Abstract
The brown planthopper (BPH), Nilaparvata lugens, is a significant agricultural pest capable of long-distance migration and transmission of viruses that cause severe disease in rice. In this study, we identified a novel segmented RNA virus in a BPH, and this virus exhibited a close relationship to members of a recently discovered virus lineage known as "quenyaviruses" within the viral kingdom Orthornavirae. This newly identified virus was named "Nilaparvata lugens quenyavirus 1" (NLQV1). NLQV1 consists of five positive-sense, single-stranded RNAs, with each segment containing a single open reading frame (ORF). The genomic characteristics and phylogenetic analysis support the classification of NLQV1 as a novel quenyavirus. Notably, all of the genome segments of NLRV contained the 5'-terminal sequence AUCUG. The characteristic virus-derived small interfering RNA (vsiRNA) profile of NLQV1 suggests that the antiviral RNAi pathway of the host BPH was activated in response to virus infection. These findings represent the first documented report of quenyaviruses in planthoppers, contributing to our understanding of quenyaviruses and expanding our knowledge of insect-specific viruses in planthoppers.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
The brown planthopper (BPH), Nilaparvata lugens (Stål, 1854) (Hemiptera: Delphacidae), is a major insect pest that inflicts damage on rice crops and is widely distributed in tropical and temperate countries within the Asian-Pacific region [1, 2]. In addition to causing direct physical damage, BPH also acts as a vector for two important rice viruses, rice ragged stunt virus (RRSV) and rice grassy stunt virus (RGSV), leading to substantial reductions in crop yield [2, 3]. Apart from plant viruses transmitted by BPH, several BPH-specific viruses belonging to the families Iflaviridae (Nilaparvata lugens honeydew virus, NLHV), Dicistroviridae (Himetobi P virus and Nilaparvata lugens C virus), and Spinareoviridae (Nilaparvata lugens reovirus) have been identified [4,5,6,7].
Over the past decade, the use of next-generation sequencing (NGS) and advanced bioinformatics methods has provided novel insights into viral evolution, leading to a new era of virus discovery [8, 9]. Most metagenomic methods rely on similarity searches to identify virus sequences based on inferred homology. However, there may be viruses that cannot be recognized solely through detectable homology, and these are referred to as "dark viruses" [10, 11]. Quenyaviruses were initially discovered in Drosophila melanogaster as "dark viruses" based on their small RNA profiles, which exhibited a characteristic pattern of virus-derived small interfering RNA (vsiRNA) in D. melanogaster, with a modal length of 21 nucleotides (nt), but these showed no identifiable similarity to sequences from known viruses or cellular organisms [10, 12]. Subsequently, through similarity searches, a series of homologous segments were identified in approximately 20 arthropod species, including insects, crustaceans, spiders, and myriapods, forming a completely new lineage named "quenyaviruses". Recent studies have classified quenyaviruses as part of a novel phylum within the viral kingdom Orthornavirae, and using artificial intelligence, they have been grouped into larger clusters of distantly related viruses [8, 13]. The quenyavirus genome consists of five positive-sense, single-stranded RNAs, each containing a single open reading frame (ORF). Proteins encoded by segments S1-S4 bear no detectable similarity to known viral proteins, while the longest segment, S5, is believed to encode the RNA-dependent RNA polymerase (RdRP) [10].
