin-responsive gene expression in the shoots and roots of transgenic plants expressing the DR5::GUS construct, and found that bacterial VOCs strongly increase auxin responsiveness in apical meristems. In addition, plants exposed to VOCs exhibited a twofold increase IAA content in shoots and threefold increase in roots. Application of the auxin transport inhibitor 1-naphthylphthalamic acid (NPA) to plants exposed to microbial VOCs reduced this phytostimulatory effect. Thus, we demonstrate that VOCs from B. methylotrophicus M4-96 enhance IAA levels in plant tissues, which likely determine its probiotic properties.
Materials and methods
Isolation and molecular characterization of M4-96 strain
M4-96 was isolated from the rhizoplane of Zea mays (maize) plants grown under greenhouse conditions on an alkaline vertisol (pH 8.6) obtained from an agricultural field termed “El Lometón,” in Mexico (19° 4646.61 N, 101° 0806.07 W). The soil was composed of 40.76% sand, 29.44% silt, 29.80% clay, 35.2% total CO3 −, and 2.63% organic matter. It was sampled from the upper layer (0 to 20 cm) in the field and air-dried. Each sample was then meshed (≤4 mm) and maintained at 4 °C until it was used as a substrate for greenhouse experiments. Plants were germinated and grown for 12 days in pots containing 1000 g soil. Plant roots were then collected and a rhizoplane bacterial suspension was obtained as previously described (Valencia-Cantero et al. 2007). Briefly, plants were carefully removed from the soil and roots were excised at the root/shoot junction. Soil was separated from the roots by vigorously shaking in 100 mL sterile water for 30 s, using a Vortex. Rhizoplanic bacteria were separated from roots by agitation at 200 rpm in 20 mL sterile phosphate-buffered saline solution for 2 h using an orbital shaker. Plates with soil agar (Ellingsoe and Johnsen, 2002) supplemented with yeast extract (1 g L−1) were inoculated with serially diluted rhizoplanic soil suspensions and incubated for 7 days. Isolates were streaked at least three times to ensure purity. The isolated bacteria were routinely cultured in nutritive agar (AN, Difco) or nutritive broth.
Genomic DNA was isolated as previously described (Hernández-Salmerón 2016). Briefly, 20 mL of an overnight culture grown at 28 °C under agitation at 250 rpm was taken and genomic DNA was isolated using the Wizard® Genomic DNA Purification Kit following the manufacturer’s instructions (Promega). DNA samples were subjected to an additional purification step using the Wizard® Genomic DNA Purification Kit (Promega). The quality and quantity of the final DNA sample were evaluated by agarose gel electrophoresis and a NanoDrop 1000 Spectrophotometer (Thermo Scientific).
Genomic DNA samples of the M4-96 strain were sequenced at LANGEBIO-Irapuato, México. Genome sequencing was performed using a MiSeq Sequencer (Illumina, Inc.) and reads were assembled employing Newbler v2.9 (Roche). A 2931-bp sequence corresponding to 23S rDNA was defined in silico employing MEGA version 6 (Tamura et al. 2013) and searching the 23S bacterial universal sequence 5′ ACCCGACAAGGAATTTCGC 3′ reported by Amann et al. (1995). The sequence was deposited in GenBank [National Center for Biotechnology Information (NCBI)] under accession number KU302815, and compared with those from the non-redundant nucleotide database at GenBank (www.ncbi.nlm.nih.gov/blast) using the blast algorithm (Altschul et al. 1990). The most scored sequences [B. amyloliquefaciens G341 (CP011686.1), B. methylotrophicus JJ-D34 (CP011346.1), B. amyloliquefaciens plantarum NJN-6 (CP007165.1), B. amyloliquefaciens L-S60 (CP011278.1), B. amyloliquefaciens L-H15 (CP010556.1), B. subtilis Bs916 (CP009611.1), B. methylotrophicus JS25R (CP009679.1), B. subtilis ATCC13952 (CP009748.1), B. amyloliquefaciens plantarum TrigoCor1448 (CP007244.1), B. amyloliquefaciens LFB112 (CP006952.1), B. amyloliquefaciens plantarum UCMB5113 (HG328254.1), B. amyloliquefaciens plantarum CAUB946 (HE617159.1)] and other relevant sequences [B. subtilis ATCC 19217 (CP009749.1), B. methylotrophicus YJ11-1-4 (CP011347.1), B. methylotrophicus KACC 13105 (NZ_JTKJ02000006.1), B. atrophaeus globigii BSS (CP007640.1), B. atrophaeus UCMB-5137 (CP011802.1), B. amyloliquefaciens DSM7 (FN597644.1), B. subtilis subtilis ATCC 6051 (NC_020507.1), B. subtilis UD1022 (CP011534.1), B. subtilis TO-A JPC (CP011882.1), B. subtilis spizizenii TU-B-10 (CP002905.1), B. paralicheniformis BL09 (CP010524.1), B. licheniformis ATCC 14580 (AE017333.1), B. pumilus TUAT1 (AP014928.1)] were used to construct a phylogenic tree using MEGA version 6 (Tamura et al. 2013) employing the “maximum parsimony,” “minimum evolution,” “maximum likelihood,” “neighbor-joining,” and unweighted pair group method with arithmetic means (UPGMA) algorithms with 1000 bootstraps repetitions.
