Abstract
Kaposi’s sarcoma-associated herpes virus (KSHV) is a gammaherpesvirus associated with Kaposi’s sarcoma and various lymphoproliferative diseases. Epithelial-to-mesenchymal transition (EMT) is an important step in the metastasis of cancer cells. Previous studies have shown an important role for EMT markers in B-cell malignancies. In the present study, we investigated the role of the KSHV latent protein LANA in the progression of EMT. Our data suggest that expression of LANA results in an increase in the migration and invasion potential of cancer cells, which is concurrent with modulation of transcriptional regulation and protein expression of several cellular genes associated with EMT. LANA expression results in upregulation of the cellular intermediate filament protein vimentin and transcription factor TCF8/ZEB1 and downregulation of tight junction protein ZO1 and adhesion protein E-cadherin. LANA co-localizes with TCF8/ZEB1, a major contributor in EMT, further suggesting an important role for LANA in epithelial-to-mesenchymal transition of KSHV-infected cancer cells.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Metastasis is one of the most important causes of cancer-related deaths [1]. Several studies have shown that epithelial-mesenchymal transition (EMT) of cancer cells, which is characterized by a switch from a polarized, epithelial phenotype to a highly motile fibroblastoid or mesenchymal phenotype is the central mechanism leading to cancer invasiveness and metastasis [2]. With currently available cancer treatment, the survival rate for certain cancer types has improved; however, there is still a long way to go until a definitive cure is found. Recent studies and advancement in our knowledge about cancer have suggested that an approach to control the metastasis of cancer cells might prove to be more effective than pushing to find a cure [3]. To develop such strategies, understanding the intricacies of intracellular processes and signaling pathways that are important for EMT could prove to be a crucial step.
Metastasis is a multi-step process [4]. The transition of cancer cells from the epithelial to mesenchymal phenotype is followed by their entry into the circulatory system after detachment from the primary site. In a later stage, they extravasate from the capillaries and establish multiple metastatic lesions in distant organs [5]. The process of EMT is modulated by the activity of many transcriptional factors, resulting in change in expression levels and/or reorganization of many cytoskeleton proteins [6]. Some of the common transcriptional factors that play a critical role in EMT include β-catenin, Snail, Slug, Twist, and transcription factor 8/Zinc finger E-box binding homeobox 1(TCF8/ZEB1) [7]. Other proteins that are known to play an essential role in EMT include N-cadherin, E-cadherin, claudin, and zonula occludens-1 (ZO-1) [8]. The changes in expression levels and intracellular localization of these molecular markers are a defining feature of EMT. Loss of cellular matrix proteins such as E-cadherin and β-catenin and upregulation of N-cadherin are commonly observed during EMT [9]. An increase in vimentin, an intermediate filament of mesenchymal origin, and overexpression of TCF8/ZEB-1, which is a transcriptional activator of many mesenchymal genes, are also indicators of EMT. Degradation of tight-junction proteins including zona occluding (ZO-1) and claudin is one of the initial events in this process. Both Snail and Slug are known to repress cellular matrix protein E-cadherin [10, 11], as well as transcriptional repression of several cell adhesion molecules including integrins by binding to their promoter sequences [12]. TCF8/ZEB-1 also downregulates transcription of E-cadherin [13]. It has been reported previously that upregulation of β-catenin expression and its nuclear localization are associated with the progression of cancer [14]. The β-catenin protein is an important downstream effector in the Wnt signaling pathway [15]. β-catenin has also been reported to activate Slug (SNAI2), which is a transcriptional repressor belonging to the Snail family of zinc finger transcription factors [16]. Slug also functions as a transcriptional repressor for E-cadherin, as it binds to E-cadherin’s promoter during early stages of development [17, 18]. Upregulation of β-catenin has been reported to result in an increase in levels of Slug and Snail, resulting in downregulation of E-cadherin and other cell adhesion molecules, thereby promoting the process of EMT [19].
Many studies have previously shown that viral antigens can have an important role in the modulation of metastasis of cancer cells [20,21,22,23]. Epstein-Barr virus (EBV) is a gammaherpesvirus that is associated with many human cancers. In a previous study, we have shown the effect of EBV nuclear antigens EBNA1 and EBNA3C on EMT of cancer cells. Expression of both of these viral latent proteins results in a reduction of expression of intracellular adhesion molecules and increased motility and invasiveness of cancer cells [24]. Kaposi’s sarcoma-associated herpesvirus (KSHV) is another gammaherpesvirus that is also associated with many malignancies and is also referred to as human herpesvirus 8 (HHV8). It has been reported to be associated as an etiologic agent of Kaposi’s sarcoma (KS), primary effusion lymphoma (PEL), and a variety of multicentric Castleman’s disease (MCD) [25]. KSHV has also been classified as a direct carcinogen by the International Agency for Research on Cancer [26]. Previous studies have shown an important role for EMT markers during B-cell malignancies. An EMT-like process has been shown to occur in B-cell lymphomas [27]. The EMT activator ZEB1 has been shown to promote tumor growth in mantle cell lymphoma, which is also a B-cell malignancy [28]. In a separate study, it has been shown that KSHV-encoded LANA can mediate regulation of Par3, which is a crucial protein in determining cell polarity, contributing to B-cell proliferation [29]. Another previous study has shown that KSHV promotes EndMT (endothelial-to-mesenchymal transition) through the Notch-dependent signaling pathway [30, 31]. Therefore it is important to study various EMT markers in KSHV-induced B-cell lymphoma and the role of specific viral proteins in the modulation of these EMT markers.
Several KSHV-encoded proteins expressed during latency have been found to play a role in the stable persistence of its genome, which is tethered to a host chromosome during latency. These include latency-associated nuclear antigen (LANA1; ORF73), viral cyclin (ORF72), kaposin A, VIRF1, IL-1 gamma-converting enzyme inhibitory protein (vFLIP; ORF 71), and LANA 2 [32]. Of these, LANA1 or LANA (latency-associated nuclear antigen) has been detected in the majority of Kaposi’s sarcoma lesions. It is a multifunctional protein that is important for transcription, replication, and anchoring of the viral genome to host chromatin during interphase and is critical for virus persistence in the infected host [33]. It has been shown to inhibit p53-mediated apoptosis [34, 35] and dysregulation of β-catenin and the Wnt signaling pathway [36]. The amino-terminal portion of LANA (LANA-N) has many essential domains, including a proline-rich domain, an acid-rich domain, a glutamine-rich domain, and a potential nuclear localization domain [37]. LANA-N binds to host chromatin through nucleosomal histones and tethers the viral genome bound to the carboxy terminus of LANA (LANA-C) to the host chromosomes. LANA-C is essential for its binding to a latent binding site through its DNA-binding domain [38]. In this manuscript, we present data that suggest that the KSHV latent antigen LANA plays a role in modulation of host cellular proteins that are important for the epithelial-to-mesenchymal transition of cancer cells.
