Abstract
Downy mildew is a highly destructive disease in grapevine production. A gene encoding pathogenesis-related (PR) thaumatin-like protein was isolated from the downy mildew-resistant grapevine “Zuoshan-1,” a clonal selection from wild Vitis amurensis Rupr. The predicted thaumatin-like protein (VaTLP) has 225 amino acids and it is acidic, with a calculated isoelectric point of 4.8. The full length of the VaTLP gene was transformed into somatic embryogenic calli of V. vinifera ‘Thompson Seedless’ via Agrobacterium tumefaciens. Real-time RT-PCR confirmed that the VaTLP gene was expressed at a high level in the transgenic grapevines. Improved resistance of the transgenic lines against downy mildew was evaluated using leaf disks and whole plants inoculated with Plasmopara viticola, the pathogen causing grapevine downy mildew disease. Bioassay of the pathogen showed that both hyphae growth and asexual reproduction were inhibited significantly among the transgenic plants. Histological analysis also confirmed this disease resistance by demonstrating the inhibition and malformation of hyphae development in leaf tissue of the transgenic plants. These results indicated that the accumulation of VaTLP could enhance resistance to P. viticola in transgenic ‘Thompson Seedless’ grapevines.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Downy mildew (DM), caused by oomycete Plasmopara viticola, is one of the most devastating diseases of grapes growing under warm and humid conditions, leading to decreases in yield and quality. The propagation of P. viticola consists of sexual and asexual sporulated stages. The asexual cycle begins in the summer when the germinated oospores penetrate the stomata via germ tubes, enter the mesophyll tissue, and form an intercellular hyphal network (Kiefer et al. 2002). Sporangiophores then emerge and asexually produce sporangia and zoospores in the susceptible grapevines. Sexual sporulation produces diploid oospores in host tissues during the autumn (Koch and Slusarenko 1990; Stark-Urnau et al. 2000). Vitis vinifera grapes are highly susceptible to downy mildew, while resistant sources are identified in Oriental and North American Vitis species (Yu et al. 2012). Integration of the resistance traits into V. vinifera grape cultivars by conventional breeding is limited due to the highly heterozygous nature and long vegetative growth cycle of grapevines and disruption of the desired phenotype for varieties especially used in wine production. Genetic engineering, as an alternative tool, has been used to introduce resistance genes through Agrobacterium-mediated transformation without altering the essential characters of the cultivar in question (Maghuly et al. 2006; Fan et al. 2008).
Plants can trigger complex defense mechanisms, including induction of pathogenesis-related (PR) genes, to counter the attack of pathogen. PR proteins are induced by pathogens, chemical elicitors, or, in some instances, environmental stresses. Overexpression of one or more PR proteins can delay disease development (Hammond-Kosack and Jones 1996). Thaumatin-like proteins (TLPs) share sequence similarity with thaumatin (Velazhahan et al. 1999), a sweet-tasting protein from Thaumatococcus daniellii (Benth.) (Van der Wel and Loewe 1972). TLPs belong to pathogenesis-related (PR) protein family 5 due to their inducible expression by pathogen/pest attack (Christensen et al. 2002; van Loon et al. 2006). PR-5 proteins are reported to have antifungal properties (Roberts and Selitrennikoff 1990; Abad et al. 1996; Cheong et al. 1997). PR proteins are secreted through the plasmalemma and present in intercellular spaces (Wagih and Coutts 1981). The mechanism of anti-oomycete activity by PR5 protein is likely based on β-1,3-glucan binding and endo-β-1,3-glucanase activity (Abad et al. 1996; Grenier et al. 1999; Roberts and Selitrennikoff 1990; Trudel et al. 1998). The extracellular PR5 proteins from barley and pea plants bind to water-insoluble β-1,3-glucans (Trudel et al. 1998) and six other PR5 proteins are also found to be active against carboxymethyl pachyman (Grenier et al. 1999). Unlike filamentous fungi that have a chitin-glucan cell wall, oomycetes can be regarded as cellulosic microorganisms because their cell walls contain little or no chitin but are rich in β-1,3-glucan polymers and cellulose (Zevenhuizen and Bartnicki-Garcia 1968). It is predicted they will not be detected by the plant’s chitin-fragment surveillance system, nor will they be affected by chitinases, but can be detected by β-1,3-glucanases secreted by the plant. These components of the cell wall could be involved in PR5 protein-oomycete interactions (Grenier et al. 1999; Trudel et al. 1998).
TLPs are expressed in different plant tissues, such as pistils, fruits, and seeds (Neale et al. 1990). They are also accumulated in fruit during ripening (Vu and Huynh 1994; Fils-Lycaon et al. 1996; Barre et al. 2000; Tattersall et al. 1997; Pocock et al. 2000). TLPs are induced by pathogen, abiotic stresses (cold, drought and salinity), wounding, and plant hormones (Liu et al. 2010). Other observations suggest that some TLPs may be involved in physiological processes, such as organ-specific or development-dependent patterns, other than various environmental stresses (Regalado and Ricardo 1996; Skadsen et al. 2000; Van Damme et al. 2002; Sassa et al. 2002). The roles of TLPs in host defense have been tested in transgenic overexpression or in vitro antifungal activity (Liu et al. 2010).
In grapevine, TLPs from V. vinifera ‘Moscatel’ have been reported to display a strong antifungal activity against Uncinula necator (powdery mildew), Phomopsis viticola (dead-arm), and Botrytis cinerea (noble rot) in vitro (Monteiro et al. 2003). A TLP (VVTL-1) from V. vinifera ‘Chardonnay’ significantly inhibits in vitro spore germination and hyphae growth of Elsinoe ampelina (Jayasankar et al. 2003). When this VVTL-1 gene was transferred to susceptible V. vinifera ‘Thompson Seedless,’ the overexpressed transgenic vines showed resistance to anthracnose and powdery mildew (Dhekney et al. 2011). The objective of this research was to isolate and characterize the VaTLP gene from ‘Zuoshan-1,’ a downy mildew-resistant clonal selection from a wild V. amurensis grapevine and to determine its potential roles against downy mildew disease in susceptible grapevines.
