Abstract
Pollen developmental pathway in plants involving synchronized transferal of cellular divisions from meiosis (microsporogenesis) to mitosis (pollen mitosis I/II) eventually offers a unique “meiosis-mitosis shift” at pollen mitosis I. Since the cell type (haploid microspore) and fate of pollen mitosis I differ from typical mitosis (in meristem cells), it is immensely important to analyze the chromosomal distribution of phosphorylated H3S10 histone during atypical pollen mitosis I to comprehend the role of histone phosphorylation in pollen development. We investigated the chromosomal phosphorylation of H3S10 histone during pollen mitosis I in orchids using immunostaining technique. The chromosomal distribution of H3S10ph during pollen mitosis I revealed differential pattern than that of typical mitosis in plants, however, eventually following the similar trends of mitosis in animals where H3S10 phosphorylation begins in the pericentromeric regions first, later extending to the whole chromosomes, and finally declining at anaphase/early cytokinesis (differentiation of vegetative and generative cells). The study suggests that the chromosomal distribution of H3S10ph during cell division is not universal and can be altered between different cell types encoded for diverse cellular processes. During pollen development, phosphorylation of histone might play a critical role in chromosome condensation events throughout pollen mitosis I in plants.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
In plants, two distinct and successive developmental phases, microsporogenesis and microgametogenesis, lead to the production of the mature pollen. Such pollen developmental pathway instigates with meiosis in pollen mother cell, which leads to the formation of a tetrad of four haploid microspores. Further, each of the four microspores undergoes first pollen mitosis (pollen mitosis I), which results in the formation of two cells of different fates and sizes, i.e., generative and vegetative cells (Tanaka 1997; Borg et al. 2009). Therefore, pollen development in plants consists of an exceptional cellular reprogramming that eventually stimulates the switching of cellular and nuclear machinery from meiosis (microsporogenesis) to mitosis (pollen mitosis I/II). Such dynamic cellular and nuclear machinery eventually serves as an excellent model to study the dynamic relationship of chromosomal histone phosphorylation at “meiosis-mitosis shift” during pollen development. Phosphorylation of H3S10 histone has been well documented and specifically correlated with cell cycle progression, chromosome condensation, and segregation processes during mitosis and meiosis (Hans and Dimitrov 2001; Johansen and Johansen 2006; Kouzarides 2007; Sharma et al. 2015). In this context, critical survey of literature suggests that during meiosis I, the hyper-phosphorylation of H3S10 is initiated at the leptotene-zygotene shift, highest during metaphase I, and terminated toward interkinesis. While, at meiosis II, it only occurs at pericentromeric regions from prophase II until telophase II and follows the same trend as evident in mitosis (Manzanero et al. 2000; Fuchs et al. 2006; Houben et al. 2007; Granot et al. 2008; Oliver et al. 2013; Marcon-Tavares et al. 2014). Although the progression of H3 phosphorylation during cell cycle is highly conserved, however, its spatial and temporal distribution was found to be altered to some extent among species, groups, cell types, and organisms (Fuchs et al. 2006; Houben et al. 2007; Sharma et al. 2015). Since the first mitosis in microspore differs from typical mitosis (in meristem cells), it is immensely important to compare the global chromatin environment associated with phosphorylated H3S10 histone in different cell types that undergo mitotic divisions for diverse cellular processes. Therefore, an attempt has been made to catalogue the cytological consequences underlying chromatin environment associated with phosphorylation of histone H3S10 during pollen mitosis I in orchids. Here, we report the differential chromosomal phosphorylation of H3S10 during pollen mitosis I division along with resulted generative and vegetative cells. The study evidently provides insight into the chromosomal histone phosphorylation pattern associated with atypical mitosis of microspores (pollen mitosis I) in plants.
