Abstract
Human immunodeficiency virus (HIV) with transmitted drug-resistance (TDR) limits the therapeutic options available for treatment-naive HIV patients. This study aimed to further our understanding of the prevalence and transmission characteristics of HIV with TDR for the application of first-line antiretroviral regimens. A total of 6578 HIV-1 protease/reverse-transcriptase sequences from treatment-naive individuals in China between 2000 and 2016 were obtained from the Los Alamos HIV Sequence Database and were analyzed for TDR. Transmission networks were constructed to determine genetic relationships. The spreading routes of large TDR clusters were identified using a Bayesian phylogeographic framework. TDR mutations were detected in 274 (4.51%) individuals, with 1.40% associated with resistance to nucleoside reverse transcriptase inhibitors, 1.52% to non-nucleoside reverse transcriptase inhibitors, and 1.87% to protease inhibitors. The most frequent mutation was M46L (58, 0.89%), followed by K103N (36, 0.55%), M46I (36, 0.55%), and M184V (26, 0.40%). The prevalence of total TDR initially decreased between 2000 and 2010 (OR = 0.83, 95% CI 0.73–0.95) and then increased thereafter (OR = 1.50, 95% CI 1.13–1.97). The proportion of sequences in a cluster (clustering rate) among HIV isolates with TDR sequences was lower than that of sequences without TDR (40.5% vs. 48.8%, P = 0.023) and increased from 27.3% in 2005–2006 to 63.6% in 2015–2016 (P < 0.001). While most TDR mutations were associated with reduced relative transmission fitness, mutation M46I was associated with higher relative transmission fitness than the wild-type strain. This study identified a low-level prevalence of TDR HIV in China during the last two decades. However, the increasing TDR HIV rate since 2010, the persistent circulation of drug resistance mutations, and the expansion of self-sustaining drug resistance reservoirs may compromise the efficacy of antiretroviral therapy programs.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The global scale-up of antiretroviral therapy (ART) for HIV-1 infections has resulted in considerable reductions in HIV/AIDS-related morbidity and mortality [1,2,3]. However, drug resistance can hamper virological suppression in patients who receive treatment. After decades of ART scale-up, the increasing HIV drug resistance in low- and middle-income countries, such as China, poses a growing threat [4].
The Chinese government officially launched the National Free Antiretroviral Treatment Program (NFATP) in 2003. By the end of 2015, approximately 70% of individuals in this country living with HIV/AIDS received ART [5]. Nearly half of ART failures in China are reported to be associated with drug-resistance-associated mutations (DRMs) [6, 7]. Primary infection with drug-resistant HIV strains limits a newly diagnosed patient’s treatment options and is associated with treatment failure. Previous studies have shown that China had a prevalence of HIV with transmitted drug resistance (TDR) of approximately 3.6% in ART-naive patients in 2015 [8, 9], which was lower than that in Western countries [10,11,12,13]. However, given the large number of HIV-infected people and the persistent circulation of HIV in China, the number of patients for whom ART failed because of drug resistance is considerably high in this country.
To achieve the UNAIDS’s goal of ending the AIDS pandemic by 2030, ever-increasing numbers of people will be enrolled in the ART program in the coming years. Thus, with the quick scale-up of treatment and the passage of time, drug resistance will require more attention. Little is known about the temporal trend of HIV with TDR or its transmission characteristics in China.
In this study, we examined the changing prevalence of HIV with TDR from 2000 to 2016 in ART-naive patients in China, utilizing pol gene sequences of HIV extracted from the Los Alamos HIV Sequence Database. We performed comprehensive phylogenetic analysis to explore the transmission characteristics of drug-resistant HIV strains. In addition, we explored the evolutionary characteristics of the largest transmission clusters by applying Bayesian coalescent-based methods.
Materials and methods
Data collection
We retrieved all available records of HIV-1 sequences from China containing the pol gene region from the Los Alamos HIV Sequence Database (http://www.hiv.lanl.gov). A total of 12,447 sequences were obtained along with basic information, including the GenBank accession number, subtype, sampling location, collection date, and ART status, if provided. For sequences without a denoted ART status, we collected information for each sequence from the original source publication. The pol gene region was extracted from these data according to the annotation information, with all sequences < 500 nt or for which the ART status was “yes” or “unknown” removed. For patients with multiple sequences, only the earliest sequence was retained.
