Abstract
Abiotic stresses, such as drought and high salinity, are major factors that limit plant growth and productivity. Late embryogenesis abundant (LEA) proteins are members of a diverse, multigene family closely associated with tolerance to abiotic stresses in numerous organisms. We examined the function of SmLEA2, previously isolated from Salvia miltiorrhiza, in defense responses to drought and high salinity. Phylogenetic analysis indicated that SmLEA2 belongs to the LEA_2 subfamily. Its overexpression in Escherichia coli improved growth performance when compared with the control under salt and drought stresses. We further characterized its roles in S. miltiorrhiza through overexpression and RNAi-mediated silencing. In response to drought and salinity treatments, transgenic plants overexpressing SmLEA2 exhibited significantly increased superoxide dismutase activity, reduced levels of lipid peroxidation, and more vigorous growth than empty-vector control plants did. However, transgenic lines in which expression was suppressed showed the opposite results. Our data demonstrate that SmLEA2 plays an important role in the abiotic stress response and its overexpression in transgenic S. miltiorrhiza improves tolerance to excess salt and drought conditions.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Abiotic stresses, such as drought and high salinity, greatly limit plant development and agricultural productivity (Munns and Tester 2008; Sengupta and Majumder 2009). Approximately 70 % of all crop yield losses may be due to abiotic challenges, especially water stress (Bray et al. 2000). As a result of global climate change and environmental pollution, plants are frequently exposed to drought and high salt conditions that cause serious tissue damage and, in extreme cases, death. Plants utilize various stress response mechanisms, including the accumulation of osmolytes (e.g., proline, raffinose, and glycine betaine) and scavengers of reactive oxygen species (ROS), as well as the synthesis of osmoprotective proteins, such as late embryogenesis abundant (LEA) proteins (Mahajan and Tuteja 2005; Munns and Tester 2008; Krasensky and Jonak 2012).
The LEA proteins were originally discovered in seeds of cotton (Gossypium hirsutum), functioning as hydrophilic polypeptides that were accumulated abundantly in the embryos during the late developmental stage (Dure et al. 1981). They have now been identified in numerous species of higher plants, and similar proteins have been found in other organisms, such as algae, fungi, bacteria, and animals (Dure 1993; Wise and Tunnacliffe 2004; Battaglia et al. 2008). These types of proteins are associated with tolerances to dehydration, low temperatures, and excess salt (Bray 1993; Close 1997; Boudet et al. 2006). They are especially prominent in plants, with up to 51 genes encoding LEA proteins in Arabidopsis (Hundertmark and Hincha 2008), 39 in rice (Oryza sativa; Wang et al. 2007), and 77 in the moss Physcomitrella patens (Komatsu et al. 2013). Despite this great diversity, most LEA proteins can be grouped into eight families in the PFAM database according to their primary sequences: dehydrin, LEA_1, LEA_2, LEA_3, LEA_4, LEA_5, LEA_6, and seed maturation protein. Those sequences have now been placed in a dedicated database, LEAPdb (Hunault and Jaspard 2010), and organized into 12 non-overlapping classes corresponding to eight PFAMs (PF00257, PF00477, PF02987, PF03168, PF03242, PF03760, PF04927, and PF10714) (Jaspard et al. 2012). This diversity and compartmentalization of LEA proteins in plant tissues invites questions about their physiological function and modes of action. Although such proteins have a broad influence on plant responses—functioning as protectants of biomolecules and membranes and also sequestering ions and ROS (Sun et al. 2013; Candat et al. 2014)—their individual roles have not been well-characterized.
Salvia miltiorrhiza (family Labiatae) is an important herbal plant. Its dried roots, also known as Danshen, are widely used in modern and traditional Chinese medicine (TCM) for treating cardiovascular/cerebrovascular diseases and various symptoms of inflammation (Wang and Wu 2010; Xu et al. 2010; Dong et al. 2011; Yan 2013). Its annual consumption in China exceeds 20 million kilograms (Kai et al. 2014). This species is emerging as a model plant for TCM studies because of its relatively small genome size (∼600 Mb), short life cycle, minimal growth requirements, and significant medicinal value (Li and Lu 2014). Plants are very adaptable to many habitats and are commonly found throughout China and other Southeast Asian countries. Although numerous reports have been made of this species, research is lacking about the molecular mechanisms involved in its response to environmental stresses. We previously showed that overexpression of SmLEA, a LEA_2 family gene from S. miltiorrhiza, enhances salt and drought tolerance in Escherichia coli and S. miltiorrhiza (Wu et al. 2014). Since then, we have isolated SmLEA2, a second member of that gene family (Wu et al. 2015). Expression of both can be significantly induced by abscisic acid (ABA). Moreover, their 5′ flanking regions contain some putative cis-elements related to abiotic stresses, such as ABRE, MBS, and a CGTCA-motif (Liu and Wang 2010; Wu et al. 2015). Bioinformatic analysis has demonstrated that SmLEA2 is a hydrophilic protein (Wu et al. 2015) while SmLEA is hydrophobic (Liu and Wang 2009). SmLEA2 is most highly expressed in stems while SmLEA transcript levels are highest in the roots and lowest in the stems (Wu et al. 2015). Therefore, we are interested in learning whether SmLEA2 has a role in conferring plant tolerance to adverse growing conditions.