Materials and methods
Plant and pre-embryogenic calli cultivation
Grapevines of V. amurensis ‘Zhuoshan-1’ and V. vinifera ‘Thompson Seedless’ were grown in the Shangzhuang Agricultural Experimental Station, China Agricultural University, Beijing, China. Pre-embryogenic calli of ‘Thompson Seedless’ originated from anther culture was provided by the Transformation Facility of the University of California, Davis, USA.
Inoculation of P. viticola and assessment of TLP expression levels
One-year-old ‘Zuoshan-1’ and ‘Thompson Seedless’ seedlings were grown in the greenhouse under a 16-h light/8-h dark photoperiod at 25 °C, 85% relative humidity. The DM inoculation operation procedure has been established and used in our lab (Wu et al. 2010). The fourth unfolded leaf from the shoot apex was harvested from each of the three vines, and the three leaves were combined to represent one replicate. Three independent replicates were collected for each sample. The infected leaves were collected at 0, 8, 12, 24, 48, and 96 h post-inoculation (hpi). Control samples were harvested from water-treated leaves incubated under the same conditions.
Total RNA was isolated from the samples mentioned above using a modified cetyltrimethylammonium bromide (CTAB) method as presented by Murray and Thompson (1980). Experiments were carried out on three independent biological replicates each containing three technical replicates. First-strand cDNA was synthesized from 1000 ng DNase (Promega, Madison, Wisconsin, USA)-treated total RNA using “ImProm-II TM Reverse Transcriptase” (Promega, Madison, Wisconsin, USA). The reactions were performed using a Roto-Gene Q Real-time PCR (QIAGEN, Hilden, Germany) in a 10-μL reaction mixture containing 5-μL SYBR Green Supermix, 0.2 μL of 10 μM primer, and 1-μL cDNA and 3.6-μL ddH2O. The primers for Vitis TLP (forward: 5′- ACCATTGCTCCTACACGGTT-3′; reverse: 5′-CTTCCCATTCCCTGACGCAT-3′) were used to detect the expression levels of total VaTLP and VvOsm in the transgenic grapevines. The primers which align to VaTLP and nos terminator (forward: 5′- TGCACTGCCGGTACCAATTA-3′; reverse: 5′- AGACCGGCAACAGGATTCAA-3′) were used to access the expression levels of exogenous VaTLP in the transgenic grapevines. The primers which align to VvOSM and its 3′UTR (forward: 5′-CAAGCACCTTCACATGCCCT-3′; reverse: 5′-AGGTGGATACCATTGCCTCCT-3′) were used to check the endogenous VvOSM expression levels. Vitis EFα (XM_002284888) f primer: 5′-TCCAAGGCAAGGTACGATG-3′; reverse primer: 5′- CAGAGATGGGGACAAATGG-3′) was used as reference gene for data normalization (Li and Wu et al. 2015). The thermal cycling conditions were as follows: 10 min at 95 °C, followed by 40 cycles of 10 s at 95 °C, 15 s at 58 °C, and 30 s at 72 °C. The specificity of the individual PCR amplifications was assessed using heat dissociation curves from 55 to 95 °C following the final cycle of the PCR. The-fold change in mRNA expression was estimated using threshold cycles, by the 2−ΔΔCT method (Livak and Schmittgen 2001).
Isolation and characterization of VaTLP
The cane tissues of ‘Zhuoshan-1’ were collected and used for the isolation and analysis of the VaTLP gene. Total genomic DNA of ‘Zhuoshan-1’ was isolated by the CTAB method (Murray and Thompson 1980). To amplify the VaTLP gene sequence, two oligonucleotide primers were designed based on the UTR region sequences identified in the Vitis genome database (http://www.genoscope.cns.fr/cgi-bin/blast_server/projet_ML/blast.pl). The forward primer UTRF was 5′-CCATCAAGCCTATCTTCCGAT-3′, and the reverse primer UTRR was 5′-CGTGTTTGGACTGCTACAAT-3′. The PCR was performed in a 50-μL reaction volume containing 50 ng of genomic DNA, 1 μM each primer, 200 μM dNTPs, 5 μL of Pfu DNA Polymerase 10× reaction buffer with MgSO4, and 1.25 U Pfu DNA polymerase (Promega, Beijing, China). The thermal cycling conditions were as follows: 95 °C for 2 min, followed by 35 cycles at 95 °C for 1 min, 58 °C for 30 s, 72 °C for 2 min, and a final extension at 72 °C for 5 min. PCR products were isolated from the agarose gel blocks after electrophoresis and purified using QIAquick Gel Extraction Kit (Qiagen, Valencia, CA, USA). The purified DNA was cloned into the pGEM T-Easy vector (Promega, Beijing, China) and sequenced. Identification of open reading frames (ORFs) in the amplified DNA sequence was carried out by using ORF finder at NCBI (www.ncbi.nlm.nih.gov). The predicted protein sequences alignments were performed with DNAman 6.0 software (Lynnon Biosoft LLC., San Ramon, CA, US).