Sequences were aligned using MAFFT (version 7) [14] and manually checked and edited using MEGA (version 7) [15]. All of the sequences were selected manually to maximize the length and the number of segments for analysis. Segments spanning 1053 nt of the pol gene, including the entire protease (PR) and partial reverse transcriptase (RT) regions (HXB2 nucleotide coordinates 2253–3305), were ultimately selected.
Drug resistance mutation analysis
The Stanford Calibrated Population Resistance Tool version 7.0 (https://hivdb.stanford.edu/) was used to screen the pol sequences for surveillance drug resistance mutations (SDRMs), according to the World Health Organization (WHO)-recommended surveillance drug resistance mutation list, which was updated in 2009 for nucleoside reverse transcriptase inhibitors (NRTIs), non-nucleoside reverse transcriptase inhibitors (NNRTIs), and protease inhibitors (PIs). HIV with TDR was defined as an HIV strain containing at least one of the standard surveillance drug resistance mutations listed by the WHO.
Transmission clusters and network construction
The transmission network was built as described previously [16]. Briefly, an approximately-maximum-likelihood phylogenetic tree was estimated for all sequences using the GTR + G + I nucleotide substitution model in FastTree 2 [17] after excising 43 codons associated with drug resistance from the alignment. The Tamura-Nei 93 (TN93) pairwise genetic distances (PGDs) were calculated for all pairs of sequences to identify potential transmission partners. Transmission clusters (TCs) were defined as sequences for which the bootstrap support value was greater than 0.9 and the PGD was less than 3.0%, and these were employed to detect longer-term transmission events. TCs were extracted from the phylogenetic tree using Cluster Picker [18]. In an undirected tree (extracted TC), there are multiple sub-trees that could be used to construct the transmission network. To select the best tree with the minimized sum of the edge weight (TN93 PGDs), a minimum genetic distance algorithm was used to define the network structure of a cluster [19]. To visualize and analyze networks, the network data were processed in the R software package utilizing the ape (version 5.4-1) [20] and qgraph packages (version 1.6.5) [21].
Bayesian phylogeographic inference
We used BEAST (Bayesian evolutionary analysis sampling trees, version 1.10.4) [22] to conduct the Bayesian phylogeographic inference for large HIV with TDR clusters after removing all of the drug-resistance-associated sites from the alignment. All of the clusters were analyzed using a suitable nucleotide substitution model (HKY for M46L clusters, JC for M46I clusters), which was selected by jModelTest (version 2.1.10) [23]. We used a strict molecular clock model to infer the timescale of HIV evolution. A Bayesian skygrid coalescent tree was used to estimate the effective infection population size. A continuous-time Markov chain (CTMC) model was used to estimate the transition rates between each pair of locations, and the Bayesian stochastic search variable selection (BSSVS) procedure was applied to identify well-supported transitions and infer the location state of the ancestral nodes of the trees.
Markov chain Monte Carlo (MCMC) analyses were run for 100 million generations, with parameter values sampled every 10,000 steps. The resulting log files were imported into the program TRACER v1.7.1, and the first 10% of the output was discarded as burn-in from each run. The convergence of the estimates was evaluated using generation vs. log probability plots in TRACER, using an effective sample size >200. When reporting the substitution rate and the time of the most recent common ancestor (tMRCA), we used the mean of the estimate parameter. The Bayes factor (BF) was calculated using SpreaD3 [24] to identify well-supported transitions by applying a cutoff of 3.0. TreeAnnotator was used to generate a summary maximum-clade-credibility (MCC) tree from the posterior distribution of the trees (after removal of the MCMC burn-in of 10%).
Statistical analysis
The temporal trends of the prevalence of HIV with TDR were evaluated with odds ratios (ORs) and 95% confidence intervals (CIs) using a logistic regression model. An OR > 1 indicates an increase in the prevalence of HIV with TDR over time, and an OR < 1 indicates a decrease over time. Furthermore, we examined the trend in a smaller window size, using a local weighted regression model with a reciprocal of the standard error of prevalence set as the weight. Fischer’s exact test was used for the analysis of differences between proportions, and Student’s t-test was used for the analysis of differences between means. Transmission fitness was defined as the ability of a pathogen to be transmitted and cause secondary cases of the disease as a means to describe the fitness of the virus at the population level [25]. To evaluate the transmission fitness of each TDR mutation, we calculated the ORs and 95% CIs from the clustering frequency by comparison with wild-type strains using a logistic regression model. An OR > 1 indicates positive fitness, and an OR < 1 indicates negative fitness. Statistical significance was defined as a p-value < 0.05. All analyses were carried out with the R program (version 3.5.3) [26].
