Abstract
Passion fruit woodiness disease (PWD), caused by cowpea aphid-borne mosaic virus (CABMV), produces socioeconomic problems in Brazil. The objectives of this study were to i) evaluate the temporal progression of PWD, ii) identify Passiflora genotypes with resistance to CABMV, and iii) detect virus infection in asymptomatic plants by reverse transcription quantitative polymerase chain reaction (RT-qPCR) in cases where standard RT-PCR detection failed. The experiment was conducted in a greenhouse using 128 genotypes belonging to 12 species and three hybrids (inter- and intraspecific) of Passiflora, evaluated at five time points after inoculation. Progression rates and disease severity were lower in P. cincinnata, P. gibertii, P. miersii, and P. mucronata than in P. edulis, P. alata, Passiflora sp., and hybrids. Of the genotypes tested, 20.31% were resistant, especially the accessions of P. suberosa, P. malacophylla, P. setacea, P. pohlii, and P. bahiensis, which remained asymptomatic throughout the experiment. The absence of symptoms does not imply immunity of plants to the virus, since RT-qPCR analysis confirmed infection by the virus in asymptomatic plants of P. cincinnata, P. gibertii, P. miersii, P. mucronata, P. setacea, P. malacophylla, and P. suberosa. Even after four inoculations, the virus was not detected by RT-qPCR in the upper leaves in plants of the species P. pohlii and P. bahiensis, indicating that these species are probably immune to CABMV.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Brazil stands out as the largest global producer of yellow passion fruit (Passiflora edulis Sims) [1, 2]. In 2018, the production was 593,429 metric tons from an area of 41,584 ha. Despite being the largest worldwide producer, the average productivity of 14.3 t ha-1 is low [3]. This low yield is partly due to the severity of passion fruit woodiness disease (PWD), caused by cowpea aphid-borne mosaic virus (CABMV) [4,5,6,7].
CABMV (genus Potyvirus, family Potyviridae) has a genome consisting of a single-stranded, positive-sense RNA, which encodes proteins affecting viral replication and accumulation, defense, viral movement, and symptoms in infected plants [8, 9]. The virus is transmitted by aphid vectors (Hemiptera: Aphididae) in a non-circulative and non-persistent manner during probing [10,11,12]. Plants infected by CABMV show inhibited growth, their leaves have a mosaic appearance, blisters, and/or deformations, and their fruits are deformed and smaller, becoming hardened [10, 13]. Diseases caused by viruses are considered to have the highest socioeconomic impact on passion fruit cultivation in Brazil because they reduce the plant’s longevity, productivity, and fruit quality, and there are no effective measures to control them [14, 15], only damage mitigation measures [16]. The use of resistant cultivars is considered the best strategy, because it does not increase production costs for labor or require chemicals to control the vector insect [17,18,19]. However, so far, there are no yellow passion fruit cultivars with this attribute [18,19,20,21]. On the other hand, studies indicate that wild passion fruit tree species carry CABMV resistance genes [19, 22,23,24,25], making them an alternative for developing resistant cultivars through interspecific crosses with susceptible species [18, 26].
The evaluation of the Passion Fruit Active Germplasm Bank of the Embrapa Cassava and Fruits (Embrapa Mandioca e Fruticultura) research unit, with the aim of identifying resistant wild genotypes is considered an indispensable step for the development of CABMV-resistant cultivars [19, 25]. However, the evaluation and accuracy of the quantification of disease severity are highly dependent on the method used, which has a direct correlation with the quality of the data generated for subsequent manipulation and analyses [27]. In this pathosystem, phytopathometric indices have been used to quantify the CABMV-induced symptoms in Passiflora species [25, 27,28,29]. However, inferring the reaction only based on leaf symptoms has not been sufficient to determine the resistance level, because wild species may not develop leaf symptoms and may therefore be classified as immune despite the presence of virus in the tissues. In this context, RT-qPCR is a sensitive technique that allows the viral titre in plants to be determined [30, 31]. Despite its relevance, there are few studies that have identified the resistance of wild passion fruit species by means of RT-qPCR [14, 32].
Thus, this study had the following objectives: i) to evaluate the progression of passion fruit woodiness disease symptoms in Passiflora spp., ii) to perform screening of Passiflora spp. genotypes for identification of CABMV resistance sources, aiming to select genotypes with high resistance for use in interspecific crosses, and iii) to validate the infection in plants with symptoms through reverse transcription polymerase chain reaction (RT-PCR), and in asymptomatic plants by RT-qPCR in cases where standard RT-PCR detection failed.
Materials and methods
Location and plant material
The study was carried out at the facilities of Embrapa Mandioca e Fruticultura, located in Cruz das Almas, Bahia, Brazil (12°40’39” S, 39°06’23” W, 226 m altitude). The region’s climate is transitional from Am to Aw type (tropical sub-humid to dry) according to the classification of Köppen and Geiger [33], with an annual average air temperature of 23.8 ºC. One hundred twenty-eight genotypes of Passiflora spp. were evaluated, using plants from the Passion Fruit Active Germplasm Bank of Embrapa Mandioca e Fruticultura, belonging to 12 species (Passiflora edulis Sims., P. cincinnata Mast., P. mucronata Lam., P. gibertii N.E Brown., P. alata Curtis., P. setacea DC., P. pohlii Mast., P. miersii Mast., P. bahiensis Klotzsch., P. malacophylla Mast., P. suberosa L., and Passiflora sp.) and three hybrids (a simple interspecific hybrid (F1), a third-generation interspecific hybrid from backcrossing – BC3 [(P. edulis × P. cincinnata) × P. edulis] and an intraspecific hybrid). Two other genotypes were used as controls, one susceptible (P. edulis, cv. BRS Gigante Amarelo) and the other resistant (P. cincinnata, BGP200) [25] (Table 1).
Biological assay and sampling
Approximately 80 seeds of each genotype were soaked in 2 mL of the growth regulator GA4+7 + N-(phenylmethyl)-aminopurine at concentration of 400 mg/L for 24 hours [34]. After this period, the seeds were sown in 162 cells of rigid polypropylene trays (50 mL vol.) filled with a combination of a mixture of coconut fiber (Gold Mix®) and a commercial substrate (Vivato®) in a ratio of 3:1 (v:v), with the addition of 50 g of slow-release fertilizer (Osmocote®) for each 10 L of substrate. After emergence (40 days after sowing), the 30 most uniform plants were selected for the assay. Subsequently, the plants were transferred to polypropylene tubes (100 cm3) and acclimatized in a greenhouse with a temperature of 28 ± 2 ºC and relative humidity (RH) of 75 ± 5%.
Plant inoculation and evaluation of disease symptoms
Leaves with severe symptoms induced by CABMV were collected from yellow passion fruit plants from the Passiflora experimental area and inoculated into plants maintained in a greenhouse as a source of inoculum. The mechanical inoculations with CABMV were performed when the plants had at least four expanded leaves, approximately 60 days after the emergence of the seedlings, as described by Gonçalves et al. [25]. To prevent escape, two inoculations per plant were performed with a four-day interval (Supplementary Fig. S1).
The intensity of symptoms in each leaf was scored using the diagrammatic scale scores proposed by Novaes and Rezende [35], ranging from 1 (without leaf symptoms) to 4 (severe leaf symptoms) (Supplementary Fig. S2). The evaluations started 12 days after the first inoculation (DAI) in all plants, using the first leaf of the fully developed apex, totaling five leaves per plant. Subsequent evaluations were done weekly until 40 DAI.
Evaluation of disease severity and prevalence
Symptom severity was measured by the McKinney disease severity index [36], where disease index (DI%) = (DS × L)/(TNL × HGS); where DS = degree of the determined scale for each leaf; L = number of leaves with each degree of symptoms (score); TNL = total number of evaluated leaves; and HGS = highest grade of the scale (maximum scale score). The prevalence of the disease in various genotypes was determined as the percentage of plants that exhibited typical disease symptoms.
