Abstract
Southern tomato virus (STV) is often found infecting healthy tomato plants (Solanum lycopersicum). In this study, we compared STV-free and STV-infected plants of cultivar M82 to determine the effect of STV infection on the host plant. STV-free plants exhibited a short and bushy phenotype, whereas STV-infected plants were taller. STV-infected plants produced more fruit than STV-free plants, and the germination rate of seeds from STV-infected plants was higher than that of seeds from STV-free plants. This phenotypic difference was also observed in progeny plants (siblings) derived from a single STV-infected plant in which the transmission rate of STV to progeny plants via the seeds was approximately 86%. These results suggest that the interaction between STV and host plants is mutualistic. Transcriptome analysis revealed that STV infection affects gene expression in the host plant and results in downregulation of genes involved in ethylene biosynthesis and signaling. STV-infected tomato plants might thus be artificially selected due to their superior traits as a crop.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Linear double-stranded RNAs (dsRNAs) are common in various crops and vegetables, such as radish (Raphanus sativus), bell pepper (Capsicum annuum) and rice (Oryza sativa) [1,2,3,4]. dsRNAs are not transcribed from the host DNA, and nucleotide sequencing and phylogenetic analysis have indicated that these dsRNAs are derived from the genomes of viruses [5, 6]. These viruses have no obvious effects on the host plant, are distributed throughout the tissues at a stable low concentration, and are efficiently transmitted to the next generation via seeds. As the properties of these viruses differ from those of conventional “acute” or “chronic” viruses, they are referred to as “persistent” viruses [7]. Most persistent viruses found in plants are members of the family Partitiviridae or Endornaviridae [7,8,9].
A putative persistent dsRNA virus, southern tomato virus (STV), has been identified in various cultivars of tomato (Solanum lycopersicum) [10]. The complete nucleotide sequence of the STV dsRNA genome and results of phylogenetic analysis have indicated that this virus contains a single genomic dsRNA encoding two partially overlapping open reading frames (ORFs) [10]. This genome organization is similar to that of fungal dsRNA viruses of the family Totiviridae, which contain two partially overlapping ORFs that encode a capsid protein and an RNA-dependent RNA polymerase [11]. Based on phylogenetic analysis, Sabanadzovic et al. [10] concluded that STV exhibits properties of members of both the family Partitiviridae and the family Totiviridae. Thus, STV has been classified by the International Committee on Taxonomy of Viruses as a new dsRNA virus in a new genus (Amalgavirus) within a new family (Amalgaviridae) [12,13,14].
STV was first detected in tomato plants grown in the USA and Mexico that exhibited symptoms of stunting, fruit discoloration, and reduced fruit size [10]. STV was subsequently detected in tomato plants exhibiting similar symptoms in Spain, Italy, France, China and Bangladesh [15,16,17,18,19]. However, because STV is frequently detected in mixed infections with other acute viruses such as pepino mosaic virus (PepMV) and tomato mosaic virus [20,21,22] as well as in asymptomatic tomato plants [22, 23], the effect of STV infection on host plants remains unclear.
In this study, we examined the effect of STV infection on host plants by comparing the characteristics of STV-infected and STV-free tomato plants of the otherwise virus-free cultivar (cv.) M82, which is a standard cultivar used in tomato research [24, 25]. STV-infected plants were taller and produced more fruit than STV-free plants. The effects of asymptomatic STV infection on gene expression, plant architecture, fruit production, and seed germination are discussed.
Materials and methods
Plant materials and growth conditions
Tomato plants were grown in pots in a glass greenhouse under natural daylight or in a controlled-environment chamber under the following conditions: 40-50 μmol m-2 sec-1, 16-h light and 8-h dark cycle, at 24 °C. Seeds of cv. M82 were kindly provided by the National BioResource Project of Japan (NBRP, Tsukuba University). Commercially available seeds of the tomato cultivars Cherry Mate, Sweet Hearts, Ponderosa and Momotaro were purchased from the seed companies Tohoku Seed, ING Corp., Noguchi Seed, and Takii Seed.
M82 seeds were also harvested from self-pollinated fruits of both STV-free and STV-infected plants (Table 1). The germination rates of harvested seeds were determined in a controlled-environment chamber.
Detection and preparation of dsRNA
dsRNA was purified from leaves using a previously reported method [26, 27]. In brief, approximately 0.2 g of leaves was pulverized in liquid nitrogen using a mortar and pestle. dsRNA was extracted using 2 × STE buffer (200 mM NaCl, 20 mM Tris-HCl [pH 8.0] and 1 mM EDTA) containing 1% SDS and phenol/chloroform/isoamyl alcohol (25:24:1) and purified by spin-column chromatography with cellulose (Advantec, Tokyo, Japan). The dsRNA was analyzed by electrophoresis on a 1.0% (w/v) agarose gel containing TAE buffer (40 mM Tris-acetate, 1 mM EDTA [pH 8.0]) and 500 ng ml−1 ethidium bromide.
Nucleotide sequencing
Full-length PCR products were generated from purified STV dsRNAs using a PrimeScript™ II High-Fidelity One-Step RT-PCR Kit (Takara, Shiga, Japan) with the primers listed in Supplemental Table S1. PCR products were purified using a QIAquick PCR Purification Kit (QIAGEN, Germany) and then sequenced using a HiSeq 2500 Sequencing System (Illumina, San Diego, CA, USA).
