Abstract
Torque teno virus (TTV) is a commensal human virus observed as a circular single-negative-strand DNA molecule in various tissues and biological samples, notably in blood serum and lymphocytes. TTV has no apparent clinical significance, although it might be very useful as a prospective tool for gene delivery or as an epidemiological marker. Human populations are ubiquitously infected with TTV; the prevalence may reach 100%. The majority of babies become spontaneously infected with TTV, so that by the end of the first year of life, the prevalence reaches 'adult' values. TTV positivity in healthy early infancy and the presence of TTV in umbilical cord blood samples have been reported. The mechanism of infection and the dynamics of TTV prevalence in infants with age remain understudied. Meanwhile, the potential diagnostic and prognostic value of TTV as a marker deserves special attention and study, along with the possibility, causes and consequences of placental transmission of TTV under normal or pathological conditions.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Until recently, discoveries of new viruses invariably occurred in the context of disease. The default pathogenicity, overt or covert, of any virus was finally challenged by convincing descriptions of commensal viruses, such as hepatitis G virus and torque teno virus (TTV), initially associated with post-transfusion hepatitis. Commensal viruses are currently regarded on a par with normal flora of the human body. Abundant in the intestine, skin, oral and nasal cavities, pharynx, saliva, urine and even blood, commensal viruses are possibly of some adaptive value to the host, which is a likely explanation for their ubiquitous presence [1,2,3].
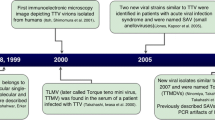
TTV was discovered in 1997 in Japan [4]. Hosted by a patient with post-transfusion hepatitis of unknown etiology, the virus was originally named after the patient's initials T.T. and renamed as torque teno (lat. ‘thin necklace’) or transfusion transmitted virus, which preserved the abbreviation.
TTV infections in humans are currently identified with members of several closely related viral species that belong to the genus Alphatorquevirus of the family Anelloviridae. The current ICTV classification of the family Anelloviridae recognizes 14 genera comprising a total of 76 species, including 29 TTV species [5]. TTVs 1, 2, 3, 6, 7, 9, 10, 25 and 27 are hosted by humans. The genomes of TTVs 1, 2 and 9 consist of 3852, 3322 and 3064 nt, respectively, and the rest of the human TTVs have genomes of very similar size, 3750–3770 nt [6]. The genomes of members of the family Anelloviridae are generally smaller than 4000 nt, so the prospects for their use as gene delivery vehicles are poor. All known anelloviruses occupy similar niches and can be found species-specifically in animals, notably in dogs, cats, apes and pigs (the latter share their TTVs with wild boars). Human-animal transmissions are possible both ways, but their contributions to viremia are negligible [7]. One of the smallest genomes in the family (2059 nt, NC_035192) belongs to giant panda anellovirus. This classification is subject to continuous update; moreover, single-genotype TTV isolates are frequently regarded as representing 'species'. Other anelloviruses hosted by humans include torque teno mini virus (TTMV, genus Betatorquevirus), torque teno midi virus (TTMDV, genus Gammatorquevirus) and small anellovirus (SAV) [8].
TTVs are small unenveloped viruses (ca. 25 nm in diameter) with icosahedral capsids. Their genomes are transmitted as single-(-)-strand circular DNA. These viruses infect mitotically active cells only. TTV genomes contain three ORFs, which encode the capsid protein VP1 and the non-structural proteins VP2 (a dual-specificity protein phosphatase) and VP3 (required for rolling-circle-type replication in the nucleus and formation of viral particles). Interestingly, VP3 is thought to exert oncolytic effects by specifically inducing apoptosis in transformed cells [9]. TTV replication can be blocked with aphidicolin [10], which reflects its dependence on cellular DNA polymerases.
The commensal nature of TTV has been repeatedly emphasized in the literature [11, 12]. The highly prevalent lifelong TTV viremia is asymptomatic and independent of age group or health status [13, 14]. The viral load of TTV may vary; early studies indicated lower TTV titers in healthy blood donors and patients, with the total prevalence under 90% [15, 16]. Such underestimation probably resulted from the exclusion of certain TTV variants, which escaped detection due to the use of insufficiently universal PCR primers. In 2009, about 94% of the healthy Russian population were living with > 1000 genomic copies of TTV per 1 ml of blood [17]. The global prevalence of TTV reaches 95% [18]. The data on TTMDV and TTMV prevalence are less consistent and rather scarce. Independent cohort studies carried out in different countries indicate TTMDV prevalence varying from 14.5% in Iran [19] to 75% in Japan [20] and TTMV prevalence varying from 29% in Korea [21] to 84.8% in Japan [22].
A recent comprehensive tissue-specific characterization of the human virome with the use of 8991 RNA-sequencing data sets corresponding to somatic tissues from 547 healthy individuals was carried out in the framework of the Genotype-Tissue Expression (GTEx) Project [23]. Contrary to expectations (based on the single-stranded nature of its genome), TTV loads showed no association with interferon-stimulated gene expression (observed for HSV-1 and some other viruses). Transcription and replication observed in a broad range of human tissues (blood, liver, myocardium, ovary, lung, etc.) is apparently controlled by the immune system in an interferon-independent manner.