Plants that did not show symptoms at 40 DAI were reinoculated to confirm their resistance to CABMV (Fig. 1). These asymptomatic plants (n = 7 to 25) of the 34 genotypes (Table 2) were pruned to 15 cm (Fig. 1c). Forty days after pruning (DAP), when the plants had at least four leaves, inoculations and leaf symptom evaluations were performed as described above.
Representative scheme of the steps for the detection of cowpea aphid-borne mosaic virus (CABMV) infection in symptomatic and asymptomatic inoculated plants. A) Inoculation (1 and 2) and evaluation of symptoms in passion fruit plants. B) Apical leaf tissue collection from inoculated plants with and without symptoms at 40 DAI. C) Separation of plants that did not show symptoms, pruning and reinoculation (inoculation 3 and 4) of plants at 40 DAP. D) Apical leaf tissue collection of symptomatic and asymptomatic reinoculated plants at 40 days after reinoculation (DARI) for detection of CABMV by reverse transcription polymerase chain reaction (RT-PCR) and reverse transcription quantitative polymerase chain reaction (RT-qPCR). Inoc., inoculation; DAI, days after inoculation.
Detection of CABMV by RT-PCR
At 40 DAI, apical leaf tissues were collected from symptomatic (S-IP) and asymptomatic (A-IP) inoculated plants, symptomatic reinoculated plants (S-RIP), asymptomatic reinoculated plants (A-RIP), and non-inoculated plants (NIP – negative controls) of the 12 Passiflora spp. and hybrids. RNA extractions were performed from pools containing five apical leaves, representative of five plants of each of the sets (S-IP, A-IP, S-RIP, A-RIP and NIP) (Fig. 1a-d), following the protocol of Ferreira et al. [37]. The treatments were performed with 10 µL of total RNA (2 µg) and 1.5 µL of DNase (2 U/μL) according to the manufacturer’s recommendations (Ambion), with the RNA concentration adjusted to 100 ng/µL.
cDNA was synthesized from 3.0 μL of total RNA, using an with M-MLV Reverse Transcriptase Kit (Invitrogen). PCR reactions were performed with 3.0 μL of cDNA (30 ng/μL) and 10 μM primers for amplification of part of the CABMV cylindrical inclusion gene (CI) to yield a product of 1311 bp [38] (Supplementary Fig. S3). The CABMV CI gene amplification program and procedures for visualization of the PCR products were described previously by Gonçalves et al. [25].
Reverse transcription quantitative polymerase chain reaction
Amplification efficiency was determined from a tenfold serial dilution series from 200 to 0.02 ng/μL. The primer pair qCABMV07_For (5’ CTGGTAGAGTGCTTCTCAATTTGG 3’) and qCABMV07_Rev (5’ CTCTCCCTTGATGGCCTCAA 3’), was used to amplify part of the CABMV coat protein (CP) gene to produce a product of 121 bp [39] (Supplementary Fig. S3). The amplification efficiency was calculated automatically using 7500 Fast v2.0.6 software, using the slope obtained by linear regression according to the following formula: % Efficiency = [10(-1/slope) -1] × 100 [40, 41]. A standard curve of purified CABMV RT-PCR product was generated using a tenfold serial dilution from 1 to 0.0001 ng/μL. The number of molecules of the CABMV (copies/μL) at each dilution point was determined using the following formula: Copy number = (Sample concentration [ng/µL] × 6.022 × 1023)/(Fragment size [bp] × 1 × 109 × 660 g/mol) [42].
The qPCR assays were performed using a 7500 Fast Real-Time PCR System (Applied Biosystems) programmed for analysis of the “Quantitation - Standard Curve” type. The reactions were performed in a MicroAmp™ Fast Optical 96-Well Reaction Plate (0.1 mL), with 10 µL reactions containing 1.0 μL of cDNA (generated from 100 ng/μL of RNA), 0.1 μM each primer, GoTaq® qPCR Master Mix (Promega), and CXR Reference Dye. A cDNA sample derived from a plant that was not inoculated with CABMV and a control without template were included as negative controls. All reactions were conducted in technical triplicate. The cycling conditions for the CABMV coat protein gene were 50 ºC for 2 min, 95 ºC for 10 min, 40 cycles at 95 ºC for 15 s, and 60 ºC for 1 min.
Design and data analysis
The experimental design used was completely randomized, considering each of the 25 plants inoculated one repetition. Another five plants that were not inoculated with CABMV were used as controls. Some plants of P. edulis that were inoculated with CABMV and remained asymptomatic (n = 1 to 4) were not included in the severity analysis because P. edulis is generally susceptible to CABMV [25], and they might have escaped infection.
The mean disease index at each time point (12, 19, 26, 33, and 40 DAI) was plotted on a logarithmic scale to examine the evolution of disease in the 12 Passiflora species and three hybrids (inter- and intraspecific). To calculate the rate of disease progression in the leaves, severity values (DI%) were used at the five evaluation time points only for species with symptoms. The original severity or linearized data were tested for some models and adjusted as described previously [43]. Using the best adjustment, the disease progression rate was estimated (r) and determined by the angular coefficient (\({\theta }_{1}\) and \({\theta }_{2}\)) of the regression equation (R2) [43].
The average DI (%) estimates at 40 DAI were compared using the Scott-Knott test (p ≤ 0.05). The genotypes were classified as [19] resistant (R; DI 0.0 – 15.9%), moderately resistant (MR; DI 16.0–31.9%), susceptible (S; DI 32.0–50.9%) or highly susceptible (HS; DI ≥ 51.0%). The analyses were performed in R, using the ‘ExpDes.pt’ package [44]. Genotypes were grouped based on the Gower index [45] and the unweighted pair group method with arithmetic mean (UPGMA). A dissimilarity matrix was made using the Genes program [46], and from the matrix, MEGA7.0 software was used to generate a dendrogram [47]. The viral titres of asymptomatic inoculated plants were determined by measuring the concentration of the viral cDNA (ng/µL) a standard curve [42].
Results
Temporal progression of PWD caused by CABMV in Passiflora species
The progression of the disease was scored as null for the species belonging to group 1 (P. suberosa, P. setacea, P. pohlii, P. malacophylla, P. bahiensis, P. gibertii, and P. miersii) and stable for those in group 2 (P. mucronata and P. cincinnata). The progression of the disease was faster in the species of group 3 (P. alata, P. edulis, the interspecific hybrid BC3, the interspecific hybrid F1, and Passiflora sp.) and of group 4 (intraspecific hybrid) (Fig. 2).
Logarithmic regression of the disease index in 12 species of Passiflora spp. and three hybrids (inter- and intraspecific) on different days after inoculation with cowpea aphid-borne mosaic virus. Group 1 (resistant – R: P. suberosa, P. setacea, P. pohlii, P. malacophylla, P. bahiensis, P. gibertii, and P. miersii); group 2 (moderately resistant – MR: P. mucronata and P. cincinnata); group 3 (susceptible – S: P. alata, interspecific hybrids BC3 and F1, P. edulis, and Passiflora sp.); group 4 (highly susceptible – HS: intraspecific hybrid). The values in parentheses represent the range of the disease index for the species in each group.
Based on the values of \({\theta }_{1}\) or \({\theta }_{2}\) between pairs of species (Supplementary Table S1), it was possible to identify significant differences, with disease progression rates being slower in P. cincinnata than in P. alata and Passiflora sp. The species P. gibertii had a slower progression rate than the intraspecific hybrid, Passiflora sp., and P. alata, but a faster rate than P. miersii and P. mucronata. On the other hand, P. miersii and P. mucronata had slower rates than the intraspecific hybrids, P. alata, P. edulis, and Passiflora sp., while the interspecific hybrid (F1) had a faster rate than P. mucronata, while P. miersii and P. mucronata had faster rates than P. cincinnata (Supplementary Table S1). Other comparisons did not reveal significant differences (Supplementary Table S1).