Materials and methods
Plasmids, cells and culturing conditions
The HEK-293T, HeLa, and human breast carcinoma MDA-MB-231T cell lines (gifted by Prof. Erle S. Robertson, University of Pennsylvania) were used in the present study. Cell lines were maintained in DMEM (LONZA, Basel, Switzerland) supplemented with 5% fetal bovine serum (Thermo Fisher Scientific, Waltham, MA USA). The expression constructs pA3F-LANA-Full, pA3F-LANA-N (1-340) and pA3F-LANA-C(945-1162) were a gift from Dr. Subhash C. Verma (University of Nevada, Reno) and were used to transiently overexpress the LANA protein in cancer cells. The cells were transfected using polyethylenimine (PEI) (Polyscience, Inc. Warrington, PA, USA) as per the protocol described earlier [39].
Real-time PCR analysis of the levels of EMT marker cellular gene transcripts
Total cellular RNA was isolated using an RNA isolation kit (Invitrogen, Carlsbad, USA), followed by cDNA preparation using a Superscript First-Strand Reverse Transcription Kit (Invitrogen, Carlsbad, USA) as per the manufacturer’s instructions. Specific primers (Table 1) described in earlier studies [24] were used for real-time PCR. Three independent experiments were performed, and data were analyzed using the ΔΔCt method for relative quantification after normalization using GAPDH as an internal control.
SDS-PAGE and Western blot
Whole-cell lysates were obtained by lysing the cells 48 hours post-transfection, and the amount of total protein was quantified using Bio-Rad Protein Assay Dye Reagent Concentrate (Bio-Rad Laboratories, USA). Approximately 100 micrograms of total cellular protein was resolved by SDS PAGE and transferred to a PVDF membrane. LANA-Full, LANA-N, and LANA-C were expressed as FLAG-tagged proteins and were probed using anti-FLAG primary antibody (Sigma, Aldrich, St. Louis, Missouri, USA). Cellular EMT markers were probed using specific primary antibodies against each protein (Cell Signaling Technology, Danvers, MA, US). GADPH was used as a loading control for normalization. Three independent experiments were performed, and images were analyzed using ImageJ software for quantification of signals [40]. The images presented in all figures are representative of three independent experiments.
Immunofluorescence assays
Approximately 1 × 105 MDA-MB-231T cells were seeded on coverslips placed in wells in 6-well plates and allowed to grow for at least 24 hours. The next day, the cells were transfected with LANA expression plasmids (pA3F-LANA-Full, pA3F-LANA-N and pA3F-LANA-C). About 24 hours post-transfection, the culture medium was removed, and the coverslips were washed with phosphate-buffered saline (PBS). The cells were fixed by dipping the coverslips into a 1:1 mixture of methanol and acetone. Subsequently, the cells were blocked using 1% fish skin gelatin in PBS for 1 hour. The cells were then incubated overnight with the appropriate primary antibody (approximately 1-2 µg of antibody in 100 µl of PBS). The next morning, the coverslips were washed three times with PBS and incubated with the secondary antibodies Alexa Fluor 488 goat anti-rabbit and Alexa Fluor 594 goat anti-mouse (1:1000 dilutions). Slides were visualized using a Zeiss LSM 510 scanning fluorescence microscope. Three independent experiments were performed, and data were analyzed using ZEN software (Carl Zeiss AG, Oberkochen, Germany). The bar diagrams presented below each figure represent the average amount of fluorescence signal detected per cell from three independent experiments. On average, about 15-20 cells randomly chosen from 5-6 microscopic fields were analyzed for quantitation in each experiment.
Scratch assay (wound-healing assay)
Approximately 1 × 105 231T cells were seeded in each well of a 6-well plate and incubated overnight. The next day, the cells were transfected with LANA expression plasmids (pA3F-LANA-Full, pA3F-LANA-N and pA3F-LANA-C). After 24 hours, one or two scratches were made in the cell monolayer using a 1-ml micropipette tip, and the cells were observed daily for healing of the wound for the next 2 days, using a Nikon TS-100F Inverted Trinocular Research Microscope (Nikon Corporation, Minato, Tokyo, Japan). All of the experiments were repeated three times. The percentage of wounds healed over a period of time was calculated and the data plotted.
Transwell cell invasion assay
Approximately 1 × 106 231T cells were transfected with LANA expression plasmids (pA3F-LANA-Full, pA3F-LANA-N and pA3F-LANA-C) and grown for 24 hours. The cells were harvested 24 hours post-transfection, and 0.5 × 106 cells were then added to the top chamber of a transwell suspended in 200 μl of serum-free medium (Corning, New York, USA). About 750 µl of medium supplemented with 5% serum was added to the bottom half of the well to act as a chemoattractant, and the whole setup was further incubated for 24 hours. The transwells were removed from 6-well plates, and the cells that migrated in Matrigel were visualized by staining with crystal violet dye and counted visually under a microscope. The images were captured using a Nikon TS-100F Inverted Trinocular Research Microscope with an attached Nikon scientific-grade CMOS camera. Three individual experiments were performed with appropriate controls.
Results
Expression of KSHV latent antigen LANA results in modulation of transcription of cellular genes that are critical for epithelial-to-mesenchymal transition
In this study, we tested the specific role of KSHV latent protein LANA-Full in the modulation of EMT-specific genes. We compared the relative number of mRNA transcript copies of nine of the EMT marker genes in MDA-MB-231T cells expressing LANA-Full to that of empty-vector-transfected control cells. 231T cells are of human breast cancer origin and are commonly used in studies in which the role of proteins in cellular migration or invasive potential is investigated [39]. 231T cells were transiently transfected, resulting in expression of LANA-Full in approximately 50% of the cells. The transient overexpression of LANA-Full resulted in a significant increase (p < 0.01) in the transcript levels of Slug (9-fold; Fig. 1A), Snail (15-fold; Fig. 1B), and β-catenin (approximately 12-fold; Fig.1C) in comparison to empty-vector-transfected control cells. Our data show that the expression of LANA-Full resulted in transcriptional downregulation of the tight junction proteins ZO-1 and claudin. The relative mRNA levels of ZO-1 and claudin were decreased by up to 8-fold, which was statistically significant (p < 0.01) in comparison to their levels in empty-vector-transfected control cells (Fig.1E and F). It has been shown previously that the disruption or diffusion of tight junction proteins is one of the earliest events during EMT of cells [41]. Our data clearly indicate that the expression of LANA is coincident with changes in the transcription profile of cellular genes, similar to what is characteristic of initiation of EMT in cancer cells.