Transcriptome analysis
Total RNA was extracted from leaves of 1-month-old tomato plants (three STV-infected and three STV-free cv. M82 plants) using TRIzol Reagent according to the manufacturer’s protocol (Thermo Fisher Scientific, Japan). An RNA sequencing library was prepared using a TruSeq Stranded mRNA Sample Prep Kit (Illumina) according to the manufacturer’s instructions. Whole transcriptome sequencing was applied to the RNA samples using an Illumina HiSeq 2500 platform in 75-base single-end mode. Illumina Casava software (ver.1.8.2) was used for base calling. Sequenced reads were mapped to the tomato genome (Tomato SL2.50 ITAG2.4; Sol Genomic Network https://solgenomics.net/) using TopHat (ver. 2.0.13) in combination with Bowtie2 (ver. 2.2.3) [28] and SAMtools (ver. 0.1.19). The number of fragments per kilobase of exon per million mapped fragments (FPKMs) was calculated using Cuffnorm (ver. 2.2.1) [29]. The STV-free (nos. 12, 432, and 433) and STV-infected (nos. 22, 412, and 422) samples produced 17,951,090, 16,284,285, 14,320,826, 19,209,433, 16,789,684, and 19,806,901 reads, respectively. Deep-sequencing data presented in this study were submitted to NCBI GEO (http://www.ncbi.nlm.nih.gov/geo/) and can be retrieved under accession number GSE137303.
Quantitative real-time PCR (qPCR)
cDNA was generated from total RNA using a PrimeScript RT Reagent Kit with gDNA Eraser (Takara), and qPCR was performed using a Thermal Cycler Dice Real Time System with a SYBR Premix Ex Taq II Kit (Takara). Relative transcript levels of the genes encoding ACO1 (1-aminocyclopropane-1-carboxylate [ACC] oxidase 1), NAC1 (NAC domain protein) and EREB (ethylene responsive element binding protein) were normalized to that of elongation factor 1α (EF1α). Primers for qPCR were selected using the Primer3Plus program (http://www.bioinformatics.nl/cgi-bin/primer3plus/primer3plus.cgi/) with Sol Genomics Network (https://solgenomics.net/). The relative copy number of STV was also determined by qPCR. Primers used in this study are listed in Supplemental Table S1.
Small RNA analysis
Raw sRNA-seq data files were obtained as SRA files (GSE23562) from NCBI (https://www.ncbi.nlm.nih.gov/) [30]. The SRA files were converted into fastq files using the SRA Toolkit (http://www.ncbi.nlm.nih.gov/Traces/sra/sra.cgi?view=toolkit_doc). Adapter sequences were removed using the CUTADAPT program [31]. Low-quality reads were removed using the “fastq_quality_trimmer” and “fastq_quality_filter” programs in the FASTX -Toolkit package (http://hannonlab.cshl.edu/fastx_toolkit/), and sequence reads shorter than 15 nt were also removed. Reads were mapped to the tomato genome (Tomato SL2.50 ITAG2.4; Sol Genomic Network https://solgenomics.net/) and STV genome (Florida strain, KX949574) using Bowtie2 (ver. 2.3.3.1) [28] to create two sam files. The data were compared using a python script, and both mapped reads were written into new sam files. After converting these files into bam files, the status of mapped sRNA reads was visualized using Integrative Genomics Viewer [32]. These analyses were performed on the Shirokane4 supercomputer provided by the Super Computer System, Human Genome Center, Institute of Medical Science, University of Tokyo. The small RNA target prediction module psRobot was used for target prediction analysis (http://omicslab.genetics.ac.cn/psRobot/) [33].
Statistical analysis
Student’s paired t-test was used to determine the significance of phenotypic differences between the two lines.
Results
Detection of STV
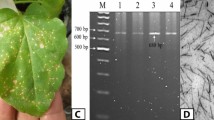
STV, which has been detected in numerous tomato cultivars, carries a single linear dsRNA genome of approximately 3.4 kbp in length [10]. We attempted to detect the dsRNA genome of STV in various tomato cultivars using the cellulose spin-column method [26, 27]. A single dsRNA molecule of approximately 3.4 kbp in length was detected in cultivar M82 (Fig. 1a). Similarly, a linear 3.4-kbp dsRNA was also detected in seedlings generated from commercially obtained seeds of tomato cultivars, such as “Cherry Mate” and “Sweet Hearts”, but not in “Ponderosa”, “Momotaro” or “Microtom”. RT-PCR (reverse transcription PCR) using previously described STV-specific primers [10] was carried out to determine whether this dsRNA was indeed the STV genome. Amplified DNA fragments with an expected size of approximately 440 bp were detected in some cv. M82 plants (Fig. 1b). The nucleotide sequence of the RT-PCR products (accession number LC429302) exhibited highest similarity to the Florida strain of STV from cv. Sweet Hearts (accession number KX949574), with only two base mismatches.