In this study, a novel insect-specific quenyavirus was identified in a BPH and named "Nilaparvata lugens quenyavirus 1" (NLQV1). The complete nucleotide sequences of the five segments of this virus were determined and characterized. BPH samples were initially collected in Vietnam as described in our previous work and subsequently maintained in a phytotron of our laboratory under the following conditions: 26 ± 1℃, 6 ± 5% relative humidity, and a 16:8 h (light/dark) photoperiod [1]. To identify RNA viruses in this population, a combination of nymph and adult insects (10 planthoppers in total) was used, and total RNA was extracted using TRIzol Reagent (Invitrogen, MA, USA). PolyA-selected RNA was used for transcriptome sequencing and whole-genome sequencing. The RNA library used for transcriptome sequencing was constructed and sequenced on an Illumina HiSeq 4000 platform (Illumina, San Diego, CA, USA). Reads were filtered to remove low-quality and adapter sequences using Trimmomatic 0.39 (LEADING:3 TRAILING:3 SLIDINGWINDOW:4:15 MINLEN:36 TOPHRED33), and de novo assembly was carried out using Trinity software (version 2.8.5) with default parameters [14], resulting in a total of 127,557 contigs. These contigs were compared to sequences in the NCBI viral RefSeq database using Diamond BLASTx. As a result, five viral contigs were identified that showed the most sequence similarity to reported quenyaviruses, with identity values ranging from 31.26% to 50.97% (Table 1). Furthermore, to validate the five segments and avoid false matches, a BLAST search against the entire NCBI nucleotide (NT) and non-redundant (NR) protein databases was conducted. The presence of NLQV1 was confirmed through reverse transcription PCR (RT-PCR), and the complete genome sequence of NLQV1 was successfully obtained using a SMARTer RACE 5'/3' Kit. The five segments of NLQV1 were then verified by Sanger sequencing. The primers used in this study are listed in Supplementary Table S1.
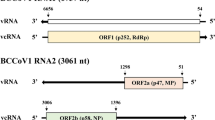
The complete genome of NLQV1 is 9,288 nt in length and comprises five segments, with each segment containing a polyA tail. Segments S1-4 range from 1,696 to 1,829 nt in length, and S5 is 2,206 nt long, excluding the polyA tail (GenBank accession numbers: PP681291-PP681295) (Table 1, Fig. 1A). Based on predictions made using NCBI ORFfinder, each segment is monocistronic and contains a single open reading frame (Fig. 1A). Segment 1 was predicted to encode an mRNA-capping enzyme domain, and segment 5 was predicted to encode an RNA-dependent RNA polymerase (RdRP), based on the presence of conserved domains identified using InterProScan and HHpred (Fig. 1A). These characteristics aligned with those of the quenyaviruses [10]. The five segments of NLQV1 have a moderate G+C content, ranging from 49.86% (S3) to 52.33% (S2) (Table 1), with an average G+C content of 50.99%. The lengths of untranslated regions (UTR) varies across the five genome segments, with the lengths of the 5'-UTRs ranging from 33 to 55 nt and those of the 3'-UTRs ranging from 154 to 449 nt. All five segments have the sequence motif AUCUG at their 5'-terminus (Table 1, Fig. 1A), suggesting that the genome segments of other quenyaviruses might also have conserved nucleotides at their 5’-termini. To assess the abundance and coverage of NLQV1, RNA-seq sequences were aligned with the reconstructed complete genome sequence of NLQV1. A total of 538,595 reads mapped perfectly to the NLQV1 genome, accounting for 2.25% of the RNA-seq reads (Fig. 1). The transcripts were distributed throughout the viral genome, indicating efficient replication of NLQV1 in the host insect.
Genome structure and phylogenetic analysis of NLQV1. A Genome structure and RNA-seq read coverage of NLQV1. UTR, untranslated region. B Relationship of the quenyaviruses to other RNA viruses of the viral kingdom Orthornavirae. An unrooted phylogenetic tree was constructed based on RNA-dependent RNA polymerase sequences, showed showing the relationships between quenyaviruses (including NLQV1) and representatives of other virus families. C Phylogenetic relationship of NLQV1 to other quenyaviruses. A close-up view of the region of the tree within the box with the dotted frame in panel B, corresponding to the quenyaviruses, is shown. Nodes with bootstrap values >50 are indicated by blue circles, with larger circles indicating higher bootstrap values
To investigate the taxonomic status of NLQV1, the predicted amino acid (aa) sequence of its RdRp and those of previously reported quenyaviruses, obtained from the NCBI database, were aligned using MAFFT (https://www.ebi.ac.uk/Tools/msa/mafft/, accessed on 12 April 2023) [10, 15]. Gaps were trimmed using trimAl, and the optimal aa substitution model was determined using ModelTest-NG with default parameters [16]. A maximum-likelihood (ML) phylogenetic tree was then constructed using IQ-tree with 1000 bootstrap replications [17, 18]. The results demonstrated that the RdRPs of quenyaviruses formed a monophyletic clade, supporting their potential inclusion in a new family (Fig. 1B), as suggested previously [8]. NLQV1 was clearly located within this clade, showing a close relationship to Sina virus, Nete virus, and Nai virus, with high bootstrap support (Fig. 1C). Therefore, our phylogenetic analysis supports the classification of NLQV1 as a new quenyavirus.