Cancer cells expressing KSHV LANA protein differentially modulate cellular genes essential for EMT at both the transcriptional and protein-expression level. MDA-MB-231T cells were transfected with pA3F LANA-Full (L-F), pA3F LANA-N (L-N) and pA3F LANA-C (L-C) along with empty vector as a control (C). Cells were harvested 24 h post-transfection, and total RNA and total protein were extracted separately. The mRNA transcript for Slug (panel A), Snail (panel B), β-catenin (panel C), N-cadherin (panel D), vimentin (panel H) and TCF8/ZEB (panel I) were found to be upregulated in cells expressing full-length or truncated LANA proteins. The mRNA transcript levels of ZO1 (panel E), claudin (panel F) and E-cadherin (panel G) were downregulated in LANA-expressing cells in comparison to control cells. The data were analyzed using the ΔΔCt method to estimate the relative number of transcripts. For normalization, GAPDH was used as an internal control. Western blot analysis of cell lysates showed that protein levels of Slug (panel J), Snail (panel K), β-catenin (panel L), vimentin (panel M), TCF8/ZEB1 (panel N), and N-cadherin (panel Q) were upregulated in LANA-expressing cells; whereas the levels of ZO1 (panel O), claudin (panel R), and E-cadherin (panel P) were significantly reduced in LANA-expressing cells (L-N, L-C and L-F) as compared to control (C). Expression of Flag-tagged LANA-N (panel S), LANA-C (panel T) and LANA-Full (panel U) was confirmed, and GAPDH (panel V) was used as internal control to confirm equal loading of cell lysates in all lanes
TCF8/ZEB1 is known to play a key role in transcriptional downregulation of E-cadherin, either directly or through activation of vimentin, a step that is critical for promotion of cell migration and increased cellular invasiveness [42]. Our data show that LANA-Full expression results in transcriptional upregulation of TCF8/ZEB1 and vimentin (Fig.1H and I) by up to 14-fold and 7-fold, respectively, which was statistically significant (p < 0.05). The transcript level of the cell adhesion molecule E-cadherin was downregulated by about 3-fold in LANA-Full-expressing cells (Fig. 1G). The loss of E-cadherin in cancer cells renders them more susceptible to mesenchymal transition and signifies a progression of cancer [6]. In addition, N-cadherin was upregulated by about 9-fold in LANA-Full-expressing cells (Fig. 1D). This shift in cadherin expression pattern from E-cadherin to N-cadherin is termed ‘cadherin switch’, which is an essential indicator of EMT [43]. The switch in cadherin expression pattern in the presence of LANA-Full clearly points to an important role of LANA in the process of EMT in KSHV-infected cells.
The LANA N- and C-termini work synergistically in LANA-mediated transcriptional modulation of cellular EMT-associated genes
Our results clearly show that LANA-Full can modulate the transcript levels of genes that are essential for EMT. The structure of LANA is like a dumbbell with two domains separated by a sequence of internal repeats [44]. The carboxy-terminal end of LANA binds to viral terminal repeats via the DNA-binding domain [38]. It has been shown previously that a mutation in the chromosome-binding domain at the C-terminal end of LANA leads to low mitotic expression indices [45]. The amino-terminal end of LANA interacts with cellular chromosomes by binding to the nucleosome at specific regions of histones H2A-H2B [46]. In order to identify which one of the two major domains of LANA is important for EMT, the truncated LANA proteins LANA-N (1-340) and LANA-C (945-1162) were expressed in MDA-MB-231T cells, and the relative change in the transcript level of EMT markers in these cells was quantified in comparison to their levels in empty-vector-transfected control cells.
Our data show that the expression of LANA-N resulted in transcriptional upregulation of Slug by 4-fold (Fig. 1A), of Snail by 9-fold (Fig. 1B), β-catenin by 5-fold (Fig. 1C), vimentin by 2-fold (Fig. 1H) and of ZEB1 by 5-fold (Fig. 1I) in comparison to empty-vector-transfected control cells, and these differences were statistically significant (p < 0.01). The transcript levels of ZO-1 (Fig. 1E) and claudin (Fig. 1F) decreased approximately 2-fold (p < 0.05), whereas E-cadherin was downregulated approximately 1.2-fold (p < 0.05) (Fig. 1G) and N-cadherin (Fig. 1D) was upregulated 4-fold (p < 0.05) in LANA-N-expressing cells. The expression of LANA-C resulted in transcriptional upregulation of Slug by 4-fold (Fig. 1A), Snail by 11-fold (Fig. 1B), β-catenin by 7-fold (Fig. 1C), vimentin by 2-fold (Fig. 1H), and TCF8/ZEB1 by 8-fold (Fig. 1I). The transcript level of E-cadherin was downregulated by approximately 1.3-fold (Fig. 1G), whereas that of N-cadherin was upregulated by 6-fold (Fig. 1D) in LANA-C-expressing cells, and these differences were statistically significant (p < 0.01). The transcript levels of ZO1 (Fig. 1E) and claudin (Fig. 1F) were downregulated by approximately 5-fold in LANA-C-expressing cells which was also significant (p < 0.01).
Our data show that expression of the C-terminal portion of LANA resulted in transcriptional downregulation of ZO-1 and claudin to a much greater extent (5-fold) than in LANA-N-expressing cells (2-fold). Slightly higher (6-fold) transcriptional upregulation was observed in the transcript level of N-cadherin in LANA-C-expressing cells in comparison to that observed in LANA-N-expressing cells (4-fold). The experiments were also repeated in 293T cells, and similar results were observed. The overview of results suggests that both termini of LANA work synergistically to modulate EMT markers.
Cancer cells expressing KSHV LANA differentially express EMT-associated transcription factors
We then tested whether changes in transcript levels of EMT-associated genes in LANA-expressing cells correlate with similar changes in their protein expression levels. MDA-MB-231T cells expressing LANA-Full, LANA-N or LANA-C and empty-vector-transfected control cells were harvested 48 hours after transfection. The cell lysates were resolved by SDS-PAGE, transferred to a PVDF membrane, and probed using specific primary antibodies to determine the expression levels of EMT marker proteins. The data are summarized in Fig. 1J to V and show that changes in transcript levels of EMT markers were largely reflected in changes in protein expression profiles.
Our data show that the transcriptional repressors Snail, Slug and β-catenin were significantly upregulated in LANA-expressing cells (Fig. 1J, K and L). The expression of LANA-Full resulted in a 2-fold increase (p < 0.05) in protein levels of Snail and Slug. Expression of LANA-N or LANA-C resulted in an approximately 1.5-fold increase (p < 0.05) in Snail and Slug. Expression of β-catenin was upregulated by approximately 3-fold (p < 0.01) in LANA-Full-expressing cells (Fig. 2C), whereas no significant changes in its protein level were observed in LANA-N-or LANA-C expressing cells. Although both LANA-N and LANA-C expression resulted in transcriptional upregulation of β-catenin (Fig. 1C), increased protein levels of β-catenin were observed only in LANA-Full-expressing cells.
Expression of KSHV LANA results in modification in localization and organization of Snail, Slug, β-catenin and E-cadherin. An immunofluorescence assay was carried out using MDA-MB-231T cells expressing LANA-Full, LANA-N and LANA-C using primary antibodies against specific EMT markers (green signal) and anti-Flag antibody to detect LANA expression (red signal). The expression of Snail (panel A), Slug (panel B) and β-catenin (panel C) was found to be upregulated. The signal for Slug and β-catenin were detected in the nucleus as well as the cytoplasm of LANA-expressing cells, whereas it was detected only in the cytoplasm in control cells. The cell adhesion molecule E-cadherin (panel D) was downregulated in LANA-expressing cells. The images were analyzed using a Zeiss LSM 510 scanning fluorescence microscope and ZEN software. The mean values and standard error of three independent experiments are presented as a bar graph at the bottom of each panel
In LANA-Full-expressing cells, TCF8/ZEB1 was upregulated more than 5-fold (p < 0.01) whereas the upregulation was only 1.8-fold in LANA-N-expressing cells and approximately 2.2-fold in LANA-C-expressing cells (Fig. 1N). Our data also show that vimentin, which is an intermediate filament of mesenchymal origin, was significantly overexpressed in cancer cells expressing LANA-Full. An approximately 4.3-fold increase (p < 0.01) in the level of vimentin was observed when LANA-Full was expressed, and 2.5-fold and 1.7-fold increases were observed in LANA-N and LANA-C expressing cells, respectively (Fig. 1M). Peripheral membrane adaptor protein ZO-1, junctional transmembrane protein claudin, and E-cadherin, which is an active suppressor of invasion and growth of cancer cells, were also significantly downregulated in LANA-expressing cells. ZO-1 was downregulated by approximately 6-fold, 5-fold and 3-fold, in LANA-Full-, LANA-C-, and LANA-N-expressing cells, respectively (Fig.10). An approximately 5-to-6-fold decrease (p < 0.01) in the expression of claudin was detected in LANA-Full- and LANA-C-expressing cells. No significant change in the expression level of claudin was observed in LANA-N-expressing cells (Fig. 1R), suggesting a role for the carboxy-terminal portion of LANA in reduced expression of claudin.