Construction of a binary vector and transformation into Agrobacterium tumefaciens
The specific primers TLP-F (5′-ATAGGATCCATGGGCCTCTGCAAAATC-3′) and TLP-R (5′ -TATGAGCTCTTATGGGCAGAAGACAAC-3′) were used to amplify the full-length ORF of VaTLP with Pfu DNA polymerase (Promega Corporation, Madison, WI, US). After double digestion with BamHI and SacI (New England Biolabs, Inc., MA, USA), the fragment was introduced into the binary vector pBI121 (Clontech Labs Inc., Palo Alto, USA) as a BamHI-SacI fragment to replace the gus reporter gene (Jefferson et al. 1987) (Fig. 1). The new construct pBI121-VaTLP contains the nptII gene driven by the nos promoter and the VaTLP gene driven by the CaMV 35S promoter. This new binary vector was transformed into Escherichia coli and then introduced into the disarmed A. tumefaciens strain EHA105 (Hood et al. 1993) using the electroporation method.
Transformation of VaTLP into ‘Thompson Seedless’
The pBI121-VaTLP constructed A. tumefaciens was cultured at 28 °C overnight in liquid Luria-Bertani (LB) medium (10 g L−1 peptone, 5 g L−1 yeast extract, 5 g L−1 sodium chloride) containing 25 mg L−1 kanamycin. The cells were centrifuged for 3 min (5000×g), and the pellet was re-suspended to approximately 1 × 108 cells mL−1 in liquid MS medium (Murashige and Skoog 1962) supplemented with 20 μM acetosyringone (PhytoTechnology Laboratories, Overland Park, KS) . The re-suspended A. tumefaciens cells were inoculated into the ‘Thompson Seedless’ pre-embryogenic calli (Agüero et al. 2006). The calli were then transferred onto co-cultivation medium (same as PT medium but lacking activated charcoal and supplemented with 4 μM picloram, 2.3 μM TDZ, and 100 μM acetosyringone). After co-cultivation for 48 h, the calli were subdivided into small clusters approximately 2 mm in diameter and selected for 4 months on the PT medium with 100 μg mL−1 kanamycin and 300 μg mL−1 cefotaxime. Plantlets were regenerated on modified WP medium (McCown and Lloyd 1981), and the established putative transformed plantlets were then transferred to the green house cultured in the 0.5-L pots with a potting mixture consisting of vermiculite and soil (1:1, v/v).
Molecular verification of transformants
To validate the putative transformed plantlets, genomic DNA was isolated from young leaves of each putative transgenic plantlet using a modified CTAB method (Murray and Thompson 1980). The primers (nos-F: 5′-ATTGCGGGACTCTAATCATA-3′, nos-R: 5′-ATCGTTCAAACATTTGGCA-3′) were chosen for detection of the 277 bp of nos terminator sequence. The plasmid of binary vector pBI121-VaTLP and the DNA of non-transformed plant were used for positive and negative control, respectively.
Southern blot was used to further validate the stable integration of the transgene. Genomic DNA (10 μg) was extracted from PCR-positive and non-transformed plants with CTAB method and further purified with the PS Erasol kit (Tiandz Inc., Beijing, China). The DNA was digested overnight with HindIII (New England Biolabs, Inc., MA, US), separated on 1% agarose gel, and transferred to a positively charged nitrocellulose membrane. The nptII probe labeled with digoxigenin-dUTP by using the DIG DNA Labeling and Detection Kit (Mylab, Beijing, China) was used for the Southern blot hybridization.
Assessment of resistance to downy mildew among the transgenic grapevines
The fourth and fifth leaves from shoot tips among the transformed and control vines were excised, rinsed with distilled water, and dried with filter paper. Leaf disks (9 mm in diameter) were obtained by a cork borer and placed (bottom side up) on 0.8% water agar in petri dishes. Fresh P. viticola was cultured on the ‘Thompson Seedless’ leaves in the growth chamber with 25 ± 1 °C and long-day condition (light 16 h, darkness 8 h). The sporangia were collected in centrifuge tubes using a small paintbrush and re-suspended in distilled water. With three biological repeats, a total of 30 leaf disks from each transformed and non-transformed plants were used for P. viticola inoculation. Each leaf disk was inoculated with a 30-μL drop of sporangia suspension (5 × 105 sporangia per mL) and incubated at 25 ± 1 °C under dark for 24 h, and then the drops on disks were removed by filter paper and cultured under long-day condition. The percentage of sporulation on leaf disks and the reproduction of P. viticola were assessed at 8 days post-inoculation (dpi) (Yu et al. 2012). The number of sporangia was counted using a hemocytometer, and the leaf disk areas exhibiting symptoms of sporulation were measured using a digital camera and Adobe Photoshop CS2 software (Adobe Systems, San Jose, USA) (Xiao et al. 2005; Yu et al. 2012).
For the whole plant, certified virus-free seedlings were grown and maintained in the greenhouse under a 16 h light/8 h dark photoperiod at 25 °C, 85% relative humidity. The fourth and fifth leaves from the apex were sprayed with sporangia suspension of P. viticola (5 × 105 sporangia per mL). The inoculated leaves were covered with plastic bags for the first night to ensure high humidity. The symptoms of infection were observed at 8 dpi. Deionized water was applied as control.
Microscopy observation
Fluorescence microscopy was used to observe the intercellular infection structures of hyphae growth. Leaf disks were collected at 0, 1, 3, 5, and 8 dpi and treated using the KOH-aniline blue fluorescence method (Hood and Shew 1996). After processing, the disks were observed under a fluorescence microscope (Olympus BX51), and the images were processed with Adobe Photoshop CS2 software. To ensure reproducibility, each specimen was observed for three times. Deionized water drops were used as control.
Statistical analysis
Data were analyzed by one-way ANOVA followed by Duncan’s multiple-range test. All statistical analysis was performed at the value of p < 0.05 using SPSS 17.0 (SPSS Inc., USA).