Classification of Passiflora spp. and genotypes based on disease severity
The disease index (DI) ranged from 0.0 to 70.80%, with 26 genotypes (20.3%) classified as resistant (R; DI 0.0 to 15.75%). Within this same class, the genotypes that should be highlighted are BGP152 (P. suberosa), BGP170 (P. malacophylla), BGP434, BGP244, BRS Pérola do Cerrado (P. setacea), BGP454 (P. pohlii), and BGP477 (P. bahiensis), which did not exhibit symptoms of PWD (DI 0.0%). Another 12 genotypes (9.4%) were moderately resistant (MR; DI 16.3 to 31.1%), 42 (32.8%) were susceptible (S; DI 33.3 to 50.9%), and 48 (37.5%) were highly susceptible (HS; DI 51.2 to 70.8%) (Figs. 3a-b and Supplementary Table S2).
Cluster and severity of passion fruit woodiness disease (PWD) caused by cowpea aphid-borne mosaic virus (CABMV) in 128 genotypes of Passiflora spp. A) Mean values and the number of genotypes in the classes resistant (R), moderately resistant (MR), susceptible (S), and highly susceptible (HS). B) Mean values of disease severity and their distribution in the classes R, MR, S and HS. C) Average severity of PWD in 12 species of Passiflora spp., an intraspecific hybrid, a simple interspecific hybrid (F1), and an interspecific hybrid of the third backcross generation – BC3 [(P. edulis × P. cincinnata) × P. edulis]. D) Prevalence of the disease in the species. 1,2P. cincinnata genotype (BGP200) and yellow passion fruit cultivar (P. edulis, cv. BRS Gigante Amarelo) used as resistant and susceptible controls during the evaluations of leaf symptoms induced by CABMV.
Regarding the P. edulis genotypes, only BGP124 was considered moderately resistant, with a DI of 31.11%. The other 53 genotypes (41.4%) showed some degree of susceptibility. Of these, 27 (21.1%) were susceptible and 26 (20.3%) were highly susceptible. All genotypes belonging to the intra-specific hybrids, interspecific hybrids (F1), and interspecific hybrids from the third generation of backcrossing (BC3) were classified as susceptible or highly susceptible (Fig. 3 and Supplementary Table S2). The genotypes used as controls for resistance (BGP200) and susceptibility (cv. BRS Gigante Amarelo) showed severity within the expected level, with mean DI values of 15.30% and 62.20%, respectively.
Of the species evaluated, seven (46.67%) were classified as resistant (P. bahiensis, P. malacophylla, P. pohlii, P. setacea, P. suberosa, P. gibertii, and P. miersii), with a DI of 0.0 to 14.80%; two (13.33%) were classified as moderately resistant (P. cincinnata and P. mucronata), with a DI of 16.10 to 18.70%; three (20%) (Passiflora sp., P. edulis and interspecific hybrids of the third backcross generation [BC3]) were classified as susceptible to CABMV, with DI values ranging from 37.60 to 50.40%; and three (20%) were considered highly susceptible to CABMV, with a mean DI of 51.90 to 64.90% (hybrids [F1], P. alata, and intraspecific hybrids) (Fig. 3c).
The average severity of 34 reinoculated genotypes ranged from 0.0 to 60.74% (Table 2). Some genotypes demonstrated typical disease symptoms, specifically those belonging to the species P. gibertii, P. cincinnata, P. mucronata, P. miersii and the interspecific hybrid (OTH-137), the latter with 100% prevalence in reinoculated plants (Table 2). Among the genotypes evaluated, 55.88% (n = 19) were considered resistant, with DI 0.0 to 15.83%. Within this same group, the genotypes BGP152 (P. suberosa), BGP170 (P. malacophylla), BRS Pérola do Cerrado, BGP434, BGP244 (P. setacea), BGP454 (P. pohlii), and BGP477 (P. bahiensis) remained asymptomatic even after reinoculation, maintaining their classification as resistant. Another 13 genotypes (38.24%) were classified as moderately resistant, with a DI of 17.50 to 30.29%, and two genotypes (BGP275 and OTH-137) were classified as susceptible (DI: 36.74%) and highly susceptible (DI: 60.74%), respectively (Table 2).
Detection of CABMV
Viral infection in symptomatic inoculated plants was confirmed by RT-PCR with amplification of a 1311-bp fragment of the CABMV CI gene (Fig. 4a). In the asymptomatic inoculated and negative control plants, the systemic replication of CABMV was not confirmed (Fig. 4b and c). Asymptomatic plants were subsequently tested by RT-qPCR to confirm the infection (Fig. 4b). The amplifications using the primer qCABMV07 were very uniform (slope = -3.53, determination coefficient [R2] = 0.997, and efficiency of 91.96%). The standard curve for the purified CABMV RT-PCR product, the R2 value was 0.998, the slope was -3.33, and the qPCR efficiency was 99.37% (Supplementary Fig. S4). The amount of the CABMV CP gene in the serial dilutions of the standard curve ranged from 7.54 × 109 to 7.54 × 105 viral copies per microliter (Supplementary Table S3). These measurements were used to determine the viral titre in the evaluated passion fruit species.
Products of amplification of the cylindrical inclusion gene (1311 bp) of cowpea aphid-borne mosaic virus by reverse transcription polymerase chain reaction analyzed by electrophoresis in a 1% agarose gel. A) Pool of leaf samples of symptomatic inoculated plants (S-IP) at 40 days after inoculation (DAI). B) Pool of leaf samples of asymptomatic inoculated plants (A-IP). C) Pool of leaf samples of plants not inoculated with CABMV (NIP, negative controls). PC: positive control (plants of P. edulis with severe PWD symptoms – cDNA 200 ng/µL). M, 1 kb DNA Marker Ladder (Invitrogen). P. ed, Passiflora edulis; Intra H., intraspecific hybrid; Inter H., interspecific hybrid; P. ala, P. alata; P. mie, P. miersii; P. muc, P. mucronata; P. cin, P. cincinnata; P. gib, P. gibertii; P. sp, Passiflora sp.; P. set, P. setacea; P. poh, P. pohlii; P. mala, P. malacophylla; P. sub, P. suberosa; P. bah, P. bahiensis
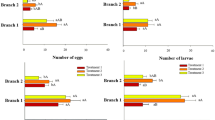
The CABMV titre in asymptomatic species ranged from 1 × 100 to 3.66 × 107 copies/µL (Fig. 5a). There was little variation in viral titre in the interspecific hybrid (OTH-137) and the species P. gibertii, P. cincinnata, and P. miersii, with 3.66 × 107, 3.24 × 107, 3.05 × 107 and 2.55 × 107 copies/µL, respectively. The species P. mucronata, P. setacea, and P. malacophylla had lower virus titres than the above-mentioned species, with 2.70 × 106, 3.07 × 105, and 9.99 × 104 copies/µL, respectively (Fig. 5a). In P. pohlii, P. suberosa, and P. bahiensis, no CABMV CP amplicons were detected, demonstrating there was no infection after two inoculation attempts (Fig. 5b). The specificity of the qPCR amplifications was confirmed by agarose gel electrophoresis, with single amplicons of the expected size of 121 bp obtained for the interspecific hybrid (OTH-137) and the species P. cincinnata, P. gibertii, P. miersii, P. mucronata, P. setacea, and P. malacophylla (Fig. 5b).