Detection of dsRNA in tomato plants of cv. M82. (a) Agarose gel electrophoresis of dsRNA purified from leaves of four plants. The black arrow indicates dsRNA of approximately 3.4 kbp in length. Molecular weight markers of linear dsDNA are indicated in lane M. (b) Detection of STV dsRNA by RT-PCR. PCR products of the predicted size (440 bp) previously reported by Sabanadzovic et al. [10] were detected in progeny plants no. 2 and no. 4 (lanes 2 and 4) but not in progeny of plants no. 1 and no. 3 (lanes 1 and 3)
Phenotypic differences between STV-free and STV-infected plants
The effect of STV infection on the phenotype of the host tomato plants was previously unclear because although STV was first detected in plants exhibiting symptoms of stunting, fruit discoloration, and reduced fruit size [10, 15,16,17,18,19], it was also detected in healthy plants [22, 23]. In this study, therefore, we identified and compared STV-infected plants (STV+) and STV-free plants (STV-) of cv. M82 to elucidate the association between STV infection and host plant phenotype. STV-infected plants and their fruit exhibited no visible disease symptoms (Figs. 2a and 2b and S1). Three-month-old STV-free plants exhibited a short stature and bushy phenotype, but 3-month-old STV-infected plants were taller than STV-free plants (Figs. 2a and S2). This different phenotype was also observed in the progeny (siblings) of a single STV-infected plant (No. 4 in Table 1, Fig. 2a). Although STV-infected plants produced more fruit than STV-free plants, the difference was not statistically significant (p = 0.06) (Table 1 and Fig. 2c). In addition, the germination rate of seeds harvested from STV-infected plants was higher than that of seeds harvested from STV-free plants (Table 1 and Fig. 2d). These results indicate that STV infection enhances the vigor of host plants, suggesting that the interaction between STV and the host plant is mutualistic.
Phenotypic differences between STV-infected and STV-free tomato plants of cv. M82. (a) STV-infected (nos. 4-1 and 4-2) and STV-free (no. 4-3) plants (approximately 3 months old) that were siblings of cv. M82 plant no. 4 (Table 1). (b) Mature fruit harvested from STV-free (nos. 1 and 3) and STV-infected (nos. 2 and 4) plants. (c) Average number of mature fruits harvested from approximately 5-month-old STV-free and STV-infected plants. Error bars indicate the standard errors of six STV-free or eight STV-infected plants, and the p-value was determined using Student’s t-test. (d) Germination rates of seeds (total 200 seeds) harvested from five STV-infected plants and seeds (total 191 seeds) harvested from four STV-free plants
Previous studies have reported mutualistic interactions between acute viruses and the host plant under conditions of stress, such as virus infection improving the drought tolerance of host plants [34, 35]. Therefore, we exposed STV-free and STV-infected plants to salt stress (100 mM NaCl for 1 week) and monitored the plants for differences in phenotype. Under normal growth conditions, 40-day-old STV-infected plants were indistinguishable from STV-free plants based on appearance, and their fresh weight was slightly lower than that of STV-free plants. However, the fresh weight of STV-infected plants exposed to salt stress was slightly higher than that of STV-free plants, but the difference was not statistically significant (Figs. 3a and b).
Comparison between STV-infected and STV-free plants of cv. M82 under salt stress conditions. (a) Forty-day-old STV-free and STV-infected plants were exposed to salt stress (100 mM NaCl) for 10 days. (b) Average fresh weight of STV-free and STV-infected plants exposed to salt stress. Approximately 40-day-old plants were grown under normal conditions or salt stress. Error bars indicate the standard errors of 15 or 16 plants
In the case of the “Cherry Mate” and “Sweet Hearts” cultivars, STV-infected plants of both cultivars appeared healthy (asymptomatic). Unlike cv. M82, STV-free and STV-infected “Cherry Mate” plants did not exhibit any significant differences in plant architecture or fruit production (Fig. S3).
Downregulation of the expression of genes involved in ethylene biosynthesis and signaling by STV infection
To examine the effect of STV infection on host gene expression, we carried out transcriptome (RNA-seq) analyses using three 1-month-old STV-infected and STV-free cv. M82 plants. In STV-infected plants, the expression of several genes encoding non-coding RNAs and pathogenesis-related proteins was upregulated, but the expression of various genes involved in ethylene biosynthesis and signaling was downregulated (Table 2 and Supplementary Table S2). Among differentially expressed genes between the two lines, we focused on genes involved in ethylene biosynthesis (ACO1, ACC oxidase 1) and ethylene signaling (NAC1 and EREB) because ethylene plays a crucial role in fruit ripening, seed germination, and plant architecture [36, 37]. ACO1 is an important enzyme that is involved in ethylene biosynthesis [38], and NAC1 and EREB encode transcription factors involved in fruit ripening [39] and the ethylene response [40], respectively. qPCR analysis confirmed that the expression of these three genes was downregulated in STV-infected plants (Fig. 4). These results suggest that the beneficial effect of STV infection on fruit production, seed germination, and plant architecture involves downregulation of ethylene biosynthesis and signaling in the host plants.
STV infection downregulates the expression of genes involved in ethylene biosynthesis and ethylene signaling. Relative transcript levels of ACO1, NAC1 and EREB in leaves of 1-month-old STV-free and STV-infected tomato plants of cv. M82 were analyzed by qRT-PCR. ACO1, NAC1 and EREB, which were selected from the results of transcriptome analyses (Tables 2 and S2), encode 1-aminocyclopropane-1-carboxylic acid (ACC) oxidase 1, NAC domain protein, and ethylene-responsive element-binding protein, respectively. Error bars indicate the standard errors of four plants, and the p-values were determined using Student’s t-test
Vertical transmission of STV to the next generation
STV genomic dsRNA was detected in approximately half of the cv. M82 plants (6/11) germinated from original seeds obtained from the National BioResource Project of Japan (first generation) (Table 1). Vertical (seed) transmission of STV was examined by harvesting seeds from self-pollinated fruits of the first generation (nos. 1 through 11 in Table 1) and the second generations (nos. 4-1 through 4-3 in Table 1). The rate of seed transmission of STV from the four infected plants (nos. 2, 4, 7 and 4-2) to the next generation was 86% (30/35, Table 1).