TTV and diseases
The data on the prevalence and genetic heterogeneity of TTV in healthy individuals and in various pathologies is abundant. A number of studies have examined TTV in the epidemiological context of liver diseases [24, 25], cancer [26, 27], thalassemia [28, 29], non-malaria fevers [30, 31], and respiratory illnesses [32, 33]. Westman et al. [34] reported significant correlations of TTV plasma loads with age, CMV IgG levels, and HLA type in patients with Alzheimer disease (but no correlation with disease status). Increased TTV viremia is common in AIDS patients [35], cancer patients [26, 36], and children with inflammatory diseases of the respiratory system [37, 38]. Increased TTV loads are also typical in patients who receive immunosuppressive therapies for transplantation of stem cells and solid organs [39, 40], notably hematopoietic stem cell transplantations [41, 42], or have severely impaired immunity otherwise, and characteristic surges of TTV replication may occur during sepsis [43]. Comparative examination of nasal fluid in children with asthma and healthy children revealed no correlation of TTV loads with the disease, but showed a significant correlation of nasal TTV loads with spirometry indices and the level of eosinophilic cationic protein in sputum [44]. Reduced TTV viral loads in blood plasma have been suggested as predictors of antibody-mediated kidney transplant rejection and as a marker of the individual net state of immunosuppression in pediatric patients with kidney grafts [45, 46]. Beland et al. observed lower TTV loads in patients with chronic hepatitis after orthotopic liver transplantation (as compared to patients with unaltered histology), apparently due to inflammatory damage to the liver as one of the sites of TTV replication [47, 48]. Some studies indicate that only certain subtypes of TTV and TTV-like viruses correlate with pathologies (for instance, TTMDV and TTMV loads correlate with respiratory illnesses [20] and periodontitis [49]). The capability of anelloviruses to exacerbate pathogenesis remains uncertain; recent studies have elucidated several mechanisms by which these viruses can influence the immune response (e.g., by activating the production of pro-inflammatory cytokines via TLR-9 [50] or making the infected lymphocytes resistant to interferon-dependent immunomodulation [51]), thus worsening the condition. Some studies suggest that replication of anelloviruses may serve as an indicator of immune dysfunction and ultimately as a marker of immunodeficiency [52, 53]. It is not appropriate to consider TTVs potential or optional hepatitis viruses because they do not meet the criteria for hepatitis pathogens [54].
The ubiquity of TTV infections in humans is hardly compatible with the concept of pathogenicity unless we specify associations of viral loads or genetic variants with particular conditions. Systematic research in this direction is justified, as many anelloviruses hosted by other animal species show distinct disease-linked behaviors. Chicken infectious anemia virus (genus Gyrovirus), a self-sufficient pathogen that causes immunosuppression via systemic atrophy of hematopoietic tissues in young chickens [55], should definitely be mentioned in this context. A porcine torque teno virus, Torque teno sus virus 1b (TTSuV 1b, genus Iotatorquevirus) has been linked to porcine circovirus type 2–induced diseases, notably to postweaning multisystemic wasting syndrome, with TTSuV 1b serum viral loads being significantly higher in affected animals [56]. This finding is consistent with the elevated TTSuV 1b viral loads in pulmonary inflammatory lesions with viral background, including interstitial and broncho-interstitial pneumonia [57].
Detection methods
TTV was discovered by representational difference analysis (RDA) based on differential representation of specific nucleotide sequences in biomaterial. The methodology is based on subtractive hybridization of two DNA samples pre-digested with a restriction endonuclease. Labeling the ends of DNA fragments with artificial adapter sequences in one of the samples (thus providing annealing sites for universal primers) allows selective exponential amplification of re-hybridized species and, ultimately, identification of differentially expressed sequences [58].
The majority of protocols for routine TTV detection are PCR-based. Positioning of PCR primers is critical given the presence of hypervariable and conserved regions in viral genomes. The untranslated region with a regulatory GC-rich 113-nucleotide sequence is conserved [59]. ORF1, the longest ORF in the TTV genome, contains hypervariable regions (responsible for diversification of the virus-host interface and viral protein functionalities) [60]. The degree of target specificity for a particular PCR test (and, accordingly, its scope) is largely determined by the choice of annealing sites. A strategy of 'nested' PCR allows phylogenetic stratification of mixed amplified samples by sequential modulation of target specificity [20, 61]. An interesting protocol of high-resolution melting analysis suggested by Spandole et al. [62] allows TTV-, TTMDV- and TTMV-derived amplicons to be distinguished at the post-amplification level.
High-throughput sequencing and metagenomics studies of blood plasma viromes are receiving increasing interest. A recent retrospective descriptive pilot study of TTV dynamics in the follow-up of kidney transplantation enrolled 15 matched donor-recipient pairs from which blood plasma samples were collected and analyzed by next-generation sequencing [63]. The study revealed a strong difference in the predominant TTV strains in recipients and donors, accompanied by significantly enhanced diversity of TTV species in recipients. A cheaper alternative to sequencing that allows the diversity of TTV isolates to be accounted for is provided by PCR analysis of restriction fragment length polymorphism distributions [64].
An immunodetection protocol for measuring TTV in human blood serum developed in India [65, 66] is based on the use of a peptide corresponding to the N22 region in ORF1 (nt 1847–2346, GenBank no. AF122916.1). This protocol represents an efficient alternative to PCR-based approaches, especially with large samples, as it requires no expensive equipment and can easily be performed in small laboratories. Immunodetection is sensitive to cleared TTV infections, which evade detection by nucleic acid-based approaches.
Possible transmission routes
Possible transmission routes for TTV have been investigated in a number of studies. The presence of TTV was revealed in breast milk [67, 68], sperm [69], cervical swabs [70], nasal fluids and tears [32, 71], feces [72], bile [73], urine [3] and saliva [74, 75]. TTV DNA detected in the liver, bone marrow, and peripheral blood mononuclear cells matches with up to four different TTV transcripts generated by alternative splicing [76,77,78].
TTV titers in saliva were found to be 100–1000 fold higher than in the corresponding plasma samples [74]. This finding underscores the impact of TTV transmission through droplets of saliva, in addition to the possible contributions of fecal-oral route and breastfeeding. The dominant route of TTV transmission through saliva is consistent with the high overall prevalence of TTV.
TTV in pregnancy and vertical transmission
Association of certain viruses with intrauterine infections is well known [79]. Viruses have developed unique adaptive mechanisms that facilitate the infection process, the development of intrauterine infection, and damage to the embryo and fetus. Primary infection with certain viruses during pregnancy poses a risk of severe fetal complications [80]. The clinically relevant neonatal viruses include respiratory viruses (rhinoviruses, picornaviruses, parainfluenza viruses, respiratory syncytial virus, metapneumo-, parecho-, entero- and adenoviruses, bocavirus 1) and herpesviruses (CMV, HSVs, Epstein-Barr virus) [81,82,83]. Persistent perinatal viral infections also include hepatitis B and hepatitis C viruses and human immunodeficiency virus transmitted parenterally. Human T-lymphotropic viruses are transmitted through breast milk [79, 84], and genital human papillomaviruses (HPV) can be transmitted through mucous membranes during labor [85]. Indolfi et al. [86] have demonstrated an absence of synergy among different viruses in the transplacental transmission of multiple infections to the fetus and confirmed the hypothesis that transmission from infected mothers results from specific interactions of each virus with the host.