Results
Characteristics of the study dataset
Ultimately, a total of 6578 HIV-1 pol gene sequences from treatment-naive patients from 24 provinces in China between 2000 and 2016 were obtained. Of these patients, 5155 were male (78.37%) and 313 were female (4.76%). The patient characteristics are shown in Table 1. The distribution of sequences in regions from each year is displayed in Supplementary Fig. S1A. The largest group of patients (3067, 46.6%) were infected with CRF01_AE, followed by CRF07_BC (2013, 30.6%) and subtype B (836, 12.7%) (Table 2). The remaining patients were infected with other pure subtypes or circulating recombinant forms (CRFs, 662, 10.1%). Subtype B infections decreased from 34.1% to 11.2% from 2000 to 2016 (p < 0.001). CRF01_AE and CRF07_BC infections increased over time (29.5% to 47.1%, p < 0.001; 11.4% to 31.6%, p = 0.0229) (Fig. 1). The infection routes constituted 4414 (67.1%) men having sex with men (MSM), 654 (9.9%) heterosexual sex, 622 (9.5%) injected drug use (IDU), and 888 (13.5%) other infection routes or data unrecorded (Table 2). The composition of infection routes of all HIV-infected patients in regions is displayed in Supplementary Fig. S2A.
Prevalence and temporal trend of transmitted drug resistance
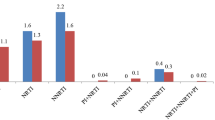
HIV with TDR was demonstrated in 274 (4.51%) patients in the period from 2000 and 2016. Supplementary Table S1 shows information for all of the HIV-1 sequences with TDR mutations. The distribution of TDR mutated sequences and TDR prevalence in regions from each year are displayed in Supplementary Figures S1B and S3. Mutations conferring resistance to NRTIs, NNRTIs, PIs, and multiclass drugs were found in 92 (1.40%), 100 (1.52%), 123 (1.87%), and 37 (0.56%) patients, respectively. The prevalence of TDR in both CRF01_AE (5.31%) and subtype B (6.58%) was higher than that in the other subtypes (Table 2). Among the subtypes, CRF01_AE had the highest prevalence of PI-TDR, while subtype B had the highest prevalence of NRTI- and NNRTI-associated TDR. Among the different infection route populations, the prevalence of HIV with TDR in heterosexual sex individuals was relatively high (7.34%). MSM had the highest prevalence of PI-TDR among the risk groups, while the prevalence of NNRTI-TDR in heterosexuals was higher than that in the IDU and MSM groups (Table 2). The composition of the infection routes of patients with TDR HIV in regions is shown in Supplementary Fig. S2B (Fig. 2).
The prevalence of drug-resistance mutations in all sequences is shown in Table 3. Overall, the most frequent mutation was M46L, which was found in 0.89% (n = 58) of patients, followed by K103N (0.55%, n = 36), M46I (0.55%, n = 36), and M184V (0.40%, n = 26). Furthermore, in patients infected with strain CRF01_AE, the most frequent mutation was M46L (1.76%), followed by M46I (1.04%), M184V (0.46%), and K103N (0.39%). The most frequent TDR mutation in patients infected with CRF07_BC was M184V (0.39%), followed by T215S (0.20%) and Y181C (0.15%). The most frequent TDR mutation in subtype B was K103N (2.27%), followed by G190A (0.96%), Y181C/Y188L/M41L/D67N (0.84%), and M184V (0.72%). The prevalence of TDR mutations in each HIV-1 subtype is shown in Fig. 2.
The prevalence of HIV with TDR varied throughout the study period. It was 2.73% in 2011–2012 and 9.09% in 2000–2003, with no clear trend observed (OR = 0.94, 95% CI 0.86–1.04, p = 0.233). The prevalence of NRTI (OR = 0.76, 95% CI 0.66–0.89, p <0.001) and NNRTI-associated (OR = 0.84, 95% CI 0.73–0.98, p = 0.020) TDR decreased slightly from 2000 to 2016, while the prevalence of PI-associated (OR = 1.09, 95% CI 0.95–1.27, p = 0.233) TDR was stable during the same period.