When we analyzed the expression levels of E-cadherin and N-cadherin, our data showed that there was a 3.5-fold increase (p < 0.01) in the N-cadherin expression level (Fig. 1Q) and about a 7-fold decrease (p < 0.01) in the E-cadherin expression level (Fig. 1P) in LANA-Full-expressing cells. The change in expression of the cadherin isoform from E-cadherin to N-cadherin (cadherin switch) in LANA-expressing cells clearly points to the role of LANA in EMT. An approximately 1.5-fold increase in the level of N-cadherin was observed in LANA-N- or LANA-C-expressing cells (Fig. 1P). E-cadherin expression was significantly downregulated by about 2- to 3-fold (p < 0.01) in LANA-N- or LANA-C-expressing cells. An experiment performed in 293T cells gave similar results.
Expression of KSHV LANA results in a change in localization and/or re-organization of EMT maker proteins
When EMT is initiated, the cell-cell junctions are remodeled and the junction proteins are either relocalized and/or degraded [47]. The intracellular localization of several other proteins that are critical for EMT is also affected. We therefore decided to investigate if the expression of LANA results in any changes in the intracellular localization and organization of EMT-associated proteins. The immunofluorescence microscopy images are presented in Figures 2, 3, 4. The bar diagrams in each figure represent the average intensity of the fluorescence signal detected per cell from three independent experiments. On average, about 15-20 cells randomly chosen from 5-6 microscopic views were analyzed for quantitation in each experiment. It should be emphasized that immunofluorescence microscopy data are representative of protein expressed in a limited number of cells and may not directly correlate with changes in relative expression levels observed in western blots, which were performed with cell lysates obtained from 10 million cells. The immunofluorescence microscopy data are more useful for studying changes in intracellular localization or organization patterns of proteins and possible co-localization with other proteins.
Expression of KSHV LANA results in modification of the location and organization of N-cadherin, ZO-1, claudin and vimentin. Expression of N-cadherin (panel A) was significantly upregulated in LANA-expressing cells. Both ZO1 (panel B) and claudin (panel C) were downregulated in LANA-expressing cells. Vimentin (panel D) was upregulated in cells expressing LANA. The images were analyzed using a Zeiss LSM 510 scanning fluorescence microscope and ZEN software. The mean values and standard error of three independent experiments are presented as a bar graph at the bottom of each panel
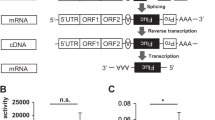
(A) Expression of KSHV LANA results in upregulation of TCF8/ZEB1 expression. Expression of the full-length LANA protein resulted in about 4.5-fold upregulation of protein expression levels of TCF8/ZEB1 compared to control cells. The images were analyzed using a Zeiss LSM 510 scanning fluorescence microscope and ZEN software. The mean values and standard error of three independent experiments are presented as a bar graph at the bottom of each panel. (B) The majority of TCF8/ZEB1 co-localizes with LANA. Scatter plot analysis of fluorescence signals show that more than 50% of the TCF8/ZEB1 protein co-localized with full-length LANA or LANA-N when co-expressed with these proteins. (C) KSHV latent protein LANA expression results in increased motility of cancer cells. A wound or scratch was made in a monolayer of cells expressing LANA-Full, LANA-N or LANA-C. The cells were examined at 24 h post-transfection, and the percentage of wound healing was calculated. The mean values and standard errors of three independent experiments are presented as a bar graph. (D) KSHV latent protein LANA expression results in increased invasiveness of cancer cells. A trans-well assay was carried out to observe the invasion potential of cells expressing LANA-Full or LANA-N or LANA-C. The cells that invaded the matrigel were visualized after staining with crystal violet dye and then counted. Images were observed and captured using a Nikon TS-100F Inverted Trinocular Research Microscope and a Nikon scientific-grade CMOS camera. The mean values and standard error of three independent experiments are presented as a bar graph at the bottom
Our data show that the transcriptional repressors Snail (Fig. 2A), Slug (Fig. 2B), and TCF8/ZEB1 (Fig. 4A) were overexpressed in LANA-Full-expressing MDA-MB-231T cells. The increased expression of these proteins was mainly in the cytoplasm. β-catenin was observed to be significantly upregulated in LANA-Full-expressing cells (Fig. 2C), and it was localized mainly around cell periphery, with a smaller fraction in the nucleus. Similar upregulation of β-catenin was also observed in LANA-N- and LANA-C-expressing cells, although the localization was more diffuse in LANA-N-expressing cells and mostly close to cell membranes in LANA-C-expressing cells. This shift in localization of β-catenin is an important regulator of cancer progression, as this is commonly observed in many cancers [48].
Downregulation of E-cadherin is an important event in the process of EMT [49]. E-cadherin and β-catenin form a complex that is linked with the actin cytoskeleton. An increase in N-cadherin is associated with the migration potential and invasion potential of cancer cells [50, 51]. In our study, we found that E-cadherin was significantly downregulated (Fig. 2D) and N-cadherin was significantly upregulated (Fig. 3A) in LANA-expressing cells. In LANA-C- and LANA-Full-expressing cells, an increase in N-cadherin was also detected in the nucleus.
Claudin and occludin proteins form a tight junction with cytoskeleton proteins of cells [52]. ZO-1 attaches claudin and occludin to the actin cytoskeleton. ZO-1 is a key peripheral membrane protein that, if downregulated, leads the cells towards the transition to a mesenchymal phenotype. Our data show that expression of the LANA protein resulted in significantly reduced expression of ZO-1 (Fig. 3B) and claudin (Fig. 3C). The effect was more profound when LANA-Full or LANA-C were expressed. In LANA-Full- and LANA-N-expressing cells, vimentin was expressed at significantly higher levels (2- to 4-fold) compared to empty-vector-transfected control cells (Fig. 3D).