Detection and quantification of CABMV in inoculated but asymptomatic plants of Passiflora spp. by reverse transcription quantitative polymerase chain reaction (RT-qPCR). A) Number of CABMV molecules (values above the columns – copies/μL) in inoculated asymptomatic plants. B) 2% agarose gel showing qPCR products from the amplification of a 121-bp fragment of the coat protein gene of CABMV. M, 100 bp DNA Marker Ladder (Ludwig). NTC (non-template control), without cDNA as template; NC (negative control), sample of P. edulis not inoculated with CABMV; Inter H, interspecific hybrid (OTH-137); P. gib, P. gibertii; P. cin, P. cincinnata; P. mie, P. miersii; P. muc, P. mucronata; P. set, P. setacea; P. mal, P. malacophylla; P. poh, P. pohlii; P. sub, P. suberosa; P. bah, P. bahiensis
Viral infection in plants that showed symptoms after the third and fourth reinoculation was confirmed by RT-PCR (Fig. 6a). However, some plants of P. cincinnata, P. gibertii, P. miersii, P. mucronata, P. setacea, P. malacophylla, P. suberosa, P. pohlii, and P. bahiensis did not exhibit PWD symptoms, so they were analyzed further by RT-qPCR. For the interspecific hybrid OTH-137, all plants had symptoms (Table 2). CP amplicons were detected by RT-qPCR in reinoculated asymptomatic plants (Fig. 1d) of P. cincinnata, P. gibertii, P. miersii, P. mucronata, P. setacea, P. malacophylla, and P. suberosa, but not in P. pohlii and P. bahiensis (Fig. 6b-c). The viral titre ranged from 1 × 100 to 4.20 × 107 copies/µL (Fig. 6b). The variation in viral titre among the species was low, with 4.20 × 107 copies/µL in P. cincinnata, 4.16 × 107 in P. gibertii, 4.09 × 107 in P. miersii, and 3.65 × 107 in P. mucronata. In turn, the viral titre in P. setacea, P. malacophylla, and P. suberosa was lower than in the preceding group, with 8.00 × 106, 4.85 × 105 and 1.79 × 105 copies/µL, respectively (Fig. 6b).
Detection and quantification of CABMV in symptomatic and asymptomatic reinoculated plants by reverse transcription polymerase chain reaction (RT-PCR) and reverse transcription quantitative polymerase chain reaction (RT-qPCR) at 40 days after reinoculation (DARI). A) 1% agarose gel showing RT-PCR amplification products of the cylindrical inclusion gene of CABMV of 1311 bp in symptomatic reinoculated plants (S-RIP). M, 1 kb DNA Marker Ladder (Invitrogen); PC, positive control (plants of P. edulis with severe PWD symptoms – cDNA 200 ng/µL). B) Number of the CABMV molecules (values above the columns – copies/μL) in asymptomatic reinoculated plants of Passiflora spp. C) 2% agarose gel showing qPCR products from the amplification of a 121-bp fragment of the coat protein gene of CABMV. M, 100 bp DNA Marker Ladder (Ludwig); NTC (non-template control), without template cDNA; NC (negative control), sample of P. edulis not inoculated with CABMV; Inter H., interspecific hybrid (OTH-137); P. mie, P. miersii; P. muc, P. mucronata; P. cin, P. cincinnata; P. gib, P. gibertii; P. set, P. setacea; P. mal, P. malacophylla; P. sub, P. suberosa; P. poh, P. pohlii; P. bah, P. bahiensis
Discussion
The differences in disease progression in the passion fruit species tested demonstrated the high degree of variability in the resistance or susceptibility of Passiflora spp. to CABMV infection (Fig. 2 and Supplementary Table S1). In the initial evaluation, plants of most of the species tested (n = 9) exhibited typical symptoms of PWD by 12 DAI, and plants of Passiflora sp. (BGP482) showed symptoms at 19 DAI (Fig. 2). Plants of P. suberosa, P. setacea, P. pohlii, P. malacophylla, and P. bahiensis did not exhibit symptoms before the end of the experiment (40 DAI) (Fig. 2). The incubation time of CABMV for most of the Passiflora spp. is not known with accuracy, and the initial expression of symptoms is dependent on the age of the plant and the genotype or isolate used, and it is affected directly by environmental conditions and the nutrition of the plants [16, 29].
Species classified as resistant and moderately resistant are uncultivated and naturally carry resistance alleles [48]. This study is a pioneer in reporting the probable immunity to CABMV of P. bahiensis and P. pohlii and the moderate resistance of P. miersii, increasing the number of wild Passiflora species evaluated for CABMV resistance. Studies have reported resistance to CABMV in genotypes belonging to P. setacea [23, 24], P. cincinnata [23, 25], P. gibertii [25], and P. suberosa [22].
Species with no symptoms or low disease severity can be used in breeding programs of Passiflora for interspecific crosses with commercial species. However, interspecific crosses may not succeed if the species differs in their number of chromosomes or belong to a different subgenus [49, 50], as has been reported in the case of P. suberosa [49] and P. pohlii [51], which can result in genetic barriers to crossings or a lack of synchronization in flowering [52]. In some cases, the interspecific compatibility barriers are relatively weak, so successful hybridization can be achieved [53]. The cytogenetic aspects involved in the crossing of P. edulis and P. cincinnata demonstrate the possibility of obtaining hybrids and thus transferring resistance alleles or other traits of the wild species [54]. Studies using P. setacea and P. cincinnata (both 2n = 18) as donors of CABMV resistance alleles for P. edulis (2n = 18) have also been successful [18,19,20,21, 26, 55]. P. malacophylla and P. bahiensis can potentially be used in breeding programs because they are resistant to CABMV and same chromosome number of P. edulis, opening the possibility of obtaining resistant commercial hybrids. However, complementary studies should be performed with these two species to confirm that viable hybrids can be obtained.
There is a relationship between prevalence and severity of disease for most of the evaluated genotypes. However, P. mucronata (BGP479) and P. edulis (BGP124) plants with disease prevalence of 100% showed only mild mosaic symptoms (score 2) and were classified as moderately resistant (Supplementary Table S2). This indicates that the prevalence is not always linked to severity. The genotypes of P. edulis showed susceptibility to CABMV (Fig. 3 and Supplementary Table S2) except for the BGP124 genotype, which was moderately resistant. Results in the literature have demonstrated different resistance levels among P. edulis genotypes from resistant to highly susceptible [23, 25, 56, 57]. This indicates intraspecific genetic variability of the resistance to CABMV. For this reason, P. edulis genotypes in active gene banks should be tested to identify those with low disease severity, thereby reducing the time required to obtain resistant cultivars [18, 25, 26].
The susceptibility observed in plants of the six intraspecific hybrids of P. edulis is directly related to the selection of parents, taking into consideration only agronomic attributes of vigor and production [58, 59]. The interspecific hybrids of the third backcross generation – BC3 [(P. edulis × P. cincinnata) × P. edulis], despite having contrasting genitors – BGP330 (susceptible) and BGP077 (resistant) [25] – did not show resistance to CABMV. This is probably due to the small number of BC3 progeny (n = 21) evaluated. For gains in resistance to CABMV, it is necessary to evaluate a much larger number of progenies, since 93.75% of the genome involved in the backcrossing belongs to the susceptible recurrent genitor (P. edulis), leading to resistance losses in the progeny. Indeed, [60] evaluated a larger number of progenies and identified CABMV resistant plants in BC3. Genotypes of the fourth and fifth backcross generation involving P. edulis × P. setacea are not resistant to CABMV due to the loss of resistance as new backcrosses are performed [61], possibly due to the polygenic heritage of the trait [18,19,20,21]. The genetic heritage for resistance to CABMV of most Passiflora species is still unknown and is therefore an open field for research in breeding programs.