As cv. M82 is the standard cultivar of cultivated tomato for researchers worldwide [24], cv. M82, a wild relative (Solanum pennellii) of cultivated tomato, and introgression lines (ILs) between M82 and S. pennellii were used to analyze the relationship between their small RNA (sRNA) profiles and phenotypes [28]. The raw sRNA datasets of M82, the F1 and F2 hybrids, and ILs analyzed by next-generation sequencing are available. Although the authors of the previous report did not mention STV [30], sRNAs that were apparently derived from STV were identified in available sRNA datasets prepared from cv. M82 (Fig. 5a). sRNAs that were apparently derived from STV were also identified in the F1 and F2 hybrids (Fig. S4), indicating that STV is efficiently transmitted to the next generation via seeds, even in interspecific hybrids.
sRNAs apparently derived from STV. (a) sRNAs that were apparently derived from STV were identified in available sRNA datasets prepared from cv. M82 tomato plant [30]. Screenshot of IGV (Integrative Genomics Viewer) visualization is shown, and the reference sequence is the STV genome of the Florida strain (KX949574). The number of sRNA reads mapped to the STV genome was 2,241, which was approximately 0.07% of the total sRNA reads mapped to the tomato genome (Tomato SL2.50 ITAG2.4). (b) sRNA apparently derived from STV homologous to the host tomato genome
sRNAs derived from STV
sRNA reads that appeared to be derived from STV accounted for only 0.05% to 0.09% of total sRNAs mapped on the tomato genome (Figs. 5a and S3), indicating that the copy number of STV was very low in comparison with that of conventional acute viruses. For instance, the viral small interfering RNAs (vsiRNAs) derived from cucumber mosaic virus (CMV) and turnip mosaic virus (TuMV) reportedly account for 3% to 15% of total sRNAs [41].
As vsiRNAs can regulate host gene expression [42], STV-derived vsiRNAs homologous to their host tomato genes were analyzed (Fig. 5b). Several specific vsiRNAs accumulated, suggesting that these vsiRNAs bind to putative proteins such as Argonautes, which protects them from ribonuclease digestion [43]. The most abundant vsiRNA was mapped to 1,048 nt from the 5’ end of the positive-sense strand of the STV genome. The sRNA-target prediction module psRobot predicted the putative target to be a gene encoding an ethylene insensitive 3 (EIN3) family protein (Solyc04g054840) (Fig. S4c) [33]. EIN3 is a central transcription factor controlling the expression of various ethylene-responsive genes, including transcription factors (e.g., ethylene response factors) [44]. However, no transcripts of Solyc04g054840 were detected in 1-month-old STV-infected and STV-free plants by either RNA-seq (Table S2) or qPCR analysis.
Recently, vsiRNAs derived from STV were detected in healthy tomato plants of cv. Merlice in a study using sRNA-seq (virome) analysis, and their accumulation was reportedly less than that of vsiRNAs derived from PepMV in the same tomato plant [22]. The accumulation pattern of STV-derived vsiRNAs in cv. Merlice plants differed from that in cv. M82 plants, as shown in Figures 5a and S4; for instance, the most abundant vsiRNA in cv. Merlice was mapped to 920 nt from the 5’ end of the positive-sense strand of the STV genome [22]. psRobot predicted the putative target to be a gene encoding nuclear transcription factor Y, subunit C-1 (Solyc03g110860) [34]. Differences in the nucleotide sequences of the two STV strains detected in cvs. M82 and Merlice (48 base substitutions in the full-length genome, approximately 1.4%) may be a cause of the different accumulation patterns of STV-derived vsiRNAs in these two cultivars.
Age-dependent increase in STV copy number
As a previous report indicated that the abundance of STV-derived vsiRNAs in cv. Merlice is higher in 6.5-month-old plants than in 1.5-month-old plants [22], the copy number of STV in cv. M82 was examined. In leaves of 3-month-old plants, the STV copy number was 4-fold higher than that in leaves of 1-month-old plants, indicating that STV accumulation increases as the plant matures (Fig. 6). If the EIN3 transcript is regulated by STV-derived vsiRNA, the EIN3 transcript level should be affected more in mature plants and fruit than in seedlings. However, no EIN3 transcripts were detected in 3-month-old leaves.
Age-dependent increase in STV copy number. (a) Agarose gel electrophoresis of dsRNAs purified from leaves of two 1-month-old (lanes 1 and 2) and two 3-month-old (lanes 3 and 4) plants of cv. M82. Black arrow indicates dsRNAs of approximately 3.4 kbp in length. Molecular weight markers of linear dsDNAs are indicated in lane M. (b) Relative levels of STV genomic RNA in leaves of 1-month-old (1 m) and 3-month-old (3 m) tomato plants of cv. M82 were analyzed by qRT-PCR. Error bars indicate the standard errors of three plants
Discussion
In order to enhance our understanding of the interaction between host plant and persistent viruses such as STV, we profiled the growth and molecular phenotypes of STV-free and STV-infected plants. STV infection did not negatively affect the growth or fruit production of host plants of the commercial cultivar “Cherry Mate” (Fig. S2). Juvenile cv. M82 plants infected with STV could not be distinguished from STV-free plants based on appearance (Fig. 3). In contrast, STV-free adult plants exhibited a short and bushy phenotype, but STV-infected adult plants exhibited increased height (Figs. 2a and S1). STV-infected plants produced more fruit than STV-free plants (Fig. 2c), and the rate of germination of seeds harvested from STV-infected plants was higher than that of seeds harvested from STV-free plants (Fig. 2d). The phenotypic differences between STV-free and STV-infected plants were also observed in the progeny (siblings) of a single STV-infected plant (Fig. 2a).
