Abstract
Pseudorabies virus (PRV) is an animal alphaherpesvirus with a wide host range. PRV has 67 protein-coding genes and several non-coding RNA molecules, which can be classified into three temporal groups, immediate early, early and late classes. The ul54 gene of PRV and its homolog icp27 of herpes simplex virus have a multitude of functions, including the regulation of viral DNA synthesis and the control of the gene expression. Therefore, abrogation of PRV ul54 function was expected to exert a significant effect on the global transcriptome and on DNA replication. Real-time PCR and real-time RT-PCR platforms were used to investigate these presumed effects. Our analyses revealed a drastic impact of the ul54 mutation on the genome-wide expression of PRV genes, especially on the transcription of the true late genes. A more than two hour delay was observed in the onset of DNA replication, and the amount of synthesized DNA molecules was significantly decreased in comparison to the wild-type virus. Furthermore, in this work, we were able to successfully demonstrate the utility of long-read SMRT sequencing for genotyping of mutant viruses.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Pseudorabies virus (PRV) - also known as Suid herpesvirus 1 - is a member of the neurotropic herpesviruses, and the causative agent of Aujeszky’s disease in pigs [1]. PRV is a widely employed model organism for studies of the pathogenesis of herpesviruses [2]. Due to the neurotropic nature of the virus, PRV is also generally used as a tracer for mapping neural circuits [3,4,5,6,7,8]. PRV can also serve as a vector for gene transfer into neurons [9, 10] and cardiomyocytes [11]. PRV has also been tested as a potential tool in gene and cancer therapy [12], as well as live vaccines against Aujeszky’s disease [13,14,15]. PRV has a double-stranded linear DNA, composed of 143,423 bp with a mean GC content of 73.59% [16]. The PRV genome encodes 67 protein-coding and at least 20 non-coding RNA genes [17]. The transcriptome of the wild-type (wt) virus has been examined using various approaches, such as real-time RT-PCR [18] and microarray [19] analyses, as well as next generation [20] and third generation [17] sequencing platforms. It has recently been shown that almost the entire viral genome is transcriptionally active and many genes produce various transcript isoforms, including length and splice variants, as well as polycistronic transcription units with varying numbers of genes [17]. PRV genes, expressed in a cascade-like manner, are divided into three major temporal classes: immediate-early (IE), early (E) and late (L). The L genes can be further subdivided into early-late (L1 or E/L) and true late (L2 or L) genes in terms of their dependency on viral DNA replication (L genes require the synthesis of viral DNA for their expression). Herpes simplex virus type 1 (HSV-1), the prototype of the Alphaherpesvirinae subfamily, has five IE genes (icp4, icp0, icp27, icp22, and icp47; [21, 22]), while in comparison, PRV has only one true IE gene, the ie180 (homologous to the icp4 of HSV-1; [23]). It has been shown that ep0 and ul54 genes of PRV (homologous to icp0 [24] and the icp27 of HSV [25], respectively) are expressed in the E phase of infection. PRV lacks the icp47 gene and no consensus has been reached as to whether the us1 gene (ICP22 in HSV) is expressed with IE [26] or E [27] kinetics. The least characterized among the above genes in PRV is the ul54 gene [25, 28, 29].
The ul54 gene is located on the unique long (UL) region of the PRV genome and it forms a tandem cluster along with 2 other genes (ul53, ul52), which produce 3’ coterminal transcripts. The PRV ul54 gene is composed of 1,164 nucleotides and encodes a protein of 361 amino acids.
Several functions of ICP27 and their homologs have been revealed thus far, including the regulation of transcription [28] and DNA replication [30,31,32,33], the shut off of host protein synthesis [34], and the usage of polyadenylation sites [35], as well as viral growth. The deletion of the ul54 gene has been shown to result in severe growth defects [36]. A previously published study [33] reported that the ul54 gene and its protein product are not essential for PRV growth and replication in tissue culture; however, the mutant virus exhibited reduced growth ability. It has also been shown that this multifunctional protein is not essential for host shut-off and that its absence causes aberrant accumulation of late proteins at the early phase of infection in a cell-type dependent manner [33]. Two recently published studies have revealed that the ICP27 also plays a role in nucleo-cytoplasmic trafficking and in nucleolar-targeting [37, 38], respectively. Despite the accumulating data on the function of the ul54 gene, its precise role in the virus lifecycle continues to remain poorly understood.
In this study, we generated an ul54-KO virus with a knockout technique based on homologous recombination, and examined the effects of the mutation on the replication and global transcription of PRV by using quantitative real-time-PCR and reverse transcription (qRT)-PCR platforms. The presence of mutation was confirmed by long-read sequencing based on the Pacific Biosciences (PacBio) Single Molecule Real-time (SMRT) sequencing technique. Similar analyses based on qRT-PCR platform have been carried out to investigate the effect of the mutation of the vhs (Tombácz et al., 2011) ep0 (Tombácz et al., 2012), and us1 (Takács et al., 2013) genes of PRV on global gene expression.
Materials and methods
Cells, viruses and infection conditions
Monolayer cultures of immortalized porcine kidney epithelial (PK-15) cells were maintained at 37°C in the presence of 5% CO2. Cells were grown in DMEM (Gibco/Thermo Fisher Scientific), supplemented with 5% fetal bovine serum (Gibco/Thermo Fisher Scientific) and 80 μg of gentamycin per ml (Gibco/Thermo Fisher Scientific). PK-15 cells were used for the propagation of pseudorabies virus. The Kaplan (wt) strain of PRV was used as the parental strain for the generation of the ul54-null mutant virus (ul54-KO). PK-15 cells were infected with a low multiplicity of infection [MOI; 0.1 plaque-forming units (pfu)/cell]. The expression of the PRV genes was monitored at the following time points: 1, 2, 4, 6, and 8 h. Three independent biological replicates were carried out for the infections at each time point (Supplementary Table 1).