Results
TLP response to P. viticola infection in grapevines
Expression profiles of TLP were measured in grapevine leaves infected with P. viticola pathogen in DM-susceptible V. vinifera ‘Thompson Seedless’ and resistant V. amurensis ‘Zuoshan-1’ at 0, 8, 12, 24, 48, and 96 hpi (Fig. 2). Expression of TLP was induced in both genotypes during the infection process. However, transcription of VvTLP in ‘Thompson Seedless’ was significantly induced (572-fold) at 8 hpi and reached a maximum level (5726-fold) at 12 hpi, then declined to 2725-fold at 24 hpi and remained very high expression at 48 hpi (2425-fold) and 96 hpi (3000-fold). In ‘Zuoshan-1,’ the expression of VaTLP also up-regulated after the P. viticola infection and reached a maximum level at 8 hpi (68-fold), then subsequently declined at 12 hpi (11-fold) and 24 hpi (1.6-fold). Its expression moderately increased again at 48 hpi (14-fold) and declined thereafter (6-fold at 96 hpi). Since the basal expression level (0 hpi) was 267-fold higher in ‘Zuoshan-1’ than that in ‘Thompson Seedless,’ the expression abundance of VaTLP was still much more than VvTLP after the pathogen infection at 8 hpi (31-fold higher), even though the-fold increase of TLP stimulated by the pathogen infection was smaller in ‘Zuoshan-1’ than in ‘Thompson Seedless.’ This result suggests that the basal expression level of TLP and fast response to P. viticola infection may play an important role in the early protection against P. viticola infection.
Expression of TLP in response to P. viticola inoculation. Real-time RT-PCR analysis of TLP expression in response to P. viticola inoculation after 0, 8, 12, 24, 48, or 96 h in ‘Zuoshan-1’ and ‘Thompson Seedless’ leaves. All values were normalized to the expression level of the Vitis EFα gene. The normalized expression level of VvTLP in ‘Thompson Seedless’ at 0 h was taken as control (expression set to 1). Bars represent standard deviation calculated from three biological replicates. Asterisks indicate that in comparison with 0 hpi for ‘Thompson Seedless’ and ‘Zuoshan-1,’ respectively, the expression levels of TLP are significantly increased at other time points by t test (P < 0.05)
Cloning and structural characterization of VaTLP
The full length of VaTLP was amplified from ‘Zuoshan-1’ gDNA and the sequence was analyzed using DNAman software. The gene was 1223 bp in length, encoding an open reading frame (ORF) of 678 nucleotides (GenBank accession number KX397573) which was 97% identical to PR5 of ‘Thompson Seedless’ (VvOSM) (Fig. 3a). VaTLP encoded a protein of 225 amino acids with a molecular weight of 23.84 kDa and a predicted isoelectric point of 4.8. The predicted protein sequence was 95% identical to VvOSM of ‘Thompson Seedless’ and was distant (61 to 80% identity) to PR5 from other plants (Musa acuminate, Nicotiana tabacum, and T. daniellii), suggesting that VaTLP is a member of a thaumatin-like protein family. All the functional domains, including TLP family signature (G-x-[GF]-x-C-x-T-[GA]-D-C-x(1,2)-[GQ]-x(2,3)-C), loop domains, pivotal amino acids in the acidic cleft and cysteine residues, were all conserved between VaTLP and VvOSM (Fig. 3b).
Sequence analysis of VaTLP. a Alignment of nucleotide sequences of VaTLP (V. amurensis) and VvOSM (V. vinifera). b Alignment of amino acid sequences among TLP proteins from different plants. The employed proteins are VaTLP (KX397573) from V. amurensis, VvOSM (Y10992.1) from V. vinifera, MaPR5 (1Z3Q_A) from M. acuminate, NtPR5 (1AUN_A) from N. tabacum, and TdTHAUMATIN (AB265690) from T. daniellii. Conserved residues of TLP family signature, G-x-[GF]-x-C-x-T-[GA]-D-C-x(1,2)-[GQ]-x(2,3)-C, are boxed. Loop domains II and III are underlined in black and gray, respectively. The five conserved amino acid positions in the acidic cleft are indicated with asterisks. The conserved cysteine residues are indicated with a black background
Development and confirmation of transgenic grapevines
The VaTLP gene was transformed into ‘Thompson Seedless’ pre-embryogenic calli by A. tumefaciens-mediated method. After 2 months of selection by kanamycin, the putative transgenic calli exhibited a white color and rapid growth on the selection medium (Fig. 4a), while the non-transgenic calli, by contrast, were black and dead (Fig. 4b). After 4 months of selection, a total of 37 transgenic calli clusters were selected from 600 clusters growing on the kanamycin-added medium. Six putative VaTLP transgenic lines were successfully regenerated from these 37 selected calli transformants (Fig. 4c, d).
Transformation process of ‘Thompson Seedless.’ a, b Proliferated putatively transformed calli on the selection medium at 2 months after transformation (bar 5.0 mm). White arrow shows dead untransformed calli. c Transformed somatic embryos growing on germination medium after 6 weeks (bar 5.0 mm). d Putative transgenic plantlets growing in pots
To confirm the true transformants among the transgenic lines, PCR was firstly used to detect the presence of the nos terminator (Fig. 5a). Three out of the six putative transgenic lines, like the positive plasmid control, showed the expected 277-bp fragment of the nos terminator. No amplification was observed for the water and the non-transformed (NT) controls.
Validation of putative transgenic plants by PCR and Southern hybridization. a PCR amplification of nos terminator. M, DL2000 marker; H 2 O, negative control; P, plasmid pBI121-VaTLP; NT, plant regenerated from the non-transformed calli; 1–6, putative transgenic grapevine lines. b Southern hybridization of transgenic lines using nptII as a digoxin-labeled probe. M, Lamda DNA/HindIII marker; P, plasmid pBI121-VaTLP; NT, plant regenerated from the non-transformed calli; 2, 4, 6, putative transgenic grapevine lines
Southern blot was performed to further validate the integration of the foreign VaTLP gene into the ‘Thompson Seedless’ genome (Fig. 5b). The DNA from the PCR-positive and non-transformed grapevines were hybridized with the 766-bp nptII gene probe. As expected, the NT vine showed no hybridization, while the pBI121-VaTLP plasmid and the PCR-positive transgenic lines generated positive hybridization bands, indicating the VaTLP gene has been successfully transformed into ‘Thompson Seedless’ genome. The banding patterns suggested that the three transgenic lines (2, 4, and 6) represented three independent transformation events with 1, 3, and 4 insertion copies, respectively.