Mutualistic interactions between various acute plant viruses and host plants have been reported under certain stress conditions, such as improved drought tolerance of virus-infected host plants [34, 35, 45]. A recent report similarly described how persistent virus infection can affect seed germination; the germination of seeds of the common bean (Phaseolus vulgaris) cultivar Black Turtle Soup infected with two endornaviruses (Phaseolus vulgaris endornavirus 1 and 2) of the family Endornaviridae was more rapid than that of seeds from virus-free plants [46]. Another persistent virus, beet cryptic virus, which is a member of the family Partitiviridae, has been reported to prevent yield losses caused by drought conditions in infected Beta vulgaris plants [47]. The interaction between STV and cv. M82 is also mutualistic.
Ethylene is a gaseous plant hormone that affects many aspects of vegetative growth, although it is best known as a ripening hormone [36, 37]. Ethylene is a major determinant of plant architecture, and, except in flooding-tolerant plants, it inhibits stem elongation [36]. Treatment of Arabidopsis plants with ethylene results in a stunted and thick inflorescence stem [48]. Judging from the plant architecture of cv. M82 (Figs. 2a and S2), the endogenous ethylene content in STV-free plants appeared to be higher than that in STV-infected plants. Transcriptome (Table 2 and S3) and qPCR (Fig. 4) analyses showed that the expression of genes encoding the key enzyme for ethylene biosynthesis (ACO1) and transcription factors for the ethylene signal transduction pathway (NAC1 and EREB) was downregulated in STV-infected plants. As the expression of ACO1 is likely to be suppressed by persistent STV infection, the concentration of ethylene in STV-infected plants might be lower, causing a decrease in the expression of the ethylene-responsive gene product (EREB) in these plants. Consequently, the effect on the height of STV-infected plants may have been alleviated. Furthermore, this change in the expression of genes involved in ethylene signaling resulting from STV infection is likely to be associated with STV-infected plants producing more fruit than STV-free plants and the higher germination rate of seeds from STV-infected plants compared with seeds from STV-free plants.
Endornaviruses are typical persistent viruses that asymptomatically infect a variety of plants [9]. Analysis of differential gene expression based on RNA-seq data from endornavirus-infected and endornavirus-free common bean has demonstrated that oxidation-reduction pathways are the primary processes affected by endornavirus infection [49]. Therefore, downregulation of genes involved in ethylene biosynthesis and signaling is not commonly associated with persistent virus infection; however, it is particularly associated with STV infection of cv. M82 tomato plants.
Although there are few reports to date of developmental-stage- and/or organ-specific regulation of the copy number of persistent viruses, the copy number of Oryza sativa endornavirus in pollen grains is reportedly 50-fold higher than in seedlings [50]. Here, we demonstrated that the STV copy number increases as plants mature (Fig. 6), suggesting that STV-derived vsiRNAs accumulate more in mature fruit-producing plants than in seedlings. This age-dependent increase in STV copy number is likely to be associated with the effect of STV-infection on fruit production and seed germination. Turco et al. reported that STV-derived vsiRNAs accumulated more in 6.5-month-old plants of cv. Merlice coinfected with two PepMV strains (CH2 and LP) than in 1.5-month-old plants infected with only one PepMV strain (CH2), suggesting that coinfection with two virus strains activated STV replication [22]. However, the age-dependent increase in STV copy number observed in our study (Fig. 6) suggests a different cause for the increased accumulation of STV-derived vsiRNAs in 6.5-month-old plants coinfected with two viruses.
If STV-derived vsiRNAs affect gene expression in the host plant, differential accumulation of vsiRNAs must also differentially affect the phenotypes of different host plants. Indeed, in this study, STV-infected cv. M82 plants were phenotypically different from STV-free plants of the same cultivar (Fig. 2), but we could not distinguish between STV-infected and STV-free plants of cv. Cherry Mate based on appearance alone (Fig. S3). Generally, acute viruses such as CMV and TuMV differentially affect phenotypes (symptoms) of different host cultivars (or ecotypes) of the same plant species; that is, these acute viruses include strains exhibiting both high and low virulence [51]. Even among persistent dsRNA viruses, one strain might affect the host cultivar(s), whereas another strain might not. The M82 strain of STV affected the host’s phenotype, but the Cherry Mate strain of STV might not affect host plants.
Although attempts to transmit STV via mechanical and graft-transmission methods have been reported, horizontal transmission of STV has never been demonstrated [10]. Turco et al. recently demonstrated that STV is not co-transmitted horizontally with PepMV in tomato plants [22]. Therefore, vertical transmission may be the only strategy used by STV to maintain and spread among host tomato plants. However, we demonstrated in the present study that the seed transmission rate of STV was not 100% in cv. M82 and that the seed transmission rate of STV from four infected plants to progeny plants was 86% (30/35, Table 1). In addition, only half of the original cv. M82 seeds (6/11) provided by the NBRP harbored STV. However, we often found STV in tomato cultivars. These conflicting results suggest at least two hypothetical mechanisms by which STV is persistently maintained in tomato plants: 1) STV may be transmitted horizontally via unknown mechanisms, and/or 2) STV-infected tomato plants might be artificially selected by breeders and/or farmers because they exhibit better crop traits than STV-free plants. STV-infected tomato plants in particular are likely to be positively selected due to their higher germination rate. In endornavirus-infected common bean plants, the higher seed germination rate of seeds from virus-infected plants has been described an important agronomic trait [46]. Acute viruses cause disease symptoms in the host plants that result in decreased agricultural yield. In contrast, persistent viruses asymptomatically infect the host plants, so they are likely to be neutral or mutualistic symbionts [45, 52].