Plant material and growth conditions
A. thaliana (ecotype Col-0) and the transgenic Arabidopsis line DR5::GUS (Ulmasov et al. 1997) were used in the different experiments. Seeds were surface sterilized with 95% (v/v) ethanol for 5 min and 20% (v/v) bleach for 7 min. After five washes in distilled water, seeds were sown on agar plates containing 0.2× Murashige and Skoog (MS) medium (basal salt mixture; M5524, Sigma, St Louis). Phytagar was purchased from Gibco-BRL. Plates were placed at a 65° angle to allow root growth along the agar surface and the unimpeded aerial growth of hypocotyls. Plants were placed in a plant-growth chamber (Percival AR-95 L) with a photoperiod of 16-h light/8-h darkness, light intensity of 100 μmol m2 s−1, and temperature of 22 °C. For auxin polar transport inhibition bioassays, nutrient medium was supplemented with 0.5, 1, 2, 4, and 8 μM NPA prepared in dimethyl sulfoxide (DMSO) (Sigma-Aldrich, St. Louis, MO, USA) (Casimiro et al. 2001). Control media contained only DMSO.
Inoculation experiments
Petri dishes (I-plates) containing 0.2× MS medium were used to assess the effect of rhizobacterial volatile emission on the growth and development of Arabidopsis. Four Arabidopsis seeds were sown on one half of the plate and 4 days after germination the rhizobacterium was inoculated at approximately 1 × 106 colony-forming unit (CFU) on the opposite half of the plate. The treatment included four replicates with a total of 16 seedlings. Axenic plants (plants in uninoculated dishes) were used as controls. The plates were incubated for 10 days in a growth chamber (Percival AR95L). At the end of this period, primary root length (PRL), lateral root number (LRN), lateral root length (LRL), and total biomass were recorded.
Histochemical analysis
For histochemical analysis of GUS activity, Arabidopsis seedlings were incubated for 12 to 14 h at 37 °C in GUS reaction buffer (0.5 mg/ml 5-bromo-4-chloro-3-indolyl-β-D-glucuronide in 100 mM sodium phosphate, pH 7). The stained seedlings were cleared using the method described by Malamy and Benfey (1997). At least 15 transgenic plants were analyzed per treatment. A representative plant was selected and photographed using a Leica MZ6 stereomicroscope.
Analysis of rhizobacterial VOCs
VOCs released by B. methylotrophicus M4-96 were analyzed in 10-day-old colonies grown in plastic Petri dishes containing 0.2× MS medium and by solid-phase microextraction (SPME) and gas chromatography-mass spectrometry (GC-MS). The compounds were collected with blue SPME fiber (PDMS/DVB) (Supelco, Inc., Bellafonte, PA, USA) and desorbed at 180 °C for 30 s in the injector port of a gas chromatograph (Agilent 6850 Series II; Agilent, Foster City, CA, USA), equipped with a MS detector 5973 from Agilent, and a free fatty acid-phase capillary column (HP-FFAP) (30 m × 0.25 mm I.D., 0.25-μm film thickness). Helium was used as the carrier gas (1 ml/min), and the detector temperature was 250 °C. The column was held for 5 min at 40 °C, and then programmed to increase by 3 °C per min to a final temperature of 220 °C, which was maintained for 5 min. Three independent measurements were conducted. Source pressure was 7 Pa and filament voltage was 70 eV, and a scan rate of 1.9 scan s−1 was employed. The compounds were identified by comparison with mass spectra from a library (NIST/EPA/NIH, “Chem Station” Agilent Technologies Rev. D.04.00 2002).
Auxin determination
IAA was extracted from Arabidopsis (Col-0) and levels were determined 6 days after inoculation with B. methylotrophicus M4-96. Plant tissues were frozen and ground in liquid N2. Then, ~200 mg ground tissue was placed in an Eppendorf tube, homogenized with 500 μl isopropanol/H2O/concentrated HCl (2:1:0.002, v/v), and shaken for 30 s. Samples were centrifuged at 11,500 rpm for 3 min, and supernatants were collected and subjected to IAA extraction with 300 μl of dichloromethane. IAA was derivatized with acetyl chloride in methanol (500 μl/2 ml), sonicated for 15 min, and heated for 1 h at 75 °C. After cooling, the derivatized sample was evaporated and resuspended in 25 μl of ethyl acetate for GC-MS analysis. GC-SIM-MS and the retention time were established for IAA-methyl ester (IAA-ME; m/z 130, 189 M+). IAA was purchased from Sigma and used as a standard, and the concentration was estimated using a calibration curve. Samples were injected into GC-MS and 5% phenyl methyl silicone capillary column (HP-5 MS) (30 m × 0.25 mm I.D., 0.25-μm film thickness). The operating conditions were 1 ml/min−1 helium as carrier gas, 250 °C detector temperature, and 180 °C injector temperature. The column was held for 3 min at 80 °C and programmed at 6 °C min−1 to a final temperature of 230 °C for 20 min.