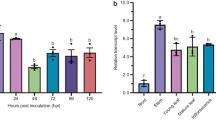
Real-time RT-PCR analysis further indicated that the total TLP expression level significantly upregulated in the transgenic line 2 (50-fold) and line 6 (32-fold) (Fig. 6). To access the expression of exogenous gene in the transgenic lines, one specific primer pair that aligns to VaTLP and the nos terminator was used to check the VaTLP expression levels in line 2 and 6. In comparison with the EFα reference gene, line 2 showed an accumulation of 56-fold of VaTLP transcripts, while line 6 was 16-fold. In addition, another primer pair specific to endogenous VvOSM (forward primer aligns to VvOSM, reverse primer aligns to its 3′UTR) was used to access the VvOSM transcription levels. It is indicated that the overexpression of exogenous VaTLP caused inhibition of endogenous VvOSM to 36% for line 2 and 72% for line 6 (Fig. 6). Unfortunately, during the review process of the paper, the transgenic line 4 was weakening. As a result, no healthy leaves of line 4 were available for the analysis of exogenous and endogenous TLP genes.
Expression analysis of the TLP gene in transgenic lines. Vitis EFα was used as an internal control. For total TLP and VvOSM expression, data represent the “fold change” in gene expression in transgenic lines (2, 4, 6) vs. non-transformed grapevine (NT). For VaTLP expression, data represent the relative fold change in VaTLP vs EFα. Bars represent standard deviation calculated from three biological replicates. Asterisks indicate that in comparison with non-transgenic (NT) vines, the fold change in transgenic lines were significantly changed by t test (P < 0.05)
Evaluation of transformed grapevines against downy mildew disease
The leaf disks of ‘Zuoshan-1,’ VaTLP transgenic, and non-transgenic ‘Thompson Seedless’ were inoculated with P. viticola at a concentration of 5 × 105 spores per mL pure water. White hyphae were observed on all the leaf disks at 3–5 dpi. However, much less severe disease symptoms were observed on the transgenic lines than in the non-transgenic ‘Thompson Seedless’ at 8 dpi (Fig. 7). The non-transgenic vines had 84% of leaf surface infected, while the transgenic ones had 42–49% of leaf surface developing the P. viticola symptom, very similar to the P. viticola-resistant V. amurensis ‘Zuoshan-1’ (44%) (Fig. 8). Consistently, the transgenic vines had significantly less number of P. viticola sporangia than did non-transgenic vines, with 0.03 × 105 spores·cm−2 in line 2, 0.05 × 105 spores·cm−2 in line 4, and 0.09 × 105 spores·cm−2 in line 6 in comparison with the 0.41 × 105 spores·cm−2 in the wild type (Fig. 9).
Percentage of the P. viticola-infected leaf area at 8 dpi in the transformed lines (2, 4, 6), non-transformed (NT), and ‘Zuoshan-1’ (ZS-1) vines. Bars represent standard deviation calculated from three biological replicates. Asterisks indicate that in comparison with non-transgenic (NT), the infected areas of transgenic lines or ‘Zuoshan-1’(ZS-1) were significantly different (P < 0.05)
Density of P. viticola spores in the transgenic, non-transgenic, and ‘Zuoshan-1’ vines. Bars represent standard deviation calculated from three biological replicates. Asterisks indicate that in comparison with non-transgenic (NT), the spore densities of transgenic lines or ‘Zuoshan-1’(ZS-1) were significantly different (P < 0.05)
The improved disease resistance of the transgenic vines was further evaluated using the potted vines growing in greenhouse. After spraying with P. viticola suspension, the transgenic lines showed less severe developing of P. viticola symptom (Fig. 7). The spore density in the non-transgenic plants was significantly higher (3.15 × 105 spores·cm−2) than that in the transgenic lines (0.98 × 105 spores·cm−2 for line 2, 1.45 × 105 spores·cm−2 for line 4, and 1.28 × 105 spores·cm−2 for line 6 (Fig. 9).
After the KOH-aniline blue staining, webbed hyphae with haustoria were observed throughout the mesophyll at 3 dpi in both control and transgenic vines (Fig. 10). During 3 to 5 dpi, complex sporangiophores with or without sporangia were observed. In contrast to the control plant (Fig. 10a, b), the growth of hyphae arrestment was only observed at 3 dpi in the transgenic lines (Fig. 10c–f). After the primary hyphae had generated, the oomycetes in the transgenic lines developed into malformed hyphae, showing incomplete, small, ball-like hyphae around their sub-stomatal vesicle.
Colonization of non-transformed leaf disks and transgenic leaf disks infected by P. viticola at 3 dpi. Mycelia were visible in both non-transgenic (NT; a, b) and transgenic (c, d, e, f) leaf disks. a, c, e Overview of hyphae development in NT (a) and transgenic (c, e) leaf disks. b, d, f Enlarged view of hyphae colonization in NT (b) and transgenic (d, f) leaf disks. The arrestment of hyphae was observed in transgenic leaf (white arrow) (bar 50 μm)
Discussion
In the process of pathogen invasion, phytoalexins, PR proteins, and other toxins are secreted to stop entrance of the pathogen accompanied by the presence of necrosis, which is regarded as a common phenomenon in the activation of systemic resistance when plants encounter pathogens. These responses form the foundation of basal resistance but their effectiveness varies depending mainly on the genetic sources of the interacting organisms. For example, different grape species and cultivars display different responses to downy mildew infection (Yu et al. 2012). In our study, the moderately resistant V. amurensis ‘Zuoshan-1’ presented local necrosis after P. viticola infection, while susceptible V. vinifera ‘Thompson Seedless’ did not.
