Abstract
Porcine epidemic diarrhea (PED) virus (PEDV) is a highly contagious virus. PED was first identified in 2008 and has greatly affected the Vietnamese pig production economy. The aim of this study was to investigate the epidemiological and genetic characteristics of PEDV in piglet herds in the Mekong Delta, Vietnam. Diarrheal stool and intestinal samples from 2262 piglets from 191 herds in five provinces were collected to test for the presence of PEDV. Ten PEDV strains were randomly selected for sequencing, and four genes encoding PEDV structural proteins were analyzed. The rates of herds and samples positive for PEDV were 27.23% and 27.72%, respectively. In positive herds, the morbidity and mortality of PEDV-positive piglets were 97.97% and 79.06%, respectively, with most of the infected piglets under 7 days of age. Phylogenetic analysis showed that the 10 PEDV strains from this study clustered with genotype G2 strains from Vietnam and neighboring countries. Many amino acid substitutions were identified in important antigenic regions in the spike protein of the 10 strains when compared to four PEDV vaccine strains. This study provides novel insights into the epidemiology and genetic diversity of circulating PEDV strains, which could facilitate the development of an appropriate and proactive strategy for controlling PED.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Porcine epidemic diarrhea (PED) virus (PEDV) is a highly contagious virus belonging to the genus Alphacoronavirus, family Coronaviridae [19]. PEDV is an enveloped virus with a diameter of approximately 95-190 nm [9]. The PEDV genome is a single-stranded positive RNA containing at least seven open reading frames (ORFs), which encode four structural proteins (S, E, M, and N) and three nonstructural proteins (replicase enzymes 1a, 1b, and ORF3) [5].
PED can spread very quickly in PEDV-negative herds and cause high morbidity (100%) and mortality (30% to 90%) in piglets. PED first occurred in Vietnam at the end of 2008 and caused heavy losses in pig herds in the Southeast provinces, with various mortality rates in piglets (50%-100%) [14]. PEDV did not cause death in mature pigs but did greatly affect reproductive performance, with a 12.6% reduction in farrowing rate, 5.7% failure to breed, 1.3% abortion rate, and 2.0% mummified fetuses [27].
The genetic and molecular epidemiological characteristics of PEDV in countries differ [17, 18]. In Vietnam, most PEDV isolates have belonged to the genotypes G1 and G2 and have very high genetic similarity to strains isolated from neighboring countries such as China and Thailand [3, 24].
The Mekong Delta is located in western Vietnam and is an important pig-production area. Although epidemiological and genetic studies of PEDV have been conducted in southeastern and northern Vietnam, a comprehensive survey of these features in the Mekong Delta has not been undertaken. In this study, we investigated the mortality, morbidity, and genetic characteristics of PEDV in the Mekong Delta, thereby providing new insights into PED in Vietnam.
Materials and methods
Study design and sample collection
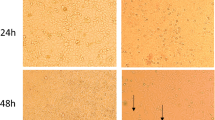
This study was conducted on 63 commercial pig farms in five provinces in the Mekong Delta of Vietnam from 2015 to 2017. On these farms, a large number of piglets showed anorexia, depression, and yellow or watery diarrhea. In addition, some gastrointestinal lesions were observed in dead piglets, which had thin intestinal walls and distended stomachs filled with undigested milk (Fig. 1). We randomly selected about three herds per farm to collect diarrheal stools and intestinal samples. A total of 2262 samples from 191 herds were obtained and tested for the presence of PEDV using RT-PCR. The morbidity and mortality were also examined based on age and locality in PEDV-infected herds. Ten PEDV strains were selected from five provinces for sequencing of the four structural genes to investigate the genetic diversity of PEDV in the Mekong Delta region.
RNA extraction and RT-PCR
Total RNA from diarrheal stools and intestinal samples was extracted using a GenJET Gel Extraction and DNA Cleanup Micro Kit (Thermo Fisher, USA) according to the manufacturer's protocol. The extracted RNA was used to synthesize cDNA using a RevertAid First Strand cDNA Synthesis Kit (Thermo Fisher, USA). A 412-bp fragment of the M gene was amplified using a pair of specific primers [12]. The thermocycling conditions included a 5-min initial denaturation step at 95°C, followed by 35 cycles of 30 s at 95°C, 30 s at 55°C, and 30 s at 72°C, and then a 10-min final elongation step at 72°C.