Conclusions
STV, a putative persistent dsRNA virus, often infects many tomato cultivars, such as cv. M82. As the effect of STV infection on host plants was unclear, we compared STV-free and STV-infected plants of cv. M82. STV infection positively affected the germination, plant establishment, and reproductive ability of cv. M82 host plants. Both the virus and the STV-associated phenotype were transmitted to the next generation, indicating that STV caused the observed growth improvement. These results suggest that the interaction between STV and host plants is mutualistic.
References
Natsuaki T, Yamashita S, Doi Y, Okuda S, Teranaka M (1983) Radish yellow edge virus, a seed-borne virus with double-stranded RNA, of a possible new group. Ann Phytopathol Soc Jpn 49:593–599. https://doi.org/10.3186/jjphytopath.49.593
Dodds JA, Morris TJ, Jordan RL (1984) Plant viral double-stranded RNA. Annu Rev Phytopathol 22:151–168. https://doi.org/10.1146/annurev.py.22.090184.001055
Boccardo G, Lisa V, Luisini E, Milne RG (1987) Cryptic plant viruses. Adv Virus Res 32:171–214. https://doi.org/10.1016/S0065-3527(08)60477-7
Brown GG, Finnegan PM (1989) RNA plasmids. Int Rev Cytol 117:1–56. https://doi.org/10.1016/S0074-7696(08)61333-9
Gibbs MJ, Koga R, Moriyama H, Pfeiffer P, Fukuhara T (2000) Phylogenetic analysis of some large double-stranded RNA replicons from plants suggests they evolved from a defective single-stranded RNA virus. J Gen Virol 81:227–233. https://doi.org/10.1099/0022-1317-81-1-227
Fukuhara T, Koga R, Aoki N, Yuki C, Yamamoto N, Oyama N, Udagawa T, Horiuchi H, Miyazaki S, Higashi Y, Takeshita M, Ikeda K, Arakawa M, Matsumoto N, Moriyama H (2006) The wide distribution of endornaviruses, large double-stranded RNA replicons with plasmid-like properties. Arch Virol 151:995–1002. https://doi.org/10.1007/s00705-005-0688-5
Roossinck MJ (2010) Lifestyles of plant viruses. Philos Trans R Soc B 365:1899–1905. https://doi.org/10.1098/rstb.2010.0057
Ghabrial SA, Nibert ML, Maiss E, Lesker T, Baker TS, Tao YJ (2012) Family Partitiviridae. In: King AMQ, Adams MJ, Carstens EB, Lefkowitz EJ (eds) Virus taxonomy: ninth report of the International Committee on Taxonomy of Viruses. Elsevier, San Diego, pp 523–534
Fukuhara T, Gibbs MJ (2012) Family Endornaviridae. In: King AMQ, Adams MJ, Carstens EB, Lefkowitz EJ (eds) Virus taxonomy: ninth report of the International Committee on Taxonomy of Viruses. Elsevier, San Diego, pp 519–521
Sabanadzovic S, Valverde RA, Brown JK, Martin RR, Tzanetakis IE (2009) Southern tomato virus: the link between the families Totiviridae and Partitiviridae. Virus Res 140:130–137. https://doi.org/10.1016/j.virusres.2008.11.018
Wickner RB, Ghabrial SA, Nibert ML, Patterson JL, Wang CC (2012) Family Totiviridae. In: King AMQ, Adams MJ, Carstens EB, Lefkowitz EJ (eds) Virus taxonomy: ninth report of the International Committee on Taxonomy of Viruses. Elsevier, San Diego, pp 639–650
Tzanetakis IE, Sabanadzovic S (2013) Establishment of the family Amalgaviridae, the genus Amalgavirus and inclusion of four species in the genus. https://talk.ictvonline.org/files/ictv_official_taxonomy_updates_since_the_8th_report/m/plant-official/4828
Krupovic M, Dolja VV, Koonin EV (2015) Plant viruses of the Amalgaviridae family evolved via recombination between viruses with double-stranded and negative-strand RNA genomes. Biol Direct 10:12. https://doi.org/10.1186/s13062-015-0047-8
Nibert ML, Pyle JD, Firth AE (2016) A +1 ribosomal frameshifting motif prevalent among plant amalgaviruses. Virology 498:201–208. https://doi.org/10.1016/j.virol.2016.07.002
Candresse T, Marais A, Faure C (2013) First report of Southern tomato virus on tomatoes in southwest France. Plant Dis 97:1124. https://doi.org/10.1094/PDIS-01-13-0017-PDN
Iacono G, Hernandez-Llopis D, Alfaro-Fernandez A, Davino M, Font M, Panno S, Galipenso L, Rubio L, Davino S (2015) First report of Southern tomato virus in tomato crops in Italy. New Dis Rep 32:27. https://doi.org/10.5197/j.2044-0588.2015.032.027
Padmanabhan C, Zheng Y, Li R, Sun SE, Zhang D, Liu Y, Fei Z, Ling KS (2015) Complete genome sequence of southern tomato virus identified in China using next-generation sequencing. Genome Announc 3:e01266–15. https://doi.org/10.1128/genomeA.01226-15
Padmanabhan C, Zheng Y, Li R, Fei Z, Ling KS (2015) Complete genome sequence of southern tomato virus naturally infecting tomatoes in Bangladesh. Genome Announc 3:e01522–15. https://doi.org/10.1128/genomeA.01522-15