Effect of acetoin on plant growth and development
Acetoin (purchased from Sigma) was dissolved in DMSO. Acetoin was added to the plant growth medium at concentrations of 0, 7, 70, and 700 μM. As a control, DMSO was added at a volume equal to that used for the highest concentration of compound. Petri dishes containing seven plants per treatment were placed in a Percival AR95L growth chamber for 10 days and used to estimate biomass production. The concentrations of acetoin used in this experiment were selected based on the findings of our previous reports, which showed the effect of pure volatiles identified from soil microorganisms on the growth of Arabidopsis in vitro (Garnica-Vergara et al. 2016; Raya-González et al. 2017).
Statistical analysis
The effects of the bacteria on parameters of growth and development in Arabidopsis seedlings were analyzed in a completely randomized experimental design with a one-way analysis of variance (ANOVA) and least significant difference (LSD) test. To assess the degree of interaction between the different parameters, a cluster analysis was performed. All statistical analyses were performed using Statistica 8.0 software (Statsoft, Inc. Tulsa, OK, USA).
Results
B. methylotrophicus M4-96, a rhizobacterium isolated from maize rhizoplane
The M4-96 strain was characterized by 23S rDNA sequence analysis. A 2931-bp sequence corresponding to 23S rDNA was used to search the NCBI GenBank database for homology. Sequences from the M4-96 isolate (Ac. KU302815) showed almost 100% homology to those from B. amyloliquefaciens strain G34, B. methylotrophicus strain JJD34, B. amyloliquefaciens subsp. plantarum strain NJN6, and B. subtilis strain ATCC 13952, among others. This places the M4-69 isolate within the B. subtilis group (Borshchevskaya et al. 2013).
In order to clarify the taxonomic position of the M4-96 isolate, a phylogenetic tree was constructed using 12 of the closest sequences, and 13 relevant sequences, including those from B. amyloliquefaciens DSM7 (Rückert et al. 2011), B. subtilis subtilis ATCC 6051 (Borshchevskaya et al. 2013), and B. methylotrophicus KACC 13105 (Madhaiyan et al. 2010). Based on the results of this analysis, the M4-96 isolate was included in the subtilis-amyloliquefaciens-methylotrophicus cluster, clearly separated from other closely related Bacillus species, such as B. licheniformis, B. paralicheniformis, and B. pumilus (Fig. 1). Deeper in the subtilis-amyloliquefaciens-methylotrophicus cluster, we found a subtilis sub-cluster defined by B. subtilis subtilis 6051 and an amyloliquefaciens-methylotrophicus sub-cluster defined by B. amyloliquefaciens DSM7 and B. methylotrophicus KACC 13105, and including the isolate M4-96. Recently, B. amyloliquefaciens subsp. plantarum was reclassified as a later heterotypic synonym of B. methylotrophicus based on the analysis of genomic data (Dunlap et al. 2015). Taking into account the data from the 23S rRNA analysis, and based on the fact that M4-96 is a plant-colonizing bacterium isolated from the Zea mays rhizoplane, we conclude that M4-96 is a B. methylotrophicus strain.
Phylogenetic tree inferred on the basis of 23S rDNA sequences (2931 bp) obtained from the M4-96 isolate. Sequence comparisons were performed with 12 of their closest species, and additional 13 relevant sequences including the type strains B. amyloliquefaciens DSM7, B. methylotrophicus KACC 13105 and B. subtilis subtilis ATCC 6051 (bold letters). The phylogenetic tree was constructed by the “maximum parsimony” algorithm. Numbers next to branches represent bootstraps in the base to 1000 repetitions
VOCs from B. methylotrophicus M4-96 modulate the growth and development of Arabidopsis seedlings
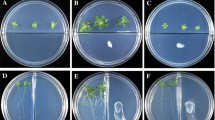
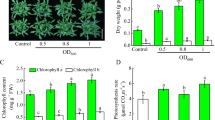
In this study, we analyzed the effect of VOCs from B. methylotrophicus M4-96 strain on the growth and development of Arabidopsis after 10 days of interaction. For this purpose, Arabidopsis seedlings were grown for 4 days on one side of a divided Petri plate supplied with MS 0.2× agar medium, and after this period, the strain was inoculated on the opposite side of the plate. Then, rosette leaf number and shoot and root fresh weight were determined every 2 days. Inoculation with B. methylotrophicus M4-96 increased the growth and development of plants (Fig. 2a), stimulating rosette leaf formation after 4 days of interaction (Fig. 2b). In addition, shoot and root fresh weight roughly increased by 6 days and doubled by 10 days (Fig. 2c, d).