Generation of the ul54 -deleted virus
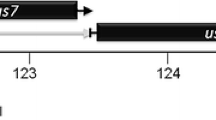
In this study, we deleted the ul54 gene of pseudorabies virus (PRV) and investigated the effect of this mutation on the transcription of 37 protein-coding and 3 non-coding viral genes (Table 1), as well as on the DNA replication. The PRV ul54-KO mutant virus was constructed as follows: as a first step, the BamHI fragment containing the entire ul54 gene was isolated from an agarose gel, and then was subcloned into the pRL525 cloning plasmid [39]. The resulting recombinant plasmid was used as a template for the PCR amplification of the two arms of the flanking sequences providing homology with the targeted genomic region of the PRV. A unique EcoRI site was inserted in place of a 1,017-bp segment (located within 2901 and 3919 bps) within the ul54 gene during the PCR reaction (Figure 1, Table 2). This was then followed by the insertion of a green-fluorescent protein (GFP; pEGFP-N1 vector, Clontech) gene expression cassette (Clontech) bracketed by EcoRI sites into the EcoRI site of the targeting sequence. The resulting recombinant plasmid was used as a transfer construct for the generation of the knockout virus. The linearized targeting plasmid was transfected along with the purified wild-type (wt, strain Kaplan) viral DNA into porcine kidney (PK-15) immortalized epithelial cells. The recombinant virus was generated by homologous recombination, followed by isolation on the basis of fluorescence of cells infected by the recombinant virus using an inverted fluorescent microscope (Olympus iX71). The first isolate was plaque-purified, which was repeated until the contaminating wt virus was completely eliminated.
Deletion of the ul54 gene of PRV. Almost the entire ul54 gene was eliminated by a technique based on homologous recombination. A: this part of the figure shows the schematic representation of the inserted GFP expression cassette (illustrated at the top), as well as the knocked-out region of the PRV genome. B: Integrative Genomics Viewer (IGV) representation showing the presence of the mutation
RNA purification
Total RNAs were isolated from infected cells by using Nucleospin RNA Kit (Macherey-Nagel) as was suggested by the manufacturer. In summary, the cells were collected by centrifugation and lysed by a buffer (kit component). Potential genomic DNA contamination was digested by RNase-free rDNase solution (supplied with the kit). Samples were eluted in nuclease-free water (part of the kit) in a total volume of 60 μl. Next, possible residual DNA contamination was eliminated by using the TURBO DNA-free Kit (Ambion/Thermo Fisher Scientific). RNA concentration was calculated using a Qubit 2.0 Fluorometer instrument (through use of the Qubit RNA BR Assay Kit (Life Technologies/Thermo Fisher Scientific). The RNA samples were stored at − 80 °C until use.
Reverse transcription
Single-stranded (ss)cDNA production was carried out by using SuperScript III reverse transcriptase (Invitrogen/Thermo Fisher Scientific) and gene-specific primers [18] (Table 3). Briefly, the reaction mixtures containing RNA, primer, SuperScript III enzyme, buffer, and dNTP mix were incubated at 55°C for 1 h. Finally, the reaction was terminated by heating at 70°C for 15 min. Samples were diluted 10-fold with nuclease-free water (Ambion/Thermo Fisher Scientific).
Real-time PCR and real-time RT PCR
The entire viral genome or single-stranded cDNAs were used as templates for the amplification of specific sequences by Rotor-Gene Q real-time PCR cycler (Qiagen) and Absolute QPCR SYBR Green Mix (Thermo Fisher Scientific) as published in our previous article [18]. To ensure the accuracy, the following controls were used: no-RT, no-primer, no-template, as well as loading control (pig 28 rRNA). Purified viral DNA was also used to verify the specificity of the primers.
Data analysis
Relative expression ratio (R) values were calculated by using the following formula, as described in our previous study [18]:
where R is the relative expression ratio; E is the efficiency of amplification; Ct is the cycle threshold value; sample refers to any particular gene at a given time point; and ref is the 28S rRNA that was used as a reference gene in each run. We used the average 6h value of ECt for each gene in the wt background as the control [61]. The R values in each independent experiment were calculated by normalizing cDNAs to 28S rRNA using the Comparative Quantitation module of the Rotor-Gene Q software (Version 2.3.1, Qiagen). The final R values represent the means of the three biological replicates (Supplementary Table 2). The net increase in the mRNA level (RΔ) and the rate of change (Ra) were calculated as was previously described [18]:
The effect of the UL54 protein on the expression of PRV genes was calculated by using the Rr value, which is the ratio between the R values of the ul54-knockout and the wt PRVs:
where R ul54Δ and R wt are the R values of a particular gene at a given time point in the ul54-knockout and wt genetic background, respectively. The average Rr for the E, E/L and L genes was also calculated for each time point:
Pearson’s correlation coefficient was calculated for the analysis of the expression dynamics, using the following equation [62]:
In this study we used the mean expression value of all examined transcript (total) in a given sample as a normalization factor for the transcripts (sample) real-time quantitative PCR to obtain Rx values. This method is similar to that used by Mestdagh et al [63]. However, instead of using the Ct values alone, we used the of ECt for the calculation of the expression values.
DNA purification
PK-15 cells were infected with ul54-KO PRV (0.1 pfu/cell) and incubated until complete cytopathic effect was observed. After three rounds of freeze-thaw treatment, cells were centrifuged at 750 xg for 5 minutes at 4°C to eliminate the cell debris. The supernatant was collected and layered beneath a 30% sucrose solution in ultracentrifuge tubes. Ultracentrifugation was carried out at 70000 xg for 1h using Sorvall WX Ultra 90 (Thermo Scientific) device. This was followed by removal of the supernatant and resuspension of the sedimented virus in a buffer solution (2% NaCl 5M, 1% Tris 1M (pH=6.7), 0.5% EDTA 0.2 M (pH=7). The proteins were removed by adding SDS (0.5%) and proteinase K (500 µg/ml) to the virus solution followed by incubation at 55°C for 1h. DNA was purified by phenol/chloroform extraction, ethanol precipitated, and resuspended in nuclease-free water.