Verbeek M, Dullemans A, Espino A, Botella M, Alfaro-Fernández A, Font M (2015) First report of Southern tomato virus in tomato in the Canary Islands, Spain. J Plant Pathol 97:392. https://doi.org/10.4454/JPP.V97I2.038
Elvira-González L, Puchades AV, Carpino C, Alfaro-Fernandez A, Font-San-Ambrosio MI, Rubio L, Galipienso L (2017) Fast detection of Southern tomato virus by one-step transcription loop-mediated isothermal amplification (RT-LAMP). J Virol Methods 241:11–14. https://doi.org/10.1016/j.jviromet.2016.12.004
Puchades AV, Carpino C, Alfaro-Fernandez A, Font-San-Ambrosio MI, Davino S, Guerri J, Rubio L, Galipienso L (2017) Detection of Southern tomato virus by molecular hybridization. Ann Appl Biol 171:172–178. https://doi.org/10.1111/aab.12367
Turco S, Golyaev V, Seguin J, Gilli C, Farinelli L, Boller T, Schumpp O, Pooggin MM (2018) Small RNA-omics for virome reconstruction and antiviral defense characterization in mixed infections of cultivated Solanum plants. Mol Plant Microbe Interact 31:707–723. https://doi.org/10.1094/MPMI-12-17-0301-R
Alcalá-Briseño RI, Coşkan S, Londoño MA, Polston JE (2017) Genome sequence of Southern tomato virus in asymptomatic tomato ‘Sweet Hearts’. Genome Announc 5:e01374–16. https://doi.org/10.1128/genomeA.01374-16
Menda N, Semel Y, Peled D, Eshed Y, Zamir D (2004) In silico screening of a saturated mutation library of tomato. Plant J 38:861–872. https://doi.org/10.1111/j.1365-313X.2004.02088.x
Galpaz N, Wang O, Menda N, Zamir D, Hirschberg J (2008) Abscisic acid deficiency in the tomato mutant high pigment 3 leading to increased plastid number and higher fruit lycopene content. Plant J 53:717–730. https://doi.org/10.1111/j.1365-313X.2007.03362.x
Morris TJ, Dodds JA (1979) Isolation and analysis of double-stranded RNA from virus-infected plant and fungal tissue. Phytopathology 69:854–858. https://doi.org/10.1094/Phyto-69-854
Okada R, Kiyota E, Moriyama H, Fukuhara T, Natsuaki T (2015) A simple and rapid method for viral dsRNA isolation from plant and fungal tissue. J Gen Plant Pathol 81:103–107. https://doi.org/10.1007/s10327-014-0575-6
Langmead B, Salzberg S (2012) Fast gapped-read alignment with Bowtie 2. Nat Methods 9:357–359. https://doi.org/10.1038/nmeth.1923
Trapnell C, Roberts A, Goff L, Pertea G, Kim D, Kelley DR, Pimentel H, Salzberg SL, Rinn JL, Pachter L (2012) Differential gene and transcript expression analysis of RNA-seq experiments with TopHat and Cufflinks. Nat Protoc 7:562–578. https://doi.org/10.1038/nprot.2012.016
Shivaprasad PV, Dunn RM, Santos BA, Bassett A, Baulcombe DC (2012) Extraordinary transgressive phenotypes of hybrid tomato are influenced by epigenetics and small silencing RNAs. EMBO J 31:257–266. https://doi.org/10.1038/emboj.2011.458
Martin M (2011) Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet J 17:10–12. https://doi.org/10.14806/ej.17.1.200
Robinson JT, Thorvaldsdóttir H, Winckler W, Guttman M, Lander ES, Getz G, Mesirov JP (2011) Integrative genomics viewer. Nat Biotechnol 29:24–26. https://doi.org/10.1038/nbt.1754
Wu HJ, Ma YK, Chen T, Wang M, Wang XJ (2012) PsRobot: a web-based plant small RNA meta-analysis toolbox. Nucleic Acids Res 40:W22–W28. https://doi.org/10.1093/nar/gks554
Xu P, Chen F, Mannas JP, Feldman T, Sumner LW, Roossinck MJ (2008) Virus infection improves drought tolerance. New Phytol 180:911–921. https://doi.org/10.1111/j.1469-8137.2008.02627.x
Westwood JH, McCann L, Naish M, Dixon H, Murphy AM, Stancombe MA, Bennett MH, Powell G, Webb AA, Carr JP (2013) A viral RNA silencing suppressor interferes with abscisic acid-mediated signalling and induces drought tolerance in Arabidopsis thaliana. Mol Plant Pathol 14:158–170. https://doi.org/10.1111/j.1364-3703.2012.00840.x
Abeles S, Morgan PW, Saltveit ME (1992) Ethylene in plant biology, 2nd edn. Academic Press, San Diego. https://doi.org/10.1016/C2009-0-03226-7
Smalle J, Van Der Straeten D (1997) Ethylene and vegetative development. Physiol Plant 100:593–605. https://doi.org/10.1111/j.1399-3054.1997.tb03065.x
Jafari Z, Haddad R, Hosseini R, Garoosi G (2013) Cloning, identification and expression analysis of ACC oxidase gene involved in ethylene production pathway. Mol Biol Rep 40:1341–1350. https://doi.org/10.1007/s11033-012-2178-7
Meng C, Yang D, Ma X, Zhao W, Liang X, Ma N, Meng Q (2016) Suppression of tomato SlNAC1 transcription factor delays fruit ripening. J Plant Physiol 193:88–96. https://doi.org/10.1016/j.jplph.2016.01.014