Effective transplacental transmission of anelloviruses in swine was discovered in 2009. Pozzuto et al. observed a high incidence of fetal infection with TTSuV (genus Iotatorquevirus) detected in 22% of serum samples from gnotobiotic piglets delivered by caesarian section [87]. Martínez-Guinó et al. reported up to 50% prevalence of TTSuV infections in stillborn pigs [88] and later on the absence of a correlation between the vertically transmitted infection with these viruses and spontaneous abortions in swine [89]. These results do not negate the possible impact of TTV transmission by the cervical route during labor or by the oral-fecal route shortly after birth, as the same team revealed a high TTV prevalence in both the milk whey and cellular fractions of colostrum [88].
Transplacental transmission of TTV in humans is not evident. A number of studies have claimed to show the presence of TTV in umbilical cord (UC) blood [90,91,92,93]. In other studies, all tested UC blood samples were TTV negative [94,95,96]. Probability estimations for TTV transplacental transmission made on the basis of UC blood sampling vary from 1% (n = 100) [97] to 33.3% (n = 57) [98]. The evidence on the possibility and incidence of TTV transmission in utero is summarized in Table 1.
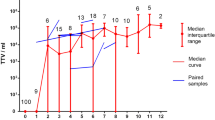
The relevance of dedicated monitoring of TTV loads in pregnancy is an open question. Our own results [94] indicate a high TTV prevalence (ca. 84%) with median plasma viral loads of 8 × 104 copies/mL and a 100-fold higher content of TTV DNA in whole blood than in plasma (which indicates high rates of TTV replication in circulating blood cells during pregnancy).
Bzhalava et al. [103] highlight the phylogenetic diversity of TTV in serum samples of pregnant women as determined by high-throughput sequencing; the team identified 40 different TTV isolates, including 29 that were previously unknown. Bagaglio et al. [101], who consider intrauterine transmission likely though not highly prevalent, report an infant infected at birth with a TTV strain different from her mother's dominant TTV genotype. This finding indicates the possibility of TTV evolution in fetuses.
TTV in neonates and infants
Although the vertical transmission of TTV has not been confirmed unequivocally [94], TTV is commonly detected in infants regardless of their clinical status [91, 92, 98, 104]. The early period of life is extremely important for immunity and in many respects determines the health of the individual in adulthood [105]. Physiological suppression of innate immunity in newborns and small infants enhances their susceptibility to bacterial and viral infections [106].
TTV infection in small infants appears clinically irrelevant. An observational study covering 30 months after birth [101] revealed an age-dependent increase in the prevalence of TTV infection in infants and toddlers. During the observation period, 13 out of 22 children became positive for TTV DNA, and, notably, the children with TTV viremia developed no symptoms of liver dysfunction, which confirms the lack of association between TTV and liver disorders. A recent study shows that TTV viral loads most rapidly increase during the first 60 days after birth and reach plateau at the age of 6 months [104]. These findings are consistent with a study by Yokozaki et al. [107], who revealed that while newborns are negative for TTV DNA, most 1-year-olds are positive. Gerner et al. demonstrated increased TTV loads in the blood plasma of > 1-week-old newborns (9 of 33 samples were TTV positive) [98]. Komatsu et al. also demonstrated an increase in the percentage of TTV-positive infants during the first months of life by analyzing the TTV DNA content of blood plasma; the authors emphasize increasing phylogenetic discrepancy in mother-infant pairs, with 85% of the pairs having the same genotype at the first PCR-positive time point and 50% of the children changing their dominant TTV genotype during follow-up [108]. Lower intrahost TTV heterogeneity in infants compared with their mothers was reported by Sugiyama et al. [109]; moreover, TTV sequences in mother-infant pairs (n = 7) differed by > 10% in all cases except one. Results of a recent study on the dynamics of TTV content in the cellular elements of peripheral blood during the first year of life [104] are generally consistent with the corresponding prevalence data obtained by the analysis of blood plasma. However, for peripheral blood cells, the percentage of positive samples is significantly higher than for the matching samples of blood plasma; the difference reflects high rates of TTV replication in lymphocytes [76].
TTV in later periods
Among healthy children of preschool and school age, TTV infection is common [67]. There are no convincing data on an association of TTV with pathologies in children. For example, in Egypt, the TTV indices for thalassemia patients and age-matched healthy children are similar [28, 29]. A number of studies indicate no correlation of TTV infection with liver disorders [110]. The prevalence of TTV and TTV-like viruses in children with acute hepatitis of unknown etiology, children with hepatitis B or C, and healthy children turned out to be very similar. Other studies have demonstrated that TTV infection prevails in healthy children from infancy [111, 112].
Conclusions
The placenta effectively serves as a barrier against various pathogens present in maternal blood, but the exact mechanisms of this defense remain largely elusive. It is likely that pathogens (and commensal microbes and viruses as well) may use different strategies to cross the placental barrier, depending on gestational age, the level of maternal infection, or the state of immune responses in the mother-fetus system. Vertical transmission of TTV appears to be mitigated by the placental barrier. However, TTV infection usually becomes detectable soon after birth, and the prevalence reaches 'adult' values by the end of the first year of life. As the possible pathogenicity of TTV is still an open question, monitoring of TTV in pregnant women, newborns, and infants is important.
