Abstract
African swine fever (ASF) is an infectious disease caused by ASF virus (ASFV), which is characterized by high infectivity, rapid onset of disease, and a high mortality rate. Outbreaks of ASFV have caused great economic losses to the global pig industry, and there is a need to develop safe and effective vaccines. In this study, two recombinant pseudorabies virus (PRV) strains, rGXGG-2016-ΔgI/ΔgE-EP364R and rGXGG-2016-ΔgI/ΔgE-B119L, expressing the EP364R and B119L protein, respectively, of ASFV, were constructed by homologous recombination technology. Western blotting and immunofluorescence analysis showed that these foreign proteins were expressed in cells infected with the recombinant strains. The strains showed good genetic stability and proliferative characteristics for 20 passages in BHK-21 cells. Both of these strains were immunogenic in mice, inducing the production of specific antibodies against the expressed ASFV proteins while providing protection against lethal challenge with PRV. Thus, the recombinant strains rGXGG-2016-ΔgI/ΔgE-EP364R and rGXGG-2016-ΔgI/ΔgE-B119L could be used as candidate vaccines for both ASFV and PRV. In addition, our study identifies two potential target genes for the development of safe and efficient ASFV vaccines, provides a reference for the construction of bivalent ASFV and PRV vaccines, and demonstrates the feasibility of developing a live ASFV vector vaccine.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
African swine fever (ASF) mainly affects domestic pigs and wild boars [1]. Its pathogenesis is short and its mortality rate is high [2]. An ASF epidemic was first reported in Kenya in 1921, after which the disease spread throughout Africa [3]. In 1957, an outbreak of ASF spread from West Africa to Portugal, marking the beginning of the European epidemic [4]. In China, ASF was first reported in 2018 on a pig farm in the city of Shenyang, Liaoning Province, and it quickly spread to various provinces across the country [5]. Although ASF has been prevalent for nearly 100 years, there is neither an effective vaccine nor a drug to prevent and control the epidemic at home or abroad. The genome of the causal agent, ASF virus (ASFV), is very large, and many of its genes play an important role in immune escape [6,7,8,9,10]. The genes encoding the viral proteins B119L and EP364R are potential candidates for use in anti-ASFV live vector vaccine design.
The EP364R protein is mainly involved in the replication and repair of viral DNA and is related to the principal Holliday junction resolvase of eukaryotes (Mus81) [11]. This protein is able to induce a strong continuous cellular immune response [12, 13]. Previous studies have shown that the EP364R protein of ASFV has the characteristics of a nuclease that can inhibit the IFN signaling pathway of the host's antiviral innate immune response. The site-directed mutations Y76S and N78A in the EP364R protein have been shown to restore IFN production in infected cells [7]. The pB119L (9GL) protein of ASFV, which belongs to the Erv1p/Alrp family of sulfhydryl oxidases, is a late non-structural protein that is necessary for the correct assembly of the ASFV virion [14]. Its gene is highly conserved among ASFV isolates. It is involved in the maturation of viral particles and the growth of the virus in macrophages in vitro, and it has been shown to affect the virulence of the virus in pigs [15]. As a gene that confers virulence, B119L is usually deleted in the development of live attenuated vaccines. For example, O'Donnell et al. [16] found that low-dose immunization of pigs with an ASFV-G strain in which the B119L gene was deleted induced effective protection against challenge with the parental strain. In contrast, strains with deletions of MGF360/505 or CD2V/EP153R did not provide protection against the parental strain [17, 18]. However, it was found that the protection was enhanced by deleting the UK gene [19], indicating that this gene plays an important role in the replication of ASFV in the host.
Pseudorabies virus (PRV) is a member of the subfamily Alphaherpesvirinae of the family Orthoherpesviridae [20]. This virus can persist in the trigeminal ganglion of pigs for their entire lifetimes, causing pseudorabies (PR) [21]. PRV can cause death in piglets and reproductive disorders in sows, and it is often encountered in mixed infections with other disease agents, resulting in the death of pigs at all developmental stages. PRV can also infect a variety of other farm animals [22]. Liu et al. [23] isolated a recombinant strain, hSD-1/2019, derived from a variant and a classical PRV strain, from cases of acute human encephalitis. This suggests that PRV can spread across species and infect humans, making it a potentially emerging zoonotic pathogen. The genome of PRV is a double-stranded linear DNA molecule with a length of approximately 142 kb [24]. There are many non-essential genes in the relatively large genome of PRV that are not related to viral replication. The gE, gI, and TK genes can be deleted in order to develop genetically engineered vaccines against PRV. For example, PRV HB2000 [25], SA215 [26], gE-/gI-/TK-PRV (HeN1) [27], and other new PR vaccine strains with gE/gI/TK triple gene deletions have shown good protection against PRV infection. Two of these, HB2000 and SA215, have been used in commercial vaccines. PRV has also been widely used as an experimental viral vector for expression of foreign proteins, including the E2 protein of classical swine fever virus (CSFV) [28], the VP2 protein of porcine parvovirus and porcine IL-6 [29], and the CD2v protein of ASFV [30].
PRV strains are currently divided into two genotypes. Genotype I strains are mainly found in European and American countries, whereas genotype II strains are mainly found in Asian countries. Genotype II strains are mainly concentrated in China and are further divided into classical and variant strains. The Fa [31], Ea [32], and SC [33] strains belong to the group of classical strains, and the HeN1 [34], TJ [35], and JS-2012 [36] strains are variant strains. The vaccine strain Bartha-K61 belongs to genotype I [37], and vaccine preparations based on this strain do not provide comprehensive protection against the PRV variants found in Asian countries [38, 39]. Hence, it is necessary to develop effective novel vaccines to prevent future outbreaks.
In this study, to investigate the applicability of a recombinant multivalent vaccine expressing an ASFV gene using PRV as a vector, the recombinant PRV strains rGXGG-2016-ΔgI/ΔgE-EP364R and rGXGG-2016-ΔgI/ΔgE-B119L, expressing the pEP364R and pB119L protein, respectively, were constructed by homologous recombination technology. The virulence, immunogenicity, and protective efficacy of these recombinant strains were then evaluated in a mouse model.