Effects of B. methylotrophicus M4-96 on biomass production in Arabidopsis. The assay was performed using divided Petri plates to test the activity of VOC emissions on plant growth. A. thaliana seeds were sown on one-half of the plate and the rhizobacterium was inoculated on the opposite half of the plate. a Representative photographs showing the effects of the M4-96 strain on Arabidopsis growth at different times of the interaction. b Leaf number in axenic control plants and during the interaction with bacterial volatiles. c, d Shoot and root fresh weights of Arabidopsis seedlings with and without inoculation with M4-96. Bars represent the mean ± standard error values (n = 16). Asterisks are used to indicate means that differ significantly by Student’s test (p ≤ 0.05)
The PGPR effect of B. methylotrophicus M4-96 was accompanied by changes in root architecture. The bacterial VOCs stimulated primary root growth and lateral root formation after 2 days of interaction (Fig. 2a and Fig. 3a, b). This effect is better observed in Fig. 3c, where an increment in root density can be clearly observed. These data indicate that VOCs from B. methylotrophicus M4-96 promote plant growth and modulate rhizogenesis in Arabidopsis.
Effects of B. methylotrophicus M4-96 inoculation on Arabidopsis root developmental parameters. a Primary root length. b Lateral root number. c Lateral root density (lateral root number/cm of primary root). Bars represent the mean ± standard error values (n = 16). Asterisks are used to indicate means that differ significantly by Student’s test (p ≤ 0.05)
B. methylotrophicus M4-96 increases DR5::GUS expression and the biosynthesis of IAA in Arabidopsis seedlings
An important property of B. methylotrophicus M4-96 is its ability to promote root growth, which is often associated with auxin signaling (Fukaki et al. 2007). To determine if VOCs from M4-96 increase lateral root formation through this hormonal mechanism, we analyzed the influence of VOCs on the expression of the auxin response marker DR5::GUS (Sabatini et al. 1999; Stepanova et al. 2005; Ulmasov et al. 1997). Changes in β-glucuronidase (GUS) activity were monitored every 2 days. Representative photographs of GUS expression in the control, non-inoculated plants are shown in Fig. 4a, c, e, g, i, and in inoculated plants in Fig. 4b, d, f, h, j. Interestingly, plants exposed to bacterial VOCs showed a more robust primary root following 2 days of interaction, and the GUS expression pattern was higher in these plants at all times. Conversely, in the lateral root formation zone, we observed higher auxin-inducible expression in the vasculature and in lateral root primordia (Fig. 4l, n, p, r, t) than in their respective controls (Fig. 4k, m, o, q, s). These findings suggest that auxin signaling is activated during the initiation of lateral root formation.
Effects of B. methylotrophicus M4-96 on auxin-regulated gene expression in roots on different days of interaction. Twelve-hour GUS staining of DR5::GUS Arabidopsis seedlings grown on agar plates containing 0.2× MS medium. a, c, e, g, i DR5::GUS expression in primary roots from uninoculated plates and b, d, f, h, j from inoculated plants. k, m, o, q, s Lateral root primordia from control plants and l, n, p, r, t from plants exposed to bacterial volatiles. Note an increase in GUS expression in meristems and lateral root primordia in the treatments exposed to volatiles from M4-96 strain. Photographs are representative individuals of at least 20 stained seedlings. The experiment was repeated twice with similar results
Next, we analyzed GUS expression in cotyledons and true leaves from the same plants (Fig. 5). The results showed that M4-96 increased GUS activity starting 4 days after inoculation. Magnified images of 10-day-old seedlings showed that the newest leaves expressed GUS in the vasculature. In addition, leaves from inoculated plants were larger than those from the control at most times, suggesting that active auxin transport might improve leaf growth. To determine whether VOCs from B. methylotrophicus M4-96 induce IAA biosynthesis in Arabidopsis-inoculated plants, shoots and roots were collected and the IAA content was determined by GC-MS. B. methylotrophicus increased the concentration of IAA two- and threefold in the shoot and root, respectively (Fig. 6a, b), which correlates with the increased DR5::GUS expression in bacterized plants; this increment might be responsible for enhanced auxin transport within the plant. To investigate this possibility, the plant growth medium was supplemented with the auxin transport inhibitor NPA at concentrations from 0.5 to 8 μM. Supplementation of media with NPA was found to inhibit the growth of primary roots and the formation of lateral roots in both control and inoculated plants (Fig. 7a, b). However, with 1 μM NPA, the bacteria conferred resistance to the inhibitor, such that plants required 2 μM or more NPA to arrest lateral root development (Supplementary Fig. S1). NPA also diminished the leaf number in bacterial-inoculated seedlings (Fig. 7d). In contrast, DR5::GUS expression increased in shoots and roots in line with the NPA increment; however, in inoculated plants, the expression was more intense (Fig. 8a, b). These data suggest that volatiles from the M4-96 strain stimulate plant growth and development through an auxin-dependent mechanism.