Here, we took an ectopic approach and found that E. coli expressing SmLEA2 is more tolerant than the control strain against high salinity and drought. In addition, transgenic plants of S. miltiorrhiza that overexpresses this gene have improved tolerance to those types of stress. To further characterize the biological function of SmLEA2, we used transgenic plants in which expression was suppressed by RNAi-mediated silencing. Physiological and biochemical characteristics were compared between the transgenic and empty-vector control plants, including rates of water loss, superoxide dismutase (SOD) activity, and concentrations of malondialdehyde (MDA) and chlorophyll. Our results suggested that SmLEA2 has a role in making these plants tolerant to dehydration and high salinity.
Materials and methods
Plant materials, growing conditions, and stress treatments
Sterile plantlets of S. miltiorrhiza were obtained and cultured as we have described previously (Yan and Wang 2007). To simulate water stress, we used polyethylene glycol (PEG-6000) as an osmotic agent. Four-leaf-stage shoots of uniform size and healthy appearance were removed from transgenic plants and were cultured on standard MS media for 4 weeks. After acclimation with a water solution for 1 week, some plantlets were moved to a 15 % PEG water solution (PEG6000) for 3 days. As the control, other plantlets were transferred from the MS solid medium to the water solution. To investigate the effects of salt stress, some selected plantlets were treated for 3 days with either a standard water solution or one containing 100 mM NaCl. After this treatment period, all plants were subjected to evaluations of their phenotype, SOD activity, and concentrations of MDA and chlorophyll. At least three replicates were done for the experiments.
Bioinformatics analysis of protein sequences
All plant LEA proteins were obtained from LEAPdb (http://forge.info.univ-angers.fr/∼gh/Leadb/index.php). A phylogenetic tree was constructed by the neighbor-joining method and Molecular Evolutionary Genetics Analysis (MEGA 5.2). The grand average of hydropathy (GRAVY) for those deduced proteins was predicted with the Expasy program (http://www.expasy.org/tools).
Constitutive expression of SmLEA2 in E. coli
Genomic DNA was extracted from whole seedlings with a DN15-Plant DNA Mini Kit (Aidlab, China). Using gDNA as template, the full-length SmLEA2 open reading frame (ORF) was amplified with primers SmLEA2Pro-F and SmLEA2Pro-R, which were designed according to the sequence we have previously reported from our laboratory (GenBank Accession Number HQ676610). Reactions were performed in a S1000™ Thermal Cycler (Bio-RAD, USA) under the following conditions: 95 °C for 8 min; then 30 cycles of 95 °C for 30 s, 54 °C for 30 s, and 72 °C for 1 min; followed by a final elongation at 72 °C for 10 min. The PCR products were digested with BamHI and EcoRI (Takara, Japan). Afterward, the fragment of SmLEA2 with sticky BamHI/EcoRI was inserted into the PET28a vector and sequenced to create recombinant plasmid PET28a–SmLEA2. Both the construct and the empty-vector PET28a were transformed into E. coli BL21 strain, which was inoculated in a lysogeny broth (LB) medium with 50 mg L−1 kanamycin (kan) and then cultured at 37 °C overnight. When the OD600 reached 0.6 to 0.8, isopropylthio-β-galactoside (IPTG) was added to the LB culture to achieve a final concentration of 1 mM. After IPIG induction for 0, 2, 4, 6, or 8 h, the E. coli cells were boiled for 5 min in the sample buffer before being loaded on 13 % (w/v) SDS-PAGE gels (Sambrook and Russell 2001).
For studying their tolerance to salinity, E. coli cells (carrying PET28a-SmLEA2 or PET28a) that had been induced by IPTG for 6 h were then diluted 10, 100, or 1000 times. Afterward, 10 μL of these different dilutions were spread onto LB solid media containing 50 mg L−1 kan plus 400 mM NaCl. The plates were incubated upside-down overnight at 37 °C. Other cells carrying PET28a–SmLEA2 or PET28a were inoculated into an LB liquid medium containing 50 mg L−1 kan (1:100 ratio). Samples were taken at 2-h intervals to produce a growth curve.
For evaluating their degree of drought tolerance, E. coli cells carrying PET28a–SmLEA2 or PET28a were incubated on an LB solid medium containing 50 mg L−1 kan plus 400 mM mannitol, as described above. Other cells carrying PET28a–SmLEA2 or PET28a were inoculated into an LB liquid medium containing 50 mg L−1 kan and 15 % PEG to produce a growth curve.
Construction of plant expression vectors and plant transformation
To construct the SmLEA2-overexpressing vectors, we used the pET28a–SmLEA2 construct. Gene-specific primers oNILF and oNILR (Table 1) were designed to amplify SmLEA2. The PCR products were digested with KpnI/BamHI and ligated into the pKANNIBAL vector to generate recombinant plasmid pKan–SmLEA2. An overexpressing box with SmLEA2 was removed with NotI (Takara, Japan) from the new plasmid and subcloned into the pART27 vector to generate overexpression plasmid pAK–SmLEA2. For constructing the RNAi plasmid, a 312-bp fragment of SmLEA2 cDNA (located from coding sequence positions 181–492 bp) was amplified with primers iN1LF/iN1LR (Table 1) and ligated into the pMD19-T vector (Takara) before sequencing. The fragments were digested with XhoI/KpnI and BamHI/HindIII, and then inserted into the pKANNIBAL vector (Wesley et al. 2001) to generate plasmid pKAN–SmLEA2. An interfering box with the pyruvate orthophosphate dikinase (PDK) intron was cloned into pART27, as we have described previously (Song and Wang 2011), to generate RNAi vector pSmLEA2–RNAi. Agrobacterium-mediated gene transfer was performed according to our protocol (see Song and Wang 2011). In parallel, plasmid pAK, constructed in our laboratory (Wu et al. 2014), was introduced into WT S. miltiorrhiza as the empty-vector control PDK. All transgenic plants were generated by Agrobacterium-mediated gene transfer according to the protocol established in our laboratory (Yan and Wang 2007) and were selected on MS medium containing kanamycin.