Pacific Biosciences Single Molecule Real-Time sequencing and data analysis
A SMRTbell Template was prepared from the isolated viral DNA as previously described [64, 65], using standard protocols for 5-kb library preparation (PacBio “Procedure & Checklist- 5 kb Template Preparation and Sequencing). Briefly, for preparation of a 5kb library, 2 μg DNA was sheared by using g-TUBEs (Covaris) according to the manufacturer´s recommendations. DNA was end-repaired and ligated to the adapters using the DNA/Polymerase Binding Kit P6 v2 (Pacific Biosciences). Annealing and binding conditions of sequencing primers and polymerase to the purified SMRTbell Template were calculated using the Binding Calculator (Pacific Biosciences). Sequencing was carried out on the PacBio RSII long-read instrument (Pacific Biosciences) taking one 240-min movie for a single SMRT Cell with P6 DNA polymerase and C4 chemistry (P6-C4) yielding a total of 6,902 reads and a high coverage (~ 120-200×) across the genome. Sequencing reads were aligned against the PRV Ka reference genome (KJ717942.1) using the BLASR long read aligner (PacBio). The sequencing reads were mapped to the nucleotide sequence of GFP (U87625.1), to ensure that the construct was inserted to the PRV genome. Integrative Genomics Viewer (IGV) [66] was used for data visualization and mapping quality assurance. Artemis (Wellcome Trust Sanger Institute) [67] was also used for the visualization and analysis of the sequencing data.
Results
Genotyping of PRV ul54-KO mutant with PacBio SMRT sequencing
The presence of the deletion (Figure 1) in the PRV ul54-KO mutant virus was confirmed with Pacific Biosciences SMRT whole-genome sequencing (WGS) technique using a PacBio RSII sequencer and P6-C4 chemistry. The wt parental strain was sequenced earlier using the same method [16]. Comparison of the sequences of wt and mutant PRVs showed that the desired deletion was the only genetic difference between the two viruses; that is, no undesired genetic alterations took place during the generation of the knockout virus. The WGS platform that was used for genotyping eliminated the need for the generation and analysis of the kinetic properties of a rescue virus.
The effect of the ul54 gene on viral growth
In this part of the study, we investigated the effect of the ul54 gene deletion on the growth of PRV on cultured cells during a 24-h examination period using both low (MOI = 0.1 pfu/cell) and high (MOI = 10 pfu/cell) multiplicity of infection. Our analysis revealed a severe effect of the ul54 mutation on the viral growth in both cases (Figure 2, Supplementary Table 3). We also observed that the mutant virus does not cause any visible cytopathic effect even after 24 h p.i. (Figure 3).
Growth curves of wt and mutant PRVs. This figure shows that abrogation of the ul54 function exerts a drastic effect on the viral growth. In the MOI experiment the mutant virus reached a lower concentration than the wt virus by 24 h p. i. The low MOI experiment revealed that the mutant virus exhibits a slower rate of growth and reaches a much lower titer by 24 hpi
Cytopathic effect exerted by the wt and ul54-deleted viruses. High-magnification (200x) photomicrograph shows that – in contrast to the wt virus (A), which results in rounding of the infected cells – the shapes of the cells infected by the mutant virus exhibit normal phenotype (B) even after 24 hpi. Arrows show GFP expression from the cells infected by the ul54-KO virus
The effect of the ul54 gene deletion on the expression of PRV genes
We also investigated the effect of the mutation on viral gene expression. PK-15 cells were infected with either the wt or PRV ul54-KO virus, using low a MOI (0.1 pfu/cell) for the infection. The expressions of PRV genes were monitored within an 8 h period of time. The reason for using a short infection period was to exclude that mature viruses released from the infected cells could initiate a new cycle of replication in the non-infected cells at later periods. We observed that the mutation exerts a drastic effect on the PRV transcriptome. Compared to the wt, the PRV ul54-KO exhibits aberrant expression of ie180 gene and several E, E/L and L genes. The effect of the mutation on individual PRV genes was examined by using the Rr values, which were calculated as the ratio of the R values of mutant and wt virus at each time point (Table 4). We also calculated the impact of mutation on the average E, E/L and L transcripts (Table 5, Figure 4). It can be seen that on average the E genes are negatively affected by the mutation at first 1 h post infection (p.i.), while the expression levels become the same in the two viral backgrounds by 2 h p.i. This is in contrast to both the E/L and L genes, which are over-expressed in the mutant virus within the same period. This latter result is consistent with the observation made by Schwartz and colleagues, who have shown the accumulation of late viral product at the early period of infection [33]. An overall decline of transcript levels was observed at 4 h p.i. which may be related to the differential effect of the initiation of DNA synthesis on the two genotypes. The genes belonging to different kinetic classes behave dissimilarly by 6 h p.i.; there is a significant fall in the rate of expression of L genes, while the E genes appear to become unaffected by the mutation. Finally, all kinetic classes of PRV genes become considerably suppressed by 8 h p.i. Note that gene expressions are significantly lower in the mutant than in the wt background at 4 h p.i. in all PRV genes, except for the latency-associated transcript (LAT), which exhibits a 7.7 fold increase of expression in PRV ul54-KO (Table 4). By far the highest elevation in gene expression is detected from the ul53 gene (8.25 fold) in the mutant genome at 6 h p.i., and furthermore, this gene is the only gene that is expressed at a higher level at 8 h p.i. in the PRV ul54-KO than in the wt background. This phenomenon may be explained by the fact that ul53 and the deleted ul54 genes are adjacent to each other on the viral genome, and ul54 might exert a cis- or trans-acting suppressive effect on ul53, which is non-existent in the mutant virus. Intriguingly, the ul52 gene, which also produces co-terminal transcripts with the ul54 gene, is transcribed at a lower level (Rr = 0.39) in the mutant virus at this stage of infection.