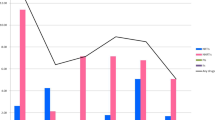
However, when we examined the trends in a smaller window, the prevalence of total TDR was found to decrease from 2000 to 2010 (OR = 0.83, 95% CI 0.73–0.95, p = 0.006), but it increased after 2010 (OR = 1.50, 95% CI 1.13–1.97, p = 0.004). The prevalence of NNRTI-associated TDR decreased from 2000 to 2010 (OR = 0.77, 95% CI 0.64–0.94, p = 0.007) and increased after 2010 (OR = 1.91, 95% CI 1.13–3.18, p = 0.014). PI-associated TDR was found to increase slightly after 2006, with no statistical significance, but it increased after 2010, with marginal significance (OR = 1.44, 95% CI 0.99–2.09, p = 0.055) (Fig. 3).
Number distribution (A) and temporal trends in the yearly proportion (B) of individuals with transmitted HIV drug resistance by drug class. Note that a local weighted regression was applied to the prevalence to smooth the curves. Abbreviations: NNRTI, non-nucleoside reverse transcriptase inhibitor-associated drug resistance; NRTI, nucleoside reverse transcriptase inhibitor-associated drug resistance; PI, protease inhibitor-associated drug resistance; Any class, any class of transmitted drug resistance
Clusters and transmission fitness of TDR mutations
Finally, Cluster Picker identified 788 transmission clusters containing 3189 of 6578 patients (48.4%). Strains CRF01_AE and CRF07_BC had higher rates of phylogenetic clustering than the other subtypes, while the rate of clustering was higher in MSMs than in the other infection route groups (Table 2).
Of these 788 clusters, 69 clusters had one or more sequences with TDR mutations, comprising 111 of 274 (40.5%) TDR sequences. The network construction of these 69 transmission clusters is shown in Supplementary Fig. S4. The clustering rate of sequences with TDR increased annually from 27.3% in 2005–2006 to 63.6% in 2015–2016 (p < 0.001). Of these 69 TDR clusters, 52 clusters had only one TDR sequence, 10 clusters had two TDR sequences, and 7 clusters had three or more TDR sequences. Among the 17 clusters with at least two TDR sequences, 15 clusters had the same TDR mutation, seven of which were M46L, followed by M46I (n = 2).
Overall, the sequences with TDR were less likely to form phylogenetic clusters than the sequences not resulting in TDR, even after adjusting for subtype and at-risk populations (40.5% vs. 48.8%, p = 0.023). The clustering rates of NNRTI- and NRTI-associated TDR mutations were significantly lower than those of the wild-type sequences (Table 3). The mutation M46L had the highest frequency in the clusters (n = 32), followed by M46I (n = 26), L210W (n = 7), M184V (n = 7), T215S (n = 5), and K103N (n = 4).
Of the 58 TDR mutations, seven had clustering frequencies that deviated significantly from the neutral expectation, most of which resulted in a significant reduction in transmission fitness (NNRTI and NRTI mutations: K103N, G190A, Y181C, K101E). Conversely, M46I had a relative fitness significantly greater than 1.0 (Table 3).
The spatiotemporal origins and epidemic dynamics of large TDR clusters
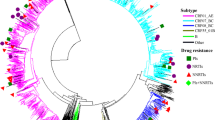
There were four large HIV with TDR clusters containing ≥ 5 sequences. All of these sequences were from patients infected with strain CRF01_AE in the MSM risk group. We extracted the two largest clusters containing the mutants M46I (cluster 300) and M46L (cluster 67) to implement Bayesian phylogeographic inference. The time of the most recent common ancestor (tMRCA) of cluster 67 and cluster 300 was 2007.2 (95% HPD 2004.1–2008.5) and 2006.9 (95% HPD 2004.1–2008.5), respectively. The substitution rates of clusters 67 and 300 were 2.144 × 10−3 (95% HPD 1.075×10-3–3.213 × 10-3) and 2.933 × 10-3 (95% HPD 1.385 × 10-3–4.873 × 10-3), respectively. From the phylogeographical analysis, the potential transmission source locations for clusters 67 and 300 were Beijing (posterior probability [PP] = 0.80) and Liaoning (PP = 0.69), respectively (Fig. 4).
Molecular dating analysis of large HIV-1 clusters with evidence of transmitted drug resistance. The phylogenies of the (A) M46L and (B) M46I clusters are colored according to the sampling location. The posterior probabilities of the potential transmission sources of clusters are annotated on the right of the node.