Materials and methods
Three orchid species viz. Cymbidium sasanami, Cymbidium sinense (2n = 40), and Dendrobium chinsai (2n = 38) and one hybrid, i.e., Dendrobium specio-kingianum ‘Red Splash’ (2n = 38), of the Orchidaceae family were used for the study. Flower buds with appropriate size for pollen mitosis were used as research materials after necessary confirmation of the desired stages. The flower buds were fixed in PHEMES buffer (50 mM PIPES, 25 mM HEPES, 5 mM MgCl2 and 5 mM EGTA, 0.35 M sorbitol, pH 6.9) containing 3 % (w/v) paraformaldehyde and 0.2 % (v/v) Triton X-100 and washed twice in phosphate-buffered saline (PBS, pH 7.4). The anthers were digested for 30–45 min at 37 °C in the mixture of 1 % (w/v) Cellulase Onozuka RS (Yakult Pharmaceutical Industry Co. Ltd), 1 % Cytohelicase (Sigma), and 0.5 % (w/v) Pectolyase Y-23 (Seishin Pharmaceuticals) prepared in PBS. The digested anthers were washed twice in PBS and then squashed in a drop of PBS on the slides coated with poly-l-lysine (Sigma). The slides were incubated for a minimum of 12 h at 4 °C with a rabbit polyclonal anti-H3S10ph (diluted 1:500 in PBS containing 3 % bovine serum albumin (BSA); Santa Cruz Biotech. SC-8656R). After washing in PBS, the slides were incubated for 1 h at 37 °C with an Alexa Fluor 546-labeled goat anti-rabbit antibody (1:100 diluted in 3 % BSA/PBS; Molecular Probes 11035). After final washes in PBS, the chromosome preparations were counterstained with 4′,6-diamidino-2-phenylindole hydrochloride (DAPI) and mounted with ProLong gold antifade mountant (Life Technologies, P10144). Immunosignals and stained chromosomes were captured using a CCD camera (Hamamatsu Photonics, model 4880) equipped with a Axioscope fluorescence microscope (Zeiss), and image analysis was performed as described previously (Suzuki et al. 2010).
Results and discussion
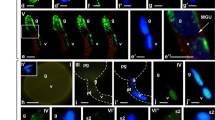
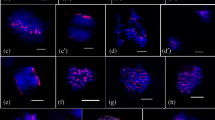
Brown and Lemmon (1991, 1992, 1994) analyzed the cytoskeleton and ultrastructure of first mitosis in the pollen of Cypripedium and Phalaenopsis orchids using immunofluorescence and electron microscopy. This classical report evidently demonstrated the structure and progression of pollen mitotic divisions in orchids. As similar to other plant taxa, microsporogenesis in orchids includes asymmetric and atypical mitosis (pollen mitosis I) in the unreleased four microspores within tetrad. The tetrad of four haploid young microspores did not release the microspore individually and collectively underwent pollen mitotic events to produce pollen grains. Eventually, pollen mitosis in orchids, different stages starting from prophase to cytokinesis, could be identified precisely. Such pollen mitotic I (PM-I) divisions in each microspore ultimately resulted in an inward large vegetative cell and outward smaller generative cell. Most of the stages associated with PM-I (prophase to anaphases) were identified precisely in conventional cytological preparations (supplementary Figs. S1–S3), as well as subsequent immunostained preparations too (Figs. 1 and 2). Further, the chromosomal distribution of H3S10ph during different stages of PM-I was determined precisely. The immunosignals of phosphorylated H3S10 were first appeared at PM-I prophase (Fig. 1a) in diffused manner and declined at PM-1 anaphases. Such observation supports the view of earlier workers (Fuchs et al. 2006; Houben et al. 2007) that phosphorylation of H3S10 begins with the entry of the cell into mitotic cell cycle and declines toward the exit of the cycle. However, in contrast, peri-centromeric distribution of H3S10ph was observed during late prophase or pro-metaphase of PM-I (Fig. 1b–d). In angiosperm, phosphorylation of H3S10 during typical mitotic-metaphase and meiosis II was found to be strongly associated with peri-centromeric regions and suggested a major role in chromosome condensation and sister centromere cohesion respectively (Manzanero et al. 2000; Houben et al. 2007; Caperta et al. 2008). However, one of the interesting features associated with the study involves intense hyper-phosphorylation along the entire chromosome length (Fig. 2a–d) during metaphase of pollen mitosis I. Notably, such uniform labeling of metaphase chromosomes was also found to be unique in typical mitosis of holokinetic plant species, i.e., Luzula luzuloides and Rhynchospora tenuis (Gernand et al. 2003; Guerra et al. 2006), and in meiosis I of angiosperms with monocentric chromosomes (Houben et al. 1999; Manzanero et al. 2000; Gernand et al. 2003; Paula et al. 2013). However, overall observation suggests that the progression of chromosomal histone phosphorylation during pollen mitosis I in orchids more or less is similar to that of mitosis in mammalian cells where phosphorylation of H3S10 firstly appeared at pericentromeric regions and then spread