Transformant selection and transcript analysis
As our template for PCR analysis of transgenic lines, DNA was extracted from the leaves of 1-month-old kan-resistant plantlets. Primers 35SF/35SR (Table 1) were designed based on the CaMV35S promoter sequence for both SmLEA2-overexpressing and SmLEA2-RNAi lines. This was done so that we could evaluate whether the overexpressing and interfering box had been integrated into the transgenic plant genome. The PCR reactions were performed as follows: preheating at 94 °C for 10 min; then 35 cycles of 94 °C for 30 s, 58 °C for 30 s, and 72 °C for 1 min; with a final extension at 72 °C for 10 min. Plasmid pAK–SmLEA2 was used as the positive control while genomic DNA from untransformed plants was our negative control.
Total RNA was isolated with a Plant RNA kit (OMEGA, USA), according to the manufacturer’s protocol, and was converted into cDNA using a primeScript® RT reagent kit (Takara). The RT products were amplified with RLEA2-S/RLEA2-A primers (Table 1) in order to evaluate gene expression via real-time quantitative PCR. A housekeeping gene, SmGAPDH (glyceraldehyde-3-phosphate dehydrogenase; CV170251), served as the control and was amplified with primers GAPDH-S/GAPDH-A (Table 1). Quantitative real-time reactions were performed in triplicate under the following conditions: 1 cycle of predenaturation at 95 °C for 10 min; then 45 cycles of amplification at 95 °C for 10 s and 60 °C for 30 s. Relative expression for each gene was calculated by the comparative CT method (Vandesompele et al. 2002). The transgenic lines of the T1 generation were used for phenotype analysis and measurement of water loss rates, chlorophyll concentrations, MDA, SOD, and K+ and Na+ content.
Measurement of water loss rates
Plants selected at uniform developmental stages were placed on filter paper and exposed to ambient air in the greenhouse. Fresh weights were recorded for various tissues after 0, 10, 30, 60, 90, 120, 150, and 180 min. Eight independent measurements were made from three separate repeats.
Calculations of malondialdehyde and chlorophyll concentrations
Concentrations of malondialdehyde (MDA) were determined as we have described previously (Wu et al. 2014). Briefly, trichloroacetic acid (TCA; 3 mL, 10 %) was mixed with powdered S. miltiorrhiza leaves (0.2 g) at 4 °C overnight. After centrifugation, the supernatants (2 mL) were added with 2 mL of 0.6 % thiobarbituric acid, and then heated at 100 °C for 15 min and centrifuged. The supernatants were measured at 532, 450, and 600 nm, with TCA as the background. Each MDA concentration was calculated as CMDA (μmol g−1) = [6.45 × (OD532 − OD600) − 0.56× OD450] / 0.2.
For measuring plant chlorophyll concentrations, the leaves were harvested, weighed, and homogenized in liquid nitrogen. Chlorophyll a/b was extracted in 95 % ethanol for 2 days at 37 °C with continuous shaking and was quantified as micrograms per gram fresh weight. The amount of chlorophyll in each sample was determined spectrophotometrically, using the formula Ch a + b = (6.63 × A665) + (18.08 × A649), where Ch is the chlorophyll concentration in micrograms per milliliter and A is absorption (Lichtenthaler 1987).
Assay for superoxide dismutase activity
Enzymes were extracted from plants treated for 3 days with 100 mM NaCl or 15 % PEG. The leaves were ground to fine powder in liquid nitrogen with a mortar and pestle. To each 0.5-g sample, we added 5 mL of 50 mmol L−1 phosphate-buffered saline containing 1 % β-mercaptoethanol and 1 % polyvinylpyrrolidone (pH 7.8), and held them for 60 min at 4 °C. After centrifugation (4 °C, 10,000 rpm, 20 min), the supernatant (enzyme extraction) was collected for estimating the activity of superoxide dismutase (SOD), based on the method described by Dhindsa and Matowe (1981).
Quantification of potassium and sodium content
The leaves were harvested from the transgenic plants treated for 3 days with 100 mM NaCl or 0 mM NaCl, rinsed twice with tap water followed by deionized H2O, and then dried at 80 °C to constant weight in an oven. Then, the dried samples were ground to fine powder. The potassium (K+) and sodium (Na+) contents of tissue powders were measured using an atomic absorption spectrophotometer (TAS-990Super AFG, Beijing, China) as described previously (Jin et al. 2016).
Results
Phylogenetic characterization of SmLEA2
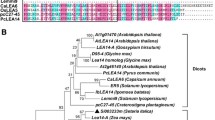
The phylogenetic tree (Fig. 1a) revealed that SmLEA2 belongs to the LEA_2 subfamily, based on the presence of a PF03168 domain (Jaspard et al. 2012). This protein is most closely related to At2g44060 of Arabidopsis thaliana, which is part of LEAPdb Class 8 (Candat et al. 2014). The hydropathy plot, as developed by Kyte and Doolittle (1982), predicted that SmLEA2 is hydrophilic and has a GRAVY score of −0.359 (Fig. 1b). In contrast, SmLEA is hydrophobic, has a GRAVY score of 0.207 (Wu et al. 2014), and shows high similarity to At2g46140, which is in Class 7. The LEA protein classes 7 and 8 have GRAVY scores close to zero and both classes are included in PF03168. Although both SmLEA2 and SmLEA belong to the LEA_2 group due to the presence of a PF03168 domain, the nucleotide sequence identity and the deduced amino acid sequence identity of SmLEA2 and SmLEA was only 31.53 and 29 %, respectively.