Abbreviations
- AIDS:
-
Acquired immune deficiency syndrome
- CMV:
-
Cytomegalovirus
- DNA:
-
Deoxyribonucleic acid
- HSV:
-
Herpes simplex virus
- IgG:
-
Immunoglobulin G
- ICTV:
-
International Committee on Taxonomy of Viruses
- ORF:
-
Open reading frame
- nt:
-
Nucleotides
- PCR:
-
Polymerase chain reaction
- RDA:
-
Representational difference analysis
- SAV:
-
Small anellovirus
- TTMDV:
-
Torque teno midi virus (genus Gammatorquevirus)
- TTMV:
-
Torque teno mini virus (genus Betatorquevirus)
- TTSuV:
-
Torque teno sus virus (genus Iotatorquevirus)
- TTV:
-
Torque teno virus (genus Alphatorquevirus)
- UC:
-
Umbilical cord
References
Haynes M, Rohwer F (2010) The human virome. In: Metagenomics of the human body. Springer, New York, pp 63–77
Pride DT, Salzman J, Haynes M et al (2011) Evidence of a robust resident bacteriophage population revealed through analysis of the human salivary virome. ISME J 6:915–926. https://doi.org/10.1038/ismej.2011.169
Santiago-Rodriguez TM, Ly M, Bonilla N et al (2015) The human urine virome in association with urinary tract infections. Front Microbiol 6:1–12. https://doi.org/10.3389/fmicb.2015.00014
Nishizawa T, Okamoto H, Konishi K, Yoshizawa H, Miyakawa Y, Mayumi M (1997) A novel DNA virus (TTV) associated with elevated transaminase levels in posttransfusion hepatitis of unknown etiology. Biochem Biophys Res Commun 241:92–97. https://doi.org/10.1006/bbrc.1997.7765
https://talk.ictvonline.org/ictv-reports/ictv_9th_report/ssdna-viruses-2011/w/ssdna_viruses/139/anelloviridae. Accessed 8 Aug 2020
https://www.ncbi.nlm.nih.gov/genomes/GenomesGroup.cgi?taxid=687329. Accessed 8 Aug 2020]
Sarairah H, Bdour S, Gharaibeh W (2020) The molecular epidemiology and phylogeny of torque teno virus (TTV) in Jordan. Viruses 12:165. https://doi.org/10.3390/v12020165
Kaczorowska J, van der Hoek L (2020) Human anelloviruses: diverse, omnipresent and commensal members of the virome. FEMS Microbiol Rev 44:305–313. https://doi.org/10.1093/femsre/fuaa007
Payne S (2017) Viruses: from understanding to investigation. Elsevier Inc, pp 243–246
Kakkola L, Tommiska J, Boele LCL, Miettinen S, Blom T, Kekarainen T et al (2007) Construction and biological activity of a full-length molecular clone of human Torque teno virus (TTV) genotype 6. FEBS J 274:4719–4730. https://doi.org/10.1111/j.1742-4658.2007.06020.x
Griffiths P (1999) Time to consider the concept of a commensal virus? Rev Med Virol 9:73–74. https://doi.org/10.1002/(sici)1099-1654(199904/06)9:2<73:aid-rmv254>3.0.co;2-5
Simmonds P, Prescott LE, Logue C, Davidson F, Thomas AE, Ludlam CA (1999) TT virus—part of the normal human flora? J Infect Dis 180:1748–1749. https://doi.org/10.1086/315105
Abe K, Inami T, Asano K, Miyoshi C, Masaki N, Hayashi S et al (1999) TT virus infection is widespread in the general populations from different geographic regions. J Clin Microbiol 37:2703–2705. https://doi.org/10.1128/JCM.37.8.2703-2705.1999
AbuOdeh R, Al-Mawlawi N, Al-Qahtani AA et al (2015) Detection and genotyping of torque teno virus (TTV) in healthy blood donors and patients infected with HBV or HCV in Qatar. J Med Virol 87:1184–1191. https://doi.org/10.1002/jmv.24146
Charlton M, Adjei P, Poterucha J et al (1998) TT-virus infection in north american blood donors, patients with fulminant hepatic failure, and cryptogenic cirrhosis. Hepatology 28:839–842. https://doi.org/10.1002/hep.510280335
Irshad M, Singh S, Irshad K, Agarwal SK, Joshi YK (2008) Torque teno virus: its prevalence and isotypes in North India. World J Gastroenterol 14:6044–6051. https://doi.org/10.3748/wjg.14.6044
Vasilyev EV, Trofimov DY, Tonevitsky AG, Ilinsky VV, Korostin DO, Rebrikov DV (2009) Torque teno virus (TTV) distribution in healthy Russian population. Virol J 6:134–137. https://doi.org/10.1186/1743-422X-6-134
Brajão de Oliveira K (2015) Torque teno virus: a ubiquitous virus. Rev Bras Hematol Hemoter 37:357–358. https://doi.org/10.1016/j.bjhh.2015.07.009
Fatholahi M, Bouzari M (2015) Torque Teno midi virus/small Anellovirus in sera of healthy, HIV/HCV and HIV infected individuals in Lorestan Province, Iran. Jundishapur J Microbiol 8:e25368. https://doi.org/10.5812/jjm.25368
Ninomiya M, Takahashi M, Nishizawa T, Shimosegawa T, Okamoto H (2008) Development of PCR assays with nested primers specific for differential detection of three human anelloviruses and early acquisition of dual or triple infection during infancy. J Clin Microbiol 46:507–514. https://doi.org/10.1128/JCM.01703-07
Han TH, Chung JY (2006) Detection of small Anellovirus DNA from blood products. Korean J Blood Transfus 17:126–134
Spandole S, Cimponeriu D, Berca LM, Mihaescu G (2015) Human anelloviruses: an update of molecular, epidemiological and clinical aspects. Arch Virol 160:893–908. https://doi.org/10.1007/s00705-015-2363-9
Kumata R, Ito J, Takahashi K, Suzuki T, Sato K (2020) A tissue level atlas of the healthy human virome. BMC Biol 18:55. https://doi.org/10.1186/s12915-020-00785-5
Hsieh SY, Wu YH, Ho YP, Tsao KC, Yeh CT, Liaw YF (1999) High prevalence of TT virus infection in healthy children and adults and in patients with liver disease in Taiwan. Am Soc Microbiol 37:1829–1831. https://doi.org/10.1128/JCM.37.6.1829-1831.1999