Genetic sequencing and analysis
Ten PEDV-positive samples were selected randomly for sequencing of the four genes encoding structural proteins (S, E, M, and N). The primer pairs used to amplify the complete S, E, M, and N genes are shown in Supplementary Table S1. The structural genes were sequenced on both strands at Macrogen (Korea), using the primers T7 and SP6. Sequence data were processed and assembled using Sequencher 5.4.6 software (Genecodes). The nucleotide sequences determined in this study were deposited in the GenBank database under the accession numbers shown in Supplementary Table S2.
Nucleotide and amino acid sequences were analyzed and compared with reference sequences (Supplementary Table S3) using Bioedit 7.2.6 software [8]. Phylogenetic trees based on the S, E, M, and N genes were built using the maximum-likelihood method in MEGA 6.06 software [23] with 1000 replicates.
Results and discussion
The prevalence of PEDV in piglets in the Mekong Delta provinces
Out of 191 herds of piglets with diarrhea, 52 (27.23%) tested positive for PEDV (Table 1), which was a higher rate than that reported previously for the southern region of Vietnam (16.96%) [15]. Tien Giang and Ben Tre had the highest rates of PEDV-positive herds, with 33.33% and 32.65%, respectively, followed by Vinh Long (24.13%). The percentage of PEDV-infected herds in Can Tho and Dong Thap was the lowest (18.75% and 20%, respectively). The proportion of PEDV-positive samples in the total samples collected was 27.72%. Tien Giang and Ben Tre had the highest proportion of infected samples, with 33.79% and 32.20%, followed by Vinh Long, Dong Thap, and Can Tho (23.42%, 20.19%, and 19.67%, respectively).
Of the 640 piglets in the positive herds, 627 (97.97%) tested positive for PEDV, and 506 (79.06%) died from PEDV infection (Table 2). This is consistent with the rates of PEDV-infected pigs found previously in the southern region (93.94% and 81.67%, respectively) and some northern provinces (96% and 68.6%, respectively) [14, 16]. A survey in the USA also showed high morbidity and mortality rates in infected piglets, with rates of 100% and 90-95%, respectively [20]. These results show that most of the piglets in the positive herd were infected with PEDV and died. Can Tho had the highest infection rate (100%) but the lowest mortality rate (73.24%). Meanwhile, the remaining provinces had similar infection and mortality rates, ranging from 97.37% to 98.48% and from 78.46% to 83.91%, respectively. The morbidity and mortality of piglets up to 28 days old in PEDV-positive herds were recorded for three age groups: under 7 days old, 7-10 days old, and over 10 days old. Piglets under 7 days of age had the highest PEDV morbidity and mortality (100% and 84.47%, respectively), and the lowest morbidity and mortality were in piglets older than 10 days (88.10% and 64.86%, respectively) (p < 0.05). In Europe, it has been reported that the mortality rate can reach 100% in suckling piglets, while it is less than 10% in piglets older than 10 days of age, and less than 5% in adult and fattening pigs [1]. This seems to be due to differences in resistance and immune status among the animals. In previous studies, piglets were found to be very sensitive to PEDV, especially those under a week of age [18, 20, 22]. In PED outbreaks in 2010, the morbidity and mortality of piglets under 7 days of age were 100% and 80-100%, respectively [22]. In PED outbreaks in Taiwan, these rates were also very high (80-100% and 90-100%, respectively) [13].
Genetic characteristics of PEDV strains
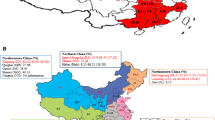
Phylogenetic analysis based on four structural gene sequences showed that the 10 PEDV strains isolated from PEDV-positive piglets in this study belonged to the G2 genotype (Fig. 2). This contrasts with previous studies conducted in southern Vietnam, where all six PEDV strains collected in three provinces in southern Vietnam in 2009-2010 belonged to the G1 genotype [4]. More recently, two strains (HUA-PED176 and HUA-PED254) isolated in the south of Vietnam in 2015-2016 also belonged to the G1 genotype [24]. However, 23 out of 25 PEDV strains collected in northern Vietnam during 2012–2015 belonged to the G2 genotype [3]. Twenty-eight (100%) PEDV strains isolated in the northern and central regions of Vietnam between 2015 and 2016 belonged to the G2 genotype [24]. Furthermore, the majority of PEDV strains collected in northern Vietnam in 2018–2019 also belonged to the G2 genotype [7]. Together, these data show that PEDV strains belonging to genotype G2 have become more prevalent in Vietnam. The PEDV strains from this study had a very close genetic relationship to three strains from Vietnam that were isolated in the northern region in 2013-2014 (VN/VAP1113-1, VN/JFP1013-1, and HUA-14PED96), the CBR1 strain from Thailand, and three Chinese strains (CHGD-01, CH/FJZZ-9/2012, and PEDV-CHZ).