Materials and methods
Cells, viruses, antibodies, animals, and ethical approval
Vero (African green monkey kidney), HEK 293T (human embryonic kidney), and BHK-21 (baby Syrian hamster kidney) cells were cultured in Dulbecco's modified Eagle medium (DMEM; Gibco, California, USA) supplemented with 10% fetal bovine serum (FBS; Gibco). The GXGG-2016 strain (GenBank accession no. OP589231) is a classical PRV strain of genotype II that was isolated in our laboratory. Its TK gene has a natural deletion of 69 amino acids. GXLB-2015 (GenBank accession no. OP605538) is a genotype II variant PRV strain that was isolated in our laboratory [40]. The rGXGG-2016-ΔgI/ΔgE-EGFP strain, which has deletions of the gI and gE genes and expresses the enhanced green fluorescent protein (EGFP), was also constructed previously in our laboratory. Twenty-eight-day-old ICR mice and New Zealand white rabbits were provided by the Laboratory Animal Center of Guangxi Medical University. Our study was approved by the Ethics Committee of Animal Experiments of Guangxi University (protocol number GXU2020-038).
Preparation of rabbit polyclonal antibodies against the ASFV EP364R and B119L proteins
EP364R and B119L gene sequences were synthesized and cloned into the pET-32a(+) and pCold-I vector, respectively. The constructs were sequenced and then used to transform competent Escherichia coli BL21(DE3) cells. Expression of the cloned genes was induced by treatment with 0.5 mM IPTG, recombinant proteins were purified by Ni2+ affinity chromatography (Cowin, Beijing, China), and their antigenicity were confirmed by western blotting. Two New Zealand white rabbits weighing about 2 kg were selected, 2 mL of blood was collected from a vein at the ear margin after 1 week of resting, and the serum was separated for use as a negative control. The recombinant protein was fully emulsified with an equal volume of Freund’s complete adjuvant (Sigma-Aldrich, Darmstadt, Germany), and a dose of 1 mg was used to immunize rabbits subcutaneously. On the 14th and 21st days after the first vaccination, the rabbits were given a booster immunization with the protein emulsified in Freund's incomplete adjuvant (Sigma-Aldrich). On the 26th day after the first immunization, blood was collected from the heart, and the sera were separated by centrifugation and stored at -80 ℃. After the blood collection was completed, the blood vessels were pressed in order to stop the bleeding, and the animals were nursed to reduce the trauma encountered.
Plasmid construction and generation of recombinant strains
rGXGG-2016-ΔgI/ΔgE-EP364R and rGXGG-2016-ΔgI/ΔgE-B119L were used to produce recombinant strains with the gI/gE double gene deletion. First, the B119L and EP364R genes (GenBank accession numbers QBH90547.1 and QBH90562.1, respectively) were chemically synthesized by GENEWIZ Co. Ltd. (New Jersey, USA) and inserted into the pMD-18T plasmid. Then, using PRV GXGG-2016 as a template, the left homologous arm of the gD site and the right homologous arm of the US9 and US2 sites were amplified to construct the parental plasmid pMD-18T-LR. Next, the transfer plasmid pMD-18T-L-EP364R/B119L-EGFP-R, containing the ASFV immunogenic gene expression cassette, was constructed using a single-step cloning technique, and sequencing was performed to verify the absence of mutations. HEK 293T cells infected with the GXGG-2016 strain were transfected with 2.5 μg of the transfer plasmid using Lipofectamine 2000 (Invitrogen, California, USA). The resulting recombinant strains were amplified and purified in Vero cells by plaque isolation. A 2% MEM medium containing 1% low-melting-point agarose was added to Vero cells infected with the virus, and the culture plates were inverted and incubated for 2-3 days. The cell plate was then viewed under a fluorescence microscope (Carl Zeiss AG, Oberkochen, Germany) to pick plaques exhibiting green fluorescence. These plaques were expanded in 24-well plates and then purified for 5-6 rounds until all of the cytopathic lesions exhibited green fluorescence.
To assess the genetic stability of the ASFV genes in the recombinant strains, the rGXGG-2016-ΔgI/ΔgE-EP364R and rGXGG-2016-ΔgI/ΔgE-B119L strains were passaged 20 times consecutively in BHK-21 cells. The viral genomes from passages 5, 10, 15, and 20 were isolated using a TIANamp Virus DNA/RNA Kit (TIANGEN, Beijing, China), the EP364R and B119L genes were amplified by PCR, and the positive viral DNA was sent to Sangon Biotech Co., Ltd. (Shanghai) for gene sequencing to confirm that the recombinant strains maintained the EP364R and B119L genes during cell passage.
Titration of recombinant strains
The recombinant strains were serially diluted tenfold with DMEM containing 2% FBS. The 101- to 1010-fold dilutions were added to Vero cells grown in 96-well plates to 80% confluence, and cells grown in DMEM containing 2% FBS were used as negative controls. The cells were cultured at 37 ℃ in a 5% CO2 incubator and observed under a microscope for 5 days, and cytopathic effects (CPEs) were recorded. The 50% tissue culture infective dose (TCID50) was determined according to the Reed–Muench method [41].
Expression of pEP364R and pB119L (9GL) proteins by the recombinant strains
The expression of foreign genes inserted into the recombinant virus was detected by western blotting and indirect immunofluorescence assay (IFA). BHK-21 cell monolayers grown in 6-well plates were infected with GXGG-2016, rGXGG-2016-ΔgI/ΔgE-EGFP, rGXGG-2016-ΔgI/ΔgE-EP364R, or rGXGG-2016-ΔgI/ΔgE-B119L at a multiplicity of infection (MOI) of 0.1. For indirect IFA, the cell culture medium was discarded, and the cells were washed twice with PBS at 24 hpi. The cells were then fixed with 1 mL of cold methanol and blocked with 1 mL of 1% fraction V bovine serum albumin (BSA; Solarbio). Subsequently, rabbit polyclonal antibodies (PAbs) against the EP364R and B119L proteins were used as the primary antibodies, and CoraLite594-conjugated goat anti-rabbit IgG (H+L; 1:1000; MO BIO, California, USA) was used as the secondary antibody. Expression of the target proteins was detected using a fluorescence microscope.
For western blotting, the supernatant was collected from the infected cells, mixed with loading buffer, and boiled at 100 °C for 5 minutes. The proteins were then separated by sodium dodecyl sulfate polyacrylamide gel electrophoresis (SDS-PAGE) and transferred to a polyvinylidene fluoride (PVDF; Millipore, Massachusetts, USA) membrane. Nonspecific binding was blocked with 5% skim milk at room temperature for 2 h, after which the membrane was incubated with rabbit PAbs against the EP364R and B119L proteins at 4 ℃ for 12 h. Subsequently, the membranes were incubated with HRP-tagged ant-rabbit IgG (1:10000) (MO BIO), and the proteins bands were visualized using an ImageQuant LAS 500 imager (Cytiva, Massachusetts, USA).