Effects of B. methylotrophicus M4-96 on auxin-regulated gene expression in shoots on different days of interaction. Twelve-hour GUS staining of DR5::GUS Arabidopsis seedlings grown on agar plates containing 0.2× MS medium with and without M4-96. Note the increased GUS expression in the apical meristem following exposure to volatiles from the M4-96 strain, principally after 8 days of interaction. In addition, the leaves treated with M4-96 were bigger and more robust. Photographs are representative individuals of at least 20 stained seedlings. The experiment was repeated twice with similar results
Quantification of indole-3-acetic acid from control or inoculated Arabidopsis seedlings by gas chromatography and mass spectrometry. Arabidopsis seedlings were germinated and grown for 10 days in 0.2× MS medium, then shoots and roots were excised and individually analyzed. a Indole-3-acetic acid content in shoot or b in root. Bars represent the mean ± standard error values (n = 6). Asterisks are used to indicate means that differ significantly by Student’s test (p ≤ 0.05)
Effect of the auxin transport inhibitor 1-naphthyphthalamic acid (NPA) on the growth and development of Arabidopsis plants exposed to B. methylotrophicus M4-96 volatiles. Plants were grown in Petri plates in MS media supplemented with the solvent dimethyl sulfoxide (DMSO) or 0.5, 1, 2, 4, and 8 μM NPA in axenic system or inoculated with M4-96. After 10 days of growth, a primary root length, b lateral root number, c lateral root length, and d leaf number per plant were recorded. Bars represent the mean ± standard error values (n = 16). Asterisks are used to indicate means that differ significantly by Student’s test (p ≤ 0.05)
Effect of the auxin transport inhibitor NPA on the expression of the auxin-responsive reporter DR5::GUS in plants exposed to B. methylotrophicus M4-96 volatiles. Twelve hour GUS staining of DR5::GUS Arabidopsis seedlings grown on agar plates containing 0.2× MS medium and DMSO or different NPA doses (0.5, 1, 2, 4, and 8 μM). a Auxin-regulated gene expression in shoot and b root. Photographs are representative individuals of at least 16 stained seedlings. The experiment was repeated twice with similar results
B. methylotrophicus M4-96 produces acetoin as a major volatile compound
To determine the VOCs released by B. methylotrophicus M4-96, we performed SPME-GC-MS analyses. Table 1 shows the identity and abundances of all compounds. We identify 23 VOCs, with 3-hydroxy-2-butanone, commonly named acetoin, being the most abundant (74.03%). All other compounds are listed in Table 1, and were found to belong mostly to four classes, ketones (ten compounds), alcohols (eight compounds), aldehyde (one compound), and hydrocarbons (two compounds). Regarding the abundance of acetoin, 2,3-butanedione and the other ketones, airborne chemical signaling from M4-96 in MS medium is found to be dominated by ketones (88.73%) and alcohols (8.05%).
Acetoin slightly induces Arabidopsis growth and development, but its effect is not comparable with the pool of VOCs emitted by B. methylotrophicus M4-96
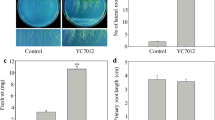
Since acetoin was the most abundant compound within the chromatographic profile of the VOCs, we investigated its effects on the growth and development of Arabidopsis seedlings. Different concentrations of acetoin (0, 7, 70, and 700 μM) were tested, according to the methodology reported for other microbial volatiles (Garnica-Vergara et al. 2016; Raya-González et al. 2017). Interestingly, 7 μM acetoin statistically increased shoot fresh weight, root growth, and lateral root formation, whereas increasing doses, such as 700 μM, diminished root fresh weight (Fig. 9a–d). Although some bioactivity could be attributed to acetoin, it is unlikely that the phenotypic effects observed in plants exposed to the pool of compounds emitted by B. methylotrophicus M4-96 can be attributed to acetoin alone.
Effect of acetoin on plant biomass production and root system architecture. A. thaliana (Col-0) seedlings were germinated and grown for 10 days under increasing acetoin concentrations. Seedlings were grown in 0.2× MS medium supplied with DMSO or 7 and 700 μM acetoin. a Shoot biomass, b root biomass, c primary root length, and d lateral root number were recorded. Bars represent the mean ± standard error values (n = 16). Asterisks are used to indicate means that differ significantly by Student’s test (p ≤ 0.05)
Discussion
A few bacterial species have been reported to modulate plant growth and development through the emission of VOCs. Here, we identified B. methylotrophicus M4-96, a bacterium isolated from maize rhizosphere as PGPR, which promotes Arabidopsis growth via VOC emission. Methylotrophy is a property of specialized bacteria that are capable of growth on reduced one-carbon compounds, such as methanol or methane, and strains from the genus Bacillus display strong resistance to high methanol concentrations (Madhaiyan et al. 2010). Few studies have investigated the plant growth-promoting activities of these species, which include P-solubilization, IAA and siderophore production, ACC deaminase, sulfur oxidation, and antifungal activities (Madhaiyan et al. 2010; Mehta et al. 2014). In the present study, we found that VOCs from B. methylotrophicus M4-96 had a strong effect on plant growth after 4 days of interaction, and such stimulation remained until the last day of interaction. The rhizobacteria promoted biomass accumulation in the shoot and roots, and induced the emergence of new leaves and more lateral roots without compromising primary root growth.