a Unrooted phylogenetic tree showing evolutionary relationships between SmLEA2 and LEA proteins from other plant species: Arabidopsis thaliana (gi|633765|, gi|79332705|, gi|15221813|, gi|556805|, gi|332645212|, AAD20140.1 AtECP63, At2g44060, At2g46140, At4g02380, At5g06760, At2g35300, At3g22500, At3g22490, At2g33690, At2g23110, At1g01470), Salvia miltiorrhiza (gi|51512229|, gi|52148123|, AAU29064.3 SmLEA, HQ676610 SmLEA2), Ipomoea batatas (ADM33676.1 IbLEA14), Gossypium hirsutum (AAA33058.1 GhD11, P09443.2 GhLEA D-19, CAA31589.1 GhLEA D-7, P13940.1 GhLEA D-29, P46518.1 GhLEA14-A, P46522.1 GhLEA5-D, CAA31590.1 GhLEA D-113, CAA31594.1 GhLEA D-34), Nicotiana tabacum (gi|46020012|, gi|153793260|, AAC06242.1 NtLEA5), Zea mays (CAA34123.1 RAB-17, gi|168467|, gi|195606062|, NP_001152646.1 ZmLEA14-A, gi|226531554|, CAA41854.1 RAB28), Capsicum annuum (gi|37905913|, gi|23507177|, AAD53078.1 ER5), Pisum sativum (gi|56709428|, CAA44787.1 DHN1), and Glycine max (gi|170020|, gi|1762955|, gi|1389897|, gi|212552204|, gi|1732556|, gi|82394552|, AAF21311.1 PM25, gi|75194027|). Different colors indicate individual LEA groups; SmLEA2 from S. miltiorrhiza is marked by red square. b Hydropathy plot of deduced SmLEA2 amino acid sequence
Expression of SmLEA2 enhanced the viability of E. coli cells under salt stress or simulated drought conditions
The physicochemical characteristics of SmLEA2 protein were studied by transforming pET28a–SmLEA2 and an empty pET28a vector into E. coli BL21. The protein profiles of that strain carrying the recombinant construct revealed overexpression of an approximately 39-kDa recombinant protein (Fig. 2), which was approximately 3 kDa larger than the predicted value of 35.34 kDa (Wu et al. 2015). The maximum accumulation of SmLEA2 occurred at 6 h after induction with IPTG (Fig. 2).
SDS-PAGE analysis of SmLEA2 overexpression in E. coli BL21. Lanes 1, 2, 3, 4, and 5: 1 mM IPTG induction for 8, 6, 4, 2, and 0 h, respectively, with pET 28a–LEA2 vector; Lanes 6, 7, and 8: 1 mM IPTG induction for 6, 4, and 0 h, respectively, with empty pET 28a vector; Lane M: low molecular-weight protein marker
To evaluate whether the recombinant SmLEA2 protein aids in conferring tolerance to salinity stress in vivo, we exposed overexpressing cells of E. coli to 400 mM NaCl and compared their viability with that of control (empty-vector) cells. Under normal conditions (without excess NaCl), performance did not differ between treatments on either solid or liquid media, regardless of IPTG induction (Fig. 3a, d). Under salt stress, the overexpressing cells produced more and larger clones than did the control when grown on solid media at dilutions of 100 and 1000 times (Fig. 3b). On the liquid media, E. coli cells expressing recombinant SmLEA2 entered the logarithmic phase after 10 h, which was approximately 8 h earlier than the control (Fig. 3e). These results indicated that salinity-induced overexpression of SmLEA2 in E. coli obviously improved its growth performance over that of the control.
Heterogeneous expression of SmLEA2 in E. coli strain BL21. Growth of E. coli harboring pET28a or pET–SmLEA2 on control medium (a) or LB solid medium supplemented with 400 mM NaCl (b) or 400 mM mannitol (c). Growth curves for IPTG-induced E. coli harboring pET28a or pET–SmLEA2 on control medium (d) or LB liquid medium supplemented with 800 mM NaCl (e) or 15 % PEG (f).
The effect of dehydration was investigated by incubating E. coli cells (with/without SmLEA2) on LB solid media supplemented with 400 mM mannitol or 15 % PEG. Cells carrying SmLEA2 produced more and larger clones than those on either the control medium or the medium with the mannitol (Fig. 3c). In the presence of 15 % PEG, cells with SmLEA2 showed a slight growth advantage over the control (Fig. 3f). These results demonstrated that SmLEA2 could increase drought tolerance in E. coli.
Generation of transgenic S. miltiorrhiza plants
Using Agrobacterium-mediated transformation, we generated transgenic plants of S. miltiorrhiza that either overexpressed or suppressed SmLEA2. After selective culturing on a kanamycin medium, those plants were confirmed through PCR amplification to contain an expected 901-bp fragment of the CaMV 35S promoter (Fig. 4a). Expression was examined in the transgenics by quantitative RT-PCR (Fig. 4b, c). Based on their transcript levels, we chose two independent overexpressing lines (OE44 and OE72) with significantly elevated SmLEA2 expression, two SmLEA2-silenced lines (Li9 and Li11) with significantly reduced expression, and the empty control (PDK) for further analysis.