Hafez MM, Shaarawy SM, Hassan AA, Salim RF, Abd El Salam FM, Ali AE (2007) Prevalence of transfusion transmitted virus (TTV) genotypes among HCC patients in Qaluobia governorate. Virol J 4:135. https://doi.org/10.1186/1743-422X-4-135
Sawata T, Bando M, Nakayama M et al (2018) Clinical significance of changes in Torque teno virus DNA titer after chemotherapy in patients with primary lung cancer. Respir Investig 56:173–178. https://doi.org/10.1016/j.resinv.2017.12.004
Kosulin K, Kernbichler S, Pichler H et al (2018) Post-transplant replication of torque teno virus in granulocytes. Front Microbiol 9:2956. https://doi.org/10.3389/fmicb.2018.02956
Kondili LA, Pisani G, Beneduce F et al (2001) Prevalence of TT virus in healthy children and thalassemic pediatric and young adult patients. J Pediatr Gastroenterol Nutr 33:629–632. https://doi.org/10.1097/00005176-200111000-00025
Hassuna NA, Naguib E, Abdel-Fatah M, Mousa SMO (2017) Phylogenetic analysis of torque teno virus in thalassemic children in Egypt. Intervirology 60:102–108. https://doi.org/10.1159/000480507
Hsu J, Rosenthal P, Ruel T, Kyohere M et al (2018) Etiology of non-malarial febrile illnesses in ugandan children. Pediatrics 141:463. https://doi.org/10.1542/peds.141.1_MeetingAbstract.463
McElvania TeKippe E, Wylie KM, Deych E, Sodergren E, Weinstock G, Storch GA (2012) Increased prevalence of anellovirus in pediatric patients with fever. PLoS ONE 7:e50937. https://doi.org/10.1371/journal.pone.0050937
Maggi F, Pifferi M, Fornai C et al (2003) TT virus in the nasal secretions of children with acute respiratory diseases: relations to viremia and disease severity. J Virol 77:2418–2425. https://doi.org/10.1128/jvi.77.4.2418-2425.2003
Jones MS, Kapoor A, Lukashov VV, Simmonds P, Hecht F, Delwart E (2005) New DNA viruses identified in patients with acute viral infection syndrome. J Virol 79:8230–8236
Westman G, Schoofs C, Ingelsson M, Järhult JD, Muradrasoli S (2020) Torque teno virus viral load is related to age, CMV infection and HLA type but not to Alzheimer's disease. PLoS ONE 15:e0227670. https://doi.org/10.1371/journal.pone.0227670
Thom K, Petrik J (2007) Progression towards AIDS leads to increased torque teno virus and torque teno minivirus titers in tissues of HIV infected individuals. J Med Virol 79:1–7. https://doi.org/10.1002/jmv.20756
De Villiers E-M, Bulajic M, Nitsch C et al (2007) TTV infection in colorectal cancer tissues and normal mucosa. Int J Cancer 121:2109–2112. https://doi.org/10.1002/ijc.22931
Maggi F, Pifferi M, Tempestini E, Lanini L, de Marco E, Fornai C et al (2004) Correlation between Torque tenovirus infection and serum levels of eosinophil cationic protein in children hospitalized for acute respiratory diseases. J Infect Dis 190:971–974. https://doi.org/10.1086/423143
Pifferi M, Maggi F, Caramella D, de Marco E, Andreoli E, Meschi S et al (2006) High torquetenovirus loads are correlated with bronchiectasis and peripheral airflow limitation in children. Pediatr Infect Dis J 25:804–808. https://doi.org/10.1097/01.inf.0000232723.58355.f4
Albert E, Solano C, Giménez E, Focosi D, Pérez A, Macera L et al (2017) The kinetics of torque teno virus plasma DNA load shortly after engraftment predicts the risk of high-level CMV DNAemia in allogeneic hematopoietic stem cell transplant recipients. Bone Marrow Transplant 53:180–187. https://doi.org/10.1038/bmt.2017.235
Abbas AA, Diamond JM, Chehoud C, Chang B, Kotzin JJ, Young JC et al (2017) The perioperative lung transplant virome: torque teno viruses are elevated in donor lungs and show divergent dynamics in primary graft dysfunction. Am J Transplant 17:1313–1324. https://doi.org/10.1111/ajt.14076
Peker BO, Daloğlu AE, Görzer I, Puchhammer-Stöckl E, Parkan ÖM, Akbaş H, Kintrup GT, Mutlu D, Küpesiz OA, Çolak D (2020) Investigation of torque teno virus (TTV) DNA as an immunological and virological marker in pediatric hematopoietic stem cell transplantation (HSCT) patients. Microb Pathog. https://doi.org/10.1016/j.micpath.2020.104397
Pradier A, Masouridi-Levrat S, Bosshard C, Dantin C, Vu DL, Zanella MC, Boely E, Tapparel C, Kaiser L, Chalandon Y, Simonetta F, Roosnek E (2020) Torque teno virus as a potential biomarker for complications and survival after allogeneic hematopoietic stem cell transplantation. Front Immunol 11:998. https://doi.org/10.3389/fimmu.2020.00998
Davila S, Halstead ES, Hall MW et al (2018) A viral DNAemia and immune suppression in pediatric sepsis. Pediatr Crit Care Med 19:14–22. https://doi.org/10.1097/PCC.0000000000001376
Pifferi M, Maggi F, Andreoli E, Lanini L, Marco ED, Fornai C et al (2005) Associations between nasal torquetenovirus load and spirometric indices in children with asthma. J Infect Dis 192:1141–1148. https://doi.org/10.1086/444389
Schiemann M, Puchhammer-Stöckl E, Eskandary F, Kohlbeck P, Rasoul-Rockenschaub S, Heilos A, Kozakowski N, Görzer I, Kikić Ž, Herkner H, Böhmig GA, Bond G (2017) Torque teno virus load-inverse association with antibody-mediated rejection after kidney transplantation. Transplantation 101:360–367. https://doi.org/10.1097/TP.0000000000001455