throughout the chromosome arms as mitotic stages proceed (Hendzel et al. 1997; Goto et al. 1999). In this context, several reports on phosphorylation of histone have successfully demonstrated its association with condensation of mitotic chromatin in mammalian cells (Gurley et al. 1974; Prigent and Dimitrov 2003; Nowak and Corces 2004; Houben et al. 2007). Finally, a significant decrease in phosphorylation with dispersed immunolabeling pattern was observed during the PM-I anaphases/early cytokinesis (differentiation of vegetative and generative cells) (Fig. 2e). The diminishing of phosphorylated immunosignals is a common phenomenon associated with the distribution of H3S10ph during typical mitosis in plants (Houben et al. 2007; Marcon-Tavares et al. 2014). Overall observation suggests that the cell cycle-dependent spatial and temporal distribution of chromosomal H3S10ph is not universal between different cell types (e.g., microspore and meristematic cell) programmed for diverse cellular processes as advocated earlier too (Fuchs et al. 2006; Houben et al. 2007; Sharma et al. 2015).
Chromosomal distribution pattern of H3S10ph during early pollen mitosis I stages in orchids. a–a” Prophase showing the early immunosignals of H3S10ph. b–b” to d–d” Pro-metaphase showing the pericentromeric distribution of H3S10ph in Cymbidium sasanami (b–b”), Cymbidium sinense (c–c”), and Dendrobium chinsai (d–d”). Counterstain DAPI and immunosignals of anti-H3S10ph are pseudocolored in blue and in red, respectively. Scale bar = 10 μm
Chromosomal distribution pattern of H3S10ph during pollen mitosis I in orchids. a–a” to d–d” Metaphase showing the distribution of H3S10ph along the entire length of the chromosomes in Cymbidium sinense (a–a”) and Cymbidium sasanami (b–b”), metaphase in single microscope which seems to be busted out from tetrad of Dendrobium chinsai (c–c”), and metaphase in single microscope of Dendrobium specio-kingianum ‘Red Splash’ (d–d”). e–e” Anaphase showing the decline in signals of anti-H3S10ph (Cymbidium sasanami). f–f” Vegetative cell (V) showing the presence of anti-H3S10ph immunosignals whereas generative cell (G) did not show the any anti-H3S10ph signals (Cymbidium sinense, arrow marks showing chromocenters). Counterstain DAPI and immunosignals of anti-H3S10ph are pseudo-colored in blue and in red, respectively. Scale bar = 10 μm
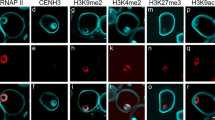
In the present study, the dispersed smaller dots like phosphorylation pattern of H3S10ph were observed in vegetative nuclei (VN) only whereas generative nuclei (GN) did not show any immunosignal during this stage (Fig. 2f). On the other hand, the accumulation of densely DAPI-stained nuclear materials in VN without any signal of H3S10 phosphorylation indicates the probable heterochromatic regions (Figs. 2f and S4; arrow marked). Earlier reports suggest that the VN have diffuse and transcriptionally active chromatin, whereas the generative cells contain highly condensed and transcriptionally inactive chromatin (Saito et al. 1997; Borg et al. 2009). Several studies have already documented the distribution of diverse histone modification marks in generative and vegetative cells of several plant species including lily (Okada et al. 2006; Sano and Tanaka 2010), Arabidopsis (Cartagena et al. 2008), oak (Ribeiro et al. 2009), rye (Houben et al. 2011), and, recently, in barley (Pandey et al. 2013). However, most of the abovementioned reports were actually focused on distribution of eu/heterochromatin-specific histone modification marks for, e.g., H3K4me2, H3K9me2; H3K27me2/3, H3K9ac, H3K36me, and H4K5/8 that suggest that the chromatin organization in vegetative and generative cells of pollen grains is not universal across angiosperms and subjected to differential expression (reviewed in Sharma et al. 2015). Notably, the H3S10 phosphorylation is a versatile histone modification that is found to be involved in diverse nuclear functions encompassing transcription activation of several genes for diverse cellular programming besides chromosome condensation events during cell cycle progression in mitosis and meiosis (Prigent and Dimitrov 2003; Nowak and Corces 2004). Though, in the framework of the present observations, it may be quite premature to suggest the plausible role of phosphorylation of H3S10 in vegetative cell; however, the co-localization of other histone modification marks along with phosphorylated histones may shed light on the precise chromatin organization of vegetative/generative cells of the pollen. In conclusion, during pollen development, phosphorylation of histone might play a critical role in chromosome condensation events throughout pollen mitosis I in plants. However, spatial and temporal distribution of H3S10ph can be altered between different cell types programmed for diverse cellular processes.