One of the key antiviral pathways utilized by insects to combat viral invaders is the small interfering RNA (siRNA)-based RNA silencing pathway. In this pathway, viral RNA is cleaved, resulting in the accumulation of vsiRNA, which is a signal for the activation of the RNAi antiviral pathway [19]. To gain a better understanding of siRNA-based immunity against NLQV1, a small RNA library from a BPH was created using an Illumina TruSeq Small RNA Sample Preparation Kit (Illumina, San Diego, California, USA) and sequenced on an Illumina HiSeq 2500 platform. The quality of the library was ensured through adapter removal and elimination of low-quality sequences, using the Cutadapt tool [20]. The quality-trimmed sRNA reads were subsequently mapped back to the complete genome of NLQV1 using the zero-mismatch Bowtie software as described previously [21]. The sRNA sequence library contained a total of 13,471 vsiRNA reads that matched NLQV1, accounting for 0.04% of the library. The vsiRNAs derived from the five genome segments of NLQV1 all exhibited a distribution pattern peaking at 21 and 22 nt and originated equally from both sense and antisense strands of the viral genome. NLQV1-derived RNAs showed a length preference similar to that of many other viruses that infect planthoppers [21,22,23,24]. These vsiRNAs were evenly distributed across all five segments of the NLQV1 genome, with certain regions showing preferential targeting by Dicer (Fig. 2). This characteristic vsiRNA profile suggests that the host antiviral RNAi pathway plays an active role in the response to NLQV1 infection.
In conclusion, we discovered a novel insect-specific quenyavirus, NLQV1, in a BPH and determined its complete genome sequence. Quenyaviruses form a novel viral clade within the viral kingdom Orthornavirae and have a broad distribution among arthropods. This study represents the first report of a quenyavirus in planthoppers, thereby enhancing our knowledge of quenyaviruses and contributing to a better understanding of insect-specific viruses in planthoppers.
References
Hu Q, Zhuo J-C, Ye Y-X et al (2024) The genomic history and global migration of a wind-borne pest. Sci Adv 10:17
Nault LR, Ammar ED (1989) Leafhopper and planthopper transmission of plant viruses. Annu Rev Entomol 34:503
Ammar E-D, Nault LR (2002) Virus transmission by leafhoppers, planthoppers and treehoppers (auchenorrhyncha, homoptera). Adv Bot Res 36:141–167. https://doi.org/10.1016/S0065-2296(02)36062-2
Qi Y-H, Ye Z-X, Zhang C-X et al (2023) Diversity of RNA viruses in agricultural insects. Comput Struct Biotechnol J 21:4312–4321. https://doi.org/10.1016/j.csbj.2023.08.036
Noda H, Ishikawa K, Hibino H, Omura T (1991) A reovirus in the brown planthopper, Nilaparvata lugens. J Gen Virol 72:2425–2430. https://doi.org/10.1099/0022-1317-72-10-2425
Nakashima N, Kawahara N, Omura T, Noda H (2006) Characterization of a novel satellite virus and a strain of Himetobi P virus (Dicistroviridae) from the brown planthopper, Nilaparvata lugens. J Invertebr Pathol 91:53–56. https://doi.org/10.1016/j.jip.2005.10.001
Murakami R, Suetsugu Y, Kobayashi T, Nakashima N (2013) The genome sequence and transmission of an iflavirus from the brown planthopper, Nilaparvata lugens. Virus Res 176:179–187. https://doi.org/10.1016/j.virusres.2013.06.005
Zayed AA, Wainaina JM, Dominguez-Huerta G et al (2022) Cryptic and abundant marine viruses at the evolutionary origins of Earth’s RNA virome. Science 376:156–162. https://doi.org/10.1126/science.abm5847
Koonin EV, Krupovic M, Dolja VV (2022) The global virome: How much diversity and how many independent origins? Environ Microbiol 1462–2920:16207. https://doi.org/10.1111/1462-2920.16207