Plaque assays
Vero cells were cultured in 6-well cell culture plates until they formed a monolayer. Virus suspensions were diluted 103-106 times in DMEM without serum, and 20 μL of each virus dilution was inoculated onto the cells. Then, 480 μL of DMEM without serum was added to a final volume of 500 μL and the cells were incubated for 1 h at 37 °C in an atmosphere of 5% CO2. The monolayers were then washed twice with phosphate-buffered saline (PBS) (Solarbio, Beijing, China) and further incubated in 2x MEM containing 1% low-melting-point agarose (Solarbio) and 2% FBS for 3-4 days. Subsequently, the medium was removed, and the cells were fixed with 4% paraformaldehyde for 6 h and stained with 5% crystal violet for 30 min. After gently rinsing the plates with distilled water, the cells were dried at room temperature. Ten plaques were randomly selected for each strain, and their surface areas were measured using ImageJ 1.8.0 software (National Institutes of Health, Bethesda, USA) and averaged.
Growth curves of the recombinant strains
To evaluate the growth properties of the recombinant strains in vitro, Vero cells were grown in 24-well plates and infected with rGXGG-2016-ΔgI/ΔgE-EP364R or rGXGG-2016-ΔgI/ΔgE-B119L at an MOI of 0.1. Cell supernatants were collected at 6, 12, 24, 36, 48, 60, 72, 84, 96, and 108 hpi, and the viral titer at each time point was determined by performing a TCID50 assay as described above. The multi-step growth curve for each recombinant virus was plotted using the infection time as the abscissa and the virus titer as the ordinate.
Establishment of indirect ELISAs
The purified target proteins pCold-I-B119L and pET-32a-EP364R were diluted with coating buffer (Solarbio) to correspond to the coating concentration and used as diagnostic antigens to coat 96-well microtiter plates. Five percent skimmed milk was used to block nonspecific binding, the serum to be tested was used as the primary antibody, and HRP-tagged anti-pig IgG (H+L) was used as the secondary antibody. 3,3′,5,5′-Tetramethylbenzidine (TMB) substrate was added to produce a blue color, and the reaction was stopped by adding a termination solution. The absorbance of each well was read at 450 nm on a microplate reader. An indirect ELISA for detecting ASFV antibodies was established by optimizing each condition in the test by titration.
Pathogenicity and immune protection assay
To investigate the pathogenicity of the virus, 28-day-old SPF ICR mice were divided into six groups (groups 1-6, eight mice per group). The mice in groups 1 to 5 were injected subcutaneously with 105 TCID50 of GXLB-2015, GXGG-2016, rGXGG-2016-ΔgI/ΔgE-EGFP, rGXGG-2016-ΔgI/ΔgE-EP364R, and rGXGG-2016-ΔgI/ΔgE-B119L, respectively. The mice in group 6 were injected with the same volume of DMEM and as a negative control. Disease signs and survival were continuously observed, and survival curves were plotted. At about 72 h after immunization, two mice in each group were randomly sacrificed, and the spleen and brain tissues were removed, fixed with 4% paraformaldehyde for 24 h, and embedded in paraffin. Sections of tissues were subjected to H&E staining, and pathological changes were observed under an optical microscope.
In order to evaluate the immune protection effect of the recombinant strains, 28-day-old SPF ICR mice were divided into five groups (groups 1-5, six mice per group). The mice in groups 1 to 4 were injected subcutaneously with 105 TCID50 of GXGG-2016, rGXGG-2016-ΔgI/ΔgE-EGFP, rGXGG-2016-ΔgI/ΔgE-EP364R, and rGXGG-2016-ΔgI/ΔgE-B119L, respectively. The mice in group 5 were injected with the same volume of DMEM and used as a negative control. The mice in each group received a second inoculation one week later, following the same procedure. Serum samples were collected at 7, 14, and 21 days after the second booster immunization dose. PRV gB- and gE-specific antibodies in blood samples were identified using a PRV gB and gE Antibody Test Kit (IDEXX, Delaware, USA) in accordance with the manufacturer’s instructions.
The presence of specific antibodies against the ASFV EP364R and B119L proteins in mice was detected by western blot and IFA, using the serum samples as the primary antibody, and HRP-tagged anti-mouse IgG (1:10,000) and CoraLite594-conjugated goat anti-mouse IgG (H+L; 1:1000) (MO BIO) was used as the secondary antibody for western blotting and IFA, respectively. The antibody levels in serum samples against the EP364R and B119L proteins were also assessed by using the indirect ELISA methods established in this study. Four weeks after the booster immunization, the mice in each group were challenged with the PRV GXLB-2015 strain by subcutaneous injection. Disease signs in the mice were observed, and their survival was recorded.
Statistical analysis
Data were analyzed using GraphPad Prism 8.0 software (GraphPad Software, Inc., California, USA), and the plaque diameters were measured using Image J 1.8.0 software (National Institutes of Health, Bethesda, USA). All experiments were repeated at least three times independently. All values were expressed as the mean ± standard deviation of three or more experiments. Statistical significance is indicated as follows: *, P < 0.05; **, P < 0.01; ***, P < 0.001; ****, P < 0.0001; n.s., P > 0.05.
Results
Generation of recombinant strains expressing the ASFV EP364R and B119L proteins
The recombinant strains rGXGG-2016-ΔgI/ΔgE-EP364R and rGXGG-2016-ΔgI/ΔgE-B119L (Fig. 1A) were continuously passaged 20 times in BHK-21 cells, and the genetic stability of these viruses was evaluated by PCR and sequencing. The results showed that the EP364R, B119L, and gB genes were present in the genome of each generation of recombinant virus, but the gE gene could not be amplified. Sequencing showed that there were no mutations in the inserted foreign genes. These results indicated that the recombinant virus had been successfully constructed, that the gI and gE genes had been stably deleted, and that the foreign EP364R and B119L genes were stably maintained during passage of the virus.
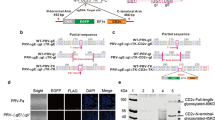
Construction of the recombinant virus. (A) The homologous recombination fragment was inserted between the gI and gE genes. The left and right homologous arms were 1236 and 1347 bp, respectively. The EGFP and ASFV antigen genes were driven by a CMV promoter. (B) Immunofluorescence images showing that BHK-21 cells infected with rGXGG-2016-ΔgE/ΔgI-EP364R and rGXGG-2016-ΔgE/ΔgI-B119L expressed EP364R and B119L protein, respectively. The white scale bars are 150 μm. (C) Western blot showing 41.4-kDa EP364R and14.4-kDa B119L bands produced in cells infected with rGXGG-2016-ΔgE/ΔgI-EP364R and rGXGG-2016-ΔgE/ΔgI-B119L, respectively
Expression of ASFV EP364R and B119L proteins by the recombinant strains
Western blot and IFA were performed using rabbit PAbs against the EP364R and B119L proteins as the primary antibodies. The IFA results showed that, after infection with the two recombinant strains, BHK-21 cells displayed red fluorescence, whereas cells infected with GXGG-2016 or rGXGG-2016-ΔgI/ΔgE-EGFP were negative (Fig. 1B). Western blot analysis also showed the presence of specific bands, of approximately 14 kDa and 41 kDa, corresponding in size to the EP364R and B119L proteins, in the supernatants of cells infected with rGXGG-2016-ΔgI/ΔgE-B119L and rGXGG-2016-ΔgI/ΔgE-EP364R, respectively. No specific bands were found when testing the supernatants of cells infected with GXGG-2016 or rGXGG-2016-ΔgI/ΔgE-EGFP or the negative control (Fig. 1C). These results indicated that the recombinant strains carried the EP364R and B119L protein expression cassettes and that they were successfully expressed in BHK-21 cells.
