Abstract
The complete genomic sequence of scorzonera virus A (SCoVA) from a Scorzonera austriaca Willd. plant in South Korea was determined by high-throughput sequencing and confirmed by Sanger sequencing. The SCoVA genome contains 9867 nucleotides, excluding the 3′-terminal poly(A) tail. The SCoVA genome structure is typical of potyviruses and contains a single open reading frame encoding a large putative polyprotein of 3168 amino acids. Pairwise comparison analysis of the complete genome and polyprotein sequences of SCoVA with those of other potyviruses showed that they shared the highest nucleotide and amino acid sequences identity (54.47% and 49.57%, respectively) with those of lettuce mosaic virus (GenBank accession number KJ161185). Phylogenetic analysis of the amino acid sequence of the polyprotein confirmed that SCoVA belongs to the genus Potyvirus. These findings suggest that SCoVA should be considered a novel member of the genus Potyvirus, family Potyviridae.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Scorzonera austriaca Willd. is a perennial herb in the family Asteraceae (also known as Compositae) that is distributed in a wide variety of habitats from subpolar to tropical regions. S. austriaca has been used widely as a traditional herbal medicine for curing various diseases [1].
The genus Potyvirus is the largest genus in family Potyviridae. Members of this genus form flexuous filamentous particles (680 to 900 nm long) that contain a monopartite positive-sense RNA genome approximately 9.7–10 kb in length with a 3′ poly(A) tail [2]. The genome contains a single large open reading frame (ORF) encoding a polyprotein that is post-translationally cleaved into 10 mature proteins: protein 1 (P1), the helper component-protease (HC-Pro), protein 3 (P3), 6 kDa peptide 1 (6K1), the cylindrical inclusion protein (CI), 6 kDa peptide 2 (6K2), the genome-linked viral protein (VPg), the nuclear inclusion-a protease (NIa-Pro), the nuclear inclusion-b protein (NIb), the capsid protein (CP), and a small frameshift-derived peptide (PIPO) from the P3 region [3]. Potyviruses are transmitted in a non-persistent manner by more than 200 species of aphids and cause serious diseases in a wide range of economically important crops and wild plants worldwide [4].
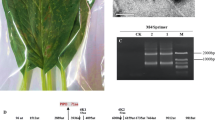
An S. austriaca plant with virus-like symptoms, including mild leaf mosaic and mild yellowing, was collected in Haenam County, South Jeolla Province, South Korea, on 24 June 2020. To determine the etiology of the virus-like symptoms, total RNA was extracted from the symptomatic leaves using an RNeasy Plant Mini Kit (QIAGEN, Hilden, Germany). A cDNA library was built from an rRNA-depleted library using a Ribo-Zero™ rRNA Removal Kit (Plant Leaf) (Illumina, CA, USA). Subsequently, the sample library was sequenced by Beijing Genomics Institute (BGI, China) using the BGISEQ platform to generate 100-bp paired-end reads. From the sequences generated, we obtained a total of 605,027,638 clean reads, from which the adaptor and low-quality sequences were trimmed. The high-quality clean reads were assembled de novo, and the resulting contig sequences were searched against the GenBank database using BLASTx. Among the assembled contigs, we detected one large potyvirus-related contig that was 9,756 nt long and shared the highest identity (50.90%) with lettuce mosaic virus (LMV, GenBank accession number KJ161185), which suggested that the nearly complete, assembled viral genome sequence was from a novel potyvirus, which we tentatively named "scorzonera virus A" (SCoVA). To confirm the full SCoVA genome sequence, total RNA was extracted from the symptomatic leaves of S. austriaca using a HiGene Total RNA Prep Kit (BIOFACT, Daejeon, South Korea). Contig-specific primers were designed based on the 9,756-nt long SCoVA contig sequence. The primer sequences are listed in Supplementary Table S1. Reverse transcription (RT)-PCR was performed with SuPrimeScript RT-PCR Premix (GeNet Bio, Daejeon, South Korea). Most of the SCoVA genome sequence was obtained by amplifying 10 partial overlapping RT-PCR fragments, and the 5′- and 3′-terminal sequences were determined by Rapid Amplification of cDNA Ends (RACE) using RACE kits (Invitrogen, Waltham, MA, USA) according to the manufacturer’s protocols [5].
All of the PCR products were cloned into an RBC T&A Cloning Vector (RBC Bioscience, Taipei, Taiwan), and at least three clones for each fragment were selected randomly and sequenced by GenoTech (Daejeon, South Korea). The sequences were assembled and analyzed using DNAMAN software (Lynnon Biosoft, Quebec, Canada). Pairwise comparisons of the nt and aa sequences were performed using DNAMAN (Lynnon Biosoft). Phylogenetic analysis was performed using the maximum-likelihood (ML) method with 1,000 bootstrap replicates in Molecular Evolutionary Genetic Analysis (MEGA) version 10.1.8 [6].