Previously, the effect of VOCs from 12 rhizobacterial isolates on the root architecture of Arabidopsis was investigated (Gutiérrez-Luna et al. 2010). None of the reported plant responses was similar to those observed in response to B. methylotrophicus M4-96. This suggests that the “molecular dialogue” between plants and bacteria is highly specific. In this case, the increased auxin fingerprint in apical meristems and leaf veins, as revealed by the reporter line DR5::GUS, is positively correlated with increases in the IAA content in the shoot and roots. This suggests that VOCs from the M4-96 strain induce IAA biosynthesis, increasing the internal IAA pool; therefore, it is possible that the plant concomitantly activates auxin transport mechanisms from source to sink tissues to induce developmental processes, such as lateral root development and shoot branching.
Our understanding of how auxin biosynthesis and degradation are regulated in the root system under normal and stress conditions is still very limited, but it is clear that auxin homeostasis plays a key role in shaping root architecture (Ljung 2013). To test the hypothesis that bacterial VOCs influence plant growth and development via an auxin-dependent mechanism, we analyzed the expression of DR5::GUS in the aerial part of inoculated plants and in the root meristem of seedlings grown in medium supplemented with 1 μM NPA, which inhibits auxin efflux. At low concentrations (1–5 μM), NPA efficiently blocks the basal polar auxin flow required for plant development, while at high (>50 μM) concentrations, NPA seems to affect cell-to-cell trafficking components (Geldner et al. 2001; Gil et al. 2001; Peer et al. 2009). We observed that inhibition of auxin efflux with low concentrations of NPA, leads to the accumulation of IAA at the sites of synthesis, and increased root thickness, with no effect on the appearance of new organs. Otherwise, inoculated plants showed increased resistance to higher doses of NPA compared with the control plants, maybe because these plants produce more IAA, which protects them from the effect of the inhibitor. These results confirm that bacterial VOCs can modulate auxin biosynthesis and transport, two critical factors involved in root growth and lateral root formation.
The finding that Azospirillum produces auxin and that roots respond by forming more lateral roots and root hairs (Spaepen et al. 2007) reveals a critical role for auxins in mediating plant-microbe interactions. Thus, root sensing of rhizobacteria via auxin release into the rhizosphere, or the detection of emitted VOCs, coordinates shoot patterns such as phyllotaxis, branching, and stem initiation; this may be dependent on the multiple feedback loops in the auxin machinery, coupled with plant phase transitions that may change the root-shoot feedback relationships as patterning proceeds (Leyser et al. 2010). The ability of auxin to promote its own transport suggests it can act as a long-distance communication signal, linking shoot patterning to root auxin pools, which largely depend on the rhizosphere microbiome.
Analysis of VOCs from the M4-96 strain showed acetoin as the major compound. Various microorganisms produce acetoin as a nonacidic end-product from sugar fermentation to prevent lethal medium acidification. Acetoin can be oxidized to 2,3-butanedione by O2 present in the fermentation medium, or enzymatically reduced (with NADH as cofactor) to form 2,3-butanediol. The ratio of these metabolites depends on the culture redox potential, initial substrate concentration, temperature, pH, and oxygen level (Sharma and Noronha 2014). Besides acetoin, 2,3-butanedione (0.77%) was identified within the VOC profile of M4-96. However, supplementation of the plant growth medium with purified acetoin did not induce the strong biostimulant effect observed in seedlings exposed to bacterial volatiles. Analysis of the type of volatiles emitted by the strain revealed mainly ketones and alcohols, which are very common among bacteria and fungi from soil. Therefore, the precise identity of the compound-(s) responsible for the strong B. methylotrophicus phytostimulation remains unknown.
Acetoin and its analogues have been previously reported as the major GC peaks in two PGPR, B. subtilis GB03 and B. amyloliquefaciens IN937a, which promote plant growth through the emission of VOCs (Ryu et al. 2003; Farag et al. 2006). Dose-response curves of 2,3-butanediol have revealed its effects as a bioactive compound in the promotion of plant growth, but the role of acetoin in the configuration of root system architecture remains uncertain. Our data strengthen these previous findings by showing the ubiquity of acetoin in PGPR volatile blends. It is possible that this molecule acts as part of the “molecular fingerprint” that plants recognize from their microbiomes, and consequently, it might act as a switch to activate root developmental programs.