Proliferation of recombinant strains in vitro
A plaque assay was performed to evaluate the ability of the recombinant strains to proliferate in vitro. As shown in Fig. 2A and B, the mean surface area of plaques produced by rGXGG-2016-ΔgI/ΔgE-B119L and rGXGG-2016-ΔgI/ΔgE-EP364R was 63,068.2 μm2 and 72,814.3 μm2, respectively. These plaques were significantly smaller than those produced by GXGG-2016 (p < 0.001). They were also smaller than those produced by rGXGG-2016-ΔgI/ΔgE-EGFP. This suggests that the pathogenicity of PRV to the cells was reduced by the gI/gE double gene deletion and that the ability of the virus to invade cells was greatly reduced after the foreign genes were inserted.
Insertion of ASFV proteins does not change the proliferation capacity or virulence of rGXGG-2016-ΔgE/ΔgI-EGFP. (A) Viruses (103-106 TCID50) were inoculated onto 1 × 105 Vero cells and cultured for 2 to 3 days. The growth of plaques was observed by crystal violet staining. (B) Plaque measurement. The areas corresponding to 10 randomly selected plaques were measure using a ruler in ImageJ 1.8.0 software. (C) Multistep growth curve Vero cells were inoculated with viruses at an MOI of 0.1, and cell supernatants were collected at different time points (6-108 hpi). The virus titers were calculated by the Reed–Muench method. The value at each time point is the mean of at least three replicates. An unpaired t-test was performed using GraphPad Prism 8.0 (San Diego, CA, USA). *, p < 0.05; ****, p < 0.0001 (n = 10 per group). The bars represent the mean ± SD of three independent experiments (n = 3).
To evaluate the in vitro growth properties of the recombinant strains, Vero cells were infected with the strains at an MOI of 0.1, and the virus titers were determined at different time points, using the Reed-Muench method. As shown in Fig. 2C, the viral titers of the GXGG-2016 were significantly higher than those of the recombinant strains at all of the time points. However, the proliferation trends were the same, indicating that the insertions of EP364R and B119L did not change the growth kinetics of the viruses in vitro.
Induction of B119L/EP364R- and PRV-gB-specific antibodies by the recombinant strains and protection against challenge by virulent viruses
To evaluate their safety, mice were infected with each of the recombinant strains. Mice infected with GXGG-2016, rGXGG-2016-ΔgI/ΔgE-EGFP, rGXGG-2016-ΔgI/ΔgE-EP364R, or rGXGG-2016-ΔgI/ΔgE-B119L were in good condition and showed no clinical signs. However, mice infected with the GXLB-2015 strain developed clinical signs typical of PRV, such as marked scratching and bite marks, and they all died within 5 days of inoculation (Fig. 3A). H&E staining of brain tissue sections showed that GXLB-2015 strain infection induced infiltration of inflammatory cells into the brain, but this was not seen in the other infected groups (Fig. 3B). These results indicated that the recombinant strains constructed in this study were safe and non-pathogenic to mice. The immune effects of the recombinant strains in the mouse model were evaluated by H&E staining of mouse spleen tissue sections. As shown in Fig. 3C, mice infected with rGXGG-2016-ΔgI/ΔgE-EGFP, rGXGG-2016-ΔgI/ΔgE-EP364R, or rGXGG-2016-ΔgI/ΔgE-B119L had obvious lymphocyte infiltration in their spleens, while the others showed no significant changes. This indicated that the attenuated and recombinant strains were immunogenic in mice.
The virulent strain GXLB-2015 causes brain inflammation and death in mice. (A) 28-day-old SPF ICR mice were injected subcutaneously with viruses, and 15-day survival curves were plotted (n = 6 per group). (B) Infection with the variant causes immune cell infiltration in the brain of mice, as indicated by black arrows. (C) Spleen lymphocytes of mice inoculated with the recombinant strains and rGXGG-2016-ΔgI/ΔgE-EGFP were observed to be clustered, as indicated by black arrows.
Using an indirect ELISA method, 4-week-old mice that had been inoculated with rGXGG-2016-ΔgI/ΔgE-EP364R, or rGXGG-2016-ΔgI/ΔgE-B119L were found to be positive for antibodies against the PRV gB protein and the ASFV EP364R or B119L protein (Figs. 4A-C). Except for the parental strain, no antibodies against gE were detected after infection with these viruses (Fig. 4D). Further verification by IFA (Fig. 5A) and western blotting (Fig. 5B) showed that antibodies specific for ASFV EP364R and B119L proteins were present in serum samples from the mice. These data indicated that the recombinant strains were immunogenic and could induce an immune response against the ASFV pEP364R and pB119L proteins in mice.
Production of specific antibodies against B119L, EP364R, and gB after immunization with the recombinant viruses, measured by ELISA. (A) Anti-B119L specific antibodies were produced by immunizing mice with rGXGG-2016-ΔgI/ΔgE-B119L. The horizontal line represents the cutoff value (0.526). (B) Anti-EP364R antibodies were produced by immunizing mice with rGXGG-2016-ΔgI/ΔgE-EP364R. The horizontal line represents the cutoff value (0.588). (C) Anti-gB antibodies were produced in all immunized mice except the control DMEM group. The horizontal line represents the cutoff value (0.6). (D) Anti-gE antibodies were produced by immunizing mice with GXGG-2016. The horizontal line represents the cutoff value (0.6). The bars represent the mean ± SD of three or more independent experiments. Significance was analyzed using an unpaired t-test. *, p < 0.05; **, p < 0.01; ***, p < 0.001; ****, p < 0.0001; n.s., p > 0.05 (n = 3)
Detection of specific antibodies against B119L and EP364R after immunization with the recombinant strains and protection against lethal challenge with variant PRV strains. (A) Immunofluorescence images showing the detection of antibodies in the sera of mice immunized with the recombinant strains. The white scale bars are 150 μm. (B) Western blot results showing the recognition of ASFV antigens by antibodies in the sera of mice infected with the recombinant strains (41.4 kDa EP364R band, 14.4 kDa B119L band). (C) Protection of mice against lethal challenge with variant PRV strains by both the recombinant and parental strains.