Zhang H, Zhang D, Chen J, Yang Y, Huang Z, Huang D, Wang XC, Huang R (2004) Tomato stress-responsive factor TSRF1 interacts with ethylene responsive element GCC box and regulates pathogen resistance to Ralstonia solanacearum. Plant Mol Biol 55:825–834. https://doi.org/10.1007/s11103-004-2140-8
Donaire L, Wang Y, Gonzalez-Ibeas D, Mayer KF, Aranda MA, Llave C (2009) Deep-sequencing of plant viral small RNAs reveals effective and widespread targeting of viral genomes. Virology 392:203–214. https://doi.org/10.1016/j.virol.2009.07.005
Ramesh SV, Williams S, Kappagantu M, Mitter N, Pappu HR (2017) Transcriptome-wide identification of host genes targeted by tomato spotted wilt virus-derived small interfering RNAs. Virus Res 238:13–23. https://doi.org/10.1016/j.virusres.2017.05.014
Mi S, Cai T, Hu Y, Chen Y, Hodges E, Ni F, Wu L, Li S, Zhou H, Long C, Chen S, Hannon GJ, Qi Y (2008) Sorting of small RNAs into Arabidopsis argonaute complexes is directed by the 5’ terminal nucleotide. Cell 133:116–127. https://doi.org/10.1016/j.cell.2008.02.034
Müller M, Munné-Bosch S (2015) Ethylene response factors: a key regulatory hub in hormone and stress signaling. Plant Physiol 169:32–41. https://doi.org/10.1104/pp.15.00677
Roossinck MJ (2011) The good viruses: viral mutualistic symbioses. Nat Rev Microbiol 9:99–108. https://doi.org/10.1038/nrmicro2491
Khankhum S, Valverde RA (2018) Physiological traits of endornavirus-infected and endornavirus-free common bean (Phaseolus vulgaris) cv Black Turtle Soup. Arch Virol 163:1051–1056. https://doi.org/10.1007/s00705-018-3702-4
Xie WS, Antoniw JF, White RF, Jolliffee TH (1994) Effects of beet cryptic virus infection on sugar beet in field trials. Ann Appl Biol 124:451–459. https://doi.org/10.1111/j.1744-7348.1994.tb04150.x
Kieber JJ, Rothenberg M, Roman G, Feldmann KA, Ecker JR (1993) Ctr1, a negative regulator of the ethylene response pathway in Arabidopsis, encodes a member of the Raf family of protein kinases. Cell 72:427–441. https://doi.org/10.1016/0092-8674(93)90119-B
Khankhum S, Sela N, Osorno JM, Valverde RA (2016) RNAseq analysis of endornavirus-infected vs. endornavirus-free common bean (Phaseolus vulgaris) cultivar Black Turtle Soup. Front Microbiol 7:1905. https://doi.org/10.3389/fmicb.2016.01905
Moriyama H, Horiuchi H, Koga R, Fukuhara T (1999) Molecular characterization of two endogenous double-stranded RNAs in rice and their inheritance by interspecific hybrids. J Biol Chem 274:6882–6888. https://doi.org/10.1074/jbc.274.11.6882
Read AF (1994) The evolution of virulence. Trends Microbiol 2:73–76. https://doi.org/10.1016/0966-842X(94)90537-1
Roossinck MJ (2015) Plants, viruses and the environment: ecology and mutualism. Virology 479–480:271–277. https://doi.org/10.1016/j.virol.2015.03.041
Acknowledgements
We thank Dr. Tomohide Natsuaki, Utsunomiya University, for valuable discussions, and the National BioResource Project of Japan, Tsukuba University, for providing tomato seeds. We acknowledge support received from the Gene Research Center at Tokyo University of Agriculture and Technology, the NGS core facility of the Genome Information Research Center at the Research Institute for Microbial Diseases of Osaka University, and the Human Genome Center at the Institute of Medical Science of the University of Tokyo.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflicts of interest.
Funding
This work was supported by the Ministry of Education, Culture, Sports, Science and Technology (MEXT) of Japan (Scientific Research on Innovative Areas [nos. 16H06435, 16H06429, and 16H21723] to T.F. and H.T.) and the Global Innovation Research Organization of Tokyo University of Agriculture and Technology (to T.F.), and the Japan Society for the Promotion of Science (JSPS) through the JSPS Core-to-Core Program (Advanced Research Networks) entitled “Establishment of international agricultural immunology research-core for a quantum improvement in food safety” (to H.T.).
Additional information
Handling Editor: Jesús Navas-Castillo.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Fukuhara, T., Tabara, M., Koiwa, H. et al. Effect of asymptomatic infection with southern tomato virus on tomato plants. Arch Virol 165, 11–20 (2020). https://doi.org/10.1007/s00705-019-04436-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00705-019-04436-1