A significant fraction of transcription factor ZEB1 co-localizes with LANA
TCF8/ZEB1 is an important contributor to the process of EMT and cancer progression [53]. Several studies have shown that upregulation in expression of ZEB1 is a key activator of EMT [54,55,56,57]. It has been shown to act as a transcriptional activator for vimentin [42], to repress the E-cadherin promoter, and to be an important regulator of cell adhesion molecules [58]. Our data show that ZEB1 was significantly upregulated (2- to 5-fold) in LANA-Full-expressing cells (Fig. 4A). We also observed that a significant fraction of ZEB1 co-localized with LANA inside cells. A scatter plot analysis performed on pixel-by-pixel basis using ZEN software (Carl Zeiss AG, Oberkochen, Germany) revealed that more than 50% of ZEB1 co-localized in same subcellular localization as LANA-Full or LANA-N (Fig. 4B), indicating that LANA might be critical for modulation of ZEB1-mediated functions through direct interaction, which will need to be investigated further.
KSHV LANA expression results in increased migration and invasion potential of cancer cells
We then tested whether LANA-mediated modulation of expression of EMT markers results in any changes in cellular phenotype or properties related to cellular migration and invasion potential. We performed a scratch wound healing assay to test the migration potential of cells. Our data show that LANA expression resulted in healing of a wound at a significantly faster rate than what was observed in empty-vector-transfected control cells (Fig. 4C). The expression of LANA-Full resulted in healing of a wound by 78% in 24 hours (p < 0.05) as compared to 25% healing in empty-vector-transfected control cells (Fig. 4C). For LANA-N- and LANA-C-expressing the cells, the wound healing was 70% and 55%, respectively (p < 0.05), after 24 hours (Fig. 4C).
To investigate the invasion potential of cells expressing LANA protein, we performed a trans-well invasion assay. Our data show that the number of cells expressing LANA-N and LANA-C that could invade the matrix was approximately 2 times greater than that observed in the empty-vector-transfected control (Fig. 4D). The number of cells invading the matrix was approximately 3.5-fold higher than in the empty-vector-transfected control when LANA-Full-expressing cells were tested, indicating that LANA expression results in a significant increase in the migration and invasion potential of cells. Our data clearly show that LANA-mediated modulation of transcription, translation, and localization of EMT-associated cellular proteins is concurrent with changes in cellular migration and invasive potential.
Discussion
EMT has been proposed previously to be classified into three different biological subtypes [6]. The type I EMT is associated with early stages of embryogenesis and fetal development, and type II EMT is associated with tissue regeneration and wound healing. In type III EMT, primary or secondary epithelial cells transform into cancerous cells, which enables them to metastasize and invade other tissues [6]. The transition of cancerous cells from epithelial to mesenchymal properties is mediated by many transcription factors in cells, including, Snail, Slug, Twist, ZEB1 and ZEB2 [7]. Changes in the way these proteins function lead to alterations in morphology, reorganization of skeletal proteins and adhesion molecules, and an increase in the migration potential of the cell. Earlier studies have shown that cancer cells expressing EBV nuclear antigens EBNA1 and EBNA3C have higher migration potential than empty-vector-transfected control cells [59]. In our previous study, we demonstrated the role of EBNA1 and EBNA 3C in the modulation of some common molecular markers that are crucial for EMT [24]. An EMT-like process has been shown earlier to occur in B-cell lymphomas [27]. Several previous studies have shown an important role for EMT markers during B-cell malignancies, including in B-cell lymphoma and mantle cell lymphoma, and in KSHV-infected B-cells [28,29,30,31]. KSHV is associated with many malignancies, amongst which Kaposi sarcoma (KS) is the most common form of cancer reported in HIV-infected immune-compromised patients [60, 61]. KS can also occur due to reactivation of the virus from its latent state. During KSHV latency, a very limited set of genes are expressed, which include ORF73 (LANA), ORF72 (vCyclin), ORF71 (vFLIP), ORFK12 (kaposins A, B, C), BCL-2 and 12 microRNAs. These are responsible for the establishment and persistence of KSHV latent infection [62,63,64]. Amongst these, LANA has been consistently detected in KSHV-related diseases. It can promote oncogenesis by repressing the tumor suppressor gene p53. LANA also activates survivin by phosphorylation, which is a major contributing factor to latent replication of KSHV [65]. LANA DNA binding domains (carboxyl terminal) also have structural similarities to EBV EBNA1 and human papillomavirus (HPV) E2 protein [66,67,68,69]. The carboxyl-terminal portion of LANA can also activate EBV latent membrane protein 1 (LMP1) [70].
In the present study, we tested our hypothesis that LANA can have a critical role in promoting EMT of cancer cells. We investigated the role of LANA in EMT by testing the effect of its expression on some important cellular genes that are critical for EMT. These included Snail, Slug, β-catenin, TCF8/ZEB1, vimentin, E-cadherin, N-cadherin, ZO1 and claudin. Our data show that expression of LANA resulted in transcriptional upregulation of transcriptional factors, including Snail, Slug, β-catenin, and TCF8/ZEB1. The protein levels of TCF8/ZEB1 and vimentin were found to be high in LANA-expressing cells, and TCF8/ZEB1 mostly co-localized with LANA in the same subcellular compartment. The amino-terminal portion of LANA (LANA-N) upregulated ZEB1 and vimentin to a higher level compared to the carboxyl-terminal portion of LANA (LANA-C), and both appeared to work synergistically to modulate EMT markers. In LANA-expressing cells, β-catenin was significantly upregulated and partially translocated to the nucleus. In addition, the expression of LANA also resulted in significant downregulation of ZO1, claudin and E-cadherin. The expression of the carboxyl-terminal portion of LANA resulted in greater downregulation of ZO1 and claudin than was observed in cells expressing the amino-terminal portion of LANA. LANA expression also resulted in a ‘cadherin switch’ in cells, whereby the expression of E-cadherin was significantly reduced and that of N-cadherin was significantly increased.
These changes in expression, organization and localization of EMT-associated cellular proteins in LANA-expressing cells clearly support a role of LANA in epithelial EMT. In a previous genome-wide study that compared the gene expression profile of KSHV-infected BCBL-1 and BC-3 cells with KSHV-negative BJAB and DG75 cells (data accessible at NCBI, accession no. GSE1880), the EMT-associated cellular genes, including E-cadherin, N-cadherin, vimentin, tight junction protein 1, claudin 1, claudin 9, ZEB1 and SNAI2 (Snail 2/ Slug), were reported to be similarly modulated in KSHV-infected cells [71, 72]. These data can be accessed at the NCBI GEO database with accession numbers GSM33134, GSM33135, GSM33155, and GSM33156. Our limited study now provides evidence for a specific role for KSHV latent antigen LANA in the modulation of cellular genes that are important for the epithelial-to-mesenchymal transition of KSHV-infected cells. It is highly probable that other KSHV latency-associated proteins could also have a role in KSHV-mediated EMT, and this needs to be investigated separately.