Interestingly, two other clusters, 65 and 68, together with clusters 62, 63, and 69, were adjacent to cluster 67 in the phylogenetic tree, and most of them shared the same TDR mutation, M46L. Analysis of the sequences from these clusters showed that the TN93 pairwise genetic distances were < 3.0% for the majority of the sequences (Supplementary Fig. S5). Therefore, we applied the same analyses to these clusters with the tMRCA estimated at 2002.9 (95% HPD 2000.4–2004.8) and its substitution rate estimated at 2.087 × 10-3 (1.467 × 10-3–2.818 × 10-3) while Beijing was estimated as the source location with the highest support (PP = 0.99) (Fig. 4). Analysis of the geographical migration as estimated from the Bayes factors indicated that the migration pathways, from Beijing to Hebei, Hubei, Hunan, and Guangdong, and from Hubei to Hunan and Shanghai, were all well supported for the M46L clusters, while in the large M46I cluster, viral migration from Beijing to Liaoning was well supported. Table 4 illustrates the best-supported pathway of migration of the large M46I/L clusters in China.
Discussion
In the present study, we identified transmitted drug-resistant HIV in ART-naive HIV-1-infected individuals in China. The overall prevalence of HIV with TDR was 4.51%, which is comparable to that reported by Zhao et al. and Su et al. (3%–5%) [8, 9]. It is not surprising that the prevalence of HIV with TDR in China was lower than that in Europe (9.4%) and North America (11.5%), where ART was initiated much earlier [27].
The prevalence of HIV with TDR varied throughout the study period and ranged from 2.73% in 2011–2012 to 9.09% in 2000–2003. The higher prevalence of HIV with TDR in the earlier phase of the NFATP may be due to the widespread use of generic agents, including zidovudine, didanosine, and nevirapine [28]. The decreasing trend in HIV with TDR from 2000 to 2010 mainly contributed to the decrease in NRTI-related resistance mutations (Fig. 3B). We speculate that the decline of the prevalence of HIV with TDR could be partially attributed to the antiretroviral drugs that were newly available after 2004, such as lamivudine and efavirenz. The toxicity and side effects of lamivudine are lower than those of didanosine, which improved patient compliance and provided better therapeutic effects [29]. Accordingly, lamivudine was used in the following years [30]. In addition, to guide the national free ART Program, four editions of the China Free ART Manual were released from 2004 to 2016. Because of standardized treatment guidance, increased availability of effective drugs, and improved patient compliance with the NFATP, the virological suppression rate increased from 78% in 2004–2005 to 90% in 2013 [8], and the prevalence of HIV with TDR declined accordingly.
However, after 2010, a slight increase in HIV with TDR prevalence was observed in our study. A similar trend between 2009 and 2012 was also observed by Su et al. [9]. The authors attributed this phenomenon to the broad expansion of access to ART during this time period and/or to the cumulative increase in prevalence of drug-resistant strains in HIV-infected patients at the onset of therapy. However, when we looked closely at the factors driving the increase, we found that PI-resistant strains were the main contributors to the growth trend since 2010. Furthermore, in both the study by Su et al. and this study, one noteworthy finding was the elevated prevalence of PI-specific TDR in the MSM group. MSM have become the most predominant risk group for HIV-1 infection in the last decade [31]. Accordingly, the high prevalence and an increasing trend of PI resistance in this population will pose a significant challenge to the prevention and control efforts in a large number of HIV-infected MSM in China.
The most frequent TDR mutation in China was M46I/L, whereas in Europe and North America, it was K103N [12, 32,33,34]. This difference might largely be attributed to the dominant HIV-1 subtypes that circulate between regions. In China, the predominant subtype of HIV-1 is CRF01_AE, in which most of the M46I/L substitutions were observed. In Europe and North America, subtype B is the most prevalent. The second-most frequent TDR mutation was K103N, which represents the most frequent TDR mutation in subtype B. This is consistent with what has been observed in European and North American isolates. Furthermore, the TDR rates overall and in NRTI/NNRTI in subtype B were higher than those in other subtypes. Subtype B was the predominant strain of the early HIV epidemic in China [35]. Therefore, it had a more prolonged exposure to ART and consequently accumulated drug resistance mutations, leading to this relatively high TDR prevalence.
In this study, we found that TDR mutant was associated with reduced transmission clustering rate, suggesting that TDR strains have lower fitness than the corresponding wild-type strains. Drug-resistance mutants can revert to wild type and then outcompete strains with the DRMs. In addition, viral competition experiments have demonstrated that DRMs generally reduce the replication rate of HIV (i.e., replicative fitness). When transmission occurs, systemic infection is typically established by a single genetic variant, and founder virus selection is biased toward certain genetic characteristics [36]. This genetic bottleneck could limit the transmission of DRMs. In the absence of antiretroviral therapy, strains containing DRMs commonly have reduced fitness compared with the wild-type strain within the population [3]. Therefore, TDR-associated strains are likely to be less active in onward transmission. This transmission disadvantage could explain the reduced clustering rate of TDR strains compared with wild-type strains. Despite this negative impact of TDR on transmission, the clustering rate of TDR strains remained at 40.5%, showing that the transmission clusters made a significant contribution to the spread of TDR. More importantly, we found that the clustering rate of TDR strains increased annually after 2004, indicating that the contribution of transmission clusters to the spread of TDR has been increasing.