The variation in the resistance to CABMV among the evaluated genotypes is associated with the genetic variability of the Passiflora species, since they are self-incompatible [52, 62, 63]. Moreover, different studies have attributed differences in the passion fruit response to CABMV to the use of different viral isolates [22, 56], the latency period [16], the individual resistance levels of genotypes [25], genetic and environmental factors (such as temperature and relative humidity) [20, 26, 64], and differences in nutritional condition and age among plants [65]. Alone or together, these factors can influence the virulence of the pathogen and the manifestation of the disease symptoms. In this study, many plants without disease symptoms were observed, especially those of wild species. However, some of these factors may not be the cause, since the evaluated genotypes were the same age and the environmental and nutritional conditions were the same.
The observation that some genotypes showed slightly lower severity after the first two inoculations but were still classified as resistant can be attributable to pre-immunization of these plants, since the same viral isolate was used in two reinoculations (third and fourth inoculation) (Fig. 1c). Crotalaria juncea plants infected with two mild strains of passion fruit woodiness virus (PWV; currently recognized as CABMV [7]) were shown previously to be protected against infection by a severe strain and/or expression of symptoms [66]. This may be related to the uniform distribution of viruses in leaf tissues, limiting the availability of infection sites for the severe strain and thus inhibiting replication and establishment of systemic infection by the severe strain [66]. The competition for replication sites between mild and severe strains of papaya ringspot virus, cucumber green mottle mosaic virus, and citrus tristeza virus has also been demonstrated [68,69,69]. The selection of passion fruit plants that allow a higher rate of multiplication of mild strains or the selection of other mild strains with greater invasive power can enable pre-immunization to control viruses causing PWD under field conditions [35]. In the specific case of the genotypes BGP275 (P. cincinnata) and OTH-137 (interspecific hybrid), after reinoculation, the plants showed increased disease severity, leading to their reclassification as susceptible and highly susceptible, respectively (Table 2). This indicates that the initial tolerance of these genotypes was overcome as the plants were challenged with a high dose of virus after the third and fourth inoculation.
Researchers have attributed the occurrence of asymptomatic plants to escape from inoculation [20, 38]. However, in this study, this explanation is unlikely, because four inoculations were performed, similar to the procedure reported by Correa et al. [14]. The occurrence of asymptomatic plants can be related to the individual resistance features of genotypes, since the passion fruit resistance to CABMV has been shown to be genotype-dependent [29] or due to a viral RNA silencing mechanism, as suggested by Correa et al. [14] and verified in another study [70, 71]. On the other hand, the manifestation of symptoms after reinoculation of initially asymptomatic plants may be due to viral suppressors of gene silencing [72], which suppress host resistance and favor viral replication and movement to the apical leaves of the plants. However, this hypothesis needs to be validated in Passiflora spp..
Despite not having sequenced the amplicons, the unique DNA bands and expected size of 1311 bp in all species with PWD symptoms (Fig. 4a) indicated that the virus used in the artificial inoculations belongs to the species Cowpea aphid-borne mosaic virus. The lack of detection of CABMV in inoculated but asymptomatic plants (Fig. 4b) may be related to the low sensitivity of the RT-PCR when the viral titre in the leaf is low [73, 74].
The small variation in viral titre among asymptomatic wild passion fruit species (Fig. 5a and Fig. 6b) indicated high resistance to CABMV is capable of restricting virus replication to basal levels. In fact, asymptomatic plants may not be free from viral infection [75]. In resistant species, the viruses accumulate to some extent without causing significant negative effects in their hosts. Although a viral titre is maintained, plant growth and fruit yield are minimally affected, and the symptoms of the disease are absent or mild [76].
The slight decrease in viral titre when asymptomatic plants were reinoculated with CABMV (Fig. 6b) may have been due to a previously active molecular signaling system conferring faster recognition and response against viral infection, probably by the effect of the systemic acquired resistance caused by the first inoculation or viral RNA silencing [77, 78]. In the specific case of P. suberosa, this species was reported to be immune to CABMV [22]. In our study, viral infection in this species was confirmed after the third and fourth inoculations, indicating that the high inoculum pressure resulted in infection. On the other hand, plants of P. bahiensis and P. pohlii were not infected by CABMV, even under these conditions, indicating probable immunity. However, it will be necessary to test this immunity with inoculation of CABMV in protoplasts to look for viral replication at the cellular level, as observed in citrus protoplasts infected with citrus tristeza virus [79]. Furthermore it is prudent to test the effective immunity of these species in the field, since the environment is more heterogeneous and the natural infection by aphid vectors is more specialized [12, 80, 81].
A study by Carvalho et al. [82] demonstrated that proteins linked to the regulation of proteasomes, heat shock proteins, and ubiquitination are involved in defense and resistance signaling of P. setacea to CABMV. It is possible that these proteins were involved in the signaling and tolerance of CABMV in the wild species evaluated in this study, but many other proteins can be involved, since they are different species and the response patterns to the virus can be distinct. Defense responses can involve numerous signaling pathways, culminating in the limitation of viral replication. These responses are varied, depending on the species, phenological phase, and environmental conditions. Collectively, the mechanisms mentioned above can be involved in the tolerance of wild species to CABMV, and further studies are needed to determine whether this is the case in Passiflora spp..
Conclusions
Mean PWD progression rates and disease severity in symptomatic plants were lower in P. cincinnata, P. gibertii, P. miersii, and P. mucronata than in P. edulis and P. alata, inter- and intraspecific hybrids, and Passiflora sp. The accessions belonging to P. suberosa, P. malacophylla, P. setacea, P. pohlii, and P. bahiensis did not show visual symptoms of the disease after mechanical inoculation. Some asymptomatic plants (P. cincinnata, P. gibertii, P. miersii, and P. mucronata), after additional inoculations, exhibited lower mean disease severity in relation to the symptoms of the two initial inoculations, which may indicate a control mechanism such as pre-immunization. The absence of visible symptoms in some plants or accessions does not indicate immunity, because asymptomatic plants can still be infected by CABMV, as demonstrated by RT-qPCR analysis. However, P. pohlii and P. bahiensis, even after four inoculations, remained asymptomatic and free of the virus, suggesting that they are probably immune to CABMV.
