Abstract
Few reports are available on HCV molecular epidemiology among IDUs in Eastern Europe, and none in Montenegro. The aim of this study was to investigate the HCV genotype distribution in Montenegro among IDUs and to perform Bayesian and evolutionary analysis of the most prevalent HCV genotype circulating in this population. Sixty-four HCV-positive IDUs in Montenegro were enrolled between 2013 and 2014, and the NS5B gene was sequenced. The Bayesian analysis showed that the most prevalent subtype was HCV-3a. Phylogenetic data showed that HCV-3a reached Montenegro in the late 1990s, causing an epidemic that exponentially grew between the 1995 and 2005. In the dated tree, four different entries, from 1990 (clade D), 1994 (clade A) to 1999 (clade B) and 2001 (clade C), were identified. In the NS5B protein model, the amino acids variations were located mainly in the palm domain, which contains most of the conserved structural elements of the active site. This study provides an analysis of the virus transmission pathway and the evolution of HCV genotype 3a among IDUs in Montenegro. These data could represent the basis for further strategies aimed to improve disease management and surveillance program development in high-risk populations.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Hepatitis C virus (HCV) is the leading cause of chronic liver disease, cirrhosis, and hepatocellular carcinoma in developed countries [1]. It is a positive-strand RNA virus that belongs to the family Flaviviridae. Like the majority of other RNA viruses, it is characterized by a high level of genetic variability that is particularly marked at the level of the E1 and E2 genes, which encode viral envelope glycoproteins [2].
At least six major genotypes and a number of subtypes with different ethno-geographical distributions have been identified. Phylogenetic studies indicate that genotypes 1, 2, 4 and 5 originated in sub-Saharan Africa; and genotypes 3 and 6, in Southeast Asia [3]. Most of the HCV infections worldwide are caused by a small subset of “epidemic” HCV subtypes, including subtypes 1a, 1b, 2b, 3a and 4a [4]. It is widely agreed that these subtypes spread rapidly throughout the world during the 20th century, mainly because of transfusions, use of blood products, unsafe medical injections, and injection drug use [5, 6]. Antiviral medicines can cure approximately 90% of individuals with hepatitis C infection, thereby reducing the risk of death from liver cancer and cirrhosis, but access to diagnosis and treatment is low (http://www.who.int/en/).
According to WHO estimates, at least 17 million people living in Eastern Mediterranean countries are HCV carriers, emphasizing the importance of HCV infection in this part of the world [1, 7, 8].
The epidemiology of hepatitis C in the different European regions is diverse, with a prevalence of anti-HCV antibodies ranging from very low (0.1% in Ireland) to high (5% in Italy and 13% in Uzbekistan), (http://www.who.int/en/).
In Europe, the main route of HCV transmission is via injecting drug use because of sharing contaminated needles (http://ecdc.europa.eu/). There is a wide variety in the reported HCV prevalence in IDUs, ranging from 25% to 70%. Italy reported the lowest prevalence (10.8–25.6%); and Norway, the highest (70%). Slovenia reported prevalence below 25% in national samples of injecting drug users (http://ecdc.europa.eu/en/healthtopics/hepatitis_C/Pages/index.aspx).
Unfortunately, little is known about the prevalence of HCV infection in Serbia, Montenegro or the Balkans in general [8, 9]. The only available information relates to the prevalence of HCV in Montenegro and Bulgaria, where the most prevalent subtypes, 1B and 3A, respectively, have been found [7–11]. Additionally, a few reports are available on the molecular epidemiology of HCV infection among injecting drug users in Eastern Europe [12, 13], but there are none on injecting drug users in Montenegro.
The aim of this study was to investigate HCV subgenotype distribution in Montenegro in a homogeneous population consisting of IDUs and to perform Bayesian and evolutionary analysis of the most prevalent HCV genotype circulating in this high-risk population. In the same population, selective pressure analysis and homology modelling were applied to identify positively selected sites and to assess their influence on NS5B protein structure and function.
Materials and methods
Patients
Sixty-four anti-HCV- and HCV-RNA-positive IDUs living in Montenegro were consecutively enrolled between 2013 and 2014 at the Institute of Public Health of Podgorica, and the serum samples were further characterized by NS5B gene sequencing. Demographic and epidemiological data were available for all patients.
Sample processing and HCV RNA sequencing
Viral RNA was extracted from the chronic patients’ serum (200 μl stored at −80°C) using NucleoMag 96 Virus (Macherey-Nagel, Düren, Germany) and automated KingFisher™ ml Magnetic Particle Processors (Thermo Scientific) in accordance with the manufacturer’s instructions. Serum samples from healthy subjects were used as negative controls. The RNA was eluted in 50 μl of nuclease-free distilled water and reverse transcribed using the SuperScript III reverse transcriptase protocol (Thermo Fisher Scientific Inc., Waltham, MA), and the cDNA was amplified by means of nested PCR using GoTaq® DNA Polymerase (Promega). The primers for the first and second rounds of NS5B amplification and the PCR conditions have been described previously [8, 14].