The impact of the ul54 mutation on the expression of PRV genes. This plot shows the average Rr values of the three kinetic classes of PRV genes. The late genes are up-regulated at the early stage, while they are down-regulated at the late stage of infection in the mutant background. The early genes are down-regulated at 1h and 8h of infection in the ul54-KO virus. Black-filled circles with a straight line indicate the measured average Rr values of the E genes; white-filled triangles with a dashed line represent the E/L genes, while the values of L genes are labeled by black-filled squares with a dense dashed line
Correlations between gene expression
We calculated the “Rx” values showing the relative contribution of a transcript or a group of transcripts to the total transcriptome, and obtained a complementary pattern of gene expression in the two viruses for the IE, E and E/L genes. This was contrary to the L genes, whose expressions retained a similar pattern of gene expression throughout the entire period of infection. However, the kinetic curves of mutant and wt viruses do intersect one another at 4 h p.i., that is, the relative proportion of L genes becomes higher in the mutant than in the wt virus after 4 h (Figure 5), despite the fact that the expression of L genes in the two backgrounds exhibits a reverse pattern for the Rr values (Figure 2). This apparent contradiction can be explained by the fact that the difference in the amount of L transcripts is much larger in the early and late stage of infection in the wt virus than in PRV ul54-KO.
The expression dynamics - based on the Rx values - of the four kinetic classes of PRV genes in the two examined genetic backgrounds. a. The expression of ie180 gene in the wt and mutant viruses. The major transactivator gene of PRV exhibits an inverse expression pattern in the two background. b. The average gene expression values of the E genes in the two viruses show inverted dynamics. c. The Rx values of the E/L genes indicate a complementary pattern of expression in the wt and mutant viruses. d. The gene expression dynamics of the L genes show the same expression pattern between the two viruses
The transcription dynamics of the ie180 gene (illustrated by the RΔ values) are similar in the two genotypes, but the expression of the ep0 gene, another transcription regulator of PRV, exhibits complementary patterns in the two viruses (Figure 6a). To compare their expression in the same background, we can observe similar runs of the curves in the wt, while it is complementary in the mutant virus (Figure 6b). Furthermore, while in the wt virus the ie180/ast (Figure 6c) and ep0/lat (Figure 6d) gene pairs exhibit a perfect inverse pattern in their transcription dynamics, this is not the case in the mutant background, where their expression becomes non-correlated (ie180/ast) or becomes similar (ep0/lat). The relationship between the antisense transcript (AST) and LAT non-coding RNA molecules (produced from the complementary DNA strand of ie180 and ep0 genes, respectively) alters in such a way that in the mutant virus, their expression becomes dissimilar (Figure 6e), in contrast to the wt virus where the two transcripts exhibit almost identical expression profiles. Figure 6f shows the genomic region of PRV encoding the two transcription factors and the overlapping non-coding RNA molecules.
Comparisons of the expression patterns of transcription factor genes and their overlapping non-coding RNAs. A) Comparison of the expression kinetics of the ie180 and ep0 genes. The expression pattern of ie180 gene is similar, while the ep0 gene is complementary in the two genetic backgrounds. B) Comparison of the expression kinetics of ie180 and ep0 genes. The expression pattern of the two transcription factor genes is similar in the wt, while exhibits an inverse dynamics in the mutant virus. C) Comparison of the expression of ie180 and its antisense RNA genes. An inverse correlation can be observed between the transcription kinetics of IE180 and AST transcripts in the wt background, while no correlation exists between these transcripts in the mutant virus. D) Comparison of the expression of ep0 and its antisense RNA genes. An inverse correlation can be observed between the transcription kinetics of EP0 and LAT transcripts in the wt background, while these transcripts exhibit similar kinetics in the mutant virus. E) Comparison of the expression of LAT and AST transcripts. These antisense transcripts exhibit a similar expression pattern in the wt virus, while they express an inverse pattern in the mutant virus. F) The genomic location of ie180 and ep0 genes and their antisense RNAs
We also showed that the expressions of the ie180 and ep0 transcriptional activator genes in the mutant background exceed that of wt expression at 2 h p.i, while the AST peaks at 1 h p.i. and the LAT has the highest value at 4 h p.i., which also refers to a complementary pattern of gene expressions (Table 4).
The ul54 gene mutation exerts an inhibitory effect on the synthesis of viral DNA
Abrogation of ul54 gene function also affected the DNA synthesis of the mutant virus. The onset of replication exhibited a more than 2-h delay in PRV ul54-KO and a low copy number of DNA was produced when compared to the wt virus (Figure 7). The expression of the cto gene encoding a non-coding transcript [55] is also delayed by two hours and expressed in a very low amount in the mutant background (Figure 8). The CTO is supposed to interact with DNA replication [55]; the correlation between the delays in the two processes may therefore not be a coincidence. The time slip in the initiation of viral replication has a great impact on gene expression, especially for the L genes from 4 h p.i., whose expression is dependent on replication. At a later stage of the viral life cycle a global repression of gene expression in the mutant background was observed. This phenomenon is explained by the low copy number of the mutant viral DNA.
Expression of CTO non-coding RNA in the wt and mutant viruses. CTO expression is significantly reduced (similar relative expression /R/ values in the first 2 h pi, a ~ two-fold difference at 4h pi, a ~ six-fold difference at 6h pi, while an almost eight-fold difference at 8h pi) in the mutant background, which may partly explain the shift in the initiation of DNA synthesis
The effect of the ul54 gene mutation on gene expression from a single PRV genome
The dynamics of gene expression in the mutant virus also show a pattern, with the same 2h delay in genes normalized to the copy number of DNA molecules (Figure 9). Before the onset of DNA synthesis, the suppressive effect on the global gene expression may be exclusively attributed to the missing ul54 function, while later the lower copy number of viral DNA may also contribute to this phenomenon. In the late stage of infection, the single copies of genes are also expressed at a lower level in the mutant virus than in the wt if we compare the relevant time points (4 h wt to 6h mutant and 6 h wt to 8 h in mutant), which suggests a role for ICP27 in the control of gene expressions.