The complete genome sequence of SCoVA (GenBank accession number MW972223) is 9,867 nt long, excluding the 3′ poly(A) tail (Fig. 1). The 5′ and 3′ untranslated regions (UTRs) are 116 nt and 244 nt long, respectively. The SCoVA genome contains a single large ORF of 9,507 nt encoding a putative polyprotein of 3,168 aa (359.28 kDa), with an AUG start codon (nt 117–119) and a UAA stop codon (nt 9,621–9,623). The SCoVA polyprotein was predicted to be cleaved to produce 10 functional proteins: P1 (355 aa), HC-Pro (458 aa), P3 (358 aa), 6K1 (52 aa), CI (647 aa), 6K2 (53 aa), VPg (194 aa), NIa-Pro (243 aa), NIb (520 aa), and CP (288 aa). The predicted cleavage sites are IIHY/S, YQVG/G, ISLQ/A, VVLQ/S, ILLQ/S, ITLQ/G, INLE/S, IRLQ/S, and VLHA/L, respectively, which are similar to those of closely related viruses (Fig. 1) [7]. Additionally, a small ORF termed PIPO, which encodes 83-aa protein, was also identified by the presence of the G1A6 motif (nt 3038-3044) [8, 9]. Analysis of the SCoVA polyprotein aa sequence revealed several conserved functional motifs: 536FRNK539, 665PTK667, 697GYCY700, and 756KAEL759 in HC-Pro; 1311GSGKS-X3-P1319, 1328VLLLEPTRPL1337, 1397DECH1400, 1426SATPP1430, 1475LVYV1478, 1525VIATNIIENGVTL1537 (instead of VATNIIENGVTL), and 1570GERIQRLGRVGR1581 in CI; 1985MYGY1988 (instead of MYGF) in VPg; 2530SLKAEL2535, 2667GNNSGQPSTVVDNT2680 (instead of GNNSGQPSTVVDTN), and 2711GDD2713 in NIb; 2889DAG2891, 3016MVWCIENGTSP3026, 3099AFDF3102, and 3119QMKIAA3124 (instead of QMKAAA) in CP. Some of these conserved motifs have particularly important functions; namely, FRNK and PTK in HC-Pro, which are involved in symptom development and aphid transmission, respectively, DECH in CI, which is important for helicase activity, GDD in NIb, which is essential for RNA polymerase activity, and DAG in CP, which is involved in aphid transmission [10, 11].
Schematic representation of the genome organization of scorzonera virus A (SCoVA). Putative protein cleavage sites for generating the 10 mature proteins are compared with those of the four most similar potyviruses. The 5′ and 3′ untranslated regions (UTRs) are represented as black bold lines, and the open reading frame, which encodes the polyprotein, is depicted by an open box. The numbers below the genome indicate the positions of the putative cleavage sites. The amino acid sequences at the cleavage sites of SCoVA (GenBank accession no. MW972223) and the other four potyviruses are given below the numbers. The small rectangle above the putative P3 protein indicates the position of PIPO. LMV, lettuce mosaic virus (KJ161185); PPV, plum pox virus (MN603425); TuMV, turnip mosaic virus (EF374098); PVA, potato virus A (NP659729)
A BLASTn search showed that the complete genome sequence of SCoVA shared the highest nt sequence identity (68.77%; 42% query coverage) with LMV. We downloaded the genome sequences of 31 potyviruses from the GenBank database for pairwise comparisons and phylogenetic analysis. The pairwise comparison analyses indicated that the SCoVA genome and polyprotein sequences shared 48.58%–54.47% nt and 38.37%–49.57% aa sequence identity with those of the known potyviruses and that SCoVA shared the highest nt (54.47%) and aa (49.57%) sequence identity with LMV. Pairwise comparisons of the cleaved proteins of SCoVA with those of the 31 potyviruses showed that the aa sequence identity was 10.94%–20.72% in P1, 40.35%–55.46% in HC-Pro, 11.73%–29.33% in P3, 23.88%–73.08% in 6K1, 45.56%–57.70% in CI, 12.96%–50.94% in 6K2, 13.47%–57.14% in VPg, 37.86%–55.56% in NIa-Pro, 56.62%–67.88% in NIb, and 46.96%–59.38% in CP (Supplementary Table S2). According to the current International Committee on Taxonomy of Viruses species demarcation criteria for the genus Potyvirus, less than 76% nucleotide (nt) and less than 82% amino acid (aa) identity in the complete polyprotein or coat protein region are required to distinguish potyvirus species [12,13,14]. Our pairwise comparison results are all much lower than the current species demarcation cutoffs and confirm that SCoVA is distinct from other recognized potyviruses. The maximum-likelihood phylogenetic tree showed that SCoVA is most closely related to LMV in the genus Potyvirus (Fig. 2).