The fragments obtained by means of PCR were purified using a commercial purification kit (QIAquick PCR Purification Kit) and then sequenced bidirectionally using a BigDye Terminator Kit version 3.1 (Applied Biosystems) according to the manufacturer’s instructions.
The sequencing products were purified from a 10-μl sample by precipitating in an ethanol/sodium acetate mixture. Finally, the sequences were determined using an automated DNA sequencer (ABI PRISM 3130 XL Genetic Analyser, Applied Biosystems). Sixty-four sequences were successfully amplified.
Sequences have been deposited at DDBJ/ENA/GenBank under the accession numbers KY379287 to KY379319.
Phylogenetic analysis
HCV datasets
For phylogenetic analysis, four datasets were built. The first dataset was used for the genotype/subtype assignment and included 131 sequences: all 64 of the Montenegrin sequences for this study and 67 representative sequences of the main genotypes/subtypes of HCV that were downloaded from GenBank (http://www.ncbi.nlm.nih.gov/Genbank).
The second dataset included 33 sequences obtained in this study, plus all of the available reference sequences (61 sequences from Australia, Belgium, Canada, France, Great Britain, the Netherlands, and Uzbekistan) downloaded from GenBank (http://www.ncbi.nlm.nih.gov/Genbank) that met the following criteria: i) there was no uncertainty about the 3a subtype assignment; ii) sampling locations were known and clearly established in the original publication; and iii) sequences from IDUs were established clearly in the original publication.
The third dataset was prepared to detect the presence of a phylogenetic signal and to investigate the phylodynamics of the HCV subtype most frequently found in Montenegrin IDU patients. This dataset included 46 HCV-NS5B sequences of HCV subtype 3a; 33 sequences were obtained in this study and 13 additional sequences of subtype 3a strains from Montenegrin IDUs were obtained from a previous study [8]. Strict and relaxed molecular clock methods as well as different demographic coalescent models were tested to infer the demographic history of HCV in Montenegrin IDUs. We evaluated two parametric estimates (constant effective population size and exponential population growth) and one nonparametric estimate (Bayesian skyline plot) of the viral population size over time.
The fourth dataset included only the 33 sequences identified in the current study with subtype 3a, and this was used to evaluate the selective pressure and for homology modelling.
Alignment, model selection and genotype/subtype characterization
All strains were aligned using both ClustalX [15] and ClustalW [16] software included in BioEdit, followed by manual editing (final alignment length = 360nt).
The JModelTest [17] was used for all datasets to select the simplest evolutionary model fitting the data, which was the GTR+G model of nucleotide substitution [18].
The genotype/subtype was determined by phylogenetic analysis of the NS5B gene sequences of the first dataset, using MrBayes [19] with the previously selected model.
A Bayesian phylogenetic tree was constructed using the second dataset with the best-fitting substitution model chosen by JModelTest [17] (GTR+G), using MrBayes [19].
A Markov chain Monte Carlo (MCMC) search was made for 5 × 106 generations using tree sampling every 100th generation and a burn-in fraction of 50%. Statistical support for specific clades was obtained by calculating the posterior probability of each monophyletic clade, and a posterior consensus tree was generated after a 50% burn-in.
The tree was displayed and edited using Figtree software v 1.4.1, which is freely available on the web (http://tree.bio.ed.ac.uk/software/figtree/).
Phylodynamics
Likelihood mapping analysis. In order to obtain an overall impression of the phylogenetic signal present in the phylodynamic dataset (the third dataset), we made a likelihood-mapping analysis of 10,000 random quartets generated using TreePuzzle [20]. A likelihood map consists of an equilateral triangle: each dot within the triangle represents the likelihood of the three possible unrooted trees for a set of four sequences (quartets) randomly selected from the dataset. The dots close to the corners or at the sides respectively represent tree-like (fully resolved phylogenies in which one tree is clearly better than the others) or network-like phylogenetic signals (three regions for which it is not possible to decide between two topologies); the central area of the map represents a star-like signal (the region where the star tree is the optimal tree).
Evolutionary demography reconstruction
For the third dataset, the time-scaled phylogeny, evolutionary rates, and demographic models were co-estimated using the Bayesian Markov chain Monte Carlo (MCMC) method implemented in the BEAST package version 1.8.0 [21]. Strict and relaxed clocks with an uncorrelated log normal rate of distribution under a less restrictive Bayesian skyline plot (BSP, a non-parametric piecewise-constant model) as coalescent prior were estimated [21]. The nucleotide substitution model (GTR+G) was selected as described previously [18].
The best-fitting models were selected using a Bayes factor (BF, using marginal likelihoods) as implemented in BEAST [22]. In accordance with Kass and Raftery [23], the strength of the evidence against H0 was evaluated as 2lnBF <2 = no evidence; 2–6 = weak evidence; 6–10 = strong evidence; and >10 = very strong evidence. A negative 2lnBF indicates evidence in favor of H0. Only values greater than 6 were considered significant. In particular, BF analysis showed that the relaxed lognormal molecular clock fitted the data better than the strict clock model (2lnBF = 59.7).