Uhl P, Heilos A, Bond G, Meyer E, Böhm M, Puchhammer-Stöckl E, Arbeiter K, Müller-Sacherer T, Csaicsich D, Aufricht C, Rusai K (2020) Torque teno viral load reflects immunosuppression in paediatric kidney-transplanted patients-a pilot study. Pediatr Nephrol. https://doi.org/10.1007/s00467-020-04606-3
Beland K, Dore-Nguyen M, Gagne MJ et al (2014) Torque teno virus in children who underwent orthotopic liver transplantation: new insights about a common pathogen. J Infect Dis 209:247–254. https://doi.org/10.1093/infdis/jit423
Reshetnyak VI, Maev IV, Burmistrov AI, Chekmazov IA, Karlovich TI (2020) Torque teno virus in liver diseases: On the way towards unity of view. World J Gastroenterol 26:1691–1707. https://doi.org/10.3748/wjg.v26.i15.1691
Zhang Y, Li F, Shan TL, Deng X, Delwart E, Feng XP (2016) A novel species of torque Teno mini virus (TTMV) in gingival tissue from chronic periodontitis patients. Sci Rep 6:26739. https://doi.org/10.1038/srep26739
Rocchi J, Ricci V, Albani M, Lanini L, Andreoli E, Macera L, Pistello M, Ceccherini-Nelli L, Bendinelli M, Maggi F (2009) Torquetenovirus DNA drives proinflammatory cytokines production and secretion by immune cells via toll-like receptor 9. Virology 394:235–242. https://doi.org/10.1016/j.virol.2009.08.036
Kincaid RP, Burke JM, Cox JC, de Villiers EM, Sullivan CS (2013) A human torque teno virus encodes a microRNA that inhibits interferon signaling. PLoS Pathog 9(12):e1003818. https://doi.org/10.1371/journal.ppat.1003818
Wohlfarth P, Leiner M, Schoergenhofer C et al (2018) Torquetenovirus dynamics and immune marker properties in patients following allogeneic hematopoietic stem cell transplantation: a prospective longitudinal study. Biol Blood Marrow Transplant 24:194–199. https://doi.org/10.1016/j.bbmt.2017.09.020
Nordén R, Magnusson J, Lundin A et al (2018) Quantification of torque teno virus and Epstein-Barr virus is of limited value for predicting the net state of immunosuppression after lung transplantation. Open Forum Infect Dis 5(4):ofy050. https://doi.org/10.1093/ofid/ofy050
Mushahwar IK (2000) Recently discovered blood-borne viruses: are they hepatitis viruses or merely endosymbionts? J Med Virol 62:399–404. https://doi.org/10.1002/1096-9071(200012)62:4<399:aid-jmv1>3.0.co;2-u
Fenner's Veterinary Virology [book, 5th Edition] (2016) Chapter 13—circoviridae and anelloviridae, pp 259–268. https://www.sciencedirect.com/book/9780128009468/fenners-veterinary-virology
Aramouni M, Segalés J, Sibila M, Martin-Valls GE, Nieto D, Kekarainen T (2011) Torque teno sus virus 1 and 2 viral loads in postweaning multisystemic wasting syndrome (PMWS) and porcine dermatitis and nephropathy syndrome (PDNS) affected pigs. Vet Microbiol 153:377–381. https://doi.org/10.1016/j.vetmic.2011.05.046
Aramouni M, Martínez J, Nieto D, Kekarainen T, Segalés J (2013) Exploratory study of Torque teno sus viruses in pulmonary inflammatory lesions in pigs. Vet Microbiol 162:338–344. https://doi.org/10.1016/j.vetmic.2012.09.011
Okamoto H, Nishizawa T, Kato N, Ukita M, Ikeda H, Iizuka H, Mayumi M (1998) Molecular cloning and characterization of a novel DNA virus (TTV) associated with posttransfusion hepatitis of unknown etiology. Hepatol Res 10:1–16. https://doi.org/10.1016/S1386-6346(97)00123-X
Hsiao KL, Wang LY, Lin CL, Liu HF (2016) New phylogenetic groups of torque teno virus identified in Eastern Taiwan Indigenes. PLoS ONE 11(2):e0149901. https://doi.org/10.1371/journal.pone.0149901
Jelcic I, Hotz-Wagenblatt A, Hunziker A, Zur Hausen H, de Villiers EM (2004) Isolation of multiple TT virus genotypes from spleen biopsy tissue from a Hodgkin's disease patient: genome reorganization and diversity in the hypervariable region. J Virol 78:7498–7507. https://doi.org/10.1128/JVI.78.14.7498-7507.2004
Wei Y, Chen M, Yang X, Zhang L, Rao L, Yuan F, Wang Y, Gong J, Li L (2015) Molecular characterization of human torque teno virus. Biomed Rep 3:821–826. https://doi.org/10.3892/br.2015.508
Spandole S, Cimponeriu D, Toma M, Radu I, Ion DA (2013) Rapid detection of human torque teno viruses using high-resolution melting analysis. Balkan J Med Genet 16:55–62. https://doi.org/10.2478/bjmg-2013-0018
Kulifaj D, Tilloy V, Scaon E, Guerin E, Essig M, Pichon N, Hantz S, De Bernardi A, Joannes M, Barranger C, Alain S (2020) Viral metagenomics analysis of kidney donors and recipients: torque teno virus genotyping and prevalence. J Med Virol. https://doi.org/10.1002/jmv.26298
Tanaka T, Kuroda K, Kobayashi M, Sato K (2001) Detection and typing of TT virus DNA genotype by the PCR-RFLP method. Mol Cell Probes 15:195–200. https://doi.org/10.1006/mcpr.2001.0358
Mankotia DS, Irshad M (2014) Cloning and expression of N22 region of torque teno virus (TTV) genome and use of peptide in developing immunoassay for TTV antibodies. Virol J 11:96. https://doi.org/10.1186/1743-422X-11-96
Mankotia DS, Irshad M (2017) Development of an immunoassay for detection of torque teno virus (TTV) antibodies using the N22 expression product from TTV genotype 2. Intervirology 60:207–216. https://doi.org/10.1159/000487481