References
Fidler IJ (2003) The pathogenesis of cancer metastasis: the ‘seed and soil’ hypothesis revisited. Nat Rev Cancer 3(6):453–458. https://doi.org/10.1038/nrc1098
Thiery JP (2002) Epithelial-mesenchymal transitions in tumour progression. Nat Rev Cancer 2(6):442–454. https://doi.org/10.1038/nrc822
Gatenby RA (2009) A change of strategy in the war on cancer. Nature 459(7246):508–509. https://doi.org/10.1038/459508a
Patel LR, Camacho DF, Shiozawa Y, Pienta KJ, Taichman RS (2011) Mechanisms of cancer cell metastasis to the bone: a multistep process. Future Oncol 7(11):1285–1297. https://doi.org/10.2217/fon.11.112
Pienta KJ, Loberg R (2005) The “emigration, migration, and immigration” of prostate cancer. Clin Prostate Cancer 4(1):24–30
Kalluri R, Weinberg RA (2009) The basics of epithelial-mesenchymal transition. J Clin Investig 119(6):1420–1428. https://doi.org/10.1172/JCI39104
Goossens S, Vandamme N, Van Vlierberghe P (1868) Berx G (2017) EMT transcription factors in cancer development re-evaluated: beyond EMT and MET. Biochim Biophys Acta 2:584–591
Lee JM, Dedhar S, Kalluri R, Thompson EW (2006) The epithelial-mesenchymal transition: new insights in signaling, development, and disease. J Cell Biol 172(7):973–981
Yang J, Mani SA, Donaher JL, Ramaswamy S, Itzykson RA, Come C, Savagner P, Gitelman I, Richardson A, Weinberg RA (2004) Twist, a master regulator of morphogenesis, plays an essential role in tumor metastasis. Cell 117(7):927–939. https://doi.org/10.1016/j.cell.2004.06.006
Bolos V, Peinado H, Perez-Moreno MA, Fraga MF, Esteller M, Cano A (2003) The transcription factor Slug represses E-cadherin expression and induces epithelial to mesenchymal transitions: a comparison with Snail and E47 repressors. J Cell Sci 116(Pt 3):499–511
Hajra KM, Chen DY, Fearon ER (2002) The SLUG zinc-finger protein represses E-cadherin in breast cancer. Cancer Res 62(6):1613–1618
Turner FE, Broad S, Khanim FL, Jeanes A, Talma S, Hughes S, Tselepis C, Hotchin NA (2006) Slug regulates integrin expression and cell proliferation in human epidermal keratinocytes. J Biol Chem 281(30):21321–21331
Zhang P, Sun Y, Ma L (2015) ZEB1: at the crossroads of epithelial-mesenchymal transition, metastasis and therapy resistance. Cell Cycle 14(4):481–487. https://doi.org/10.1080/15384101.2015.1006048
Ishida K, Ito S, Wada N, Deguchi H, Hata T, Hosoda M, Nohno T (2007) Nuclear localization of beta-catenin involved in precancerous change in oral leukoplakia. Mol Cancer 6:62
Cadigan KM, Nusse R (1997) Wnt signaling: a common theme in animal development. Genes Dev 11(24):3286–3305
Inukai T, Inoue A, Kurosawa H, Goi K, Shinjyo T, Ozawa K, Mao M, Inaba T, Look AT (1999) SLUG, a ces-1-related zinc finger transcription factor gene with antiapoptotic activity, is a downstream target of the E2A-HLF oncoprotein. Mol Cell 4(3):343–352
Schmidt CR, Gi YJ, Patel TA, Coffey RJ, Beauchamp RD, Pearson AS (2005) E-cadherin is regulated by the transcriptional repressor SLUG during Ras-mediated transformation of intestinal epithelial cells. Surgery 138(2):306–312
Adhikary A, Chakraborty S, Mazumdar M, Ghosh S, Mukherjee S, Manna A, Mohanty S, Nakka KK, Joshi S, De A, Chattopadhyay S, Sa G, Das T (2014) Inhibition of epithelial to mesenchymal transition by E-cadherin up-regulation via repression of slug transcription and inhibition of E-cadherin degradation: dual role of scaffold/matrix attachment region-binding protein 1 (SMAR1) in breast cancer cells. J Biol Chem 289(37):25431–25444. https://doi.org/10.1074/jbc.M113.527267M
Sakai D, Tanaka Y, Endo Y, Osumi N, Okamoto H, Wakamatsu Y (2005) Regulation of Slug transcription in embryonic ectoderm by beta-catenin-Lef/Tcf and BMP-Smad signaling. Dev Growth Differ 47(7):471–482
Cai LM, Lyu XM, Luo WR, Cui XF, Ye YF, Yuan CC, Peng QX, Wu DH, Liu TF, Wang E, Marincola FM, Yao KT, Fang WY, Cai HB, Li X (2015) EBV-miR-BART7-3p promotes the EMT and metastasis of nasopharyngeal carcinoma cells by suppressing the tumor suppressor PTEN. Oncogene 34(17):2156–2166. https://doi.org/10.1038/onc.2014.341
Chung TW, Kim SJ, Choi HJ, Song KH, Jin UH, Yu DY, Seong JK, Kim JG, Kim KJ, Ko JH, Ha KT, Lee YC, Kim CH (2014) Hepatitis B virus X protein specially regulates the sialyl lewis a synthesis among glycosylation events for metastasis. Mol Cancer 13:222. https://doi.org/10.1186/1476-4598-13-222
Harakeh S, Abou-Khouzam R, Damanhouri GA, Al-Hejin A, Kumosani T, Niedzwiecki A, Rath M, Barbour E, Diab-Assaf M, Azar R (2014) Effects of nutrients on matrix metalloproteinases in human T-lymphotropic virus type 1 positive and negative malignant T-lymphocytes. Int J Oncol 45(5):2159–2166. https://doi.org/10.3892/ijo.2014.2638
Knight LM, Stakaityte G, Wood JJ, Abdul-Sada H, Griffiths DA, Howell GJ, Wheat R, Blair GE, Steven NM, Macdonald A, Blackbourn DJ, Whitehouse A (2015) Merkel cell polyomavirus small T antigen mediates microtubule destabilization to promote cell motility and migration. J Virol 89(1):35–47. https://doi.org/10.1128/JVI.02317-14
Gaur N, Gandhi J, Robertson ES, Verma SC, Kaul R (2015) Epstein-Barr virus latent antigens EBNA3C and EBNA1 modulate epithelial to mesenchymal transition of cancer cells associated with tumor metastasis. Tumour Biol 36(4):3051–3060. https://doi.org/10.1007/s13277-014-2941-6
Moore PS, Chang Y (2003) Kaposi’s sarcoma-associated herpesvirus immunoevasion and tumorigenesis: two sides of the same coin? Annu Rev Microbiol 57:609–639. https://doi.org/10.1146/annurev.micro.57.030502.090824
Ignatovich IA, Dizhe EB, Akif’ev BN, Burov SV, Boiarchuk EA, Perevozchikov AP (2002) Delivery of “suicide” thymidine kinase gene of herpes virus in the complex with cationic peptide into human hepatoma cells in vitro. Tsitologiia 44(5):455–462
Lemma S, Karihtala P, Haapasaari KM, Jantunen E, Soini Y, Bloigu R, Pasanen AK, Turpeenniemi-Hujanen T, Kuittinen O (2013) Biological roles and prognostic values of the epithelial-mesenchymal transition-mediating transcription factors Twist, ZEB1 and Slug in diffuse large B-cell lymphoma. Histopathology 62(2):326–333. https://doi.org/10.1111/his.12000