Although there have been a large number of papers reporting expression of PR genes for enhancing disease resistance (Velazhahan and Muthukrishnan 2003; Schestibratov and Dolgov 2005; Mackintosh et al. 2007; Rajam et al. 2007), there remain only a few reports on transgenic plants including tobacco, orange, and potato expressing PR genes to enhance resistance against oomycete pathogens (Alexander et al. 1993; Liu et al. 1994; Fagoaga et al. 2001). The present study reports for the first time the resistant function of TLP gene on oomycete in grapevine. V. amurensis VaTLP, a PR5 cisgene of Vitis, may play an important role during the establishment of DM resistance in ‘Zuoshan-1.’ When ‘Zuoshan-1’ was inoculated with P. viticola, VaTLP was highly induced, especially at the early infection stage (8 hpi). Similarly, induction of VvTLP was also detected in the ‘Thompson Seedless’-P. viticola interaction. It is indicated that VvTLP and VaTLP were both involved in the basic response to downy mildew infection. However, it is noticed that the basal expression level (0 hpi) and early induction (8 hpi) of TLP were much higher in ‘Zuoshan-1’ than those in ‘Thompson Seedless.’ It is supposed to be an important self-defense mechanism to help the host degrade pathogen cell wall due to β-1,3-glucanase activity and suppress the preliminary colonization of the spores, ultimately protecting the vines from further infection and damage by the pathogen.
When VaTLP was transformed to V. vinifera ‘Thompson Seedless,’ the transgenic vines showed improved downy mildew resistance. In comparison with the non-transgenic vines, the transgenic lines showed remarkably reduced intensity of sporulation and limited spread of the pathogen, indicating the pathogen was inhibited and ceased to develop to other healthy leaf areas due to VaTLP accumulation. The transgenic line 2, with a single copy of the insertion gene, presented the highest VaTLP expression and disease resistance among the three transgenic lines. The transgenic lines presented developmental malformations of the pathogen, showing hyphae arrestment and numerous incomplete, small, ball-like hyphae around their sub-stomatal vesicle. It is indicated that accumulation of VaTLP protein could strongly improve disease resistance of transgenic vines through induction of pathogen malformations. The increased expression levels of TLP was observed in the transgenic plants which may trigger the TLP-oomycete interactions to degrade the cell wall of the P. viticola, leading to the hyphae arrestment and suppression of the pathogen. Enhancement of fungal resistance by overexpressing TLP genes in transgenic plants has been reported in previous studies. For example, overexpression of the rice TLP gene in tobacco plants shows an enhanced resistance to A. alternata (Velazhahan and Muthukrishnan 2003); transgenic strawberry plants expressing the thaumatin II gene shows a higher level of resistance to B. cinerea (Schestibratov and Dolgov 2005); the bulbs of transgenic hyacinth lines present more resistance to fungi than do non-transgenic plants (Popowich et al. 2007); V. vinifera ‘Chardonnay’ overexpressing a TLP gene (VVTL-1) displays high resistance to anthracnose and powdery mildew (Dhekney et al. 2011). However, there are limited reports on the PR genes to improve resistance against oomycetes. Our study clearly indicated the effect of TLP on oomycete in grapevines, even though the suppression of P. viticola was moderate, probably due to the complex network of the disease resistance in plant, in which the accumulation of only one protein could not be sufficient to establish a broad systemic resistance.
In the present study, three transformants were analyzed for gene copy number through Southern blot. Line 2 with the lowest transgene copy number and the highest transgene expression level exhibited the highest downy mildew resistance level in comparison with that for lines 4 and 6. Similar results are reported in potato and citrus where a negative correlation exists between transgene copy number and expression levels (Chan et al. 1996; Cervera et al. 2000). On the other hand, in some studies, even though a positive correlation is found between transcript expression levels and downy mildew resistance, neither positive nor negative correlations could be found between copy number and gene expression levels (Li et al. 2003; Zanek et al. 2008; Joshi et al. 2011). The reason for this phenomenon has not been clarified. The insertion of T-DNA is random within the plant genome and the activity of the introduced genes may be affected by adjacent plant DNA (position effect). In addition, truncation, rearrangement, or repetition of the introduced T-DNA may also affect gene expression and there has not always been a direct correlation shown between copy number and gene expression.