In contrast, people infected with a virus containing the mutation M46I were involved in increased rates of transmission. The enhanced transmission of M46I strains is noteworthy, as it has implications for second-line regimens containing lopinavir or atazanavir in resource-limited countries [37]. Another mutation at the same site, M46L, is also noteworthy, as it had the highest prevalence in the study, with a high rate of clustering. Using Bayesian phylogeographic models, we found that Beijing was a critical transmission location for M46I/L. Interestingly, all of the sequences of M46I/L were derived from CRF01_AE-infected MSM patients. Considering the high prevalence of CRF01_AE in China and the increasing proportion of newly diagnosed HIV-infected patients within MSM populations [31], concern about the rapid onward transmission of the M46I/L variant within the Chinese MSM community is warranted. Given that M46I/L is one of the most commonly transmitted PI mutations globally, especially in South/Southeast Asia [27], there is an urgent need to better understand its clinical impact on the efficacy of the current second-line therapy, as well as its effects combined with other mutations.
Some limitations should be noted here. First, because data on the infection history and disease progression were limited, a proportion of the individuals might not have been infected recently. The overall trend of many TDR-associated mutations is toward a gradual reversion to the wild type in the years following the initial infection. Therefore, the drug-resistant strains in plasma would become minor quasispecies after a period of infection. Accordingly, the prevalence of HIV with TDR in chronically infected individuals is generally lower than that in acutely infected individuals, which would have impacted the estimated TDR rate determined in this study. Second, the population heterogeneity in the study was noteworthy. Due to the strict filter condition, the number of eligible sequences from females was relatively small, and the majority of the sequences were from MSM. Nevertheless, no significant difference in clustering rates was detected between females and males, either in heterosexual sex or IDU (data not shown). Therefore, we adjusted the infection route in the subsequent transmission analysis. Third, like all genetic clustering studies, we were unable to determine whether the genetic linkage results were due to direct or indirect transmissions.
In conclusion, the overall prevalence of HIV with TDR among pretreatment HIV-infected individuals in China has remained at a low level during the last two decades. However, the onward transmission of drug resistance mutations may compromise the efficacy of ART programs. We also found self-sustaining reservoirs of TDR mutations among transmission networks that may have important implications for the success of ART programs. We thus suggest that effective measures are still needed to strengthen monitoring and guide ART usage, such as drug-resistance testing for patients enrolling in ART programs.
References
Sterne JA, Hernan MA, Ledergerber B, Tilling K, Weber R, Sendi P et al (2005) Long-term effectiveness of potent antiretroviral therapy in preventing AIDS and death: a prospective cohort study. Lancet 366(9483):378–384. https://doi.org/10.1016/s0140-6736(05)67022-5
Grinsztejn B, Hosseinipour MC, Ribaudo HJ, Swindells S, Eron J, Chen YQ et al (2014) Effects of early versus delayed initiation of antiretroviral treatment on clinical outcomes of HIV-1 infection: results from the phase 3 HPTN 052 randomised controlled trial. Lancet Infect Dis 14(4):281–290. https://doi.org/10.1016/s1473-3099(13)70692-3
Coffin JM (1995) HIV population dynamics in vivo: implications for genetic variation, pathogenesis, and therapy. Science 267(5197):483–489. https://doi.org/10.1126/science.7824947
Gupta RK, Gregson J, Parkin N, Haile-Selassie H, Tanuri A, Andrade Forero L et al (2018) HIV-1 drug resistance before initiation or re-initiation of first-line antiretroviral therapy in low-income and middle-income countries: a systematic review and meta-regression analysis. Lancet Infect Dis 18(3):346–355. https://doi.org/10.1016/s1473-3099(17)30702-8