Maximum-likelihood (ML) phylogenetic tree of the polyprotein amino acid sequences of SCoVA and representative members of the family Potyviridae. The tree was constructed using the maximum-likelihood algorithm in MEGA. Bootstrap analysis was applied using 1,000 replicates. Cucumber vein yellowing virus (CVYV), genus Ipomovirus, family Potyviridae (GenBank accession number AY578085), was used as an outgroup.
In summary, we have determined and characterized the complete genome sequence of a novel virus, SCoVA, from S. austriaca. On the basis of pairwise comparisons and phylogenetic analysis with other potyviruses and the current species demarcation criteria, we consider SCoVA to be a novel member of the genus Potyvirus in the family Potyviridae.
References
Xie Y, Guo Q-S, Wang G-S (2016) Flavonoid glycosides and their derivatives from the Herbs of Scorzonera austriaca Wild. Molecules 21:803
Urcuqui-Inchima S, Haenni A-L, Bernardi F (2001) Potyvirus proteins: a wealth of functions. Virus Res 74:157–175
Deng P, Wu Z, Wang A (2015) The multifunctional protein CI of potyviruses plays interlinked and distinct roles in viral genome replication and intercellular movement. Virol J 12:1–11
Gadhave KR, Gautam S, Rasmussen DA, Srinivasan R (2020) Aphid transmission of Potyvirus: the largest plant-infecting RNA virus genus. Viruses 12:773
Lim S, Lee S-H, Moon JS (2019) Complete genome sequence of a putative novel potyvirus isolated from Platycodon grandiflorum. Adv Virol 164:621–624
Kumar S, Stecher G, Li M, Knyaz C, Tamura K (2018) MEGA X: molecular evolutionary genetics analysis across computing platforms. Mol Biol Evol 35:1547–1549
Goh CJ, Hahn Y (2021) Analysis of proteolytic processing sites in potyvirus polyproteins revealed differential amino acid preferences of NIa-Pro protease in each of seven cleavage sites. PLoS ONE 16:0245853
Rodamilans B, Valli A, Mingot A, San León D, Baulcombe D, López-Moya JJ, García JA (2015) RNA polymerase slippage as a mechanism for the production of frameshift gene products in plant viruses of the Potyviridae family. J Virol 89:6965–6967
Olspert A, Chung BYW, Atkins JF, Carr JP, Firth AE (2015) Transcriptional slippage in the positive-sense RNA virus family Potyviridae. EMBO Rep 16:995–1004
Shiboleth YM, Haronsky E, Leibman D, Arazi T, Wassenegger M, Whitham SA, Gaba V, Gal-On A (2007) The conserved FRNK box in HC-Pro, a plant viral suppressor of gene silencing, is required for small RNA binding and mediates symptom development. J Virol 81:13135–13148
Wu X, Lai Y, Lv L, Han K, Chen Z, Lu Y, Peng J, Lin L, Chen J, Zheng H (2021) Complete genome sequence of a new achyranthes virus A isolate from Achyranthes bidentata in China. Adv Virol 166:287–290
Adams M, Antoniw J, Fauquet C (2005) Molecular criteria for genus and species discrimination within the family Potyviridae. Adv Virol 150:459–479
Adams MJ, Lefkowitz EJ, King AM, Harrach B, Harrison RL, Knowles NJ, Kropinski AM, Krupovic M, Kuhn JH, Mushegian AR (2017) Changes to taxonomy and the International Code of Virus Classification and Nomenclature ratified by the International Committee on Taxonomy of Viruses (2017). Adv Virol 162:2505–2538
King AM, Lefkowitz E, Adams MJ, Carstens EB (2011) Virus taxonomy: ninth report of the International Committee on Taxonomy of Viruses. Elsevier
Acknowledgements
This study was supported by the KRIBB Research Initiative Program (No. KGM5372113), which is funded by the Ministry of Science and ICT, Republic of Korea. We thank Margaret Biswas, PhD, from Edanz Group (https://en-author-services.edanz.com/ac) for editing a draft of this manuscript. We are thankful to the Office of Research and Innovation at the Mongolian National University of Education.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
This article does not contain any studies of involving human participants or animals.
Additional information
Handling Editor: Ralf Georg Dietzgen.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Igori, D., Cho, H.S., Kim, H. et al. Complete genome sequence and genome organization of scorzonera virus A (SCoVA), a novel member of the genus Potyvirus. Arch Virol 166, 2901–2904 (2021). https://doi.org/10.1007/s00705-021-05182-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00705-021-05182-z