To evaluate whether immunization with the recombinant strains induced protective immunity against PRV in mice, each animal was challenged with 5 × 105 TCID50 of the PRV GXLB-2015 strain 4 weeks after the booster immunization. After challenge, all of the immunized mice survived, and no clinical signs were observed in mice that had been immunized with GXGG-2016, rGXGG-2016-ΔgI/gE-EGFP, rGXGG-2016-ΔgI/ΔgE-EP364R, or rGXGG-2016-ΔgI/ΔgE-B119L. The mice in the control group displayed typical signs of PRV infection, and all of them died (Fig. 5C). The data indicate that the recombinant strains rGXGG-2016-ΔgI/ΔgE-EP364R and rGXGG-2016-ΔgI/ΔgE-B119L, constructed in this study, not only induced the production of antibodies against the ASFV EP364R and B119L proteins in mice but also protected mice against lethal challenge with a virulent PRV strain.
Discussion
The outbreaks of ASF have caused huge economic losses to the global pig-breeding industry and have seriously affected international trade related to the pig industry. Since 2021, ASF outbreaks have been reported in 41 countries around the world, and seven of these countries reported their first outbreak of the disease since then [42]. This has resulted in the loss of more than 1 million animals worldwide.
ASFV has been prevalent for nearly a hundred years since its discovery, and many researchers are working to develop an effective vaccine against the virus. The team led by Gladue and Borca deleted the I177L gene from the genome of ASFV-G and developed a candidate attenuated vaccine as well as evaluating its safety [43, 44]. It was found that it could effectively protect the host from challenge with the parental strain, although mild clinical signs were still observed. Takamatsu’s team immunized local African pigs with a naturally attenuated strain, OURT88/3, which can induce cross-protective immunity against some virulent isolates [45, 46]. However, in practice, the attenuated vaccines still need to be carefully evaluated with respect to safety issues such as virulence reversion, adverse reactions, and persistent infections. Cadenas-Fernández et al. [47] used high doses of inactivated strains to immunize pigs and found that this did not provide adequate protection against challenge with the parental strains. Tamas et al. [48] immunized pigs with viruses produced by deleting the MGF-110 gene of strain Lv17/WB/Rie1 and found that the deletion reduced pathogenicity of this strain at high doses while having essentially no effect on its protective capacity. These studies also emphasized that the currently available inactivated vaccine did not effectively protect pigs against challenge with a virulent ASFV strain.
Lokhandwala et al. [49] showed that immunizing pigs with ASFV antigens (composed of p32, p54, p72, and pp62) that had been produced using adenovirus vectors induced specific antibodies as well as cytotoxic T lymphocyte responses. Goatley et al. [50] used a recombinant adenovirus for primary immunization and a modified vaccinia virus Ankara strain as a booster for expression of ASFV immunogens, and found that the immunized pigs were protected from lethal doses of ASFV. These studies have shown that recombinant viral vectors are a promising vaccine platform, combining the advantages of live attenuated vaccines and subunit vaccines and avoiding some of the shortcomings of each. They are relatively safe and inexpensive and are being actively developed. In the present study, genes encoding immunogens from ASFV were inserted into PRV by homologous recombination, resulting in two recombinant viruses that were able to induce an effective immune response in mice.
Since 2011, there have been PR outbreaks on several large-scale pig farms in China where the animals had been immunized with the Bartha-K61 vaccine. The epidemic has spread rapidly throughout the country, causing serious economic losses to the pig industry in China. Through epidemiological investigations and genetic analysis of pigs on farms in different regions of China, it was found that the positive rate for PRV gE antibodies had generally shown a downward trend, but the rate in some areas was still as high as 40-60% [51, 52]. In addition, the virulence of PRV variants has increased significantly, with infected pigs showing more obvious clinical signs and higher mortality rates, resulting in huge economic losses to many pig farms [53, 54]. gE mediates the entry of the virus into the central nervous system of the host, often playing a synergistic role with gI [55]. Deletion of the gI and gE genes or insertion of foreign genes in their place does not affect viral replication. Therefore, the gI and gE genes were selected as the modification sites for the PRV GXGG-2016 construct in this study. Although the attenuated and recombinant strains in this study showed lower virus titers than the parental strain, the virus titer at 60 h could nevertheless approach or even exceed 107 TCID50. There was no significant difference in proliferation between the recombinant and attenuated strains. Our results are consistent with those of previous studies, which also showed that gI/gE double gene deletions resulted in a reduction in the replicative ability of the virus in vitro [33, 56].
Adenovirus, vaccinia virus, baculovirus, alphavirus, and Newcastle disease virus are commonly used viral vectors for ASFV vaccines. The advantages of the PRV expression vector system are similar to those of other viral expression vectors, including a good safety record, a large capacity for exogenous genes, low production costs, and a relatively simple process for its use. Pathological observation of sections of brain and spleen tissues showed that the recombinant strains constructed in this study did not induce a lethal inflammatory response, further confirming their safety. Interestingly, the spleen tissues of mice infected with GXLB-2015 or GXGG-2016 did not show significant histopathological changes, which may be related to the lack of an inflammatory response at the early stages of PRV infection or to the extent of tissue damage and cellular disruption caused by higher concentrations of virus, which is consistent with previous observations [33, 56]. It is worth noting that the expression products of recombinant PRV retain their antigenicity, immunogenicity, and function. Our recombinant strains induced antibodies against the EP364R and B119L proteins in immunized mice and provided protection against lethal PRV infection. These results suggest that PRV can be used safely for efficient expression of foreign genes for the development of multivalent genetically engineered live vaccines.
In summary, we constructed PRV recombinant strains expressing the ASFV EP364R and B119L genes, rGXGG-2016-ΔgI/ΔgE-EP364R, and rGXGG-2016-ΔgI/ΔgE-B119L, using homologous recombination. The recombinant strains showed good safety and immunogenicity in mice, and they were 100% protective against a virulent virus challenge. Therefore, this study provides a reference for the construction of ASFV and PRV bivalent vaccines. It also verifies the feasibility of using live virus vector vaccines for the future development of an effective novel ASV vaccine. Our study has so far only verified the safety and efficacy of the recombinant strains in mice. They now need to be tested in pigs to determine whether they can induce an effective immune response and provide protection in order to assess their suitability for practical applications.