Phylogenetic trees based on the nucleotide sequences of the genes encoding the S (A), E (B), M (C), and N (D) proteins of 10 PEDV strains. The neighbor-joining method was used for the construction of the phylogenetic trees in MEGA 6.06 software. Numbers at branches are bootstrap values >50% (1000 replicates). Red triangles and black circles indicate PEDV strains from this study and vaccine strains, respectively. Scale bars represent nucleotide substitutions per site
Multiple sequence alignments of the S, E, M, and N genes showed that the 10 PEDV strains from this study shared a very high degree of similarity at the nucleotide and amino acid levels (97.2-100% and 93.0-100% identity, respectively) (Supplementary Table S4). They also had very high sequence similarity to three reference strains isolated in northern Vietnam (96.0-99.8% and 89.2-100% identity, respectively). Although the 10 PEDV strains exhibited high similarity at the nucleotide and amino acid levels in the E, M, and N genes to reference strains isolated from Korea, Japan, Thailand, China, and the USA (94.6-99.8% and 97.2-100% identity, respectively), the S gene had low sequence identity (81.5-98.3% and 73.5-95.5%, respectively). A comparison of the S genes of the 10 strains from this study and four PEDV vaccine strains (CV777, SM98, DR13/virulent, and DR13/attenuated) revealed that the sequence identity was only 92.3-94.3% at the nucleotide level and 83.5-88.1% at the amino acid level. In particular, these strains lacked 15 nucleotides (167, 176-186, and 430-432) and had an extra six nucleotides (492-497) when compared to the CV777, DR13/Attenuated, and DR13/Virulent strains, and they lacked 32 nucleotides (167, 176-186, 430-432, 4167-4183) and had an extra 18 nucleotides (344-355, and 492-497) when compared to the SM98 strain. Meanwhile, these 10 strains and the four vaccine strains showed low similarity at the nucleotide and amino acid levels in the E, M, and N genes (Supplementary Table S4).
Because of the large differences at the amino acid level, the predicted amino acid sequences of four major epitopes in the spike proteins of the 10 PEDV strains and four PEDV vaccine strains were compared (Fig. 3). There were many amino acid substitutions in the S gene of the 10 PEDV strains compared to the four PEDV vaccine strains. The S protein consists of 1383 amino acids and is divided into two subunits: S1 (1-789) and S2 (790-1383) [6]. In the S1 subunit, there were amino acid substitutions (S508A, L531I, A533S, Q550R, K568T, V581A, V592L, G599S, Y608H, A/E610D, G614S, and L/F617Y) in the core neutralizing epitope (aa 504-643), which contains the antigen-determining group that induces neutralizing antibodies and is also a variable region [10]. The specific mutation Y/D771S has been identified in a B cell epitope (aa 769-776) of the S1 region [21]. No amino acid substitutions were found in the epitope 1373GPRLQPY1379, which induces anti-PEDV neutralizing antibodies [2]. Although the S protein plays an important role in receptor binding and viral entry, the amino acid mutation rate of the PEDV S gene is very high, up to 10% [11, 25], and multiple epitopes that induce neutralizing antibodies have been identified in this protein. Thus, mutations in the antigenic determinant regions of PEDV can lead to a decrease in the ability of vaccines to protect against field strains [26]. It is therefore important to select vaccines with high similarity to the strains of PEDV that are circulating on farms for PED prevention and mitigation.