Statistical support for specific clades was obtained by calculating the posterior probability of each monophyletic clade. Accordingly, with our previous estimate [8], the phylogeny was calibrated by adjusting the mean substitution rate to the mean external evolutionary rate estimates (1.3 × 10−3 sub/site/years, with a credibility interval of 1.04–1.48 × 10−3).
The MCMC chains were run for at least 50 million generations and sampled every 5,000 steps. Convergence was assessed on the basis of the effective sampling size (ESS) after a 10% burn-in [21], using Tracer software version 1.5 (http://tree.bio.ed.ac.uk/software/tracer/). Only ESS values of >250 were accepted, and uncertainty in the estimates was indicated by 95% highest posterior density (95% HPD) intervals. The final tree was manipulated in FigTree version 1.4.1 for display purposes.
Selective pressure analysis
The CODEML program implemented in the PAML 3.14 software package (http://abacus.gene.ucl.ac.uk/software/paml.html) [24] was used to investigate the adaptive evolution of the NS5B protein using the fourth dataset including 33 Montenegrin isolates. Six models of codon substitution: M0 (one-ratio), M1a (nearly neutral), M2a (positive selection), M3 (discrete), M7 (beta), and M8 (beta and omega) were used in this analysis [25]. Since these models are nested, we used codon-substitution models to fit the model to the data using the likelihood ratio test (LRT) [26]. The dN/dS rate (ω) was also estimated by the ML approach implemented in the program HyPhy [27]. Site-specific positive and negative selection was estimated by two different algorithms: the fixed-effects likelihood (FEL) algorithm, which fits an ω rate to every site and uses the likelihood ratio to test if dN = dS, and the random-effects likelihood algorithm (REL), a variant of the Nielsen–Yang approach [26] that assumes that a discrete distribution of rates exists across sites and allows both dS and dN to vary independently site by site. The three methods have been described in more detail elsewhere [28]. In order to select sites that are under selective pressure and keep our test conservative, a P-value of ≤ 0.1 or a posterior probability of ≥ 0.9 as relaxed critical values was assumed. For evolutionary analysis, the reference sequence with the accession number YP_001491557 was used to trace the exact position of the amino acids found to be under selection.
Residue conservation analysis and homology modelling
The nucleotide sequence alignment of the NS5B gene sequence dataset was used to generate a conceptual translation to the corresponding peptide sequences, using UGene software [29]. The amino acid sequences were then aligned, using Clustal Omega [30], to the sequence of the reference RNA-dependent RNA polymerase sequence from hepatitis C virus genotype 3 subtype 3a (NS5B-3a, RefSeq accession no.: YP_001491557). The variability of NS5B polymerase protein sequences was assessed by calculating the prevalence of the most common wild-type amino acid at each position of the alignment. The amino acid conservation was defined as the percentage of sites with <10% residue variability.
Since the NS5B-3a structure was not available, homology modelling was used to generate a three-dimensional model. The availability of structural templates in the Protein Data Bank was checked using BLASTp [31]. The structure showing the highest quality, resolution and degree of sequence similarity to HCV NS5B-3a polymerase was selected. The alignment of the target sequence with the selected template was calculated using Clustal Omega. A total of ten homology models was generated and optimized using Modeller 9.13 [32]. The model with the best values of the Modeller scoring function was selected for subsequent analysis. The model was validated using the standard programs ProsaII [33] and Procheck [34]. Residue conservation was evaluated using the Consurf server [35]. Alignment display and editing were done using UGene. Protein structure analysis, in silico mutagenesis, and figure design were performed using PyMOL [36]. Two similar and independent tools, I-Mutant-2.0 [37] and CUPSAT [38], were used to analyze the overall impact of each amino acid variation on protein stability. Both programs predict the stability of a point-mutated protein, based on its protein sequence or 3-D structure, respectively. The output file shows the predicted free energy change (ΔΔG), which is calculated as the unfolding Gibbs free energy change of the mutated protein minus the unfolding free energy value of the native protein (units, kcal/mol). The overall impact of a point mutation on protein stability was evaluated using a consensus criterion: a result was accepted only if both methods predicted the same effect on the protein structure, i.e., destabilizing or stabilizing.
Results
Characteristics of patients and HCV genotype/subtype distribution
Among 64 IDUs patients with persistent HCV infection living in Montenegro, 53 were males (82.8%) and 11 females (17.2%). The median age was 31 years (range 19–51).
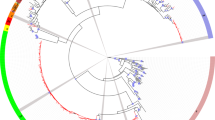
A phylogenetic tree (Fig. 1) constructed using the first dataset showed different statistically supported clades, corresponding to different HCV genotype/subgenotypes. A Bayesian tree showed the following HCV subtype distribution in Montenegro (Fig. 1): 33 HCV-3a (51.6%), 15 1a (23.4%), 13 4d (20.3%) and three 1b (4.7%). The only subtype not found in females was 1b. Furthermore, for each Montenegrin HCV genotype, statistically supported monophyletic clades (pp ranging from 0.80 to 1.0) were identified (Fig. 1). Moreover, when the population was divided into three age groups (19-30, 31–40 and 41–51 years), no differences were observed in the subtype distribution.