Obbard DJ, Shi M, Roberts KE et al (2020) A new lineage of segmented RNA viruses infecting animals. Virus Evolution 6:vez061. https://doi.org/10.1093/ve/vez061
Powerful Sequence Similarity Search Methods and In-Depth Manual Analyses Can Identify Remote Homologs in Many Apparently “Orphan” Viral Proteins. https://journals.asm.org/doi/epub/https://doi.org/10.1128/jvi.02595-13. Accessed 3 Feb 2024
Webster CL, Waldron FM, Robertson S et al (2015) The discovery, distribution, and evolution of viruses associated with drosophila melanogaster. PLoS Biol 13:e1002210. https://doi.org/10.1371/journal.pbio.1002210
Hou X, He Y, Fang P et al (2023) Using artificial intelligence to document the hidden RNA virosphere (preprint). https://doi.org/10.1101/2023.04.18.537342
Grabherr MG, Haas BJ, Yassour M et al (2011) Full-length transcriptome assembly from RNA-Seq data without a reference genome. Nat Biotechnol 29:644
Katoh K, Standley DM (2013) MAFFT Multiple sequence alignment software version 7: improvements in performance and usability. Mol Biol Evol 30:772–780. https://doi.org/10.1093/molbev/mst010
Capella-Gutiérrez S, Silla-Martínez JM, Gabaldón T (2009) trimAl: a tool for automated alignment trimming in large-scale phylogenetic analyses. Bioinformatics 25:1972–1973. https://doi.org/10.1093/bioinformatics/btp348
Kozlov AM, Darriba D, Flouri T et al (2019) RAxML-NG: a fast, scalable and user-friendly tool for maximum likelihood phylogenetic inference. Bioinformatics 35:4453–4455. https://doi.org/10.1093/bioinformatics/btz305
Nguyen L-T, Schmidt HA, Von Haeseler A, Minh BQ (2015) IQ-TREE: A fast and effective stochastic algorithm for estimating maximum-likelihood phylogenies. Mol Biol Evol 32:268–274. https://doi.org/10.1093/molbev/msu300
Gammon DB, Mello CC (2015) RNA interference-mediated antiviral defense in insects. Curr Opin Insect Sci 8:111–120. https://doi.org/10.1016/j.cois.2015.01.006
Kechin A, Boyarskikh U, Kel A, Filipenko M (2017) cutPrimers: A new tool for accurate cutting of primers from reads of targeted next generation sequencing. J Comput Biol 24:1138–1143. https://doi.org/10.1089/cmb.2017.0096
Li J, Andika IB, Shen J et al (2013) Characterization of rice black-streaked dwarf virus- and rice stripe virus-derived siRNAs in singly and doubly infected insect vector laodelphax striatellus. PLoS ONE 8:e66007. https://doi.org/10.1371/journal.pone.0066007
Li T, Ye Z-X, Feng K-H et al (2024) Molecular and biological characterization of a bunyavirus infecting the brown planthopper (Nilaparvata lugens). J Gen Virol 105:001977. https://doi.org/10.1099/jgv.0.001977
Lu G, Zhang X-D, Xu Z-T et al (2022) Complete sequence and genetic characterization of a novel insect-specific reovirus discovered from Laodelphax striatellus. Virology 570:117–122. https://doi.org/10.1016/j.virol.2022.03.011
Chao S, Wang H, Yan Q et al (2021) Metatranscriptomic Sequencing Suggests the Presence of Novel RNA Viruses in Rice Transmitted by Brown Planthopper. Viruses 13:2464. https://doi.org/10.3390/v13122464
Funding
This work was supported by the National Natural Science Foundation of China (32270150, 32270146, U20A2036) and the Ningbo Youth Leading Talent Program for Scientific and Technological Innovation (2023QL035).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Ethic approval
No experimental work with humans was done in this study.
Conflict of interest
The authors declare that there are no conflicts of interest.
Additional information
Handling Editor: Simona Abba
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Ning, C., Ye, ZX., Xu, ZT. et al. Complete genome sequence and genetic characterization of a novel segmented RNA virus infecting Nilaparvata lugens. Arch Virol 169, 141 (2024). https://doi.org/10.1007/s00705-024-06066-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00705-024-06066-8