Data availability
All data generated or analyzed during this study are included in this published article.
References
Gaudreault NN, Madden DW, Wilson WC, Trujillo JD, Richt JA (2020) African swine fever virus: an emerging DNA arbovirus. Front Vet Sci 7:215. https://doi.org/10.3389/fvets.2020.00215
Inmaculada G, Covadonga A, Linda D, Simon G (2017) African swine fever virus: a review. Viruses 9(5):103
Wang N, Zhao D, Wang J, Zhang Y, Wang M, Gao Y, Li F, Wang J, Bu Z, Rao Z, Wang X (2019) Architecture of African swine fever virus and implications for viral assembly. Science 366:6465
Bao J, Wang Q, Lin P, Liu C, Li L, Wu X, Chi T, Xu T, Ge S, Liu Y, Li J, Wang S, Qu H, Jin T, Wang Z (2019) Genome comparison of African swine fever virus China/2018/Anhui XCGQ strain and related European p72 genotype ii strains. Transbound Emerg Dis 66(3):1167–1176
Zhou X, Li N, Luo Y, Liu Y, Miao F, Chen T, Zhang S, Cao P, Li X, Tian K, Qiu H, Hu R (2018) Emergence of African swine fever in China, 2018. Transbound Emerg Dis 65(6):1482–1484
Li L, Fu J, Li J, Guo S, Chen Q, Zhang Y, Liu Z, Tan C, Chen H, Wang X (2022) African swine fever virus pI215L inhibits type I interferon signaling by targeting interferon regulatory factor 9 for autophagic degradation. J Virol 96(17):e94422. https://doi.org/10.1128/jvi.00944-22
Dodantenna N, Ranathunga L, Chathuranga W, Weerawardhana A, Cha JW, Subasinghe A, Gamage N, Haluwana DK, Kim Y, Jheong W, Poo H, Lee JS (2022) African swine fever virus EP364R and C129R target cyclic GMP-AMP to inhibit the cGAS-STING signaling pathway. J Virol 96(15):e102222. https://doi.org/10.1128/jvi.01022-22
Li YH, Peng JL, Xu ZS, Xiong MG, Wu HN, Wang SY, Li D, Zhu GQ, Ran Y, Wang YY (2023) African swine fever virus cysteine protease ps273R inhibits type I interferon signaling by mediating STAT2 degradation. J Virol 97(3):e194222. https://doi.org/10.1128/jvi.01942-22
Gao Q, Yang Y, Quan W, Zheng J, Luo Y, Wang H, Chen X, Huang Z, Chen X, Xu R, Zhang G, Gong L (2021) The African swine fever virus with MGF360 and MGF505 deleted reduces the apoptosis of porcine alveolar macrophages by inhibiting the NF-kappaB signaling pathway and interleukin-1beta. Vaccines (Basel) 9(11):1371. https://doi.org/10.3390/vaccines9111371
Yang K, Xue Y, Niu T, Li X, Cheng M, Bao M, Zou B, Shi C, Wang J, Yang W, Wang N, Jiang Y, Yang G, Zeng Y, Cao X, Wang C (2022) African swine fever virus MGF505-7R protein interacted with IRF7 and TBK1 to inhibit type I interferon production. Virus Res 322:198931. https://doi.org/10.1016/j.virusres.2022.198931
Lakshminarayan MI, Balaji S, Eugene VK, Aravind L (2006) Evolutionary genomics of nucleo-cytoplasmic large DNA viruses. Virus Res 117(1):156-184
Jancovich JK, Chapman D, Hansen DT, Robida MD, Loskutov A, Craciunescu F, Borovkov A, Kibler K, Goatley L, King K, Netherton CL, Taylor G, Jacobs B, Sykes K, Dixon LK (2018) Immunization of pigs by DNA prime and recombinant vaccinia virus boost to identify and rank African swine fever virus immunogenic and protective proteins. J Virol 92(8):10–1128. https://doi.org/10.1128/JVI.02219-17
Lynnette CG, Ana LR, Raquel P, Hannah G, Gareth LS, Zoe H, Chak-Sum H, María M, Pedro JS, Geraldine T, Linda KD, Christopher LN (2020) A pool of eight virally vectored African swine fever antigens protect pigs against fatal disease. Vaccines 8(2):234
Rodriguez I, Redrejo-Rodriguez M, Rodriguez JM, Alejo A, Salas J, Salas ML (2006) African swine fever virus pB119L protein is a flavin adenine dinucleotide-linked sulfhydryl oxidase. J Virol 80(7):3157–3166. https://doi.org/10.1128/JVI.80.7.3157-3166.2006
Lewis T, Zsak L, Burrage TG, Lu Z, Kutish GF, Neilan JG, Rock DL (2000) An African swine fever virus ERV1-ALR homologue, 9GL, affects virion maturation and viral growth in macrophages and viral virulence in swine. J Virol 74(3):1275–1285. https://doi.org/10.1128/jvi.74.3.1275-1285.2000
O'Donnell V, Holinka LG, Krug PW, Gladue DP, Carlson J, Sanford B, Alfano M, Kramer E, Lu Z, Arzt J, Reese B, ***Carrillo C, Risatti GR, Borca MV (2015) African swine fever virus Georgia 2007 with a deletion of virulence-associated gene 9GL (B119L), when administered at low doses, leads to virus attenuation in swine and induces an effective protection against homologous challenge. J Virol 89(16):8556–8566
Gladue DP, O'Donnell V, Ramirez-Medina E, Rai A, Pruitt S, Vuono EA, Silva E, Velazquez-Salinas L, Borca MV (2020) Deletion of CD2-like (CD2v) and C-type lectin-like (EP153R) genes from African swine fever virus Georgia-∆9GL abrogates its effectiveness as an experimental vaccine. Viruses 12(10):1185. https://doi.org/10.3390/v12101185
O’Donnell V, Holinka LG, Sanford B, Krug PW, Carlson J, Pacheco JM, Reese B, Risatti GR, Gladue DP, Borca MV (2016) African swine fever virus Georgia isolate harboring deletions of 9GL and MGF360/505 genes is highly attenuated in swine but does not confer protection against parental virus challenge. Virus Res 221:8–14. https://doi.org/10.1016/j.virusres.2016.05.014
Vivian O, Guillermo RR, Lauren GH, Peter WK, Jolene C, Lauro VS, Paul AA, Douglas PG, Manuel VB (2017) Simultaneous deletion of the 9GL and UK genes from the African swine fever virus Georgia 2007 isolate offers increased safety and protection against homologous challenge. J Virol 91(1):10–1128