Bayesian tree of 64 HCV NS5B Montenegrin sequences. The Montenegrin strains are indicated in red. The country of origin for different reference sequences is shown in blue. The clades corresponding to the main identified subtypes are marked. Posterior probabilities > 0.8 are shown at nodes. Horizontal branch lengths are drawn to scale, with the bar at the bottom indicating 0.2 nucleotide substitutions per site (colour figure online)
A phylogenetic tree (Fig. 2) based on the second dataset showed several supported clusters. Four main statistically supported clades were identified (clades A, B, C and D). These clades included only Montegrin isolates. The tree also showed intermixing clusters including Montegrin and foreign sequences, but these were not statistically supported.
Phylodynamic analysis of HCV-3a
Likelihood mapping analysis of the third dataset of HCV-3a sequences from IDUs showed that the percentage of dots falling in the central area of the triangles was 6.8%, indicating a fully resolved phylogenetic signal (Fig. 1S). The Bayes Factor (BF) strongly favored the relaxed over the strict molecular clock model (BF = 39.4), indicating that different viral strains evolved at significantly different rates. In addition, analysis of the three demographic models showed positive evidence against the null hypothesis of a constant virus population size in favor of the exponential growth model (BF = 3.7), which also outperformed the Bayesian skyline plot model (BF = 4.9).
Based on the dated tree, calibrated on the external evolutionary rate (see Materials and methods) (Fig. 3), the time to the most recent common ancestor (TMRCA) of the root was estimated to be 54 years (95% HPD = 25-93 years), corresponding to 1960.
Phylodynamic reconstruction revealed four highly supported clades including more than three strains each. The HCV 3a strains retrieved from a previous study were scattered among sequences obtained in this study. Clade A (pp = 0.98), including nine sequences (five obtained in 2014, two in 2008, one in 2007 and one in 2009), showed a TMRCA of 20 years (95% HPD = 12-29 years), corresponding to 1994. Clade B (pp = 1) included four strains (three sampled in 2014 and one in 2008) with a TMRCA of 15 years (95% HPD = 8-23 years), corresponding to 1999. Clade C (pp = 0.98) included seven isolates (four obtained in 2014, two from 2008 and one from 2009) showed a TMRCA of 13 years (95% HPD = 7-19 years), corresponding to 2001. The largest clade D (pp = 1) was the oldest, with a TMRCA of 24 years, corresponding to 1990 (95% HPD = 14-36 years). This clade included 17 strains, 14 of which were sampled in 2014, two in 2007, and one in 2009.
A skyline plot of the Montenegrin strains (Fig. 4) showed that the number of infections remained relatively constant until 1995, after which it increased exponentially, reaching a plateau in 2005.
Bayesian skyline plot representing the estimates of the effective number of HCV-3a infections (y-axis; log 10 scale) over time (x-axis; calendar year), together with the median estimate (solid line) and 95% HPD confidence interval (grey area). A broken line represents the exponential starting growth
Selective pressure analysis
Selective pressure analysis of the 33 HCV NS5B isolates from Montenegro revealed only one statistically supported positively selected site at amino acid position 334 of NS5B (A; T; E; R) numbered according to the reference sequence with accession number YP_001491557 (by using both HYPHY and PAML software). The average ω ratio ranged from 0.1234 to 0.1978 among all models, indicating that a non-synonymous mutation has only 12.34% - 19.78% as much chance of being fixed in the population as a synonymous mutation. The discrete model (M3) fits the data significantly better than the one-ratio model (M0) with the LRT statistic (2 Dl = 102.54; p < 0.05 d.f. = 4). The beta model (M7) was rejected when compared with the beta & ω model (M8) with the LRT statistic (2 Dl = 17.686; p < 0.05 d.f. = 2). The discrete model (M3) suggests a small proportion of sites (about 1%) under positive selection with ω2 = 7.35. Similarly, the M8 model also suggests a small proportion of sites (about 1%) under positive selection with ω = 7.02. Negative and positive selective sites were identified. Specifically, the FEL algorithm identified 38 statistically supported negatively selected sites.
Homology modelling
Overall, the NS5B protein sequences of our dataset are rather conserved, with 99 out of 120 amino acid positions (82.5%) completely conserved with respect to the reference sequence. Among the non-conserved positions of the alignment, eight are highly variable, showing a frequency higher than 0.1 (Table 1). This information was then evaluated based on the three-dimensional structure of the polymerase protein, in order to map the variable residues and to predict the impact of each variation on the protein structure. The model was obtained using a the homology modelling procedure, using as a template the structure identified with PDB ID 3HKW, corresponding to HCV NS5B subtype 1a, which shares 75% sequence identity with HCV 3a NS5B. This structure was chosen for its higher resolution and absence of mutations or deletions. The homology model, shown in Fig. 5, reveals the presence of the three subdomains characteristic of all polymerases: the finger, palm and thumb domains. According to this conventional partitioning of the NS5B domains, the variations are mostly located in the so-called inner palm domain (comprising the regions from residues 188 to 227 and from 287 to 370 of the protein sequence). In particular, residues at positions 219 and 221 are located in proximity to the β6 sheet, while the residues at positions 304, 305 and 307 belong to helix α16, and residues 330 and 334 belong to helix α17. The only exception is represented by position 250, which resides in the α15 helix, belonging to the so-called palm-proximal region of the finger domain.