Toyoda H, Naruse M, Yokozaki S, Morita K, Nakano I, Itakura A, Okamura M, Fukuda Y, Hayakawa T (1999) Prevalence of infection with TT virus (TTV), a novel DNA virus, in healthy Japanese subjects, newborn infants, cord blood and breast milk. J Infect 38(3):198–199. https://doi.org/10.1016/s0163-4453(99)90254-2
El-Esnawy NA, Bacheer HH, Ezzat E, Mostafa A, Ashour E et al (2007) TT virus in infants sera and mother breast milks. JGEB 5:19–26
Inami T, Konomi N, Arakawa Y, Abe K (2000) High prevalence of TT virus DNA in human saliva and semen. J Clin Microbiol 38(6):2407–2408
Fornai C, Maggi F, Vatteroni ML, Pistello M, Bendinelli M (2001) High prevalence of TT virus (TTV) and TTV-like minivirus in cervical swabs. J Clin Microbiol 39(5):2022–2024. https://doi.org/10.1128/JCM.39.5.2022-2024.2001
Matsubara H, Michitaka K, Horiike N, Yano M, Akbar SM, Torisu M, Onji M (2000) Existence of TT virus DNA in extracellular body fluids from normal healthy Japanese subjects. Intervirology 43(1):16–19. https://doi.org/10.1159/000025018
Pinho-Nascimento CA, Leite JP, Niel C, Diniz-Mendes L (2011) Torque teno virus in fecal samples of patients with gastroenteritis: prevalence, genogroups distribution, and viral load. J Med Virol 83(6):1107–1111. https://doi.org/10.1002/jmv.22024
Itoh M, Shimomura H, Fujioka S et al (2001) High prevalence of TT virus in human bile juice samples: importance of secretion through bile into feces. Dig Dis Sci 46(3):457–462. https://doi.org/10.1023/a:1005618308943
Gallian P, Biagini P, Zhong S et al (2000) TT virus: a study of molecular epidemiology and transmission of genotypes 1, 2 and 3. J Clin Virol 17(1):43–49. https://doi.org/10.1016/s1386-6532(00)00066-4
Deng X, Terunuma H, Handema R et al (2000) Higher prevalence and viral load of TT virus in saliva than in the corresponding serum: another possible transmission route and replication site of TT virus. J Med Virol 62(4):531–537
Zhong S, Yeo W, Tang M, Liu C, Lin XR, Ho WM, Hui P, Johnson PJ (2002) Frequent detection of the replicative form of TT virus DNA in peripheral blood mononuclear cells and bone marrow cells in cancer patients. J Med Virol 66(3):428–434. https://doi.org/10.1002/jmv.2163
Mueller B, Maerz A, Doberstein K, Finsterbusch T, Mankertz A (2008) Gene expression of the human Torque Teno Virus isolate P/1C1. Virology 381(1):36–45. https://doi.org/10.1016/j.virol.2008.08.017
Okamoto H, Nishizawa T, Takahashi M, Asabe S, Tsuda F, Yoshikawa A (2001) Heterogeneous distribution of TT virus of distinct genotypes in multiple tissues from infected humans. Virology 288(2):358–368
Silasi M, Cardenas I, Racicot K, Kwon JY, Aldo P, Mor G (2015) Viral infections during pregnancy. Am J Reprod Immunol 73(3):199–213. https://doi.org/10.1111/aji.12355
Stagno S, Pass RF, Dworsky ME, Alford CA (1982) Maternal cytomegalovirus infection and perinatal transmission. Clin Obstet Gynecol 25:563–576
James SH, Kimberlin DW (2015) Neonatal herpes simplex virus infection. Infect Dis Clin N Am 29(3):391–400. https://doi.org/10.1016/j.idc.2015.05.001
Luoto R, Jartti T, Ruuskanen O, Waris M (2016) Review of the clinical significance of respiratory virus infections in newborn infants. Acta Paediatr 105:1132–1139. https://doi.org/10.1111/apa.13519
Pinninti SG, Kimberlin DW (2018) Neonatal herpes simplex virus infections. Semin Perinatol 42(3):168–175. https://doi.org/10.1053/j.semperi.2018.02.004
Carneiro-Proietti ABF, Amaranto-Damasio MS, Leal-Horiguchi CF et al (2014) Mother-to-child transmission of human T-cell lymphotropic viruses-1/2: what we know, and what are the gaps in understanding and preventing this route of infection. J Pediatric Infect Dis Soc 3(1):24–29. https://doi.org/10.1093/jpids/piu070
Smith EM, Parker MA, Rubenstein LM, Haugen TH, Hamsikova E, Turek LP (2010) Evidence for vertical transmission of HPV from mothers to infants. Infect Dis Obstet Gynecol 2010:326369. https://doi.org/10.1155/2010/326369
Indolfi G, Moriondo M, Galli L, Azzari C, Poggi GM et al (2007) Mother-to-infant transmission of multiple blood-borne viral infections from multi-infected mothers. J Med Virol 79(6):743–747. https://doi.org/10.1002/jmv.20885
Pozzuto T, Mueller B, Meehan B, Ringler SS, McIntosh KA, Ellis JA, Mankertz A, Krakowka S (2009) In utero transmission of porcine torque teno viruses. Vet Microbiol 137:375–379. https://doi.org/10.1016/j.vetmic.2009.02.001
Martínez-Guinó L, Kekarainen T, Segalés J (2009) Evidence of torque teno virus (TTV) vertical transmission in swine. Theriogenology 71:1390–1395. https://doi.org/10.1016/j.theriogenology.2009.01.010
Martínez-Guinó L, Kekarainen T, Maldonado J, Aramouni M, Llorens A, Segalés J (2010) Torque teno sus virus (TTV) detection in aborted and slaughterhouse collected foetuses. Theriogenology 74:277–281. https://doi.org/10.1016/j.theriogenology.2010.02.011]
Matsubara H, Michitaka K, Horiike N, Kihana T, Yano M, Mori T, Onji M (2001) Existence of TT virus DNA and TTV-like mini virus DNA in infant cord blood: mother-to-neonatal transmission. Hepatol Res 21(3):280–287. https://doi.org/10.1016/s1386-6346(01)00115-2
Saback FL, Gomes SA, de Paula VS, da Silva RR, Lewis-Ximenez LL, Niel C (1999) Age-specific prevalence and transmission of TT virus. J Med Virol 59(3):318–322
