Abstract
A new virus, named Mutum virus, related to members of the family Tymoviridae, was isolated from mosquitoes (Mansonia spp.) in clone C6/36 cells, and its complete genome was sequenced. Its genome is 6494 nt in size with an organization resembling that of tymovirids. The isolated virus is phylogenetically related to two viruses isolated from Culex spp. mosquitoes: Ek Balam virus, reported in Mexico, and Culex-originated Tymoviridae-like virus, isolated in China. The results of this study suggest that this virus is a new member of the family Tymoviridae.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
The family Tymoviridae is composed of three genera of viruses with positive-sense RNA genomes: Maculavirus, Marafivirus, and Tymovirus [1]. Initially, the family Tymoviridae comprised only plant-infecting viruses; however, viruses related to members of this family have been isolated from mosquitoes in recent years, such as Culex-originated Tymoviridae-like virus (CuTLV) and the Ek Balam virus (EkBV) [1,2,3].
Here, we report genome sequencing of another distinct tymo-like virus, Mutum virus (MUTV), named after the mosquito collection area. MUTV was detected in female Mansonia sp. mosquitoes collected in 2018 in the vicinity of the Jirau hydroelectric dams in Nova Mutum Paraná, a rural village in the municipality of Porto Velho in the state of Rondônia, Brazil (Supplementary Fig. S1). MUTV was isolated from four pools of female Mansonia mosquitoes in Aedes albopictus cells (C6/36) [4]. The viruses were lab coded as BE-AR-855909, BE-AR-855928, BE-AR-855911, and BE-AR-855922.
Four MUTV genomes were sequenced on an Ion Torrent PGM platform (Thermo Fisher Scientific, MA, USA) according to the manufacturer's recommendations. A phylogenetic tree was constructed using the maximum-likelihood (ML) method with 1000 bootstrap replicates [5] in the RaxML v.8.0 program [6].
The sequences of four MUTV isolates have been deposited in the GenBank database with the following accession numbers: MUTV-BE-AR-855909 (MT656586), MUTV-BE-AR-855928 (MT656589), MUTV-BE-AR-855911 (MT656587), and MUTV-BE-AR-855922 (MT656588). The four MUTV isolates showed nucleotide sequence identity ranging from 99.9–100% and amino acid sequence identity of 100%, indicating that they are closely related isolates of the same virus. The MUTV genome is 6,494 nt in size, with 5′ and 3′ non-coding regions of 37 nt and 53 nt, respectively. The MUTV genome has three ORFs. ORF1 is 5,331 nt (nucleotide position 38 to 5,368) in size and encodes a 1,776-aa-long protein with an estimated mass of 201,061 kDa that is involved in virus replication. ORF2 spans 732 nt (nucleotide position 5,400 to 6,131) and codes for a 243-aa-long protein with an estimated mass of 26,134 kDa identified as a viral coat protein, while the 3’-proximal ORF 3 is 285 nt long (nucleotide position 6,182 to 6,466) and codes for a 10,729-kDa protein of unknown function. The total readings and the coverage of the genome reads ranged from 55,312 to 139,811 and from 16.10x to 48.47x, respectively.
Searches of the Interproscan databases revealed conserved protein domains for Vmethyltransf, peptidase, helicase 1, and RdRP in the product of ORF1 and a Tymo_coat motif in the ORF2-encoded protein (Fig. 1). As already indicated, no similarities were found between the protein encoded by ORF3 and currently available proteins in the GenBank/NCBI database.