Homology model of the HCV 3a NS5B protein. The protein is shown by a cartoon representation, while variant positions with respect to reference sequence, discussed in the text, are displayed as spheres coloured by atomic elements and labeled according to the numbering of the NS5B reference sequence. The different colors denote the three subdomains of NS5B, as indicated in the image
All of the amino acid positions that passed the frequency threshold were analyzed in order to calculate differences in Gibbs free energy compared to the reference NS5B protein.
As listed in Table 1, the majority of the variations were predicted to be destabilizing for the NS5B protein structure; however, in some cases, the free energy values of the wild-type and variant structures were very similar. Discarding variations for which there was no agreement between the two methods, two “stabilizing” mutation were detected, namely Tyr to Leu at position 219 and Thr to Tyr at position 221. Furthermore, all variations were analyzed by in silico mutagenesis in which the NS5B model was mutated at the indicated positions. After the selection of the best rotamer for each residue side chain, non-covalent interactions with other surrounding atoms established by the replacing residues were compared to those of the original amino acids.
In Fig. 6, part A, the positions occupied by the reference residues Tyr219 and Thr221 are shown in detail: apparently, the wild-type and the variant residues are not involved in interactions (Fig. 6, part B and part C). However, it should be noted that substitution from Thr to Tyr at position 221 could cause steric effects arising from the larger size of the side chain of the variant residue.
Detail of the variant positions 219 and 221. The protein is represented as a cartoon, while the side chains of the variable amino acids are shown by sticks. In A, reference amino acids Tyr219 and Thr221 are shown, while the Leu219 and Tyr221 substitutes are represented by grey sticks in B, and C. Small green disks are shown when atoms are almost in contact or slightly overlapping, while large red disks indicate significant van der Waals overlap (colour figure online)
Regarding the other substitutions, as shown in Figs. S2 and S3, none of them appear to alter the network of contacts observed in the reference model structure. As an example, at position 250, a substitution from a positively charged amino acid (Fig.S2 part A) to one with similar properties (Fig. S2 part B) (Lys250Arg) allows the maintenance of an H-bond to the side chain of Gln241. Similarly, substitutions at positions 330 (Asp to Glu) (Fig. S2, part C) and 304 (Lys to Arg) (Fig. S3, part A and part B) help to preserve the noncovalent interactions and the net local charge. The variants Ala305Val (Fig. S3, part C), Asn307Gly (Fig. S3, part D) and Ala334Thr (Fig. S2 part D) have side chains that are projected onto the surface of the protein and are not involved in interactions with surrounding atoms. Moreover, no steric effect is observed with the variant amino acids.
Finally, it should be noted that residue conservation analysis carried out using the Consurf server revealed that four of the variant positions are extremely variable among the NS5B protein sequences available in the UniProt database (Fig. S4). The positions 219 and 221 are particularly interesting because they belong to the most conserved Consurf classes. The residue at position 219 is predicted to be exposed to solvent and to play a putative functional role, while the other at 221 is probably buried and is expected to have a structural role.
Discussion
Countries of the Southeastern Mediterranean area have the highest anti-HCV prevalence rates in the world [39]. In the WHO European region, the highest HCV prevalence (from 1.3% to >5.2% in the general population) has been recorded in Romania, Russia (in particular in Central Asian Republics), Turkey, Bulgaria and Italy [39]. The most frequent routes of transmission are injecting drug use and iatrogenic transmission, such as blood transfusion or unsafe medical procedures [40, 41].
Drug use has been associated with increased spread of HCV infection in countries of Eastern Europe [42, 43], where factors such as unemployment, poverty and drug abuse represent a background risk for viral epidemics such as the IDU-HIV outbreaks recently reported in Russia and Ukraine [44, 45].
Epidemiological studies performed in Southeastern Europe have suggested that among the former Yugoslavian Countries, communicable diseases in IDUs are also common. Among these countries, Montenegro, sharing borders with Albania, Kosovo, Croatia, Serbia, and Bosnia and Herzegovina, can be relevant to study [46, 47]. In a first surveillance among IDUs, performed in Podgorica during the year 2005, the reported HCV prevalence was 22%. In a recent second survey, it had increased to 53.7% [48].
Reluctance among IDUs to access local testing centers, due to a lack of trust in their services and expertise as well as a stigma associated with interacting with health care and other institutions and a culture of silence and non-disclosure regarding viral infections among some sectors of the IDU population, have been documented [49].
We report the virus transmission pathway and the evolution of HCV genotype 3a among IDUs in Montenegro.