Discussion
In this study, we described the generation of a mutant PRV strain with a deletion at the ul54 locus and the transcriptional characterization of this virus in cultured cells using a real-time RT-PCR technique. We also analyzed the dynamics of viral DNA synthesis and correlated the obtained data on replication with the transcription patterns of the viral genes. Additionally, in this work, we also demonstrated the usefulness of long-read SMRT sequencing for genotyping of mutant viruses.
Here we show that the ul54 gene is not essential for the growth of PRV in tissue culture. However, our analysis showed that the abrogation of the ul54 function leads to a severe defect on viral replication. The deletion of ul54 gene exhibits a differential effect on the expression of PRV genes belonging to different kinetic classes. We observed an elevated level of L transcripts at the early stage of viral infection, as has also been shown by Schwartz and colleagues [33]. We also demonstrated that the average expression of E genes is suppressed at the very early stage of infection. The global gene expression is also suppressed by 8 h p.i. but at a different rate among the kinetic classes of viral genes; the expression of L genes was especially affected by the mutation. We explain this result by the dependency of L gene expression on DNA replication, which was also severely affected in the mutant virus. We can only speculate about the reason for the higher expression of L genes from the mutant background in the first 2 h p.i. The E genes are supposed to be under the direct control of the UL54 protein, which normally exerts an enhancing effect on the transcription of E genes, and hence the absence of enhancement results in a reduced expression of these genes at the early stage of viral infection. This event may account for the selective upregulation of the expression of L genes, perhaps through a cis-regulatory effect based on transcription interference between the transcriptional machineries [68]. The global decrease of gene expression from the mutant genome compared to that of wt is explained by the joint effect of the missing ul54 trans-activation of transcription and the low copy number of the mutant DNA molecules.
Intriguingly, the relative contributions of the ie180 and the average E and E/L genes exhibit a complementary pattern in the two genetic backgrounds throughout the 8 h examination period, while the expression curves of the L genes are similar in the two genotypes, except that the relative amounts of these gene products becomes higher in the mutant than in the wt virus from 4 h p.i. Theoretically, these results mean that the IE180 transactivator, on average, exerts a stimulatory effects on the E and E/L genes, while having a suppressive effect on the L genes. We obtained the same results in our analysis of the dynamic PRV transcripts using PacBio SMRT isoform sequencing [51].
The shape of the expression curves of the ie180 gene is very similar in the two genotypes, but they cross each other three times during the 8 h examination period (Figure 6a). This is in contrast to ep0 genes, where the RΔ expression curves show a complementary pattern, and in addition, the amount of EP0 transcript is drastically decreased in the mutant background throughout the entire infection period, which suggests that the ICP27 protein may have an important role in control of the expression of the ep0 gene (Figure 6b). In the wt virus, the two transactivator genes are expressed with similar dynamics, while gene expression becomes complementary in the mutant virus. The loss of synchronicity between the expression of ie180 and ep0 genes suggests a role for ICP27 in the expression kinetics of the ep0 gene, which is lost in the mutant virus.
We compared the relationships of the two transactivator genes (ie180 and ep0) with their antisense partners (ast and lat, respectively) in the 2 backgrounds and obtained inverse expression curves in the wt, and similar curves in the mutant virus. Since the two transcripts exhibit a full-length overlap, their transcription may interfere with one other, which explains the inverse expression pattern in the wt virus. However, this relationship collapses in the ul54-deleted virus. The reason for this phenomenon may be the very low level of transcription from both the ep0 and lat genes, which could result in a decrease of the transcriptional interaction between the two overlapping partners.
We compared the kinetics of LAT and AST antisense transcripts, and observed that they are expressed synchronously in the wt, while they are transcribed in an inverse pattern in the mutant virus (Figure 6e). This phenomenon may be related to altered expression of the transactivator genes encoded in the same genomic region.
The onset of replication exhibits a significant delay in, and a lower copy number of, the DNA in the mutant virus, which exerts a significant effect on gene expression. The E genes encoding the synthetic machinery of PRV are expressed at a low level in the ul54 deletion mutant. This phenomenon may be the reason for the late onset of DNA replication. Furthermore, there is also a time slip in the expression of CTO non-coding transcript, which is proposed to play a role in the control of DNA synthesis through an interaction between the transcription and replication machineries [61]. Perhaps, this putative mechanism also contributes to the delayed kinetics of DNA replication. Low amounts of E genes produces few viral DNAs, while low levels of L genes produces few building blocks for the capsids and envelopes; these two phenomena could underlie the growth defect of the mutant PRV reported in this study and also by Schwartz and colleagues [33]. These two reports had obtained very similar results in the growth analyses for both wt and ul54-KO viruses at both low and high MOI. The only significant difference between the two studies was that we included the 18 h post infection period for the analysis, which proved to be the time point where the wt virus produced the highest number of viral particles in the high MOI experiment. Earlier, we had already undertaken an investigation of the impact of gene deletions on the global transcriptome. For example, we have demonstrated that in the early stage of infection, the de novo VHS protein exerts a negative effect on the expression level of early mRNAs [61]. We also have shown that the effect of VHS protein on the amount of E/L gene transcripts is intermediate, while the expression of the L viral transcripts is slightly affected by the VHS. We have shown [69] that EP0 protein exerts a selective negative effect on the expression level of early mRNAs in the very early stage of infection and affects the amounts of mRNAs without bias toward any kinetic classes in the late phase. In another study, we have shown [70] that the deletion of the us1 gene leads to a general, but significant reduction of PRV gene expression in the first hour p.i. However, in the mid-phase of infection (between 1-6h pi), the L genes are selectively down-regulated, while later the early genes are up-regulated in the us1-deleted virus.