References
Bernacci LC, Soares-Scott MD, Junqueira NTV, Passos IRDS, Meletti LMM (2008) Passiflora edulis Sims: the correct taxonomic way to cite the yellow passion fruit (and of others colors). Rev Bras Frutic 30:566–576. https://doi.org/10.1590/S0100-29452008000200053
Coelho EM, Azevêdo LC, Umza-Guez MA (2016) Fruto do maracujá: Importância econômica e industrial, produção, subprodutos e prospecção tecnológica. Cad Prospec 9:347. https://doi.org/10.9771/S.CPROSP.2016.009.037
IBGE (Instituto Brasileiro de Geografia e Estatística) (2020) Banco de dados agregados. Sistema IBGE de Recuperação Automática – SIDRA. Disponível em: http://www.ibge.gov.br
Melo JRF, Figueira AR, Moreira CN, Oliveira AC (2015) Recent characterization of cowpea aphid-borne mosaic virus (CABMV) in Bahia State, Brazil, suggests potential regional isolation. Afr J Biotechnol 14:735–744. https://doi.org/10.5897/AJB2015.14409
Rodrigues LK, Silva LA, Garcêz RM, Chaves AL, Duarte LM, Giampani JS, Eiras M (2015) Phylogeny and recombination analysis of Brazilian yellow passion fruit isolates of Cowpea aphid-borne mosaic virus: origin and relationship with hosts. Australasian Plant Pathol 44:31–41. https://doi.org/10.1007/s13313-014-0308-5
Costa AP, Nogueira I, Peixoto JR, Blum LEB (2020) Screening of sour passion fruit for reaction to bacterial spot and passion fruit woodiness disease. J Agric Sci 12(2):130–137. https://doi.org/10.5539/jas.v12n2p130
Preisigke SC, Viana AP, Santos EA, Santos PR, Santos VO, Ambrósio M, Silva FA, Walter FHB (2020) Selection strategies in a segregating passion fruit population aided by classic and molecular techniques. Bragantia 79:47–61. https://doi.org/10.1590/1678-4499.20190291
Wylie SJ, Jones MG (2011) The complete genome sequence of a passion fruit woodiness virus isolate from Australia determined using deep sequencing, and its relationship to other potyviruses. Arch Virol 156:479–482. https://doi.org/10.1007/s00705-010-0845-3
Wylie SJ, Adams M, Chalam C, Kreuze J, López-Moya JJ, Ohshima K, Zerbini FM (2017) ICTV virus taxonomy profile: Potyviridae. J Gen Virol 98(3):352. https://doi.org/10.1099/jgv.0.000740
Fischer IH, Rezende JA (2008) Diseases of passion flower (Passiflora spp.). Pest Tech 2:1–19
Bragard C, Caciagli P, Lemaire O, Lopez-Moya JJ, MacFarlane S, Peters D, Torrance L (2013) Status and prospects of plant virus control through interference with vector transmission. Annu Rev Phytopathol 51:177–201. https://doi.org/10.1146/annurev-phyto-082712-102346
Dáder B, Then C, Berthelot E, Ducousso M, Ng JC, Drucker M (2017) Insect transmission of plant viruses: multilayered interactions optimize viral propagation. Insect Sci 24:929–946. https://doi.org/10.1111/1744-7917.12470
Nascimento AVS, Santana EN, Braz ASK, Alfenas PF, Pio-Ribeiro G, Andrade GP, Zerbini FM (2006) Cowpea aphid-borne mosaic virus (CABMV) is widespread in passionfruit in Brazil and causes passionfruit woodiness disease. Arch Virol 151:1797–1809. https://doi.org/10.1007/s00705-006-0755-6
Correa MF, Pinto APC, Rezende JAM, Harakava R, Mendes BMJ (2015) Genetic transformation of sweet passion fruit (Passiflora alata) and reactions of the transgenic plants to Cowpea aphid-borne mosaic virus. Eur J Plant Pathol 143:813–821. https://doi.org/10.1007/s10658-015-0733-5
Rodrigues LK, Chaves ALR, Damatto ER, Eiras M (2016) Epidemiological aspects of the transmission and management of Cowpea aphid-borne mosaic virus in a passion fruit orchard. J Plant Pathol 98:531–539. https://doi.org/10.4454/JPP.V98I3.037
Spadotti DMDA, Favara GM, Novaes QS, Mello APOA, Freitas DMS, Edwards Molina JP, Rezende JAM (2019) Long lasting systematic roguing for effective management of CABMV in passion flower orchards through maintenance of separated plants. Plant Pathol 68:1259–1267. https://doi.org/10.1111/ppa.13054
Silva FHL, Viana AP, Santos EA, Freitas JCO, Rodrigues DL, Júnior ATA (2017) Prediction of genetic gains by selection indexes and REML/BLUP methodology in a population of sour passion fruit under recurrent selection. Acta Sci Agron 39:183–190. https://doi.org/10.4025/actasciagron.v39i2.32554
Santos EA, Viana AP, Walter FHB, Freitas JCO, Ramos HCC, Boechat MSB (2019) First report of a genetic map and evidence of QTL for resistance to CABMV in a segregating population of Passiflora. Eur J Plant Pathol 155:903–915. https://doi.org/10.1007/s10658-019-01822-y
Jesus ON, Santos IS, Lima LKS, Soares TL, Oliveira EJ (2021) Field assessment of a second generation backcross (BC1 × Passiflora edulis) of passion fruit for agronomic performance and resistance to CABMV. Plant Breed. https://doi.org/10.1111/pbr.12888
Freitas JCO, Viana AP, Santos EA, Silva FH, Paiva CL, Rodrigues R, Eiras M (2015) Genetic basis of the resistance of a passion fruit segregant population to Cowpea aphid-borne mosaic virus (CABMV). Trop Plant Pathol 40:291–297. https://doi.org/10.1007/s40858-015-0048-2
Freitas JCO, Viana AP, Santos EA, Paiva CL, Silva FHL, Souza MM (2016) Sour passion fruit breeding: Strategy applied to individual selection in segregating population of Passiflora resistant to Cowpea aphid-borne mosaic virus (CABMV). Sci Hortic 211:241–247. https://doi.org/10.1016/j.scienta.2016.09.002
Maciel SC, Nakano DH, Rezende JAM, Vieira MLC (2009) Screening of Passiflora species for reaction to Cowpea aphid-borne mosaic virus reveals an immune wild species. Sci Agric 66:414–418. https://doi.org/10.1590/S0103-90162009000300018
Oliveira EJ, Soares TL, Barbosa CJ, Santos-Filho HP, Jesus ON (2013) Disease severity from passion fruit to identify sources of resistance in field conditions. Rev Bras Frutic 35:485–492. https://doi.org/10.1590/S0100-29452013000200018
Sacoman NN, Viana AP, Carvalho VS, Santos EA, Rodrigues R (2018) Resistance to Cowpea aphid-borne mosaic virus in in vitro germinated genotypes of Passiflora setacea. Rev Bras Frut 40:1–10. https://doi.org/10.1590/0100-29452017607
Gonçalves ZS, Lima LKS, Soares TL, Abreu EFM, Barbosa CJ, Cerqueira-Silva CBM, Jesus ON, Oliveira EJ (2018) Identification of Passiflora spp. genotypes resistant to Cowpea aphid-borne mosaic virus and leaf anatomical response under controlled conditions. Sci Hortic 231:166–178. https://doi.org/10.1016/j.scienta.2017.12.008
Santos EA, Viana AP, Freitas JCO, Silva FHL, Rodrigues R, Eiras M (2015) Resistance to Cowpea aphid-borne mosaic virus in species and hybrids of Passiflora: advances for the control of the passion fruit woodiness disease in Brazil. Eur J Plant Pathol 143:85–98. https://doi.org/10.1007/s10658-015-0667-y
Porto ACM, Santos ML, Oliveira AC (2017) Quality of phytopathometric variables generated from a ranking scale for the CABMV-passionfruit pathosystem. Rev Agro@mbiente 12:58–67. https://doi.org/10.18227/1982-8470ragro.v12i1.4247
Cerqueira-Silva CBM, Melo JRF, Corrêa RX, Oliveira AC (2012) Selection of pathometric variables to assess resistance and infectivity in the passion fruit woodiness pathosystem. Eur J Plant Pathol 134:489–495. https://doi.org/10.1007/s10658-012-0030-5
Gonçalves ZS, Jesus ON, Cerqueira-Silva CBM, Diniz RP, Soares TL, Oliveira EJ (2017) Methodological approaches to assess passion fruit resistance (Passiflora spp.) to passionfruit woodiness disease. Biosci J. https://doi.org/10.14393/BJ-v33n6a2017-36619