References
Abad LR, D’Urzo MP, Liu D et al (1996) Antifungal activity of tobacco osmotin has specificity and involves plasma membrane permeabilization. Plant Sci 118:11–23
Agüero CB, Meredith CP, Dandekar AM (2006) Genetic transformation of Vitis vinifera L. cv. Thompson Seedless and Chardonnay with the pear PGIP and GFP encoding genes. Vitis 45:1–8
Alexander D, Goodman RM, Gut-Rella M et al (1993) Increased tolerance to two oomycete pathogens in transgenic tobacco expressing pathogenesis-related protein la. Proc Nati Acad Sci USA 90:7327–7331
Barre A, Peumans WJ, Menu-Bouaouiche L et al (2000) Purification and structural analysis of an abundant thaumatin-like protein from ripe banana fruit. Planta 211:791–799
Boso Alonso S, Blanco S, Luis J et al (2005) A method to evaluate downy mildew resistance in grapevine. Agron Sustain Dev 25:163–165
Cervera M, Pina JA, Juárez J et al (2000) A broad exploration of a transgenic population of citrus: stability of gene expression and phenotype. Theor Appl Genet 100:670–677
Chan MT, Chen LJ, Chang HH (1996) Expression of Bacillus thuringiensis (B.t.) insecticidal crystal protein gene in transgenic potato. Bot Bull Acad Sin 37:17–23
Cheong NE, Choi YO, Kim WY et al (1997) Purification of an antifungal PR-5 protein from flower buds of Brassica campestris and cloning of its gene. Physiol Plantarum 101:583–590
Christensen AB, Cho BH, Naesby M et al (2002) The molecular characterization of two barley proteins establishes the novel PR-17 family of pathogenesis-related proteins. Mol Plant Pathol 3:135–144
Dhekney SA, Li ZT, Gray DJ (2011) Grapevines engineered to express cisgenic Vitis vinifera thaumatin-like protein exhibit fungal disease resistance. In vitro Cell Dev 47:458–466
Fagoaga C, Rodrigo I, Conejero V et al (2001) Increased tolerance to Phytophthora citrophthora in transgenic orange plants constitutively expressing a tomato pathogenesis related protein PR-5. Mol Breeding 7:175–185
Fan C, Pu N, Wang X et al (2008) Agrobacterium-mediated genetic transformation of grapevine (Vitis vinifera L.) with a novel stilbene synthase gene from Chinese wild Vitis pseudoreticulata. Plant Cell Tiss Organ Cult 92:197–206
Fils-Lycaon BR, Wiersma PA, Eastwell KC et al (1996) A cherry protein and its gene, abundantly expressed in ripening fruit have been identified as thaumatin-like. Plant Physiol 111:269–273
Grenier J, Potvin C, Trudel J et al (1999) Some thaumatin-like proteins hydrolyse polymeric β-1,3-glucans. Plant J 19:473–480
Hammond-Kosack KE, Jones JD (1996) Resistance gene-dependent plant defense responses. Plant Cell 8:1773–1791
Hood EE, Gelvin SB, Melchers LS et al (1993) New Agrobacterium helper plasmids for gene transfer to plants. Transgenic Res 2:208–218
Hood ME, Shew HD (1996) Applications of KOH-aniline blue fluorescence in the study of plant-fungal interactions. Phytopathology 86:704–708
Jayasankar S, Li Z, Gray DJ (2003) Constitutive expression of Vitis vinifera thaumatin-like protein after in vitro selection and its role in anthracnose resistance. Funct Plant Biol 30:1105–1115
Jefferson RA (1987) Assaying chimeric genes in plants: the GUS gene fusion system. Pl Mol Biol Rep 5:387–405
Joshi SG, Schaart JG, Groenwold R et al (2011) Functional analysis and expression profiling of HcrVf1 and HcrVf2 for development of scab resistant cisgenic and intragenic apples. Plant Mol Biol 75:579–591
Jürges G, Kassemeyer HH, Dürrenberger M et al (2009) The mode of interaction between Vitis and Plasmopara viticola Berk. & Curt. Ex de Bary depends on the host species. Plant Biol 11:886–898
Kiefer B, Riemann M, Büche C et al (2002) The host guides morphogenesis and stomatal targeting in the grapevine pathogen Plasmopara viticola. Planta 215:387–393
Koch E, Slusarenko A (1990) Arabidopsis is susceptible to infection by a downy mildew fungus. Plant Cell 2:437–445
Li X, Gasic K, Cammue B et al (2003) Transgenic rose lines harboring an antimicrobial protein gene, ace-AMP1, demonstrate enhanced resistance to powdery mildew (Sphaerotheca pannosa). Planta 218:226–232
Li X, Wu J, Yin L et al (2015) Comparative transcriptome analysis reveals defense-related genes and pathways against downy mildew in Vitis amurensis grapevine. Plant Physiol Bioch 95:1–14
Liu D, Raghothama KG, Hasegawa PM et al (1994) Osmotin overexpression in potato delays development of disease symptoms. Proc Nati Acad Sci USA 91:1888–1892
Liu JJ, Sturrock R, Ekramoddoullah AKM (2010) The super family of thaumatin-like proteins: its origin, evolution, and expression towards biological function. Plant Cell Rep 29:419–436
Liu SM, Sykes SR, Clingeleffer PR (2003) A method using leafed single-node cuttings to evaluate downy mildew resistance in grapevine. Vitis 42:173–180
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using realtime quantitative PCR and the 2-ΔΔCt method. Methods 25:402–408
Mackintosh CA, Lewis J, Radmer LE et al (2007) Overexpression of defense response genes in transgenic wheat enhances resistance to Fusarium head blight. Plant Cell Rep 26:479–488
Maghuly F, Leopold S, da Camara MA et al (2006) Molecular characterization of grapevine plants transformed with GFLV resistance genes: II. Plant Cell Rep 25:546–553
McCown BH, Lloyd G (1981) Woody plant medium (WPM)—a mineral nutrient formulation for microculture of woody plant species. Hortscience 16:453
Monteiro S, Barakat M, Piçarra-Pereira MA et al (2003) Osmotin and thaumatin from grape: a putative general defense mechanism against pathogenic fungi. Phytopathology 93:1505–1512
Murashige T, Skoog F (1962) A revised medium for rapid growth and bio assays with tobacco tissue cultures. Physiol Plant 15:473–479