In summary, we have established that the abrogation of ul54 function leads to a differential effect on the various kinetic classes of the PRV genes. This effect may be direct at the early phase of gene expression, but later this mutation likely exerts its influence on global gene expression at least partly through DNA replication, which is impeded in comparison to the wt virus, and through an effect on other transcription factors, such as IE180 and EP0. In future studies the mechanism of action has to be investigated for each individual gene.
References
Aujeszky A (1902) A contagious disease, not readily distinguishable from rabies, with unknown origin. Veterinarius 25:387–396 (in Hungarian)
Szpara ML, Kobiler O, Enquist LW (2010) A common neuronal response to alphaherpesvirus infection. J Neuroimmune Pharmacol 5(3):418–427
Card JP, Enquist LW (2001) Transneuronal circuit analysis with pseudorabies viruses. Curr Protoc Neurosci. doi:10.1002/0471142301.ns0105s68
Boldogkői Z, Sík A, Dénes A, Reichart A, Toldi J, Gerendai I, Kovács KJ, Palkovits M (2004) Novel tracing paradigms-genetically engineered herpesviruses as tools for mapping functional circuits within the CNS: present status and future prospects. Prog Neurobiol 72(6):417–445
Yang M, Card JP, Tirabassi RS, Miselis RR, Enquist LW (1999) Retrograde, transneuronal spread of pseudorabies virus in defined neuronal circuitry of the rat brain is facilitated by gE mutations that reduce virulence. J Virol 73:4350–4359
Song CK, Enquist LW, Bartness TJ (2005) New developments in tracing neural circuits with herpesviruses. Virus Res 111:235–249
Granstedt AE, Kuhn B, Wang SS, Enquist LW (2010) Calcium imaging of neuronal circuits in vivo using a circuit-tracing pseudorabies virus. Cold Spring Harb Protoc. doi:10.1101/pdb.prot5410
Card JP, Kobiler O, Ludmir EB, Desai V, Sved AF, Enquist LW (2011) A dual infection pseudorabies virus conditional reporter approach to identify projections to collateralized neurons in complex neural circuits. PLoS One. doi:10.1371/journal.pone.0021141
Boldogkői Z, Balint K, Awatramani GB, Balya D, Busskamp V, Viney TJ, Lagali PS, Duebel J, Pásti E, Tombácz D, Tóth JS, Takács IF, Scherf BG, Roska B (2009) Genetically timed, activity-sensor and rainbow transsynaptic viral tools. Nat Methods 6:127–130
Granstedt AE, Szpara ML, Kuhn B, Wang SS, Enquist LW (2009) Fluorescence-based monitoring of in vivo neural activity using a circuit-tracing pseudorabies virus. PLoS One. doi:10.1371/journal.pone.0006923
Prorok J, Kovács PP, Kristóf AA, Nagy N, Tombácz D, Tóth JS, Ördög B, Jost N, Virág L, Papp JG, Varró A, Tóth A, Boldogkői Z (2009) Herpesvirus-mediated delivery of a genetically encoded fluorescent Ca(2+) sensor to canine cardiomyocytes. J Biomed Biotechnol. doi:10.1155/2009/361795
Boldogkői Z, Nógrádi A (2003) Gene and cancer therapy—pseudorabies virus: a novel research and therapeutic tool? Curr Gene Ther 3:155–182
Zhu L, Yi Y, Xu Z, Cheng L, Tang S, Guo W (2011) Growth, physicochemical properties, and morphogenesis of Chinese wild-type PRV Fa and its gene-deleted mutant strain PRV SA215. Virol J 8:272
Maresch C, Lange E, Teifke JP, Fuchs W, Klupp B, Müller T, Mettenleiter TC, Vahlenkamp TW (2012) Oral immunization of wild boar and domestic pigs with attenuated live vaccine protects against pseudorabies virus infection. Vet Microbiol 161:20–25
Klingbeil K, Lange E, Teifke JP, Mettenleiter TC, Fuchs W (2014) Immunization of pigs with an attenuated pseudorabies virus recombinant expressing the hemagglutinin of pandemic swine origin H1N1 influenza A virus. J Gen Virol 95:948–959
Tombácz D, Sharon D, Oláh P, Csabai Z, Snyder M, Boldogkői Z (2014) Strain kaplan of pseudorabies virus genome sequenced by PacBio single-molecule real-time sequencing technology. Genome Announc. doi:10.1128/genomeA.00628-14
Tombácz D, Csabai Z, Oláh P, Balázs Z, Likó I, Zsigmond L, Sharon D, Snyder M, Boldogkői Z (2016) Full-length isoform sequencing reveals novel transcripts and substantial transcriptional overlaps in a herpesvirus. PLoS One. doi:10.1371/journal.pone.0162868
Tombácz D, Tóth JS, Petrovszki P, Boldogkoi Z (2009) Whole-genome analysis of pseudorabies virus gene expression by real-time quantitative RT-PCR assay. BMC Genomics. doi:10.1186/1471-2164-10-491
Flori L, Rogel-Gaillard C, Cochet M, Lemonnier G, Hugot K, Chardon P, Robin S, Lefèvre F (2008) Transcriptomic analysis of the dialogue between Pseudorabies virus and porcine epithelial cells during infection. BMC Genomics. doi:10.1186/1471-2164-9-123
Oláh P, Tombácz D, Póka N, Csabai Z, Prazsák I, Boldogkői Z (2015) Characterization of pseudorabies virus transcriptome by Illumina sequencing. BMC Microbiol. doi:10.1186/s12866-015-0470-0
Anderson K, Costa RH, Holland LE, Wagner EK (1980) Characterization of herpes simplex virus type 1 RNA. Present in the absence of de novo protein synthesis. J Virol 34(1):9–27
Mackem S, Roizman B (1980) Regulation of herpesvirus macromolecular synthesis: transcription-initiation sites and domains of α genes. Proc Natl Acad Sci USA 77(12):7122–7126
Ihara S, Feldman L, Watanabe S, Ben-Porat T (1983) Characterization of the immediate-early functions of pseudorabies virus. Virology 131:437–454