Saponari M, Loconsole G, Liao HH, Jiang B, Savino V, Yokomi RK (2013) Validation of high-throughput real time polymerase chain reaction assays for simultaneous detection of invasive citrus pathogens. J Virol Methods 193:478–486. https://doi.org/10.1016/j.jviromet.2013.07.002
Osman F, Hodzic E, Kwon SJ, Wang J, Vidalakis G (2015) Development and validation of a multiplex reverse transcription quantitative PCR (RT-qPCR) assay for the rapid detection of Citrus tristeza virus, Citrus psorosis virus, and Citrus leaf blotch virus. J Virol Methods 220:64–75. https://doi.org/10.1016/j.jviromet.2015.04.013
Riska NM, Iwai H (2020) Effects of coinfection with East Asian Passiflora virus and East Asian Passiflora distortion virus on Passiflora foetida. J Gen Plant Pathol 86:211–218. https://doi.org/10.1007/s10327-020-00913-7
Köppen W, Geiger R (1928) Klimate der Erde. Verlag Justus Perthes, Gotha. Wall-map 150cmx200cm
Moura RS, Filho MAC, Gheyi HR, Jesus ON, Lima LKS, Junghans TG (2018) Overcoming dormancy in stored and recently harvested Passiflora cincinnata Mast. seeds. Biosci J 34:1158–1166. https://doi.org/10.14393/BJ-v34n5a2018-39451
Novaes QS, Rezende JAM (2003) Selected mild strains of Passion fruit woodiness virus (PWV) fail to protect preimmunized vines in Brazil. Sci Agric 60:699–708. https://doi.org/10.1590/S0103-90162003000400014
Mckinney HH (1923) Influence of soil temperature and moisture on infection of wheat seedlings by Helminthosporium sativum. J Agric Res 26:195–218
Ferreira CF, Gutierrez DL, Kreuze JF, Iskra-Caruana ML, Chabannes M, Barbosa ACO, Jesus ON (2019) Rapid plant DNA and RNA extraction protocol using a bench drill. Genet Mol Res 18:1–8. https://doi.org/10.4238/gmr18394
Fontenele R, Abreu R, Lamas N, Alves-Freitas D, Vidal A, Poppiel R, Varsani A (2018) Passion fruit chlorotic mottle virus: molecular characterization of a new divergent geminivirus in Brazil. Viruses. https://doi.org/10.3390/v10040169
Freitas MS (2013) Patossistema Cowpea aphid-borne mosaic virus (CABMV)/maracujazeiro: titulação ‘real time’ do patógeno, sistema de classificação de reação genética diferencial de genótipos do hospedeiro e indução de resistência genética. Dissertação, Universidade Estadual do Sudoeste da Bahia, UESB, Jequié, BA
Ruiz-Ruiz S, Moreno P, Guerri J, Ambrós S (2007) A real-time RT-PCR assay for detection and absolute quantitation of Citrus tristeza virus in different plant tissues. J Virol Methods 145:96–105. https://doi.org/10.1016/j.jviromet.2007.05.011
Bustin SA, Benes V, Garson JA, Hellemans J, Huggett J, Kubista M, Vandesompele J (2009) The MIQE guidelines: minimum information for publication of quantitative real-time PCR experiments. Clin Chem 55:611–622. https://doi.org/10.1373/clinchem.2008.112797
Wang J, Zhang Y, Wang J, Liu L, Pang X, Yuan W (2017) Development of a TaqMan-based real-time PCR assay for the specific detection of porcine circovirus 3. J Virol Methods 248:77–180. https://doi.org/10.1016/j.jviromet.2017.07.007
Campbell CL, Madden LV (1990) Introducyion to plant disease epidemiology. Wiley, New York, p 532
R Development Core Team (2020) R: A Language and Environment for Statistical Computing. R Foundation for Statistical Computing, Vienna
Gower JC (1971) A general coefficient of similarity and some of its properties. Biometrics 27:857–874. https://doi.org/10.2307/2528823
Cruz CD (2013) Genes: a software package for analysis in experimental statistics and quantitative genetics. Acta Sci Agron 35:271–276. https://doi.org/10.4025/actasciagron.v35i3.21251
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S (2011) MEGA 5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28:2731–2739. https://doi.org/10.1093/molbev/msr121
Paula MS, Fonseca MEN, Boiteux LS, Peixoto JR (2010) Genetic characterization of Passiflora species via resistance genes analog markers. Rev Bras Frutic 32:222–229. https://doi.org/10.1590/S0100-29452010005000021
Soares TL, Jesus ON, Souza EH, Rossi ML, Oliveira EJ (2018) Comparative pollen morphological analysis in the subgenera Passiflora and Decaloba. An Acad Bras Ciênc 90:2381–2396. https://doi.org/10.1590/0001-3765201720170248
Richardo J, Silvério A (2019) New trends in Passiflora L. pollen grains: morphological/aperture aspects and wall layer considerations. Protoplasma 256:923–939. https://doi.org/10.1007/s00709-019-01350-w
Mäder G, Zamberlan PM, Fagundes NJ, Magnus T, Salzano FM, Bonatto SL, Freitas LB (2010) The use and limits of ITS data in the analysis of intraspecific variation in Passiflora L. (Passifloraceae). Genet mol biol 33:99–108. https://doi.org/10.1590/S1415-47572009005000101
Soares TL, Jesus ON, Souza EH, Oliveira EJ (2018) Floral development stage and its implications for the reproductive success of Passiflora L. Sci Hortic 238:333–342. https://doi.org/10.1016/j.scienta.2018.04.034
Soares TL, Jesus ON, Santos-Serejo JA, Oliveira EJ (2013) In vitro pollen germination and pollen viability in passion fruit (Passiflora spp.). Rev Bras Frutic 35:1116–1126. https://doi.org/10.1590/S0100-29452013000400023
Coelho MSE, Bortoleti KCA, Araújo FP, Melo NF (2016) Cytogenetic characterization of the Passiflora edulis Sims x Passiflora cincinnata Mast. interspecific hybrid and its parents. Euphytica 210:93–104. https://doi.org/10.1007/s10681-016-1704-4
Jesus ON, Soares TL, Oliveira EJ, Santos TCP, Farias DH, Bruckner CH, Novaes QS (2016) Dissimilarity based on morphological characterization and evaluation of pollen viability and in vitro germination in Passiflora hybrids and backcrosses. Acta Hortic 1127:401–408. https://doi.org/10.17660/ActaHortic.2016.1127.62
Cerqueira-Silva CBM, Moreira CN, Figueira AR, Corrêa RX, Oliveira AC (2008) Detection of a resistance gradient to Passion fruit woodiness virus and selection of ‘yellow’ passion fruit plants under field conditions. Genet Mol Res 7:1209–1216. https://doi.org/10.4238/vol7-4gmr484
Viana CDS, Pires MDC, Peixoto JR, Junqueira NTV, Blum LEB (2014) Partial resistance of passion fruit genotypes to the virose of the woodiness of the fruit (Cowpea aphid-borne mosaic virus-CABMV). Biosci J 30:338–345
Cruz Neto AJ, Rosa RCC, Oliveira EJ, Sampaio SR, Santos IS, Souza PU, Jesus ON (2016) Genetic parameters, adaptability and stability to selection of yellow passion fruit hybrids. Crop Breed Appl Biotechnol 16:321–329. https://doi.org/10.1590/1984-70332016v16n4a48
Jesus CASD, Carvalho EVD, Girardi EA, Rosa RCC, Jesus ON (2018) Fruit quality and production of yellow and sweet Passion fruits in northern state of São Paulo. Rev Bras Frutic 40:1–7. https://doi.org/10.1590/0100-29452018968
Santos IS, Lima LKS, Sampaio SR, Soares TL, Jesus ON (2021) Phenological precocity and resistance to CABMV in passion fruit progenies of the third generation backcross [(P. edulis × P. cincinnata) × P. edulis]. Euphytica 217:6. https://doi.org/10.1007/s10681-021-02842-8
Fonseca KG, Faleiro FG, Peixoto JR, Junqueira NTV, Silva MS, Bellon G, FariaVaz C (2009) Recovery analysis of recurrent genitor in sour passion fruit through RAPD markers. Rev Bras Frutic 31:145–153. https://doi.org/10.1590/S0100-29452009000100021
Suassuna TMF, Bruckner CH, Carvalho CR, Borém A (2003) Self-incompatibility in passion fruit: evidence of gametophytic-sporophytic control. Theor Appl Genet 106:298–302. https://doi.org/10.1007/s00122-002-1103-1