Genome organization of Mutum virus (MUTV) compared to those of members of the three recognized genera in the family Tymoviridae and few related but still unclassified viruses. The ORFs are indicated by yellow arrows. The following putative conserved domains are shown in boxes with different colors and were identified using the InterProScan database: MTR, methyltranferase; PRO, papain-like protease; HEL, helicase; RdRp, RNA-dependent RNA polymerase protein; CP, coat protein; OP, movement protein; hyp, hypothetical protein; p43, p31 and p16, proline-rich proteins. MRFV, maize rayado fino vírus; TYMV, turnip yellow mosaic vírus; GFkV, grapevine fleck vírus; EkBV, Ek Balam virus
MUTV is phylogenetically related to two viruses isolated from mosquitoes, EkBV and CuTLV, along with a bat tymo-like virus, forming a clade with bootstrap support of 100%. This clade is most closely related to a group of viruses isolated from bees or from varroa mites, together forming a distinct lineage composed only of insect viruses (bootstrap support, 85%), distinct from the plant-infecting members of the family Tymoviridae (Fig. 2). The results of pairwise comparisons show amino acid sequence identity of MUTV with EkBV, CuTLV, and bat tymo-like ranging from 39.0–70.6% and nucleotide sequence identity of 67.0%, 56.3%, and 50.9%, respectively. These values are below the species demarcation threshold for creation of new species in the family Tymoviridae.
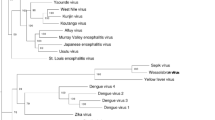
Phylogenetic tree of the family Tymoviridae using the ML method based on the complete nucleotide sequences of ORF1. The clusters are labeled in color. The numerical values presented at the nodes of the tree correspond to percent bootstrap values. The scale bar corresponds to genetic divergence. Selected members of the family Alphaflexiviridae were used as an outgroup
In summary, here we describe a new mosquito-associated virus named Mutum virus that is related to other tymovirid-like insect specific viruses (ISVs). The viruses isolated from mosquitoes are currently unclassified. The final position of MUTV and other mosquito-infecting tymovirid-like viruses within the ICTV framework of virus classification, in particular within the family Tymoviridae, is yet to be determined and may involve establishment of new genera to officially classify these viruses.
References
Martelli GP, Sabanadzovic S, Abou Ghanerm-Sabanadzovic N, Edwards MC, Dreher T (2002) The family Tymoviridae. Arch Virol 147(9):1837–1846. https://doi.org/10.1007/s007050200045
Wang L, Lv X, Zhai Y et al (2012) Genomic characterization of a novel virus of the family Tymoviridae isolated from mosquitoes. PLoS One 7(7):e39845. https://doi.org/10.1007/s00705-018-4098-x
Charles J, Tangudu CS, Hurt SL et al (2019) Discovery of a novel Tymoviridae-like virus in mosquitoes from Mexico. Arch Virol 164(2):649–652. https://doi.org/10.1007/s00705-018-4098-x
da Rosa APAT, da Rosa ES, Travassos da Rosa JFS et al (1998) Os arbovírus no Brasil: Generalidades, Métodos e Técnicas de Estudo. Documento Técnico n° 2. Instituto Evandro Chagas, Fundação Nacional de Saúde, Ministério da Saúde, Belém
Myung IJ (2003) Tutorial on maximum likelihood estimation. J Math Psychol. 47(1):90–100. https://doi.org/10.1016/S0022-2496(02)00028-7
Stamatakis A (2014) RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30(9):1312–3. https://doi.org/10.1093/bioinformatics/btu033
Acknowledgements
We thank the Evandro Chagas Institute for supporting the study, and CNPq for providing an MSc scholarship to the first author. We thank the National Institute of Amazonian Research for logistical support and fundraising, and Energia Sustentável do Brasil for financial aid.
Funding
This study was funded by Energia Sustentável do Brasil – ESBR, P&D ANEEL - (PD − 06631-0005/2017) and Evandro Chagas Institute/Ministry of Health of Brazil.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that there is no conflict of interest.
Ethics declaration
The samples were made available by project "Development of a methodology for monitoring behavioral dynamics for Mansonia Spp. and its relevance in hydroelectric use in the Amazon” (PD − 06631-0005/2017) with approval of Chico Mendes Institute for Biodiversity, protocol No. 37174–1.
Additional information
Handling Editor: Sead Sabanadzovic
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic Supplementary Material
Below is the link to the electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Miranda, K.K. ., Galvão, G.J.P., da Silva Araújo, P.A. et al. Discovery and genome sequencing of a new virus related to members of the family Tymoviridae, isolated from mosquitoes of the genus Mansonia in Brazil. Arch Virol 167, 1889–1892 (2022). https://doi.org/10.1007/s00705-022-05475-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00705-022-05475-x