References
Borg M, Brownfield L, Twell D (2009) Male gametophyte development: a molecular perspective. J Exp Bot 60:1465–1478
Brown RC, Lemmon BE (1991) Pollen development in orchids. 3. A novel generative pole microtubule system predicts unequal pollen mitosis. J Cell Sci 99:273–281
Brown RC, Lemmon BE (1992) Pollen development in orchids. 4. Cytoskeleton and ultrastructure of the unequal pollen mitosis in Phalaenopsis. Protoplasma 167:183–192
Brown RC, Lemmon BE (1994) Pollen mitosis in the slipper orchid Cypripedium fasciculatum. Sex Plant Reprod 7:87–94
Caperta AD, Rosa M, Delgado M, Karimi R, Demidov D, Viegas W, Houben A (2008) Distribution patterns of phosphorylated Thr 3 and Thr 32 of histone H3 in plant mitosis and meiosis. Cytogenet Genome Res 122:73–79
Cartagena JA, Matsunaga S, Seki M, Kurihara D, Yokoyama M, Shinozaki K, Fujimoto S, Azumi Y, Uchiyama S, Fukui K (2008) The Arabidopsis SDG4 contributes to the regulation of pollen tube growth by methylation of histone H3 lysines 4 and 36 in mature pollen. Dev Biol 315:355–368
Fuchs J, Demidov D, Houben A, Schubert I (2006) Chromosomal histone modification patterns—from conservation to diversity. Trends Plant Sci 11:199–208
Gernand D, Demidov D, Houben A (2003) The temporal and spatial pattern of histone H3 phosphorylation at serine 28 and serine 10 is similar in plants but differs between mono- and polycentric chromosomes. Cytogenet Genome Res 101:172–176
Goto H, Tomono Y, Ajiro K, Kosako H, Fujita M et al (1999) Identification of a novel phosphorylation site on histone H3 coupled with mitotic chromosome condensation. J Biol Chem 274:25543–25549
Granot G, Sikron-Persi N, Li Y, Grafi G (2008) Phosphorylated H3S10 occurs in distinct regions of the nucleolus in differentiated leaf cells. Biochim Biophys Acta 1789:220–224
Guerra M, Brasileiro-Vidal AC, Arana P, Puertas MJ (2006) Mitotic microtubule development and histone H3 phosphorylation in the holocentric chromosomes of Rhynchospora tenuis (Cyperaceae). Genetica 126:33–41
Gurley LR, Walters RA, Tobey RA (1974) Cell cycle-specific changes in histone phosphorylation associated with cell proliferation and chromosome condensation. J Cell Biol 60:356–364
Hans F, Dimitrov S (2001) Histone H3 phosphorylation and cell division. Oncogene 20:3021–3027
Hendzel MJ, Wei Y, Mancini MA, Van Hooser A, Ranalli T, Brinkley BR, Bazett-Jones DP, Allis CD (1997) Mitosis-specific phosphorylation of histone H3 initiates primarily within pericentromeric heterochromatin during G2 and spreads in an ordered fashion coincident with mitotic chromosome condensation. Chromosoma 106:348–360
Houben A, Wako T, Furushima-Shimogawara R, Presting G, Kunzel G, Schubert I, Fukui K (1999) The cell cycle dependent phosphorylation of histone H3 is correlated with the condensation of plant mitotic chromosomes. Plant J 18:675–679
Houben A, Demidov D, Caperta AD, Karimi R, Agueci F, Vlasenko L (2007) Phosphorylation of histone H3 in plants—a dynamic affair. Biochim Biophys Acta 1769:308–315
Houben A, Kumke K, Nagaki K, Hause G (2011) CENH3 distribution and differential chromatin modifications during pollen development in rye (Secale cereale L.). Chromosome Res 19:471–480