Detection and identification of transgenic lines. a PCR-amplification product of 35S promoter from transgenic S. miltiorrhiza gDNA. Positive control, amplified with plasmid as template; WT, wild-type plants. b qRT-PCR detection of SmLEA2 in overexpression of transgenic lines. c qRT-PCR detection of SmLEA2 in RNAi transgenic lines. Error bars show significant differences at P < 0.05 (*) or P < 0.01 (**)
Salinity and drought tolerance by transgenic S. miltiorrhiza plants
After 4 weeks of growth on standard MS media, phenotypes did not differ significantly between transgenic S. miltiorrhiza seedlings and PDK plants. To determine whether SmLEA2 improved their tolerance to stress as well as LEA proteins in other species, we treated with standard water or solutions supplemented with either 100 mM NaCl or 15 % PEG after plants had been acclimated for 1 week in the standard water solution. By day 3, leaves from the SmLEA2-silenced transgenic lines were the most severely wilted under both stress conditions while those from the SmLEA2-overexpressing lines showed the least inhibition of development (Fig. 5a–d).
Comparisons of phenotypes and physiology among transgenic SmLEA2-overexpressing lines OE72 and OE44, SmLEA2-silenced lines Li9 and Li11, and PDK control plants in response to 15 % PEG (a, b) or 100 mM NaCl (c, d). Changes in chlorophyll concentrations (e), MDA concentrations (f), and SOD activity (g) in control plants and transgenic lines OE72, OE44, Li9, and Li11 in response to NaCl or PEG. Error bars show significant differences at P < 0.05 (*) or P < 0.01 (**)
Chlorophyll concentrations were lower in SmLEA2-silenced transgenic lines than in the PDK plants or SmLEA2-overexpressing lines. When treated with NaCl or PEG, more chlorophyll pigment was lost from leaves of the silenced lines than from the other types (Fig. 5e).
MDA concentrations did not differ significantly between the transgenic and PDK plants under normal conditions (Fig. 5f), but levels did rise to varying degrees in all plants in response to NaCl or PEG. In particular, concentrations increased significantly more rapidly in the SmLEA2-silenced transgenic lines, thereby implying that the LEA protein could alleviate damage to cell membranes by suppressing the accumulation of MDA under both drought and salt stress.
As a key enzyme, SOD scavenges ROS that can accumulate when stress is imposed. Here, SOD activity under normal conditions followed the order (high to low) of SmLEA2-overexpressing transgenic lines > PDK > SmLEA2-silenced transgenic lines. When plants were exposed to 100 mM NaCl or 15 % PEG, this activity rose significantly in all plant types, albeit to varying degrees (Fig. 5g). These results indicated that overexpression of SmLEA2 leads to enhanced SOD activity in S. miltiorrhiza.
Salt stress imposes ionic imbalance on plants due to elevated Na+ levels. The cellular K+/Na+ ratio determines salt tolerances in plants cells, rather than the absolute intracellular Na+ concentration (Horie et al. 2001). Thus, we determined Na+ and K+ relative contents in the leaves of transgenic lines and the PDK (Fig. 6a, b). Neither Na+ nor K+ relative contents showed significant differences in all plant types under normal conditions but levels did change to varying degrees in response to NaCl. The cellular K+/Na+ ratio were higher in SmLEA2-overexpressing transgenic lines than in the PDK plants or SmLEA2-silenced lines when treated with 100 mM NaCl for 3 days (Fig. 6c). The data implied that SmLEA2 functions in the maintenance of cellular ion homeostasis in S. miltiorrhiza under salt stress.
Measurement of water loss rates in transgenic plants
Whole-plant water losses were slower from the SmLEA2-overexpressing lines than from PDK plants, while the highest loss rate was from SmLEA2-silenced lines (Fig. 7). This demonstrated that the transgenic lines in which SmLEA2 was overexpressed were more tolerant of drought.
Discussion
LEA proteins are a diverse group of heterogeneous proteins associated with cellular tolerance to dehydration. The expression of LEA genes can be induced by ABA, dehydration, low temperatures, and osmotic stress in vegetative tissues, implying that they function not only in seed maturation but also in conferring tolerance to abiotic challenges (Zhang et al. 2014). Although production of these proteins is positively correlated with tolerance, especially to drought (Tunnacliffe and Wise 2007), not all LEA proteins make a significant contribution. For example, when expressed in Nicotiana tabacum (tobacco), two cold-induced LEA proteins from Spinacia oleracea (Kaye et al. 1998) and three desiccation-induced LEA proteins from Craterostigma plantagineum (Iturriaga et al. 1992) do not significantly alter the degree to which respective transgenic plants are tolerant of freezing or drought. Taking an approach that monitored ectopic expression, we found that SmLEA2 is most abundantly expressed in plant stems as well as in the cells of E. coli. The system of expression in the latter species provides a simple, convenient, and effective model for determining the function of a heterogeneous protein in cells subjected to stress treatments (Liu and Zheng 2005). Salt tolerance was also enhanced in our recombinant bacterial cells. This role by SmLEA2 proteins in E. coli is similar to that reported previously for SmLEA (Wu et al. 2014). Analysis of SmLEA2-silenced and SmLEA2-overexpressing lines of S. miltiorrhiza also showed that SmLEA2 positively regulates plant defense responses to drought and high salinity. Therefore, overexpression of this gene in either E. coli or S. miltiorrhiza directly increases stress tolerance by the host, implying that some protective mechanisms might be common to both prokaryotes and eukaryotes under adverse conditions.