The Bayesian tree showed that the HCV subtype/genotypes circulating in Montenegro in the IDUs population were 3a, 1a, 4d and 1b. The predominating subtype in our study was 3a. The prevalence of HCV genotypes in our investigation is mostly similar to the genotype distribution in injecting drug users found in other parts of Europe [50–52]. The difference between our results and those of some other European investigators, suggesting a lower prevalence of the subtype 1b in our cohort (only present in three patients), is probably due to the specific characteristics of the IDU population. Moreover, the reported prevalence by other investigators of genotype 2, particularly in advanced liver disease, was more common than we found in our study [53]. However, a comparison of the prevalence of genotypes between our studies and previous reports must take into consideration the time of these investigations and the selected population of IDUs.
In the past decade, a shift in genotype distribution has been observed in many countries, mostly consisting of an increase in the prevalence of the genotypes 3a, 1a and 4 and a decrease in the prevalence of genotypes 1b and 2 [51, 52]. One of the reasons for these changes is still related to intravenous drug abuse, which has now become the major risk factor for HCV infection and is associated with subtypes 3a and 1a [50–52].
Evaluating the association of the genotypes and demographic data of the patients, we did not find differences in genotype distribution, age or gender of the patients. Our results showed that the subtype 3a is the most important predictive factor for IDUs, also in Montenegro.
Phylogenetic analysis confirmed four different epidemic waves in Montenegro grouped together in a monophyletic lineage. Only three sequenced intermixed with sequences from other countries in clades that were not supported statistically.
Bayesian analysis showed that the HCV-3a subtype probably reached Montenegro in the 1990s, causing an epidemic that grew exponentially in a very short time period, between 1995 and 2005. According to a recent global epidemiological investigation of hepatitis viruses among IDUs, the largest populations of HCV-positive IDUs in 2011 were living in Eastern Europe and Asia, thus explaining the continuing spread of HCV-3a in these regions.
Based on the dated tree, HCV-3a showed four different entries dating from 1990 (clade D), 1994 (clade A), 1999 (clade B) and 2001 (clade C). This temporal reconstruction is in agreement with what has been reported by other authors and our previous data reporting that Indian HCV subtype-3a sequences rooted closely with United Kingdom sequences, thus suggesting a movement of the virus from the Indian subcontinent to United Kingdom and later to other European countries, probably also involving Southeastern Europe [8, 54].
The HCV genome shows remarkable sequence variation, as demonstrated by the large number of identified genotypes [55]. However, conservation of certain amino acids or of their physiochemical properties among different HCV genotypes, strains or quasispecies variants, indicates that they are needed to maintain a vital protein function, or that they confer a significant survival advantage. Residues within the hydrophobic protein core that are required for protein stability and folding are mostly conserved. Amino acids located at the protein surface are more variable and have the potential to mediate interactions with the host. In this work, structural analysis of an NS5B model protein has shown that the variant amino acids are located mainly in the so-called palm domain, which contains most of the conserved structural elements of the active site that are common to all polymerases. This subdomain is important for the formation of the catalytic cavity and provides conformational changes of the polymerase molecules at different stages of replication [56].
Of the eight variant positions, positions 219 and 221 are predicted to be important for the maintenance of a proper protein fold. Interestingly, as shown in the protein sequence alignment in Fig. S5, all but three sequences have the variant amino acid at both positions. The same is true at position 307, where the presence of Gly (and, in one sequence, Ser) instead of the reference residue Gln can be observed.
The HCV NS5B contains all of the sequence motifs, designated as motifs A-F, that are highly conserved in all known RNA-dependent RNA polymerases [57]. Variant positions 219 and 221 are included in the A motif (from residue 213 to 228), which includes the catalytic pocket of the enzyme (from Asp220 to Asp225). The A motif is well described in the literature [58, 59], and the residues Asp220 and Thr221 (variant position in this work) are strictly conserved in the seven HCV genotypes. HCV NS5B requires divalent metal cations, Mg2+ or Mn2+, for the ligation of ribonucleoside triphosphate. The essential residues for metal binding are Asp220 and the carbonyl group of the Thr221 peptide backbone, which coordinate metal ions in the structure of ternary complexes with nucleic acid polymers and nucleotide substrates [60]. Moreover, two other variant positions (330 and 334) are located in a conserved motif termed D (residues from 326 to 347), forming the core structure of the palm subdomain. Motif D is highly conserved at the amino acid level within the HCV-1 genotype, and to a lesser extent within the other genotypes. Indeed, in recent work [61] investigating the natural polymorphism among different HCV genotypes, it was found that some variant positions characteristic of our dataset are, instead, reference positions in other HCV genotypes. For example, at position 304 we observed an Arg in place of a Lys that is the reference amino acid for HCV subtype 1b. Similarly, for position 307, our variant Gly is present in HCV genotypes 1a, 2 and 7, while the variant Glu found at position 330 is found among sequences from HCV genotypes 2, 4 and 7. Although in silico mutagenesis and structural inspection do not allow us to predict with reasonable certainty the biological effect of these mutations, these variations are in regions of NS5B that are already known to be involved in protein function. Therefore, all of the presented results can represent a framework for future site-directed mutagenesis experiments aimed at investigating the effect of point mutations on NS5B function. This study provides an estimate of the evolutionary history of HCV genotype 3a, the most prevalent genotype in Montenegro among IDUs. These data could represent the basis for further strategies aimed to improve disease management and development of surveillance programs in high-risk populations.