Johansen KM, Johansen J (2006) Regulation of chromatin structure by histone H3S10 phosphorylation. Chromosome Res 14:393–404
Kouzarides T (2007) Chromatin modifications and their function. Cell 128:693–705
Manzanero S, Arana P, Puertas MJ, Houben A (2000) The chromosomal distribution of phosphorylated histone H3 differs between plants and animals at meiosis. Chromosoma 109:308–317
Marcon-Tavares AB, Felinto F, Feitoza L, Silva AE B e, Guerra M (2014) Different patterns of chromosomal histone H3 phosphorylation in land plants. Cytogenet Genome Res 143:136–143
Nowak SJ, Corces VG (2004) Phosphorylation of histone H3: a balancing act between chromosome condensation and transcriptional activation. Trends Genet 20:214–220
Okada M, Cheeseman IM, Hori T (2006) The CENP-H-I complex is required for the efficient incorporation of newly synthesized CENP-A into centromeres. Nat Cell Biol 8:446–457
Oliver C, Pradillo M, Corredor E, Cuñado N (2013) The dynamics of histone H3 modifications is species-specific in plant meiosis. Planta 238:23–33
Pandey P, Houben A, Kumlehn J, Melzer M, Rutten T (2013) Chromatin alterations during pollen development in Hordeum vulgare. Cytogenet Genome Res 141:50–57
Paula CMP, Techio VH, Sobrinho FS, Freitas AS (2013) Distribution pattern of histone H3 phosphorylation at serine 10 during mitosis and meiosis in Brachiaria species. J Genet 92:259–266
Prigent C, Dimitrov S (2003) Phosphorylation of serine 10 in histone H3, what for? J Cell Sci 116:3677–3685
Ribeiro T, Viegas W, Morais-Cecílio L (2009) Epigenetic marks in the mature pollen of Quercus suber L. (Fagaceae). Sex Plant Reprod 22:1–7
Saito C, Fujie M, Sakai A, Kuroiwa H, Kuroiwa T (1997) Changes in the extent of the condensation of nuclear chromatin and the localization of RNA during pollen development in Nicotiana tabacum. Cytologia 62:121–132
Sano Y, Tanaka I (2010) Distinct localization of histone H3 methylation in the vegetative nucleus of lily pollen. Cell Biol Int 34:253–259
Sharma SK, Yamamoto M, Mukai Y (2015) Immuno-cytogenetic manifestation of epigenetic chromatin modification marks in plants. Planta 241:291–301
Suzuki G, Shiomi M, Morihana S, Yamamoto M, Mukai Y (2010) DNA methylation and histone modification in onion chromosomes. Genes Genet Syst 85:377–382
Tanaka I (1997) Differentiation of generative and vegetative cells in angiosperm pollen. Sex Plant Reprod 10:1–7
Acknowledgments
We thank Japan Society for the Promotion of Science (JSPS), Japan, for providing postdoctoral fellowship and research grant (SKS, no. P13399/2013). Sincere thanks to Prof. Go Suzuki and all members of Plant Molecular Genetics Laboratory and Osaka Kyoiku University, Osaka, Japan, for their constant encouragement and help.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Additional information
Handling Editor: Anne-Catherine Schmit
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
(PDF 19794 kb)
Rights and permissions
About this article
Cite this article
Sharma, S.K., Yamamoto, M. & Mukai, Y. Distinct chromatin environment associated with phosphorylated H3S10 histone during pollen mitosis I in orchids. Protoplasma 254, 161–165 (2017). https://doi.org/10.1007/s00709-015-0925-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00709-015-0925-z