Cheung AK (1991) Cloning of the latency gene and the early protein 0 gene of pseudorabies virus. J Virol 65:5260–5271
Huang C, Wu CY (2004) Characterization and expression of the pseudorabies virus early gene UL54. J Virol Methods 119:129–136
Fuchs W, Ehrlich C, Klupp BG, Mettenleiter TC (2000) Characterization of the replication origin (Ori(S)) and adjoining parts of the inverted repeat sequences of the pseudorabies virus genome. J Gen Virol 81:1539–1543
Zhang G, Leader DP (1990) The structure of the pseudorabies virus genome at the end of the inverted repeat sequences proximal to the junction with the short unique region. J Gen Virol 71:2433–2441
Baumeister J, Klupp BG, Mettenleiter TC (1995) Pseudorabies virus and equine herpesvirus 1 share a nonessential gene which is absent in other herpesviruses and located adjacent to a highly conserved gene cluster. J Virol 6(9):5560–5567
Huang YJ, Chien MS, Wu CY, Huang C (2005) Mapping of functional regions conferring nuclear localization and RNA-binding activity of pseudorabies virus early protein UL54. J Virol Methods 130(1–2):102–107
Sacks WR, Greene CC, Aschman DP, Schaffer PA (1985) Herpes simplex virus type 1 ICP27 is an essential regulatory protein. J Virol 55:796–805
Gruffat H, Batisse J, Pich D, Neuhierl B, Manet E, HammerschmidtW Sergeant A (2002) Epstein-Barr virus mRNA export factor EB2 is essential for production of infectious virus. J Virol 76(19):9635–9644
Sato B, Sommer M, Ito H, Arvin AM (2003) Requirement of varicella-zoster virus immediate-early 4 protein for viral replication. J Virol 77(22):12369–12372
Schwartz JA, Brittle EE, Reynolds AE, Enquist LW, Silverstein SJ (2006) UL54-null pseudorabies virus is attenuated in mice but productively infects cells in culture. J Virol 80(2):769–784
Hardwicke MA, Sandri-Goldin RM (1994) The herpes simplex virus regulatory protein ICP27 contributes to the decrease in cellular mRNA levels during infection. J Virol 68:4797–4810
McGregor F, Phelan A, Dunlop J, Clements JB (1996) Regulation of herpes simplex virus poly(A) site usage and the action of immediate-early protein IE63 in the early-late switch. J Virol 70(3):1931–1940
Hayashi ML, Blankenship C, Shenk T (2000) Human cytomegalovirus UL69 protein is required for efficient accumulation of infected cells in the G1 phase of the cell cycle. Proc Natl Acad Sci USA 97:2692–2696
Li M, Wang S, Cai M, Guo H, Zheng C (2011) Characterization of molecular determinants for nucleocytoplasmic shuttling of PRV UL54. Virology 417(2):385–393
Li M, Wang S, Cai M, Zheng C (2011) Identification of nuclear and nucleolar localization signals of pseudorabies virus (PRV) early protein UL54 reveals that its nuclear targeting is required for efficient production of PRV. J Virol 85(19):10239–10251
Elhai J, Wolk CP (1988) A versatile class of positive-selection vectors based on the nonviability of palindrome-containing plasmids that allows cloning into long polylinkers. Gene 68(1):119–138
Some genetic functions encoded by herpes simplex virus type 1. http://darwin.bio.uci.edu/~faculty/wagner/table.html
The genetic and transcription map of the HSV-1 genome. http://darwin.bio.uci.edu/~faculty/wagner/hsvimg04z.jpg
Roizman B (1996) The function of herpes simplex virus genes: A primer for genetic engineering of novel vectors. Proc Nat Acad Sci USA 93:11307–11312
Roizman B, Campadelli-Fiume G (2007) Alphaherpes viral genes and their functions. In: Arvin A, Campadelli-Fiume G, Mocarski E, Moore PS, Roizman B, Whitley R, Yamanishi K (eds) Human herpesviruses—biology, therapy and immunoprophylaxis, 1st edn. Cambridge University Press, Cambridge, pp 70–92
Tombácz D, Balázs Z, Csabai Z, Moldován N, Szűcs A, Sharon D, Snyder M, Boldogkői Z (2017) Characterization of the dynamic transcriptome of a herpesvirus with long-read single molecule real-time sequencing. Sci Rep. doi:10.1038/srep43751
Klupp BG, Hengartner CJ, Mettenleiter TC, Enquist LW (2004) Complete, annotated sequence of the pseudorabies virus genome. J Virol 78:424–440
Bras F, Dezelee S, Simonet B, Nguyen X, Vende P, Flamand A, Masse MJ (1999) The left border of the genomic inversion of pseudorabies virus contains genes homologous to the UL46 and UL47 genes of herpes simplex virus type 1, but no UL45 gene. Virus Res 60:29–40
Pomeranz LE, Reynolds AE, Hengartner CJ (2005) Molecular biology of pseudorabies virus: impact on neurovirology and veterinary medicine. Microbiol Mol Biol Rev 69(3):462–500
Reynolds AE, Fan Y, Baines JD (2000) Characterization of the UL33 gene product of herpes simplex virus 1. Virology 266:310–318
Klupp BG, Fuchs W, Granzow H, Nixdorf R, Mettenleiter TC (2002) Pseudorabies virus UL36 tegument protein physically interacts with the UL37 protein. J Virol 76:3065–3071
Ambagala AP, Hinkley S, Srikumaran S (2000) An early pseudorabies virus protein down-regulates porcine MHC class I expression by inhibition of transporter associated with antigen processing (TAP). J Immunol 164:93–99
de Wind N, Berns A, Gielkens A, Kimman T (1993) Ribonucleotide reductase-deficient mutants of pseudorabies virus are avirulent for pigs and induce partial protective immunity. J Gen Virol 74:351–359