Xin X, Xiaoguang Z, Ninghu Z, Youtong L, Liumei X, Boping Z (2004) Mother-to-infant vertical transmission of the transfusion-transmitted virus in South China. J Perinat Med 32(5):404–406. https://doi.org/10.1515/JPM.2004.136
Morrica A, Maggi F, Vatteroni ML et al (2000) TT virus: evidence for transplacental transmission. J Infect Dis 181:803–804. https://doi.org/10.1086/315296
Tyschik EA, Sherbakova SM, Ibragimov RR, Rebrikov DV (2017) Transplacental transmission of torque teno virus. Virol J 14(1):92–94. https://doi.org/10.1186/s12985-017-0762-0
Ohto H, Ujiie N, Takeuchi C et al (2002) TT virus infection during childhood. Transfusion 42:892–898. https://doi.org/10.1046/j.1537-2995.2002.00150.x
Fang F, Zhong W, Wang H (2001) Study on infection of transfusion transmitted virus in serum, breast milk of pregnant women and umbilical venous blood from their newborns [Article in Chinese]. Zhonghua Fu Chan Ke Za Zhi 36:330–332
Kazi A, Miyata H, Kurokawa K, Khan MA, Kamahora T, Katamine S, Hino S (2000) High frequency of postnatal transmission of TT virus in infancy. Arch Virol 145:535–540. https://doi.org/10.1007/s007050050044
Gerner P, Oettinger R, Gerner W, Falbrede J, Wirth S (2000) Mother-to-infant transmission of TT virus: prevalence, extent, and mechanism of vertical transmission. Pediatr Infect Dis J 19(11):1074–1077. https://doi.org/10.1097/00006454-200011000-00009
Iso K, Suzuki Y, Takayama M (2001) Mother-to-infant transmission of TT virus in Japan. Int J Gynaecol Obstet 75:11–19. https://doi.org/10.1016/s0020-7292(01)00450-7
Chan PK, Tam WH, Yeo W, Cheung JL, Zhong S, Cheng AF (2001) High carriage rate of TT virus in the cervices of pregnant women. Clin Infect Dis 32:1376–1377. https://doi.org/10.1086/319983
Bagaglio S, Sitia G, Prati D et al (2002) Mother-to-child transmission of TT virus: sequence analysis of the non-coding region of TT virus in infected mother-infant pairs. Arch Virol 147(4):803–812. https://doi.org/10.1007/s007050200027
Sloan P, Rodriguez C, Bedell BA, Murray J, Dagle J, Ryckman K, Holtz L (2020) Alphatorquevirus is the most prevalent virus identified in blood from a matched maternal-infant preterm cohort. J Matern Fetal Neonatal Med. https://doi.org/10.1080/14767058.2020.1763298
Bzhalava D, Ekström J, Lysholm F, Hultin E, Faust H, Persson B, Lehtinen M, de Villiers EM, Dillner J (2012) Phylogenetically diverse TT virus viremia among pregnant women. Virology 432:427–434. https://doi.org/10.1016/j.virol.2012.06.022
Tyschik EA, Rasskazova AS, Degtyareva AV, Rebrikov DV, Sukhikh GT (2018) Torque teno virus dynamics during the first year of life. Virol J 15(1):96–99. https://doi.org/10.1186/s12985-018-1007-6
Lim ES, Zhou Y, Zhao G, Bauer IK, Droit L (2015) Early life dynamics of the human gut virome and bacterial microbiome in infants. Nat Med 21(10):1228–1234. https://doi.org/10.1038/nm.3950
Yu JC, Khodadadi H, Malik A, Davidson B, Salles ÉDSL, Bhatia J, Hale VL, Baban B (2018) Innate immunity of neonates and infants. Front Immunol 9:1759. https://doi.org/10.3389/fimmu.2018.01759
Yokozaki S, Fukuda Y, Nakano I, Katano Y (1999) TT virus: a mother to child transmitted rather than blood borne virus. Blood 93(10):3569–3570
Komatsu H, Inui A, Sogo T, Kuroda K, Tanaka T, Fujisawa T (2004) TTV infection in children born to mothers infected with TTV but not with HBV, HCV, or HIV. J Med Virol 74(3):499–506. https://doi.org/10.1002/jmv.20204
Sugiyama K, Goto K, Ando T, Mizutani F, Terabe K, Yokoyama T (2001) Highly diverse TTV population in infants and their mothers. Virus Res 73:183–188. https://doi.org/10.1016/s0168-1702(00)00242-2
Abdalla NM, Galal A, Fatouh AA, Eid K, Salama EEE, Gomma HE (2006) Transfusion transmitted virus (TTV) infection in polytransfused egyptian thalassemic children. J Med Sci 6(5):833–837. https://doi.org/10.3923/jms.2006.833.837
Peng YH, Nishizawa T, Takahashi M, Ishikawa T, Yoshikawa A, Okamoto H (2002) Analysis of the entire genomes of thirteen TT virus variants classifiable into the fourth and fifth genetic groups, isolated from viremic infants. Arch Virol 147(1):21–41. https://doi.org/10.1007/s705-002-8301-7
Lin HH, Kao JH, Lee PI, Chen DS (2002) Early acquisition of TT virus in infants: possible minor role of maternal transmission. J Med Virol 66(2):285–290. https://doi.org/10.1002/jmv.2143
Acknowledgements
We acknowledge Natalia Usman for helpful discussions.
Funding
This work was supported by Grant No. 075-15-2019-1789 from the Ministry of Science and Higher Education of the Russian Federation, allocated to the Center for Precision Genome Editing and Genetic Technologies for Biomedicine.
Author information
Authors and Affiliations
Contributions
EAL and DVR drafted the manuscript. All authors read and approved the final version of the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no competing interests.
Additional information
Handling Editor: Zhongjie Shi.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Lolomadze, E.A., Rebrikov, D.V. Constant companion: clinical and developmental aspects of torque teno virus infections. Arch Virol 165, 2749–2757 (2020). https://doi.org/10.1007/s00705-020-04841-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00705-020-04841-x