Amino acid sequences of antigenic regions in the S protein of PEDV. Amino acid sequence alignments of GP5 from the 10 isolates from this study and four reference vaccine isolates are shown. Dots indicate identical amino acids, and letters indicate amino acid changes. The red, purple, black, and green boxes indicate the amino acid positions of the core neutralizing epitope, two B-cell epitopes, and anti-PEDV neutralizing antibodies, respectively
Conclusions
In this study, 27.23% of the 191 herds and 27.72% of the 2262 samples from five provinces in the Mekong Delta, Vietnam, that we tested were positive for PEDV, 97.97% of the piglets in the positive herds were PEDV-positive, and 79.06% died from PEDV infection. Ten PEDV strains from this study belonged to the G2 genotype and had a close relationship to PEDV strains isolated in Vietnam and neighboring countries. Fourteen amino acid substitutions were found in key antigenic regions of the spike protein.
Availability of data and material
The complete genome sequences of the PEDV strains identified in this study have been deposited in the GenBank database under the accession numbers MG373530-34 and OP723421-25 for the S gene, MG373535-39 and OP723406-10 for the E gene, MG373540-44 and OP723411-15 for the M gene, and MG373545-49 and OP723416-20 for the N gene.
References
Antas M, Woźniakowski G (2019) Current status of porcine epidemic diarrhoea (PED) in European pigs. J Vet Res 63:465–470
Cruz DJM, Kim C-J, Shin H-J (2008) The GPRLQPY motif located at the carboxy-terminal of the spike protein induces antibodies that neutralize Porcine epidemic diarrhea virus. Virus Res 132:192–196
Diep N, Sueyoshi M, Izzati U, Fuke N, Teh A, Lan N, Yamaguchi R (2018) Appearance of US-like porcine epidemic diarrhoea virus (PEDV) strains before US outbreaks and genetic heterogeneity of PEDV s collected in Northern Vietnam during 2012–2015. Transbound Emerg Dis 65:e83–e93
Do DT, Nguyen TT, Suphasawatt P, Roongroje T (2011) Genetic characterization of porcine epidemic diarrhea virus (PEDV) isolates from southern Vietnam during 2009–2010 outbreaks. Thai J Vet Med 41:55
Duarte M, Gelfi J, Lambert P, Rasschaert D, Laude H (1994) Genome organization of porcine epidemic diarrhoea virus. Coronaviruses. Springer, Berlin, pp 55–60
Duarte M, Laude H (1994) Sequence of the spike protein of the porcine epidemic diarrhoea virus. J Gen Virol 75:1195–1200
Duong BTT, Thao PTP, Hoa NT, Thu HT, Phuoc MH, Le TH, Van Quyen D (2022) Molecular analysis reveals a distinct subgenogroup of porcine epidemic diarrhea virus in northern Vietnam in 2018–2019. Adv Virol 167:2337–2346
Hall TA (1999) BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. In: Nucleic acids symposium series, pp 95–98
Jung K, Saif LJ, Wang Q (2020) Porcine epidemic diarrhea virus (PEDV): an update on etiology, transmission, pathogenesis, and prevention and control. Virus Res 286:198045
Kang T-J, Han S-C, Yang M-S, Jang Y-S (2006) Expression of synthetic neutralizing epitope of porcine epidemic diarrhea virus fused with synthetic B subunit of Escherichia coli heat-labile enterotoxin in tobacco plants. Protein Expr Purif 46:16–22
Kim S-H, Lee J-M, Jung J, Kim I-J, Hyun B-H, Kim H-I, Park C-K, Oem J-K, Kim Y-H, Lee M-H (2015) Genetic characterization of porcine epidemic diarrhea virus in Korea from 1998 to 2013. Adv Virol 160:1055–1064
Li R-F, Tian X-Q, Liu Y, Xu J, Liu D-Y (2016) Isolation and genetic analysis of a variant porcine epidemic diarrhea virus in China. Pol J Vet Sci 19:65–73
Lin C-N, Chung W-B, Chang S-W, Wen C-C, Liu H, Chien C-H, Chiou M-T (2014) US-like strain of porcine epidemic diarrhea virus outbreaks in Taiwan, 2013–2014. J Vet Med Sci 76:14–00498
Nguyen TT, Do DT (2012) Some related factors and pathological characteristics of Porcine Epidemic Diarrhoea in piglets in some southern provinces. J Vet Sci Tech XX:5–11