Powers L, Wilkinson KS, Ryan P (1994) Characterization of the prv43 gene of pseudorabies virus and demonstration that it is not required for virus growth in cell culture. Virology 199:81–88
Robbins AK, Watson RJ, Whealy ME, Hays WW, Enquist LW (1986) Characterization of a pseudorabies virus glycoprotein gene with homology to herpes simplex virus type 1 and type 2 glycoprotein. J Virol 58(2):339–347
Dezélée S, Bras F, Vende P, Simonet B, Nguyen X, Flamand A, Masse MJ (1996) The BamHI fragment 9 of pseudorabies virus contains genes homologous to the UL24, UL25, UL26, and UL 26.5 genes of herpes simplex virus type 1. Virus Res 42:27–39
Tombácz D, Csabai Z, Oláh P, Havelda Z, Sharon D, Snyder M, Boldogkői Z (2015) Characterization of novel transcripts in pseudorabies virus. Viruses 7(5):2727–2744
Yamada S, Imada T, Watanabe W, Honda Y, Nakajima-Iijima S, Shimizu Y, Sekikawa K (1991) Nucleotide sequence and transcriptional mapping of the major capsid protein gene of pseudorabies virus. Virology 185:56–66
Dijkstra JM, Fuchs W, Mettenleiter TC, Klupp BG (1997) Identification and transcriptional analysis of pseudorabies virus UL6 to UL12 genes. Arch Virol 142:17–35
Dean H, Cheung AK (1993) A 3’coterminal gene cluster in pseudorabies virus contains herpes simplex virus UL1, UL2, UL3 gene homologs and a unique UL3.5 open reading frame. J Virol 67:5955–5961
Krause PR, Croen KD, Ostrove JM, Straus SE (1990) Structural and kinetic analyses of herpes simplex virus type I latencyassociated transcripts in human trigeminal ganglia and in cell culture. J Clin Invest 86(1):235–241
Brideau AD, Banfield BW, Enquist LW (1998) The Us9 gene product of pseudorabies virus, an alphaherpesvirus, is a phosphorylated, tail-anchored type II membrane protein. J Virol 72:4560–4570
Tombácz D, Tóth JS, Boldogkoi Z (2011) Deletion of the virion host shut: off gene of pseudorabies virus results in selective upregulation of the expression of early viral genes in the late stage of infection. Genomics 98(1):15–25
Campbell AM, Heyer LJ (2007) Basic research with DNA microarray. In: Winslow S (ed) Discovering genomics proteomics and bioinformatics, 2nd edn. CSHL Press, San Francisco, pp 238–241
Mestdagh P, Van Vlierberghe P, De Weer A, Muth D, Westermann F, Speleman F, Vandesompele J (2009) A novel and universal method for microRNA RT-qPCR data normalization. Genome Biol. doi:10.1186/gb-2009-10-6-r64
Travers KJ, Chin CS, Rank DR, Eid JS, Turner SW (2010) A flexible and efficient template format for circular consensus sequencing and SNP detection. Nucleic Acids Res. doi:10.1093/nar/gkq543
Clark TA, Murray IA, Morgan RD, Kislyuk AO, Spittle KE, Boitano M, Fomenkov A, Roberts RJ, Korlach J (2012) Characterization of DNA methyltransferase specificities using single-molecule, real-time DNA sequencing. Nucleic Acids Res. doi:10.1093/nar/gkr1146
Thorvaldsdóttir H, Robinson JT, Mesirov JP (2013) Integrative genomics viewer (IGV): high-performance genomics data visualization and exploration. Brief Bioinform 14(2):178–192
Carver T, Harris SR, Berriman M, Parkhill J, McQuillan JA (2012) Artemis: an integrated platform for visualization and analysis of high-throughput sequence-based experimental data. Bioinformatics 28(4):464–469
Boldogkői Z (2012) Transcriptional interference networks coordinate the expression of functionally related genes clustered in the same genomic loci. Front Genet. doi:10.3389/fgene.2012.00122
Tombácz D, Tóth JS, Boldogkői Z (2012) Effects of deletion of the early protein 0 gene of pseudorabies virus on the overall viral gene expression. Gene. doi:10.1016/j.gene.2011.11.049
Takács IF, Tombácz D, Berta B, Prazsák I, Póka N, Boldogkői Z (2013) The ICP22 protein selectively modifies the transcription of different kinetic classes of pseudorabies virus genes. BMC Mol Biol. doi:10.1186/1471-2199-14-2
Acknowledgements
This research was supported by the Swiss–Hungarian Cooperation Programme SH/7/2/8 to ZB, by the European Social Fund in the framework of TÁMOP 4.2.4. A/2-11-1-2012-0001 “National Excellence Program” to DT. This research was also supported by the Bolyai János Scholarship of the Hungarian Academy of Sciences: 2015-18 to DT. This study was also supported by the NIH CEGS Grant 5P50HG00773502 to MS.
Author information
Authors and Affiliations
Contributions
ZB, DT and MS conceived and designed the study; ZC, IFT, DT and ZB performed the research; DT, ZC and ZB analyzed the data; ZB and DT wrote the paper.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no conflict of interest. The founding sponsors had no role in the design of the study; in the collection, analyses, or interpretation of data; in the writing of the manuscript, and in the decision to publish the results.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Csabai, Z., Takács, I.F., Snyder, M. et al. Evaluation of the impact of ul54 gene-deletion on the global transcription and DNA replication of pseudorabies virus. Arch Virol 162, 2679–2694 (2017). https://doi.org/10.1007/s00705-017-3420-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00705-017-3420-3