References
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ (1990) Basic local alignment search tool. J Mol Biol 215:403–410. doi:10.1016/S0022-2836(05)80360-2
Amann RI, Ludwig W, Schleifer KH (1995) Phylogenetic identification and in situ detection of individual microbial cells without cultivation. Microbiol Rev 59:143–169
Borshchevskaya LN, Kalinina AN, Sineokii SP (2013) Design of a PCR test based on the gyrA gene sequence for the identification of closely related species of the Bacillus subtilis group. Appl Biochem Microbiol 49:646–655. doi:10.1134/S0003683813070028
Brink S (2016) Unlocking the secrets of the rhizosphere. Trends Plant Sci 21:169–170. doi:10.1016/j.tplants.2016.01.020
Casimiro I, Marchant A, Bhalerao RP, Beeckman T, Dhooge S, Swarup R, Graham N, Inzé D, Sandberg G, Casero PJ, Bennett M (2001) Auxin transport promotes Arabidopsis lateral root initiation. Plant Cell 13:843–852. doi:10.1105/tpc.13.4.843
Dunlap CA, Kim SJ, Kwon SW, Rooney AP (2015) Phylogenetic analysis shows that Bacillus amyloliquefaciens subsp. plantarum is a later heterotypic synonym of Bacillus methylotrophicus. Int J Syst Evol Microbiol 65:2104–2109. doi:10.1099/ijs.0.000226
Ellingsoe P, Johnsen K (2002) Influence of soil sample sizes on the assessment of bacterial community structure. Soil Biol Biochem 34:1701–1707. doi:10.1016/S0038-0717(02)00156-6
Farag M, Ryu C, Summer L, Paré P (2006) GC-MS SPME profiling of rhizobacterial volatiles reveals prospective inducers of growth promotion and induced systemic resistance in plants. Phytochemistry 67:2262–2268. doi:10.1016/j.phytochem.2006.07.021
Fukaki H, Okushima Y, Tasaka M (2007) Auxin-mediated lateral root formation in higher plants. Int Rev Cytol 256:111–137. doi:10.1016/S0074-7696(07)56004-3
Garnica-Vergara A, Barrera-Ortiz S, Muñoz-Parra E, Raya-González J, Macías-Rodríguez L, Ruiz-Herrera L, López-Bucio J (2016) The volatile 6-pentyl-2H-pyran-2-one from Trichoderma atroviride regulates Arabidopsis root morphogenesis via auxin signaling and ethylene insensitive 2 functioning. New Phytol 209:1496–1512. doi:10.1111/nph.13725
Geldner N, Friml J, Stierhof YD, Jurgens G, Palme K (2001) Auxin transport inhibitors block PIN1 cycling and vesicle trafficking. Nature 413:425–428. doi:10.1038/35096571
Gil P, Dewey E, Friml J, Zhao Y, Snowden KC, Putterill J, Palme K, Estelle M, Chory J (2001) BIG: a calossin-like protein required for polar auxin transport in Arabidopsis. Genes Dev 15:1985–1997. doi:10.1101/gad.905201
Gutiérrez-Luna FM, López-Bucio J, Altamirano-Hernández J, Valencia-Cantero E, Reyes de la Cruz H, Macías-Rodríguez L (2010) Plant growth-promoting rhizobacteria modulate root system architecture in Arabidopsis thaliana through volatile organic compound emission. Symbiosis 51:75–83. doi:10.1007/s13199-010-0066-2
Hernández-Salmerón J, Hernández-León R, Orozco-Mosqueda MC, Valencia-Cantero E, Moreno-Hagelsieb G, Santoyo G (2016) Draft genome sequence of the biocontrol and plant growth-promoting rhizobacteria Pseudomonas fluorescens strain UM270. Stand Genomic Sci 11:5. doi:10.1186/s40793-015-0123-9
Ljung K (2013) Auxin metabolism and homeostasis during plant development. Development 140:943–950. doi:10.1242/dev.086363
Lambrecht M, Okon Y, Van de Broek A, Vanderleyden J (2000) Indole-3-acetic acid: a reciprocal signalling molecule in bacteria-plant interaction. Trends Microbiol 8:298–300. doi:10.1016/S0966-842X(00)01732-7
Leyser O (2010) The power of auxin in plants. Plant Physiol 154:501–505. doi:10.1104/pp.110.161323
Madhaiyan M, Poonguzhali S, Kwon SW, Sa TM (2010) Bacillus methylotrophicus sp. nov., a methanol-utilizing, plant-growth-promoting bacterium isolated from rice rhizosphere soil. Int J Syst Evol Microbiol 60:2490–2495. doi:10.1099/ijs.0.015487-0
Malamy JE, Benfey PN (1997) Organization and cell differentiation in lateral root of Arabidopsis thaliana. Development 124:33–44
Mehta P, Walia A, Kakkar N, Shirkot C (2014) Tricalcium phosphate solubilisation by new endophyte Bacillus methylotrophicus CKAM isolated from apple root endosphere and its plant growth-promoting activities. Acta Physiol Plant 36:2033–2045. doi:10.1007/s11738-014-1581-1