Murray MG, Thompson WF (1980) Rapid isolation of high molecular weight plant DNA. Nucleic Acids Res 8:4321–4325
Neale AD, Wahleithner JA, Lund M et al (1990) Chitinase, β-1,3-glucanase, osmotin and extensin are expressed in tobacco explants during flower formation. Plant Cell 2:673–684
Pocock KF, Hayasaka Y, McCarthy MG et al (2000) Thaumatin-like proteins and chitinases, the haze-forming proteins of wine, accumulate during ripening of grape (Vitis vinifera) berries and drought stress does not affect the final levels per berry at maturity. J Agric Food Chem 48:1637–1643
Popowich EA, Firsov AP, Mitiouchkina TY et al (2007) Agrobacterium-mediated transformation of Hyacinthus orientalis with thaumatin II gene to control fungal diseases. Plant Cell Tissue Organ Cult 90:237–244
Rajam MV, Chandola N, Goud PS et al (2007) Thaumatin gene confers resistance to fungal pathogens as well as tolerance to abiotic stresses in transgenic tobacco plants. Biol Plant 51:135–141
Regalado AP, Ricardo CPP (1996) Study of intercellular fluid in healthy Lupinus albus organs. Plant Physiol 110:227–232
Roberts WK, Selitrennikoff CP (1990) Zeamatin, an antifungal protein from maize with membrane-permeabilizing activity. J Gen Microbiol 136:1771–1778
Schestibratov KA, Dolgov SV (2005) Transgenic strawberry plants expressing a thaumatin II gene demonstrate enhanced resistance to Botrytis cinerea. Sci Horticult 106:177–189
Sassa H, Ushijima K, Hirano H (2002) A pistil-specific thaumatin/PR5-like protein gene of Japanese pear (Pyrus serotina): sequence and promoter activity of the 5′ region in transgenic tobacco. Plant Mol Biol 50:371–377
Skadsen RW, Sathish P, Kaeppler HF (2000) Expression of thaumatin-like permatin PR-5 genes switches from the ovary wall to the aleurone in developing barley and oat seeds. Plant Sci 156:11–22
Stark-Urnau M, Seidel M, Kast WK et al (2000) Studies on the genetic diversity of primary and secondary infections of Plasmopara viticola using RAPD/PCR. Vitis 39:163–166
Staudt G, Kassemeyer HH (1995) Evaluation of downy mildew resistance in various accessions of wild Vitis species. Vitis 34:225–228
Tattersall DB, Van Heeswijck R, Hoj PB (1997) Identification and characterization of a fruit-specific, thaumatin-like protein that accumulates at very high levels in conjunction with the onset of sugar accumulation and berry softening in grapes. Plant Physiol 114:759–769
Trudel J, Grenier J, Potvin C et al (1998) Several thaumatin like proteins bind to beta-1,3-glucans. Plant Physiol 118:1431–1438
Van Damme EJ, Charels D, Menu-Bouaouiche L et al (2002) Biochemical, molecular and structural analysis of multiple thaumatin-like proteins from the elderberry tree (Sambucus nigra L.). Planta 214:853–862
Van der Wel H, Loewe K (1972) Isolation and characterization of thaumatin I and II, the sweet-tasting proteins from Thaumato-coccusdaniellii Benth. Eur J Biochem 31:221–225
Van Loon LC, Rep M, Pieterse CM (2006) Significance of inducible defense-related proteins in infected plants. Annu Rev Phytopathol 44:135–162
Velazhahan R, Datta SK, Muthukrishnan S (1999) The PR-5 family:thaumatin-like proteins in plants. In: Datta SK, Muthukrishnan S (eds) Pathogenesis-related proteins in plants. CRC Press, Boca Raton, pp. 107–129
Velazhahan R, Muthukrishnan S (2003) Transgenic tobacco plants constitutively overexpressing a rice thaumatin-like protein (PR-5) show enhanced resistance to Alternaria alternata. Biol Plant 47:347–354
Vu L, Huynh QK (1994) Isolation and characterization of a 27 kDa antifungal protein from the fruits of Diospyros texana. Biochem and Bioph Res Co 202:666–672
Wagih EE, Coutts RHA (1981) Similarities in the soluble protein profiles of leaf tissue following either a hypersensitive reaction to virus infection or plasmolysis. Plant Sci Lett 21:61–69
Wu J, Zhang Y, Zhang H et al (2010) Whole genome wide expression profiles of Vitis amurensis grape responding to downy mildew by using Solexa sequencing technology. BMC Plant Biol 10:234–250
Xiao Q, Ye W, Zhu Z et al (2005) A simple non-destructive method to measure leaf area using digital camera and Photoshop software. Chin J Eco 6:711–714
Yu Y, Zhang Y, Yin L et al (2012) The mode of host resistance to Plasmopara viticola infection of grapevines. Phytopathology 102:1094–1101
Zanek MC, Reyes CA, Cervera M et al (2008) Genetic transformation of sweet orange with the coat protein gene of Citrus psorosis virus and evaluation of resistance against the virus. Plant Cell Rep 27:57–66
Zevenhuizen LPTM, Bartnicki-Garcia S (1968) Structure of the insoluble hyphal wall glucan of Phytophthora cinnamomi. Biochemistry 8:1496–1502
Acknowledgements
This research was supported by China Agriculture Research System (grant no. CARS-30-yz-2) and China Agricultural University Scientific Fund (grant no. 2012RC019). We are grateful to the friends in UC Davis who helped make this research possible: Qingfeng Huang for assistance in vector construction, Abhaya Dandekar for providing Agrobacterium tumefaciens and Kim J. Carney for supplying the embryogenic culture. We also express our gratitude to Wentao Xu in China Agricultural University for their microscopy facility.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflicts of interest..
Additional information
Rongrong He and Jiao Wu are contributed equally to this work.
Handling Editor: Hanns H. Kassemeyer
Rights and permissions
About this article
Cite this article
He, R., Wu, J., Zhang, Y. et al. Overexpression of a thaumatin-like protein gene from Vitis amurensis improves downy mildew resistance in Vitis vinifera grapevine. Protoplasma 254, 1579–1589 (2017). https://doi.org/10.1007/s00709-016-1047-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00709-016-1047-y