Homeostasis of ROS is important for preserving normal plant metabolism; excessive ROS accumulations cause severe oxidative damage to cells, in turn harming the health and viability of plants. High levels of salt as well as drought conditions can lead to ROS formation, resulting in the accumulation of MDA in plants due to membrane lipid peroxidation (Neill et al. 2002). To avoid severe phytotoxicity and oxidative damage, plants utilize several strategies to scavenge and eliminate those toxic compounds. For example, SOD is an important antioxidant that can catalyze superoxide radicals to H2O2 (Asada 2000). Thus, enhancement of antioxidative defenses in a plant can improve its tolerance to various environmental factors (Bowler et al. 1992; Jaspers et al. 2010; Mullineaux and Baker 2010). Here, we noted that overexpression of SmLEA2 reduced the accumulation of MDA and increased SOD activity when compared with the control, while the opposite was observed when SmLEA2 expression was suppressed in our transgenic plants. Those overexpressing lines also experienced less oxidative damage and cell membranes were more stable following both salt and drought treatments, thereby making the plants more stress-tolerant. Our data also further confirmed that hydrophilic and hydrophobic SmLEA proteins share identical functions under drought and high salinity (Wu et al. 2014).
To date, hundreds of LEA genes have been isolated from plants, algae, fungi, and bacteria (Dure 1993; Wise and Tunnacliffe 2004; Battaglia et al. 2008). As more LEA proteins are discovered and new methods of computational analysis are introduced, LEA protein classifications must to be updated (Dure 1993; Close 1997; Wise 2003; Shao et al. 2005; Battaglia et al. 2008; Bies-Ethève et al. 2008; Hundertmark and Hincha 2008; Hunault and Jaspard 2010). Unfortunately, this grouping of proteins and their nomenclature are not consistent in the literature. For example, At1g01470.1 in Arabidopsis, which had important protective functions under salt stress conditions in yeast and Arabidopsis (Jia et al. 2014), has been named LEA14 and placed in LEA_2, Group 7, PF03168, or Class 7 (Bies-Ethève et al. 2008; Hundertmark and Hincha 2008), as well as into various PFAM and LEAPDB classifications (Candat et al. 2014). For our research purposes, however, we have chosen to follow that of Hundertmark and Hincha (2008), in which these proteins are classified into eight groups on the basis of sequence similarity. Therefore, we conclude that both SmLEA2 and SmLEA belong to the LEA_2 group due to the presence of a PF03168 domain, even though we have previously categorized SmLEA as a LEA14 protein (Wu et al. 2014).
In summary, our findings provide evidence that SmLEA2 plays an important role in conferring drought and salt tolerance in prokaryotes and eukaryotes. However, the molecular mechanism(s) by which this is achieved has not been completely elucidated and further experiments are necessary to explore how SmLEA2 is induced in these abiotic stress responses. In general, LEA genes are highly represented in plant genomes, and global analyses of transcriptome data from Arabidopsis and rice have revealed that they have diverse functions. For example, expression of the 51 LEA genes in Arabidopsis can be detected in various tissue types and at different developmental stages under a variety of experimental conditions (Hundertmark and Hincha 2008). Furthermore, a genome-wide examination of the rice genome has identified 39 members of the LEA family, and their functions and expression patterns appear to be highly diverse (Wang et al. 2007). Therefore, genome-wide identification and analysis of LEA genes is critical to improving our understanding of their precise biological roles in S. miltiorrhiza.
References
Asada K (2000) The water-water cycle as alternative photon and electron sinks. Phil Trans Roy Soc Lond B 355:1419–1431. doi:10.1098/rstb.2000.0703
Battaglia M, Olvera-Carrillo Y, Garciarrubio A, Campos F, Covarrubias AA (2008) The enigmatic LEA proteins and other hydrophilins. Plant Physiol 148(1):6–24. doi:10.1104/pp.108.120725
Bies-Ethève N, Gaubier-Comella P, Debures A, Lasserre E, Jobet E, Raynal M, Cooke R, Delseny M (2008) Inventory, evolution and expression profiling diversity of the LEA (late embryogenesis abundant) protein gene family in Arabidopsis thaliana. Plant Mol Biol 67:107–124. doi:10.1007/s11103-008-9304-x
Boudet J, Buitink J, Hoekstra FA, Rogniaux H, Larré C, Satour P, Leprince O (2006) Comparative analysis of the heat stable proteome of radicals of Medicago truncatula seeds during germination identifies late embryogenesis abundant proteins associated with desiccation tolerance. Plant Physiol 140(4):1418–1436. doi:10.1104/pp.105.074039
Bowler C, Van Montagu M, Inze D (1992) Superoxide dismutase and stress tolerance. Annu Rev Plant Biol 43:83–116. doi:10.1146/annurev.pp.43.060192.000503