Madureira HC, Pereira TNS, Cunha MD, Klein DE (2012) Histological analysis of pollen-pistil interactions in sour passion fruit plants (Passiflora edulis Sims). Biocell 36:83–90
Obrępalska-Stęplowska A, Renaut J, Planchon S, Przybylska A, Wieczorek P, Barylski J, Palukaitis P (2015) Effect of temperature on the pathogenesis, accumulation of viral and satellite RNAs and on plant proteome in peanut stunt virus and satellite RNA-infected plants. Front Plant Sci 6:903. https://doi.org/10.3389/fpls.2015.00903
Pinto PHD, Peixoto JR, Junqueira NTV, Resende RDO, Mattos JKDA, Melo BD (2008) Reaction of passionfruit genotypes to Cowpea aphid-borne mosaic virus (cabmv). Biosci J 24:19–26
Novaes QS, Rezende JA (2005) Protection between strains of Passion fruit woodiness virus in sunnhemp. Fitopatol Bras 30:307–311. https://doi.org/10.1590/S0100-41582005000300017
Freitas DMS, Rezende JAM (2008) Protection between strains of Papaya ringspot virus: Type W in zucchini squash involves competition for viral replication sites. Sci Agric 65:183–189. https://doi.org/10.1590/S0103-90162008000200012
Liu J, Li XD, Xu S (2020) Single amino acid substitutions in the coat protein and RNA-dependent RNA polymerase alleviated the virulence of Cucumber green mottle mosaic virus and conferred cross protection against severe infection. Virus Genes. https://doi.org/10.1007/s11262-019-01726-3
Folimonova SY (2012) Superinfection exclusion is an active virus-controlled function that requires a specific viral protein. J Virol 86:5554–5561. https://doi.org/10.1128/JVI.00310-12
Kumar S, Tanti B, Patil BL, Mukherjee SK, Sahoo L (2017) RNAi-derived transgenic resistance to Mungbean yellow mosaic India virus in cowpea. PLoS ONE 12:1–20. https://doi.org/10.1371/journal.pone.0186786
Deng Y, Wang J, Tung J, Liu D, Zhou Y, He S, Li F (2018) A role for small RNA in regulating innate immunity during plant growth. PLoS pathog 14:1–22. https://doi.org/10.1371/journal.ppat.1006756
Csorba T, Kontra L, Burgyán J (2015) Viral silencing suppressors: tools forged to fine-tune host-pathogen coexistence. Virology 479:85–103. https://doi.org/10.1016/j.virol.2015.02.028
Roossinck MJ, Martin DP, Roumagnac P (2015) Plant virus metagenomics: advances in virus discovery. Phytopathology 105:716–727. https://doi.org/10.1094/PHYTO-12-14-0356-RVW
Sun SR, Ahmad K, Wu XB, Chen JS, Fu HY, Huang MT, Gao SJ (2018) Development of quantitative real-time PCR assays for rapid and sensitive detection of two badnavirus species in sugarcane. Biomed Res Int. https://doi.org/10.1155/2018/8678242
Tabara M, Nagashima Y, He K, Qian X, Crosby KM, Jifon J, Fukuhara T (2021) Frequent asymptomatic infection with tobacco ringspot virus on melon fruit. Virus Res 293:198266. https://doi.org/10.1016/j.virusres.2020.198266
Doumayrou J, Leblaye S, Froissart R, Michalakis Y (2013) Reduction of leaf area and symptom severity as proxies of disease-induced plant mortality: the example of the Cauliflower mosaic virus infecting two Brassicaceae hosts. Virus Res 176:91–100. https://doi.org/10.1016/j.virusres.2013.05.008
Gouveia BC, Calil IP, Machado JPB, Santos AA, Fontes EP (2017) Immune receptors and co-receptors in antiviral innate immunity in plants. Front Microbiol 2139:1–14. https://doi.org/10.3389/fmicb.2016.02139
Cruz ARR, Aragão FJL (2014) RNAi based enhanced resistance to Cowpea severe mosaic virus and cowpea aphid borne mosaic virus in transgenic cowpea. Plant pathol 63:831–837. https://doi.org/10.1111/ppa.12178
Albiach-Marti MR, Grosser JW, Gowda S, Mawassi M, Satyanarayana T, Garnsey SM, Dawson WO (2004) Citrus tristeza virus replicates and forms infectious virions in protoplasts of resistant citrus relatives. Mol Breed 14:117–128. https://doi.org/10.1023/B:MOLB.0000038000.51218.a7
Chirinos DT, Geraud-Pouey F, Fernandez CE, Bragard C, Romay G (2020) Genomic characterization and transmission efficiency by its vector Bemisia tabaci of a novel recombinant strain of potato yellow mosaic virus. Trop Plant Pathol 45:91–95. https://doi.org/10.1007/s40858-019-00316-w
Kondo H, Fujita M, Hisano H, Hyodo K, Andika IB, Suzuki N (2020) Virome analysis of aphid populations that infest the barley field: the discovery of two novel groups of nege/kita-like viruses and other novel RNA viruses. Front Microbiol 11:1–19. https://doi.org/10.3389/fmicb.2020.00509
Carvalho BM, Viana AP, Santos PHD, Generoso AL, Corrêa CCG, Silveira V, Santos EA (2019) Proteome of resistant and susceptible Passiflora species in the interaction with cowpea aphid-borne mosaic virus reveals distinct responses to pathogenesis. Euphytica 215:167. https://doi.org/10.1007/s10681-019-2491-5
Acknowledgements
The Coordenação de Aperfeiçoamento de Pessoal de Nível Superior (CAPES) provided a doctoral research grant to the first author (ZSG). The Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq) provided a postdoctoral scholarship to the third author (LKSL – PDJ 152109/2019-6) and a research productivity fellowship to the second (ONJ – PQ 312774/2018-4) and fourth author (RXC). We acknowledge the research unit of Embrapa Mandioca e Fruticultura for providing the plant material, infrastructure, and technical support for the execution of the research, Dr. Saulo Alves S. de Oliveira for supporting the analysis of data on disease progress rates, and Dr. Antônio Vargas de O. Figueira and the Centro de Energia Nuclear na Agricultura (CENA - Esalq/USP) for providing laboratory space and technical support for training in RT-qPCR of the first author (ZSG).
Funding
This work was funded by the Conselho Nacional de Desenvolvimento Cientifico e Tecnológico (CNPq – Process 421033/2018-5), Embrapa Mandioca e Fruticultura (Process Embrapa 22.16.04.007.00.00) and Fundação de Amparo à Pesquisa do Estado da Bahia (FAPESB – TO DTE0001/2016).
Author information
Authors and Affiliations
Contributions
All authors contributed to writing, as well as to interpreting the results, revising, and improving the paper. ZSG carried out the installation of the experiment, assessment of the severity of the disease, molecular analysis of virus detection and quantification, and writing of the paper. ZSG, ONJ, and LKSL participated in the statistical analysis, organization, and elaboration of tables and figures, as well as data interpretation. ONJ, LKSL, and RXC corrected the paper. ONJ and RXC were the creators of this research.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflicts of interest.
Ethical standards
The authors declare that the present work complies with the ethical standards of the Committee on Publication Ethics (COPE) and complies with the ethical standards the Universidade Estadual de Santa Cruz and Embrapa Mandioca e Fruticultura.
Additional information
Handling Editor: Ralf Georg Dietzgen.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Gonçalves, Z.S., Jesus, O.N., Lima, L.K.S. et al. Responses of Passiflora spp. to cowpea aphid-borne mosaic virus reveal infection in asymptomatic plants and new species with probable immunity. Arch Virol 166, 2419–2434 (2021). https://doi.org/10.1007/s00705-021-05131-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00705-021-05131-w