Peer WA, Hosein FN, Bandyopadhyay A, Bandyopadhyay A, Makam SN, Otegui MS, Lee GJ, Blakeslee JJ, Cheng Y, Titapiwatanakun B, Yakubov B, Bangari B, Murphy AS (2009) Mutation of the membrane-associated M1 protease APM1 results in distinct embryonic and seedling developmental defects in Arabidopsis. Plant Cell 21:1693–1721. doi:10.1105/tpc.108.059634
Pieterse CMJ, de Jonge R, Berendsen RL (2016) The soil-borne supremacy. Trends Plant Sci 21:171–173. doi:10.1016/j.tplants.2016.01.018
Raya-González J, Velázquez-Becerra C, Barrera-Ortiz S, López-Bucio J, Valencia-Cantero E (2017) N, N-dimethyl hexadecylamine and related amines regulate root morphogenesis via jasmonic acid signaling in Arabidopsis thaliana. Protoplasma 254:1399–11410. doi:10.1007/s00709-016-1031-6
Rückert C, Blom J, Chen X, Reva O, Borriss R (2011) Genome sequence of B. amyloliquefaciens type strain DSM7T reveals differences to plant-associated B. amyloliquefaciens FZB42. J Biotechnol 155:8–85. doi:10.1016/j.jbiotec.2011.01.006
Ryu CM, Farag MA, Hu CH, Reddy MS, Wei HX, Paré PW, Kloepper JW (2003) Bacterial volatiles promote growth in Arabidopsis. Proc Natl Acad Sci U S A 100:4927–4932. doi:10.1073/pnas.0730845100
Sabatini S, Beis D, Wolkenfelt H, Murfett J, Guilfoyle T, Malamy J, Benfey P, Leyser O, Bechtold N, Weisbeek P, Scheres B (1999) An auxin-dependent distal organizer of pattern and polarity in the Arabidopsis root. Cell 99:463–472. doi:10.1016/S0092-8674(00)81535-4
Sharma P, Noronha S (2014) Comparative assessment of factors involved in acetoin synthesis by Bacillus subtilis 168. ISRN Microbiol 2014. doi:10.1155/2014/578682
Spaepen S, Vanderleyden J, Remans R (2007) Indole-3-acetic acid in microbial and microorganism-plant signaling. FEMS Microbiol Rev 31:425–448. doi:10.1111/j.1574-6976.2007.00072.x
Stepanova A, Hoyt J, Hamilton A, Alonso JM (2005) A link between ethylene and auxin uncovered by the characterization of two root-specific ethylene-insensitive mutants in Arabidopsis. Plant Cell 17:2230–2242. doi:10.1105/tpc.105.033365
Tamura K, Stecher G, Peterson D, Filipski A, Kumar S (2013) MEGA6: molecular evolutionary genetics analysis version 6.0. Mol Biol Evol 30:2725–2729. doi:10.1093/molbev/mst197
Ulmasov T, Murfett J, Hagen G, Guilfoyle T (1997) Aux/IAA proteins repress expression of reporter genes containing natural and highly active synthetic auxin response elements. Plant Cell 9:1963–1971. doi:10.1105/tpc.9.11.1963
Valencia-Cantero E, Hernandez-Calderón E, Velásquez-Becerra C, López-Meza JE, Alfaro-Cuevas R, López-Bucio J (2007) Role of dissimilatory fermentative iron-reducing bacteria in Fe uptake by common bean (Phaseolus vulgaris L.) plants grown in alkaline soil. Plant Soil 291:263–273. doi:10.1007/s11104-007-9191-y
Vogel G (2006) Auxin begins to give up its secrets. Science 313:1230–1231. doi:10.1007/s11240-008-9419-4
Woodward AW, Bartel B (2005) Auxin: regulation, action, and interaction. Ann Bot 95:707–735. doi:10.1093/aob/mci083
Zhang H, Kim M, Krishnamachari V, Payton P, Sun Y, Grimson M, Farag M, Ryu C, Allen R, Melo I, Paré P (2007) Rhizobacterial volatile emissions regulate auxin homeostasis and cell expansión in Arabidopsis. Planta 226:839–851. doi:10.1007/s00425-007-0530-2
Acknowledgements
We gratefully acknowledge the Consejo Nacional de Ciencia y Tecnología (CONACYT) (grant number 165738) and the Consejo de la Investigación Científica (UMSNH) (grant number 2.24) for financial support. P. Pérez-Flores is indebted to CONACYT for providing a Master’s fellowship.
Author information
Authors and Affiliations
Corresponding author
Additional information
Handling Editor: Néstor Carrillo
Electronic supplementary material
Fig S1
(TIFF 1566 kb)
Rights and permissions
About this article
Cite this article
Pérez-Flores, P., Valencia-Cantero, E., Altamirano-Hernández, J. et al. Bacillus methylotrophicus M4-96 isolated from maize (Zea mays) rhizoplane increases growth and auxin content in Arabidopsis thaliana via emission of volatiles. Protoplasma 254, 2201–2213 (2017). https://doi.org/10.1007/s00709-017-1109-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00709-017-1109-9