Bray EA (1993) Molecular responses to water deficit. Plant Physiol 103(4):1035–1040
Bray EA, Bailey-Serres J, Weretilnyk E (2000) Responses to abiotic stresses. In: Gruissem W, Buchannan B, Jones R (eds) Biochemistry and molecular biology of plants. American Society of Plant Physiologists, Rockville, MD, USA, pp 1158–1249
Candat A, Paszkiewicz G, Neveu M, Gautier R, Logan DC, Avelange-Macherel MH, Macherel D (2014) The ubiquitous distribution of late embryogenesis abundant proteins across cell compartments in Arabidopsis offers tailored protection against abiotic stress. Plant Cell 26:3148–3166. doi:10.1105/tpc.114.127316
Close TJ (1997) Dehydrins: a commonality in the response of plants to dehydration and low temperature. Physiol Plant 100(2):291–296. doi:10.1111/j.1399-3054.1997.tb04785.x
Dhindsa RS, Matowe W (1981) Drought tolerance in two mosses: correlated with enzymatic defence against lipid peroxidation. J Exp Bot 32(1):79–91. doi:10.1093/jxb/32.1.79
Dong Y, Morris-Natschke SL, Lee KH (2011) Biosynthesis, total syntheses, and antitumor activity of tanshinones and their analogs as potential therapeutic agents. Nat Prod Rep 28(3):529–542. doi:10.1039/c0np00035c
Dure L (1993) A repeating 11-mer amino acid motif and plant desiccation. Plant J 3(3):363–369
Dure L III, Greenway SC, Galau GA (1981) Developmental biochemistry of cottonseed embryogenesis and germination: changing messenger ribonucleic acid populations as shown by in vitro and in vivo protein synthesis. Biochemistry 20(14):4162–4168
Horie T, Yoshida K, Nakayama H, Yamada K, Oiki S, Shinmyo A (2001) Two types of HKT transporters with different properties of Na+ and K+ transport in Oryza sativa. Plant J 27(2):129–138. doi:10.1046/j.1365-313x.2001.01077.x
Hunault G, Jaspard E (2010) LEAPdb: a database for the late embryogenesis abundant proteins. BMC Genom 11:221. doi:10.1186/1471-2164-11-221
Hundertmark M, Hincha DK (2008) LEA (late embryogenesis abundant) proteins and their encoding genes in Arabidopsis thaliana. BMC Genom 9:118. doi:10.1186/1471-2164-9-118
Iturriaga G, Schneider K, Salamini F, Bartels D (1992) Expression of desiccation-related proteins from the resurrection plant Craterostigma plantagineum in transgenic tobacco. Plant Mol Biol 20(3):555–558. doi:10.1007/BF00040614
Jaspard E, Macherel D, Hunault G (2012) Computational and statistical analyses of amino acid usage and physico-chemical properties of the twelve late embryogenesis abundant protein classes. PLoS One 7(5), e36968. doi:10.1371/journal.pone.0036968
Jaspers P, Brosché M, Overmyer K, Kangasjärvi J (2010) The transcription factor interacting protein RCD1 contains a novel conserved domain. Plant Signal Behav 5(1):78–80
Jia F, Qi S, Li H, Liu P, Li P, Wu C, Zheng C, Huang J (2014) Overexpression of late embryogenesis abundant 14 enhances arabidopsis salt stress tolerance. Biochem Biophys Res Commun 454(4):505–511. doi:10.1016/j.bbrc
Jin H, Dong D, Yang Q, Zhu D (2016) Salt-responsive transcriptome profiling of Suaeda glauca via RNA sequencing. PLoS One 11(3), e0150504. doi:10.1371/journal.pone.0150504
Kai G, Hao X, Cui L, Ni X, Zekria D, Wu J (2014) Metabolic engineering and biotechnological approaches for production of bioactive diterpene tanshinones in Salvia miltiorrhiza. Biotechnol Adv. doi:10.1016/j.biotechadv.2014.10.001
Kaye C, Neven L, Hofig A, Li Q-B, Haskell D, Guy C (1998) Characterization of a gene for spinach CAP160 and expression of two spinach cold-acclimation proteins in tobacco. Plant Physiol 116(4):1367–1377
Komatsu K, Suzuki N, Kuwamura M, Nishikawa Y, Nakatani M, Ohtawa H, Takezawa D, Seki M, Tanaka M, Taji T, Hayashi T, Sakata Y (2013) Group A PP2Cs evolved in land plants as key regulators of intrinsic desiccation tolerance. Nat Commun 4:2219. doi:10.1038/ncomms3219
Krasensky J, Jonak C (2012) Drought, salt and temperature stress-induced metabolic rearrangements and regulatory networks. J Exp Bot 63:1593–1608. doi:10.1093/jxb/err460
Kyte J, Doolittle RF (1982) A simple method for displaying the hydropathic character of a protein. J Mol Biol 157:105–132. doi:10.1016/0022-2836(82)90515-0
Li C, Lu S (2014) Genome-wide characterization and comparative analysis of R2R3-MYB transcription factors shows the complexity of MYB-associated regulatory networks in Salvia miltiorrhiza. BMC Genom 15:277. doi:10.1186/1471-2164-15-277
Lichtenthaler HK (1987) Chlorophylls and carotenoids: pigments of photosynthetic biomembranes. Methods Enzymol 148(1):350–382
Liu CL, Wang ZZ (2009) Cloning and expression analysis of SmLEA from Salvia miltiorrhiza Bunge. Biotechnol Bull 5:80–84
Liu CL, Wang ZZ (2010) Gene expression analysis of a late embryogenesis abundant gene from Salvia miltiorrhiza Bunge. China Biotechnology 30(1):51–55
Liu Y, Zheng Y (2005) PM2, a group 3 LEA protein from soybean, and its 22-mer repeating region confer salt tolerance in Escherichia coli. Biochem Biophys Res Comm 331(1):325–332. doi:10.1016/j.bbrc.2005.03.165