Sanchez-Tillo E, Fanlo L, Siles L, Montes-Moreno S, Moros A, Chiva-Blanch G, Estruch R, Martinez A, Colomer D, Gyorffy B, Roue G, Postigo A (2014) The EMT activator ZEB1 promotes tumor growth and determines differential response to chemotherapy in mantle cell lymphoma. Cell Death Differ 21(2):247–257. https://doi.org/10.1038/cdd.2013.123
Jha HC, Sun Z, Upadhyay SK, El-Naccache DW, Singh RK, Sahu SK, Robertson ES (2016) KSHV-mediated regulation of Par3 and SNAIL contributes to B-Cell proliferation. PLoS Pathog 12(7):e1005801. https://doi.org/10.1371/journal.ppat.1005801
Gasperini P, Espigol-Frigole G, McCormick PJ, Salvucci O, Maric D, Uldrick TS, Polizzotto MN, Yarchoan R, Tosato G (2012) Kaposi sarcoma herpesvirus promotes endothelial-to-mesenchymal transition through Notch-dependent signaling. Cancer Res 72(5):1157–1169. https://doi.org/10.1158/0008-5472.CAN-11-3067
Cheng F, Pekkonen P, Laurinavicius S, Sugiyama N, Henderson S, Gunther T, Rantanen V, Kaivanto E, Aavikko M, Sarek G, Hautaniemi S, Biberfeld P, Aaltonen L, Grundhoff A, Boshoff C, Alitalo K, Lehti K, Ojala PM (2011) KSHV-initiated notch activation leads to membrane-type-1 matrix metalloproteinase-dependent lymphatic endothelial-to-mesenchymal transition. Cell Host Microbe 10(6):577–590. https://doi.org/10.1016/j.chom.2011.10.011S1931-3128(11)00364-7
Dourmishev LA, Dourmishev AL, Palmeri D, Schwartz RA, Lukac DM (2003) Molecular genetics of Kaposi’s sarcoma-associated herpesvirus (human herpesvirus-8) epidemiology and pathogenesis. Microbiol Mol Biol Rev 67(2):175–212 (table of contents)
Ballestas ME, Chatis PA, Kaye KM (1999) Efficient persistence of extrachromosomal KSHV DNA mediated by latency-associated nuclear antigen. Science 284(5414):641–644
Borah S, Verma SC, Robertson ES (2004) ORF73 of herpesvirus saimiri, a viral homolog of Kaposi’s sarcoma-associated herpesvirus, modulates the two cellular tumor suppressor proteins p53 and pRb. J Virol 78(19):10336–10347. https://doi.org/10.1128/JVI.78.19.10336-10347.2004
Friborg J Jr, Kong W, Hottiger MO, Nabel GJ (1999) p53 inhibition by the LANA protein of KSHV protects against cell death. Nature 402(6764):889–894. https://doi.org/10.1038/47266
Fujimuro M, Hayward SD (2003) The latency-associated nuclear antigen of Kaposi’s sarcoma-associated herpesvirus manipulates the activity of glycogen synthase kinase-3beta. J Virol 77(14):8019–8030
Verma SC, Lan K, Choudhuri T, Cotter MA, Robertson ES (2007) An autonomous replicating element within the KSHV genome. Cell Host Microbe 2(2):106–118
Verma SC, Choudhuri T, Kaul R, Robertson ES (2006) Latency-associated nuclear antigen (LANA) of Kaposi’s sarcoma-associated herpesvirus interacts with origin recognition complexes at the LANA binding sequence within the terminal repeats. J Virol 80(5):2243–2256
Khera L, Paul C, Kaul R (2017) Hepatitis C Virus E1 protein promotes cell migration and invasion by modulating cellular metastasis suppressor Nm23-H1. Virology 506:110–120
Rueden CT, Schindelin J, Hiner MC, DeZonia BE, Walter AE, Arena ET, Eliceiri KW (2017) Image J2: ImageJ for the next generation of scientific image data. BMC Bioinform 18(1):529. https://doi.org/10.1186/s12859-017-1934-z
Huang RY, Guilford P, Thiery JP (2012) Early events in cell adhesion and polarity during epithelial-mesenchymal transition. J Cell Sci 125(Pt 19):4417–4422. https://doi.org/10.1242/jcs.099697
Nishimura G, Manabe I, Tsushima K, Fujiu K, Oishi Y, Imai Y, Maemura K, Miyagishi M, Higashi Y, Kondoh H, Nagai R (2006) DeltaEF1 mediates TGF-beta signaling in vascular smooth muscle cell differentiation. Dev Cell 11(1):93–104
Gravdal K, Halvorsen OJ, Haukaas SA, Akslen LA (2007) A switch from E-cadherin to N-cadherin expression indicates epithelial to mesenchymal transition and is of strong and independent importance for the progress of prostate cancer. Clin Cancer Res 13(23):7003–7011
Dittmer DP, Damania B (2016) Kaposi sarcoma-associated herpesvirus: immunobiology, oncogenesis, and therapy. J Clin Investig 126(9):3165–3175. https://doi.org/10.1172/JCI84418
Kelley-Clarke B, De Leon-Vazquez E, Slain K, Barbera AJ, Kaye KM (2009) Role of Kaposi’s sarcoma-associated herpesvirus C-terminal LANA chromosome binding in episome persistence. J Virol 83(9):4326–4337. https://doi.org/10.1128/JVI.02395-08
Barbera AJ, Chodaparambil JV, Kelley-Clarke B, Joukov V, Walter JC, Luger K, Kaye KM (2006) The nucleosomal surface as a docking station for Kaposi’s sarcoma herpesvirus LANA. Science 311(5762):856–861
Lamouille S, Xu J, Derynck R (2014) Molecular mechanisms of epithelial-mesenchymal transition. Nat Rev Mol Cell Biol 15(3):178–196. https://doi.org/10.1038/nrm3758
Whitaker HC, Girling J, Warren AY, Leung H, Mills IG, Neal DE (2008) Alterations in beta-catenin expression and localization in prostate cancer. Prostate 68(11):1196–1205. https://doi.org/10.1002/pros.20780
Schmalhofer O, Brabletz S, Brabletz T (2009) E-cadherin, beta-catenin, and ZEB1 in malignant progression of cancer. Cancer Metastasis Rev 28(1–2):151–166. https://doi.org/10.1007/s10555-008-9179-y
Wheelock MJ, Johnson KR (2003) Cadherins as modulators of cellular phenotype. Annu Rev Cell Dev Biol 19:207–235. https://doi.org/10.1146/annurev.cellbio.19.011102.111135
Wells A, Chao YL, Grahovac J, Wu Q, Lauffenburger DA (2011) Epithelial and mesenchymal phenotypic switchings modulate cell motility in metastasis. Front Biosci (Landmark Ed) 16:815–837
Matter K, Balda MS (2007) Epithelial tight junctions, gene expression and nucleo-junctional interplay. J Cell Sci 120(Pt 9):1505–1511
Liu Y, El-Naggar S, Darling DS, Higashi Y, Dean DC (2008) Zeb1 links epithelial-mesenchymal transition and cellular senescence. Development 135(3):579–588. https://doi.org/10.1242/dev.007047