Pomeranz LE, Reynolds AE, Hengartner CJ (2005) Molecular biology of pseudorabies virus: impact on neurovirology and veterinary medicine. Microbiol Mol Biol Rev 69(3):462–500. https://doi.org/10.1128/MMBR.69.3.462-500.2005
Cheung AK (1989) Detection of pseudorabies virus transcripts in trigeminal ganglia of latently infected swine. J Virol 63(7):2908–2913. https://doi.org/10.1128/JVI.63.7.2908-2913.1989
Zheng HH, Fu PF, Chen HY, Wang ZY (2022) Pseudorabies virus: from pathogenesis to prevention strategies. Viruses 14(8):1638. https://doi.org/10.3390/v14081638
Liu Q, Wang X, Xie C, Ding S, Yang H, Guo S, Li J, Qin L, Ban F, Wang D, Wang C, Feng L, Ma H, Wu B, Zhang L, Dong C, Xing L, Zhang J, Chen H, Yan R, Wang X, Li W (2021) A novel human acute encephalitis caused by pseudorabies virus variant strain. Clin Infect Dis 73(11):e3690–e3700. https://doi.org/10.1093/cid/ciaa987
Klupp BG, Hengartner CJ, Mettenleiter TC, Enquist LW (2004) Complete, annotated sequence of the pseudorabies virus genome. J Virol 78(1):424–440
Jiang C, Ma Z, Bai J, Sun Y, Cao M, Wang X, Jiang P, Liu X (2023) Comparison of the protective efficacy between the candidate vaccine Zj01R carrying gE/gI/TK deletion and three commercial vaccines against an emerging pseudorabies virus variant. Vet Microbiol 276:109623. https://doi.org/10.1016/j.vetmic.2022.109623
Zhu L, Yi Y, Xu Z, Cheng L, Tang S, Guo W (2011) Growth, physicochemical properties, and morphogenesis of Chinese wild-type PRV Fa and its gene-deleted mutant strain PRV SA215. Virol J 8:272. https://doi.org/10.1186/1743-422X-8-272
Tang YD, Liu JT, Wang TY, An TQ, Sun MX, Wang SJ, Fang QQ, Hou LL, Tian ZJ, Cai XH (2016) Live attenuated pseudorabies virus developed using the CRISPR/Cas9 system. Virus Res 225:33–39. https://doi.org/10.1016/j.virusres.2016.09.004
Wu T, Hao Z, Guo-Xin L, Fei G, Tong-Ling S, Yan-Jun Z, Hai Y, Yi-Feng J, Ling-Xue Y, Li-Wei L, Ning K, Guang-Zhi T, Ji-Chang L (2020) Recombinant pseudorabies virus expressing E2 of classical swine fever virus (CSFV) protects against both virulent pseudorabies virus and CSFV. Antivir Res 173(C):104652
Zheng HH, Wang LQ, Fu PF, Zheng LL, Chen HY, Liu F (2020) Characterization of a recombinant pseudorabies virus expressing porcine parvovirus VP2 protein and porcine IL-6. Virol J 17(1):19. https://doi.org/10.1186/s12985-020-1292-8
Feng Z, Chen J, Liang W, Chen W, Li Z, Chen Q, Cai S (2020) The recombinant pseudorabies virus expressing African swine fever virus CD2v protein is safe and effective in mice. Virol J 17(1):180. https://doi.org/10.1186/s12985-020-01450-7
Tan L, Yao J, Lei L, Xu K, Liao F, Yang S, Yang L, Shu X, Duan D, Wang A (2022) Emergence of a novel recombinant pseudorabies virus derived from the field virus and its attenuated vaccine in China. Front Vet Sci 9:872002. https://doi.org/10.3389/fvets.2022.872002
Yao L, Hu Q, Zhang C, Ghonaim AH, Cheng Y, Ma H, Yu X, Wang J, Fan X, He Q (2021) Untargeted LC–MS based metabolomic profiling of iPAMs to investigate lipid metabolic pathways alternations induced by different pseudorabies virus strains. Vet Microbiol 256:109041. https://doi.org/10.1016/j.vetmic.2021.109041
Tong W, Liu F, Zheng H, Liang C, Zhou YJ, Jiang YF, Shan TL, Gao F, Li GX, Tong GZ (2015) Emergence of a pseudorabies virus variant with increased virulence to piglets. Vet Microbiol 181(3–4):236–240. https://doi.org/10.1016/j.vetmic.2015.09.021
An TQ, Peng JM, Tian ZJ, Zhao HY, Li N, Liu YM, Chen JZ, Leng CL, Sun Y, Chang D, Tong GZ (2013) Pseudorabies virus variant in Bartha-K61-vaccinated pigs, China. Emerg Infect Dis 19(11):1749–1755. https://doi.org/10.3201/eid1911.130177
Luo Y, Li N, Cong X, Wang CH, Du M, Li L, Zhao B, Yuan J, Liu DD, Li S, Li Y, Sun Y, Qiu HJ (2014) Pathogenicity and genomic characterization of a pseudorabies virus variant isolated from Bartha-K61-vaccinated swine population in China. Vet Microbiol 174(1–2):107–115. https://doi.org/10.1016/j.vetmic.2014.09.003
Yu ZQ, Tong W, Zheng H, Li LW, Li GX, Gao F, Wang T, Liang C, Ye C, Wu JQ, Huang Q, Tong GZ (2017) Variations in glycoprotein B contribute to immunogenic difference between PRV variant JS-2012 and Bartha-K61. Vet Microbiol 208:97–105. https://doi.org/10.1016/j.vetmic.2017.07.019
Ye C, Zhang QZ, Tian ZJ, Zheng H, Zhao K, Liu F, Guo JC, Tong W, Jiang CG, Wang SJ, Shi M, Chang XB, Jiang YF, Peng JM, Zhou YJ, Tang YD, Sun MX, Cai XH, An TQ, Tong GZ (2015) Genomic characterization of emergent pseudorabies virus in China reveals marked sequence divergence: evidence for the existence of two major genotypes. Virology 483:32–43. https://doi.org/10.1016/j.virol.2015.04.013
Nie Z, Zhu S, Wu L, Sun R, Shu J, He Y, Feng H (2023) Progress on innate immune evasion and live attenuated vaccine of pseudorabies virus. Front Microbiol 14:1138016. https://doi.org/10.3389/fmicb.2023.1138016
Ren Q, Ren H, Gu J, Wang J, Jiang L, Gao S (2022) The epidemiological analysis of pseudorabies virus and pathogenicity of the variant strain in Shandong province. Front Vet Sci 9:806824. https://doi.org/10.3389/fvets.2022.806824
Qin Y, Qin S, Huang X, Xu L, Ouyang K, Chen Y, Wei Z, Huang W (2023) Isolation and identification of two novel pseudorabies viruses with natural recombination or TK gene deletion in China. Vet Microbiol 280:109703. https://doi.org/10.1016/j.vetmic.2023.109703
Reed LJ, Muench H (1938) A simple method of estimating fifty per cent endpoints. Am J Epidemiol 27:493–497
Woah (2023) African swine fever (ASF)—situation report 29