Ncaids N (2016) Update on the AIDS/STD epidemic in China and main response in control and prevention in December, 2015. J Chin J AIDS STD 22(2):69
Luo XL, Mo LD, Su GS, Huang JP, Wu JY, Su HZ et al (2019) Incidence and types of HIV-1 drug resistance mutation among patients failing first-line antiretroviral therapy. J Pharmacol Sci 139(4):275–279. https://doi.org/10.1016/j.jphs.2018.11.016
Kan W, Teng T, Liang S, Ma Y, Tang H, Zuohela T et al (2017) Predictors of HIV virological failure and drug resistance in Chinese patients after 48 months of antiretroviral treatment, 2008–2012: a prospective cohort study. BMJ Open 7(9):e016012. https://doi.org/10.1136/bmjopen-2017-016012
Zhao S, Feng Y, Hu J, Li Y, Zuo Z, Yan J et al (2018) Prevalence of transmitted HIV drug resistance in antiretroviral treatment naive newly diagnosed individuals in China. Sci Rep 8(1):12273. https://doi.org/10.1038/s41598-018-29202-2
Su Y, Zhang F, Liu H, Smith MK, Zhu L, Wu J et al (2014) The prevalence of HIV-1 drug resistance among antiretroviral treatment naive individuals in mainland China: a meta-analysis. PLoS ONE 9(10):e110652. https://doi.org/10.1371/journal.pone.0110652
Petersen A, Cowan SA, Nielsen J, Fischer TK, Fonager J (2018) Characterisation of HIV-1 transmission clusters and drug-resistant mutations in Denmark, 2004 to 2016. Euro Surveill 23:44. https://doi.org/10.2807/1560-7917.Es.2018.23.44.1700633
Rhee SY, Clutter D, Fessel WJ, Klein D, Slome S, Pinsky BA et al (2019) Trends in the molecular epidemiology and genetic mechanisms of transmitted human immunodeficiency virus type 1 drug resistance in a large US clinic population. Clin Infect Dis 68(2):213–221. https://doi.org/10.1093/cid/ciy453
Rossetti B, Di Giambenedetto S, Torti C, Postorino MC, Punzi G, Saladini F et al (2018) Evolution of transmitted HIV-1 drug resistance and viral subtypes circulation in Italy from 2006 to 2016. HIV Med 19(9):619–628. https://doi.org/10.1111/hiv.12640
McFaul K, Lim C, Jones R, Asboe D, Pozniak A, Sonecha S et al (2014) Transmitted antiretroviral drug resistance in treatment naive HIV-infected persons in London in 2011 to 2013. J Int AIDS Soc 17(4 Suppl 3):19747. https://doi.org/10.7448/ias.17.4.19747
Katoh K, Rozewicki J, Yamada KD (2017) MAFFT online service: multiple sequence alignment, interactive sequence choice and visualization. Brief Bioinform 20(4):1160–1166. https://doi.org/10.1093/bib/bbx108
Kumar S, Stecher G, Tamura K (2016) MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol Biol Evol 33(7):1870–1874. https://doi.org/10.1093/molbev/msw054
Yuan H, Liu Z, Wu X, Wu M, Fang Q, Tully DC et al (2019) Evolutionary characteristics and genetic transmission patterns of predominant HIV-1 subtypes among MSM in China. Int J Infect Dis 90:125–131. https://doi.org/10.1016/j.ijid.2019.10.035
Price MN, Dehal PS, Arkin AP (2010) FastTree 2-approximately maximum-likelihood trees for large alignments. PLoS ONE 5(3):e9490. https://doi.org/10.1371/journal.pone.0009490
Ragonnet-Cronin M, Hodcroft E, Hue S, Fearnhill E, Delpech V, Brown AJL et al (2013) Automated analysis of phylogenetic clusters. BMC Bioinformatics 14:317. https://doi.org/10.1186/1471-2105-14-317
Li X, Xue Y, Lin Y, Gai J, Zhang L, Cheng H et al (2016) Evolutionary dynamics and complicated genetic transmission network patterns of HIV-1 CRF01_AE among MSM in Shanghai. China. Sci Rep 6:34729. https://doi.org/10.1038/srep34729
Paradis E, Schliep K (2019) ape 5.0: an environment for modern phylogenetics and evolutionary analyses in R. Bioinformatics 35(3):526–528. https://doi.org/10.1093/bioinformatics/bty633
Epskamp S, Cramer AOJ, Waldorp LJ, Schmittmann VD, Borsboom D (2012) qgraph: network visualizations of relationships in psychometric data. J Stat Softw 48(4):1–18. https://doi.org/10.18637/jss.v048.i04
Bouckaert R, Heled J, Kuhnert D, Vaughan T, Wu CH, Xie D et al (2014) BEAST 2: a software platform for Bayesian evolutionary analysis. PLoS Comput Biol 10(4):e1003537. https://doi.org/10.1371/journal.pcbi.1003537