Mahajan S, Tuteja N (2005) Cold, salinity and drought stresses: an overview. Arch Biochem Biophys 444(2):139–158. doi:10.1016/j.abb.2005.10.018
Mullineaux PM, Baker NR (2010) Oxidative stress: antagonistic signaling for acclimation or cell death? Plant Physiol 154(2):521–525. doi:10.1104/pp.110.161406
Munns R, Tester M (2008) Mechanisms of salinity tolerance. Annu Rev Plant Biol 59(1):651–681. doi:10.1146/annurev.arplant.59.032607.092911
Neill SJ, Desikan R, Clarke A, Hurst RD, Hancock JT (2002) Hydrogen peroxide and nitric oxide as signalling molecules in plants. J Exp Bot 53(372):1237–1247
Sambrook J, Russell DW (2001) Molecular cloning: a laboratory manual, 3rd edn. Cold Spring Harbor Laboratory Press, New York
Sengupta S, Majumder AL (2009) Insight into the salt tolerance factors of a wild halophytic rice, Porteresia coarctata: a physiological and proteomic approach. Planta 229(4):911–929. doi:10.1007/s00425-008-0878-y
Shao H-B, Liang Z-S, Shao M-A (2005) LEA proteins in higher plants: structure, function, gene expression and regulation. Colloid Surface B 45:131–135. doi:10.1016/j.colsurfb.2005.07.017
Song J, Wang ZZ (2011) RNAi-mediated suppression of the phenylalanine ammonia-lyase gene in Salvia miltiorrhiza causes abnormal phenotypes and a reduction in rosmarinic acid biosynthesis. J Plant Res 124(1):183–192. doi:10.1007/s10265-010-0350-5
Sun X, Rikkerink EH, Jones WT, Uversky VN (2013) Multifarious roles of intrinsic disorder in proteins illustrate its broad impact on plant biology. Plant Cell 25(1):38–55. doi:10.1105/tpc.112.106062
Tunnacliffe A, Wise MJ (2007) The continuing conundrum of the LEA proteins. Naturwissenschaften 94(10):791–812. doi:10.1007/s00114-007-0254-y
Vandesompele J, De Preter K, Pattyn F, Poppe B, Van Roy N, De Paepe A, Speleman F (2002) Accurate normalization of real-time quantitative RT-PCR data by geometric averaging of multiple internal control genes. Genome Biol 3(7): RESEARCH0034
Wang JW, Wu JY (2010) Tanshinone biosynthesis in Salvia miltiorrhiza and production in plant tissue cultures. Appl Microbiol Biotechnol 88(2):437–449. doi:10.1007/s00253-010-2797-7
Wang XS, Zhu HB, Jin GL, Liu HL, Wu WR, Zhu J (2007) Genome-scale identification and analysis of LEA genes in rice (Oryza sativa L.). Plant Sci 172(2):414–420. doi:10.1016/j.plantsci.2006.10.004
Wesley SV, Helliwell CA, Smith NA, Wang MB, Rouse DT, Liu Q, Gooding PS, Singh SP, Abbott D, Stoutjesdijk PA, Robinson SP, Gleave AP, Green AG, Waterhouse PM (2001) Construct design for efficient, effective and high throughput gene silencing in plants. Plant J 27(6):581–590. doi:10.1046/j.1365-313X.2001.01105.x
Wise MJ (2003) LEAping to conclusions: a computational reanalysis of late embryogenesis abundant proteins and their possible roles. BMC Bioinform 4:52. doi:10.1186/1471-2105-4-52
Wise MJ, Tunnacliffe A (2004) POPP the question: what do LEA proteins do? Trends Plant Sci 9(1):13–17. doi:10.1016/j.tplants.2003.10.012
Wu Y, Liu C, Kuang J, Ge Q, Zhang Y, Wang Z (2014) Overexpression of SmLEA enhances salt and drought tolerance in Escherichia coli and Salvia miltiorrhiza. Protoplasma 251(5):1191–1199. doi:10.1007/s00709-014-0626-z
Wu Y, Ge Q, Zhou H, Jin X, Wang Z (2015) Molecular cloning and expression analysis of a late embryogenesis abundant protein gene (SmLEA2) from Salvia miltiorrhiza. Chin Trad Herb Drugs 46:1966–1974
Xu H, Zhang L, Zhou CC, Xiao JB, Liao P, Kai GY (2010) Metabolic regulation and genetic engineering of pharmaceutical component tanshinone biosynthesis in Salvia miltiorrhiza. J Med Plants Res 4(24):2591–2597
Yan XJ (2013) Cyclopedia of Danshen. People’s Medical Publishing House, Beijing, China
Yan Y, Wang Z (2007) Genetic transformation of the medicinal plant Salvia miltiorrhiza by Agrobacterium tumefaciens-mediated method. Plant Cell Tiss Org Cult 88:175–184. doi:10.1007/s11240-006-9187-y
Zhang X, Lu S, Jiang C, Wang Y, Lv B, Shen J, Ming F (2014) RcLEA, a late embryogenesis abundant protein gene isolated from Rosa chinensis, confers tolerance to Escherichia coli and Arabidopsis thaliana and stabilizes enzyme activity under diverse stresses. Plant Mol Biol 85(4–5):333–347. doi:10.1007/s11103-014-0192-y
Acknowledgments
This work benefited from financial support from the Natural Science Foundation of Shaanxi Province, China (2014JQ3107) and the Fundamental Research Funds for the Central Universities (GK201302043).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Handling Editor: Néstor Carrillo
Huaiqin Wang and Yucui Wu contributed equally to this work.
Rights and permissions
About this article
Cite this article
Wang, H., Wu, Y., Yang, X. et al. SmLEA2, a gene for late embryogenesis abundant protein isolated from Salvia miltiorrhiza, confers tolerance to drought and salt stress in Escherichia coli and S. miltiorrhiza . Protoplasma 254, 685–696 (2017). https://doi.org/10.1007/s00709-016-0981-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00709-016-0981-z