Tran XH, Phuong L, Huy NQ, Thuy DT, Nguyen VD, Quang PH, Ngon QV, Rai A, Gay CG, Gladue DP, Borca MV (2022) Evaluation of the safety profile of the ASFV vaccine candidate ASFV-G-∆I177L. Viruses 14(5):896. https://doi.org/10.3390/v14050896
Borca MV, Ramirez-Medina E, Silva E, Vuono E, Rai A, Pruitt S, Holinka LG, Velazquez-Salinas L, Zhu J, Gladue DP (2020) Development of a highly effective African swine fever virus vaccine by deletion of the I177L gene results in sterile immunity against the current epidemic Eurasia strain. J Virol 94(7):10–1128. https://doi.org/10.1128/JVI.02017-19
King K, Chapman D, Argilaguet JM, Fishbourne E, Hutet E, Cariolet R, Hutchings G, Oura CA, Netherton CL, Moffat K, Taylor G, Le Potier MF, Dixon LK, Takamatsu HH (2011) Protection of European domestic pigs from virulent African isolates of African swine fever virus by experimental immunisation. Vaccine 29(28):4593–4600. https://doi.org/10.1016/j.vaccine.2011.04.052
Mulumba-Mfumu LK, Goatley LC, Saegerman C, Takamatsu HH, Dixon LK (2016) Immunization of African indigenous pigs with attenuated genotype I African swine fever virus OURT 88/3 induces protection against challenge with virulent strains of genotype I. Transbound Emerg Dis 63(5):e323–e327. https://doi.org/10.1111/tbed.12303
Cadenas-Fernandez E, Sanchez-Vizcaino JM, van den Born E, Kosowska A, van Kilsdonk E, Fernandez-Pacheco P, Gallardo C, Arias M, Barasona JA (2021) High doses of inactivated African swine fever virus are safe, but do not confer protection against a virulent challenge. Vaccines (Basel) 9(3):242. https://doi.org/10.3390/vaccines9030242
Tamas V, Righi C, Meszaros I, D'Errico F, Olasz F, Casciari C, Zadori Z, Magyar T, Petrini S, Feliziani F (2023) Involvement of the MGF 110-11L gene in the African swine fever replication and virulence. Vaccines (Basel) 11(4):846. https://doi.org/10.3390/vaccines11040846
Lokhandwala S, Waghela SD, Bray J, Martin CL, Sangewar N, Charendoff C, Shetti R, Ashley C, Chen CH, Berghman LR, Mwangi D, Dominowski PJ, Foss DL, Rai S, Vora S, Gabbert L, Burrage TG, Brake D, Neilan J, Mwangi W (2016) Induction of robust immune responses in swine by using a cocktail of adenovirus-vectored African swine fever virus antigens. Clin Vaccine Immunol 23(11):888–900. https://doi.org/10.1128/CVI.00395-16
Goatley LC, Reis AL, Portugal R, Goldswain H, Shimmon GL, Hargreaves Z, Ho CS, Montoya M, Sanchez-Cordon PJ, Taylor G, Dixon LK, Netherton CL (2020) A pool of eight virally vectored African swine fever antigens protect pigs against fatal disease. Vaccines (Basel) 8(2):234. https://doi.org/10.3390/vaccines8020234
Chen X, Li H, Zhu Q, Chen H, Wang Z, Zheng L, Liu F, Wei Z (2022) Serological investigation and genetic characteristics of pseudorabies virus between 2019 and 2021 in Henan province of China. Viruses 14(8):1685. https://doi.org/10.3390/v14081685
Yao J, Li J, Gao L, He Y, Xie J, Zhu P, Zhang Y, Zhang X, Duan L, Yang S, Song C, Shu X (2022) Epidemiological investigation and genetic analysis of pseudorabies*** virus in Yunnan province of China from 2017 to 2021. Viruses 14(5):895. https://doi.org/10.3390/v14050895
Liu Y, Zhang S, Xu Q, Wu J, Zhai X, Li S, Wang J, Ni J, Yuan L, Song X, Zhao B, Zhou Z, Wang C, Yang L (2018) Investigation on pseudorabies prevalence in Chinese swine breeding farms in 2013–2016. Trop Anim Health Prod 50(6):1279–1285. https://doi.org/10.1007/s11250-018-1555-1
Sun L, Tang Y, Yan K, Zhang H (2022) Construction of a quadruple gene-deleted vaccine confers complete protective immunity against emerging PRV variant challenge in piglets. Virol J 19(1):19. https://doi.org/10.1186/s12985-022-01748-8
Babic N, Klupp B, Brack A, Mettenleiter TC, Ugolini G, Flamand A (1996) Deletion of glycoprotein gE reduces the propagation of pseudorabies virus in the nervous system of mice after intranasal inoculation. Virology 219(1):279–284. https://doi.org/10.1006/viro.1996.0247
Yin Y, Xu Z, Liu X, Li P, Yang F, Zhao J, Fan Y, Sun X, Zhu L (2017) A live gI/gE-deleted pseudorabies virus (prv) protects weaned piglets against lethal variant PRV challenge. Virus Genes 53(4):565–572. https://doi.org/10.1007/s11262-017-1454-y
Acknowledgements
The authors thank Dr. Dev Sooranna, Imperial College London, for editing the English text of a draft of this manuscript.
Funding
This study was supported by the National Natural Science Foundation of China (32260875).
Author information
Authors and Affiliations
Contributions
Xin-Mei Geng performed the experiments and wrote the manuscript. Ying-Mu Xi and Xiang-Mei Huang supervised data collection and analysis. Yang-Lin Wang and Xu-Ying Wang were responsible for sample collection. Kang Ouyang, Ying Chen, and Zuzhang Wei checked and finalized the manuscript. Yi-feng Qin and Wei-jian Huang initiated the study, designed the experiments, and supplied the manuscript. All authors contributed to the article and approved the submitted version.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Additional information
Handling Editor: William G. Dundon.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Geng, XM., Xi, YM., Huang, XM. et al. Construction of and evaluation of the immune response to two recombinant pseudorabies viruses expressing the B119L and EP364R proteins of African swine fever virus. Arch Virol 169, 22 (2024). https://doi.org/10.1007/s00705-023-05935-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00705-023-05935-y