Weber KL, Doucet M, Price JE (2003) Renal cell carcinoma bone metastasis: epidermal growth factor receptor targeting. Clin Orthop Relat Res 415(Suppl):S86–S94. https://doi.org/10.1097/01.blo.0000093050.96273.35
Singh M, Spoelstra NS, Jean A, Howe E, Torkko KC, Clark HR, Darling DS, Shroyer KR, Horwitz KB, Broaddus RR, Richer JK (2008) ZEB1 expression in type I vs type II endometrial cancers: a marker of aggressive disease. Mod Pathol 21(7):912–923. https://doi.org/10.1038/modpathol.2008.82
Karihtala P, Auvinen P, Kauppila S, Haapasaari KM, Jukkola-Vuorinen A, Soini Y (2013) Vimentin, zeb1 and Sip1 are up-regulated in triple-negative and basal-like breast cancers: association with an aggressive tumour phenotype. Breast Cancer Res Treat 138(1):81–90. https://doi.org/10.1007/s10549-013-2442-0
Ohira T, Gemmill RM, Ferguson K, Kusy S, Roche J, Brambilla E, Zeng C, Baron A, Bemis L, Erickson P, Wilder E, Rustgi A, Kitajewski J, Gabrielson E, Bremnes R, Franklin W, Drabkin HA (2003) WNT7a induces E-cadherin in lung cancer cells. Proc Natl Acad Sci USA 100(18):10429–10434. https://doi.org/10.1073/pnas.1734137100
Witta SE, Gemmill RM, Hirsch FR, Coldren CD, Hedman K, Ravdel L, Helfrich B, Dziadziuszko R, Chan DC, Sugita M, Chan Z, Baron A, Franklin W, Drabkin HA, Girard L, Gazdar AF, Minna JD, Bunn PA Jr (2006) Restoring E-cadherin expression increases sensitivity to epidermal growth factor receptor inhibitors in lung cancer cell lines. Cancer Res 66(2):944–950
Kaul R, Murakami M, Choudhuri T, Robertson ES (2007) Epstein-Barr virus latent nuclear antigens can induce metastasis in a nude mouse model. J Virol 81(19):10352–10361
Silverberg MJ, Lau B, Achenbach CJ, Jing Y, Althoff KN, D’Souza G, Engels EA, Hessol NA, Brooks JT, Burchell AN, Gill MJ, Goedert JJ, Hogg R, Horberg MA, Kirk GD, Kitahata MM, Korthuis PT, Mathews WC, Mayor A, Modur SP, Napravnik S, Novak RM, Patel P, Rachlis AR, Sterling TR, Willig JH, Justice AC, Moore RD, Dubrow R (2015) Cumulative incidence of cancer among persons with HIV in North America: a cohort study. Ann Intern Med 163(7):507–518. https://doi.org/10.7326/M14-2768
Robbins HA, Pfeiffer RM, Shiels MS, Li J, Hall HI, Engels EA (2015) Excess cancers among HIV-infected people in the United States. J Natl Cancer Inst. https://doi.org/10.1093/jnci/dju503
Staskus KA, Zhong W, Gebhard K, Herndier B, Wang H, Renne R, Beneke J, Pudney J, Anderson DJ, Ganem D, Haase AT (1997) Kaposi’s sarcoma-associated herpesvirus gene expression in endothelial (spindle) tumor cells. J Virol 71(1):715–719
Dittmer DP, Richards KL, Damania B (2012) Treatment of Kaposi sarcoma-associated herpesvirus-associated cancers. Front Microbiol 3:141. https://doi.org/10.3389/fmicb.2012.00141
McClure LV, Sullivan CS (2008) Kaposi’s sarcoma herpes virus taps into a host microRNA regulatory network. Cell Host Microbe 3(1):1–3. https://doi.org/10.1016/j.chom.2007.12.002
Lu J, Jha HC, Verma SC, Sun Z, Banerjee S, Dzeng R, Robertson ES (2014) Kaposi’s sarcoma-associated herpesvirus-encoded LANA contributes to viral latent replication by activating phosphorylation of survivin. J Virol 88(8):4204–4217. https://doi.org/10.1128/JVI.03855-13
Domsic JF, Chen HS, Lu F, Marmorstein R, Lieberman PM (2013) Molecular basis for oligomeric-DNA binding and episome maintenance by KSHV LANA. PLoS Pathog 9(10):e1003672. https://doi.org/10.1371/journal.ppat.1003672
Hellert J, Weidner-Glunde M, Krausze J, Richter U, Adler H, Fedorov R, Pietrek M, Ruckert J, Ritter C, Schulz TF, Luhrs T (2013) A structural basis for BRD2/4-mediated host chromatin interaction and oligomer assembly of Kaposi sarcoma-associated herpesvirus and murine gammaherpesvirus LANA proteins. PLoS Pathog 9(10):e1003640. https://doi.org/10.1371/journal.ppat.1003640
Hellert J, Weidner-Glunde M, Krausze J, Lunsdorf H, Ritter C, Schulz TF, Luhrs T (2015) The 3D structure of Kaposi sarcoma herpesvirus LANA C-terminal domain bound to DNA. Proc Natl Acad Sci USA 112(21):6694–6699. https://doi.org/10.1073/pnas.1421804112
Correia B, Cerqueira SA, Beauchemin C, Pires de Miranda M, Li S, Ponnusamy R, Rodrigues L, Schneider TR, Carrondo MA, Kaye KM, Simas JP, McVey CE (2013) Crystal structure of the gamma-2 herpesvirus LANA DNA binding domain identifies charged surface residues which impact viral latency. PLoS Pathog 9(10):e1003673. https://doi.org/10.1371/journal.ppat.1003673
Groves AK, Cotter MA, Subramanian C, Robertson ES (2001) The latency-associated nuclear antigen encoded by Kaposi’s sarcoma-associated herpesvirus activates two major essential Epstein-Barr virus latent promoters. J Virol 75(19):9446–9457. https://doi.org/10.1128/JVI.75.19.9446-9457.2001
An FQ, Folarin HM, Compitello N, Roth J, Gerson SL, McCrae KR, Fakhari FD, Dittmer DP, Renne R (2006) Long-term-infected telomerase-immortalized endothelial cells: a model for Kaposi’s sarcoma-associated herpesvirus latency in vitro and in vivo. J Virol 80(10):4833–4846
An FQ, Compitello N, Horwitz E, Sramkoski M, Knudsen ES, Renne R (2005) The latency-associated nuclear antigen of Kaposi’s sarcoma-associated herpesvirus modulates cellular gene expression and protects lymphoid cells from p16 INK4A-induced cell cycle arrest. J Biol Chem 280(5):3862–3874
Acknowledgements
This work was supported by grants from the Department of Biotechnology of the Government of India (BT/PR15109/GBD/27/320/2011), an MRP Grant from UGC (FN-41-1144/2012), an R&D Grant from the University of Delhi, and a PURSE Grant from DST. NG is a project fellow funded by UGC.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Handling Editor: Scott Schmid.
Rights and permissions
About this article
Cite this article
Gaur, N., Tikla, T. & Kaul, R. Kaposi sarcoma-associated herpes virus (KSHV) latent protein LANA modulates cellular genes associated with epithelial-to-mesenchymal transition. Arch Virol 164, 91–104 (2019). https://doi.org/10.1007/s00705-018-4060-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00705-018-4060-y