Darriba D, Taboada GL, Doallo R, Posada D (2012) jModelTest 2: more models, new heuristics and parallel computing. Nat Methods 9(8):772–772. https://doi.org/10.1038/nmeth.2109
Bielejec F, Baele G, Vrancken B, Suchard MA, Rambaut A, Lemey P (2016) Sprea D3: interactive visualization of spatiotemporal history and trait evolutionary processes. Mol Biol Evol 33(8):2167–2169. https://doi.org/10.1093/molbev/msw082
Biesinger T, Kimata JT (2008) HIV-1 transmission replication fitness and disease progression. Virol (Auckl) 2008(1):49–63
Team RC (2020) R: a language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria. https://www.R-project.org/
Rhee SY, Blanco JL, Jordan MR, Taylor J, Lemey P, Varghese V et al (2015) Geographic and temporal trends in the molecular epidemiology and genetic mechanisms of transmitted HIV-1 drug resistance: an individual-patient- and sequence-level meta-analysis. PLoS Med 12(4):e1001810. https://doi.org/10.1371/journal.pmed.1001810
Zhang F, Haberer JE, Wang Y, Zhao Y, Ma Y, Zhao D et al (2007) The Chinese free antiretroviral treatment program: challenges and responses. AIDS 21(Suppl 8):S143-148. https://doi.org/10.1097/01.aids.0000304710.10036.2b
Cao YZ, Lu HZ (2005) Care of HIV-infected patients in China. Cell Res 15(11–12):883–890. https://doi.org/10.1038/sj.cr.7290363
Dou Z, Zhang F, Zhao Y, Jin C, Zhao D, Gan X et al (2015) Progress on China’ s national free antiretroviral therapy strategy in 2002–2014. Zhonghua Liu Xing Bing Xue Za Zhi 36(12):1345–1350
2015 China AIDS response progress report (2015) National Health Family Planning Commission of the People’s Republic of China, Beijing, China. Available from: http://www.commuhealtibet.org/wp-content/uploads/2016/11/CHN_narrative_report_2015.pdf
Hofstra LM, Sauvageot N, Albert J, Alexiev I, Garcia F, Struck D et al (2016) Transmission of HIV drug resistance and the predicted effect on current first-line regimens in Europe. Clin Infect Dis 62(5):655–663. https://doi.org/10.1093/cid/civ963
Tostevin A, White E, Dunn D, Croxford S, Delpech V, Williams I et al (2017) Recent trends and patterns in HIV-1 transmitted drug resistance in the United Kingdom. HIV Med 18(3):204–213. https://doi.org/10.1111/hiv.12414
Panichsillapakit T, Smith DM, Wertheim JO, Richman DD, Little SJ, Mehta SR (2016) Prevalence of transmitted HIV drug resistance among recently infected persons in San Diego, CA 1996–2013. J Acquir Immune Defic Syndr 71(2):228–236. https://doi.org/10.1097/qai.0000000000000831
Li X, Li W, Zhong P, Fang K, Zhu K, Musa TH et al (2016) Nationwide trends in molecular epidemiology of HIV-1 in China. AIDS Res Hum Retroviruses 32(9):851–859. https://doi.org/10.1089/aid.2016.0029
Carlson JM, Schaefer M, Monaco DC, Batorsky R, Claiborne DT, Prince J et al (2014) HIV transmission. Selection bias at the heterosexual HIV-1 transmission bottleneck. Science 345(6193):1254031. https://doi.org/10.1126/science.1254031
Young TP, Parkin NT, Stawiski E, Pilot-Matias T, Trinh R, Kempf DJ et al (2010) Prevalence, mutation patterns, and effects on protease inhibitor susceptibility of the L76V mutation in HIV-1 protease. Antimicrob Agents Chemother 54(11):4903–4906. https://doi.org/10.1128/aac.00906-10
Funding
This study was supported by the Natural Science Foundation of China (81772170) and by the National Key Research and Development Program of China (No. 2017YFC0211704).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no competing interests.
Ethical approval
This study did not include experiments with human participants or animals performed by any of the authors.
Additional information
Handling Editor: Zhongjie Shi.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Yuan, H., Liu, Z., Wu, X. et al. Prevalence of transmitted HIV-1 drug resistance among treatment-naive individuals in China, 2000-2016. Arch Virol 166, 2451–2460 (2021). https://doi.org/10.1007/s00705-021-05140-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00705-021-05140-9