Nguyen TT, Nguyen ĐQ, Trinh TTH, Do TD, Tran TD, Nguyen TPN, Nguyen TTN (2012) Detection and identification of porcine epidemic diarrhea virus (PEDV) isolates from Southern-Eastern provinces of Vietnam. J Vet Sci Tech XIX:26–30
Nguyen VD, Nguyen TL, Nguyen TH, Yamaguchi H (2014) Some epidemic and pathological characteristics of the porcine epidemic diarrhea in some northern provinces, Vietnam. Vet Sci Tech 11:43–55
Park S-J, Moon H-J, Yang J-S, Lee C-S, Song D-S, Kang B-K, Park B-K (2007) Sequence analysis of the partial spike glycoprotein gene of porcine epidemic diarrhea viruses isolated in Korea. Virus Genes 35:321–332
Puranaveja S, Poolperm P, Lertwatcharasarakul P, Kesdaengsakonwut S, Boonsoongnern A, Urairong K, Kitikoon P, Choojai P, Kedkovid R, Teankum K (2009) Chinese-like strain of porcine epidemic diarrhea virus, Thailand. Emerg Infect Dis 15:1112
Schmidt A, Wolff MH, Weber O (2005) Coronaviruses with special emphasis on first insights concerning SARS. Springer, New York
Stevenson GW, Hoang H, Schwartz KJ, Burrough ER, Sun D, Madson D, Cooper VL, Pillatzki A, Gauger P, Schmitt BJ (2013) Emergence of porcine epidemic diarrhea virus in the United States: clinical signs, lesions, and viral genomic sequences. J Vet Diagn Investig 25:649–654
Sun D, Feng L, Shi H, Chen J, Cui X, Chen H, Liu S, Tong Y, Wang Y, Tong G (2008) Identification of two novel B cell epitopes on porcine epidemic diarrhea virus spike protein. Vet Microbiol 131:73–81
Sun R-Q, Cai R-J, Chen Y-Q, Liang P-S, Chen D-K, Song C-X (2012) Outbreak of porcine epidemic diarrhea in suckling piglets, China. Emerg Infect Dis 18:161
Tamura K, Stecher G, Peterson D, Filipski A, Kumar S (2013) MEGA6: molecular evolutionary genetics analysis version 6.0. Mol Biol Evol 30:2725–2729
Than VT, Choe SE, Vu TT, Do TD, Nguyen TL, Bui TT, Mai TN, Cha RM, Song D, An DJ (2020) Genetic characterization of the spike gene of porcine epidemic diarrhea viruses (PEDVs) circulating in Vietnam from 2015 to 2016. Vet Med Sci 6:535–542
Tortorici M, Veesler D (2019) Chapter four-structural insights into coronavirus entry. Adv Virus Res 105:93–116
Tran TX, Lien NT, Thu HT, Duy ND, Duong BT, Van Quyen D (2021) Changes in the spike and nucleocapsid protein of porcine epidemic diarrhea virus strain in Vietnam—a molecular potential for the vaccine development? PeerJ 9:e12329
Weng L, Weersink A, Poljak Z, de Lange K, von Massow M (2016) An economic evaluation of intervention strategies for porcine epidemic diarrhea (PED). Prev Vet Med 134:58–68
Acknowledgements
We are grateful for technical support that we received from the HanViet Veterinary Diagnosis Laboratory at Nong Lam University.
Author information
Authors and Affiliations
Contributions
NHN and MNN designed the study. TMH and HDN performed experiments. NHN, TMH, HDN, DCL, and MNN analysed the data. NHN, DCL, and MNN wrote the manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
All authors have read the journal's policy on disclosure of potential conflicts of interest, and we declare no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Ethical approval
The study was conducted in compliance with the institutional rules for the care and use of laboratory animals and using a protocol approved by the Ministry of Agriculture and Rural Development (MARD), Vietnam (TCVN 8402:2010).
Additional information
Handling Editor: Pablo Pineyro.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Nguyen, N., Huynh, T.M., Nguyen, H.D. et al. Epidemiological and genetic characterization of porcine epidemic diarrhea virus in the Mekong Delta, Vietnam, from 2015 to 2017. Arch Virol 168, 152 (2023). https://doi.org/10.1007/s00705-023-05779-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00705-023-05779-6