References
World Health Organization (2009) The growing threats of hepatitis B and C in the Eastern Mediterranean region: a call for action. Technical paper EM/RC56/3
Bernini F, Ebranati E, De Maddalena C, Shkjezi R, Milazzo L, Lo Presti A, Ciccozzi M, Galli M, Zehender G (2011) Within-host dynamics of the hepatitis C virus quasispecies population in HIV-1/HCV coinfected patients. PLoS One 6:e16551
Simmonds P, Bukh J, Combet C, Deléage G, Enomoto N, Feinstone S, Halfon P, Inchauspé G, Kuiken C, Maertens G, Mizokami M, Murphy DG, Okamoto H, Pawlotsky JM, Penin F, Sablon E, Shin-I T, Stuyver LJ, Thiel HJ, Viazov S, Weiner AJ, Widell A (2005) Consensus proposals for a unified system of nomenclature of hepatitis C virus genotypes. Hepatology 42:962–973
Pybus OG, Charleston MA, Gupta S, Rambaut A, Holmes EC, Harvey PH (2001) The epidemic behavior of the hepatitis C virus. Science 292:2323–2325
Hauri AM, Armstrong GL, Hutin YJ (2004) The global burden of disease attributable to contaminated injections given in health care settings. Int J STD AIDS 15:7–16
Pybus OG, Markov PV, Wu A, Tatem AJ (2007) Investigating the endemic transmission of the hepatitis C virus. Int J Parasitol 37:839–849
Ciccozzi M, Zehender G, Cento V, Lo Presti A, Teoharov P, Pavlov I, Bogdanova V, Perno CF, Ciotti M (2011) Molecular analysis of hepatitis C virus infection in Bulgarian injecting drug users. J Med Virol 83:1565–1570
Zehender G, Sorrentino C, Lai A, Ebranati E, Gabanelli E, Lo Presti A, Vujoševic D, Lauševic D, Terzić D, Shkjezi R, Bino S, Vratnica Z, Mugosa B, Galli M, Ciccozzi M (2013) Reconstruction of the evolutionary dynamics of hepatitis C virus subtypes in Montenegro and the Balkan region. Infect Genet Evol 17:223–230
Judd A, Rhodes T, Johnston LG, Platt L, Andjelkovic V, Simić D, Mugosa B, Simić M, Zerjav S, Parry RP, Parry JV (2009) Improving survey methods in sero-epidemiological studies of injecting drug users: a case example of two cross sectional surveys in Serbia and Montenegro. BMC Infect Dis 9:14
Stamenkovic G, Zerjav S, Velickovic ZM, Krtolica K, Samardzija VL, Jemuovic L, Nozic D, Dimitrijevic B (2000) Distribution of HCV genotypes among risk groups in Serbia. Eur J Epidemiol 16:949–954
Svirtlih N, Delic D, Simonovic J, Jevtovic D, Dokic L, Gvozdenovic E, Boricic I, Terzic D, Pavic S, Neskovic G, Zerjav S, Urban V (2007) Hepatitis C virus genotypes in Serbia and Montenegro: the prevalence and clinical significance. World J Gastroenterol 13:355–360
Kalinina O, Norder H, Vetrov T, Zhdanov K, Barzunova M, Plotnikova V, Mukomolov S, Magnius LO (2001) Shift in predominating subtype of HCV from 1b to 3a in St. Petersburg mediated by increase in injecting drug use. J Med Virol 65:517–524
Krekulova L, Rehak V, Madrigal N, Johnson M, Killoran P, Riley LW (2001) Genotypic and epidemiologic characteristics of hepatitis C virus infections among recent injection drug user and nonuser populations. Clin Infect Dis 33:1435–1438
Lu L, Nakano T, He Y, Fu Y, Hagedorn CH, Robertson BH (2005) Hepatitis C virus genotype distribution in China: predominance of closely related subtype 1b isolates and existence of new genotype 6 variants. J Med Virol 75:538–549
Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG (1997) The CLUSTAL-X windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res 2:4876–4882
Thompson JD, Higgins DG, Gibson TJ (1994) CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res 22:4673–4680
Posada D, Buckley TR (2004) Model selection and model averaging in phylogenetics: advantages of Akaike information criterion and Bayesian approaches over likelihood ratio tests. Syst Biol 53:793–808
Lanave C, Preparata G, Saccone C, Serio G (1984) A new method for calculating evolutionary substitution rates. J Mol Evol 20:86–93
Huelsenbeck JP, Ronquist F (2001) MRBAYES: Bayesian inference of phylogenetic trees. Bioinformatics 17:754–755
Schmidt HA, Strimmer K, Vingron M, von Haeseler A (2002) TREE-PUZZLE: maximum likelihood phylogenetic analysis using quartets and parallel computing. Bioinformatics 18:502–504
Drummond AJ, Rambaut A (2007) BEAST: Bayesian evolutionary analysis by sampling trees. BMC Evol Biol 7:214
Drummond AJ, Rambaut A, Shapiro B, Pybus OG (2005) Bayesian coalescent inference of past population dynamics from molecular sequences. Mol Biol Evol 22:1185–1192
Kass RE, Raftery AE (1995) Bayes factors. J Am Stat Assoc 90:773–795
Yang Z (1997) PAML: a program package for phylogenetic analysis by maximum likelihood. Comput Appl Biosci 13:555–556
Yang Z, Nielsen R (2000) Estimating synonymous and nonsynonymous substitution rates under realistic evolutionary models. Mol Biol Evol 17:32–43
Nielsen R, Yang Z (1998) Likelihood models for detecting positively selected amino acid sites and applications to the HIV-1 envelope gene. Genetics 148:929–936
Pond SL, Frost SD, Muse SV (2005) HyPhy: hypothesis testing using phylogenies. Bioinformatics 21:676–679
Kosakovsky Pond SL, Frost SD (2005) Not so different after all: a comparison of methods for detecting amino acid sites under selection. Mol Biol Evol 22:1208–1222
Okonechnikov K, Golosova O, Fursov M, UGENE team, (2012) Unipro UGENE: a unified bioinformatics toolkit. Bioinformatics 28:1166–1167
Sievers F, Wilm A, Dineen D, Gibson TJ, Karplus K, Li W, Lopez R, McWilliam H, Remmert M, Soding J, Thompson JD, Higgins DG (2011) Fast, scalable generation of high-quality protein multiple sequence alignments using Clustal Omega. Mol Syst Biol 7:539
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ (1990) Basic local alignment search tool. J Mol Biol 215:403–410
Sali A, Blundell TL (1993) Comparative protein modelling by satisfaction of spatial restraints. J Mol Biol 234:779–815
Wiederstein M, Sippl MJ (2007) ProSA-web: interactive web service for the recognition of errors in three-dimensional structures of proteins. Nucleic Acids Res 35:W407–W410
Laskowski RA, Rullmannn JA, MacArthur MW, Kaptein R, Thornton JM (1996) AQUA and PROCHECK-NMR: programs for checking the quality of protein structures solved by NMR. J Biomol NMR 8:477–486
Ashkenazy H, Erez E, Martz E, Pupko T, Ben-Tal N (2010) ConSurf 2010: calculating evolutionary conservation in sequence and structure of proteins and nucleic acids. Nucleic Acids Res 38:W529–W533
Schroedinger L (2015) In: The PyMOL molecular graphics system, Version 1.7.4, LLC. Schrödinger
Capriotti E, Fariselli P, Casadio R (2005) I-Mutant2.0: predicting stability changes upon mutation from the protein sequence or structure. Nucleic Acids Res 33:W306–W310
Parthiban V, Gromiha MM, Schomburg D (2006) CUPSAT: prediction of protein stability upon point mutations. Nucleic Acids Res 34:W239–W242
World Health Organization (2010) Sixty-third World health assembly. Viral hepatitis, report by the Secretariat A63/15
Alter MJ (2002) Prevention of spread of hepatitis C. Hepatology 36:S93–S98
Lauer GM, Walker BD (2001) Hepatitis C virus infection. N Engl J Med 345:41–52
Hamers FF, Downs AM (2003) HIV in central and eastern Europe. Lancet 361:1035–1044
Kelly JA, Amirkhanian YA (2003) The newest epidemic: a review of HIV/AIDS in central and eastern Europe. Int J STD AIDS 14:361–371
Cohen J (2010) Late for the epidemic: HIV/AIDS in eastern Europe. Science 329:160–164
Kruglov YV, Kobyshcha YV, Salyuk T, Varetska O, Shakarishvili A, Saldanha VP (2008) The most severe HIV epidemic in Europe: Ukraine’s national HIV prevalence estimates for 2007. Sex Transm Infect 84:37–41
Shapatava E, Nelson KE, Tsertsvadze T, del Rio C (2006) Risk behaviors and HIV, hepatitis B, and hepatitis C seroprevalence among injection drug users in Georgia. Drug Alcohol Depend 82:35–38
Djurić D (2003) The economic development of Montenegro. In: Bieber F (ed) Montenegro in transition: problems of identity and statehood. Nomos Verlagsgesellschaft, Baden-Baden, pp 139–158
Baćak V, Laušević D, Mugoša B, Vratnica Z, Terzić N (2013) Hepatitis C virus infection and related riskfactors among injection drug users in Montenegro. Eur Addict Res 19:68–73
Bernays S (2008) Trust, disruption and responsibility in accounts of injecting equipment sharing and hepatitis C risk. Health Risk Soc 10:221–240
Paintsil E, Verevochkin SV, Dukhovlinova E, Niccolai L, Barbour R, White E, Toussova OV, Alexander L, Kozlov AP, Heimer R (2009) Hepatitis C virus infection among drug injectors in St Petersburg, Russia: social and molecular epidemiology of an endemic infection. Addiction 104:1881–1890
van Asten L, Verhaest I, Lamzira S, Hernandez-Aguado I, Zangerle R, Boufassa F, Rezza G, Broers B, Robertson JR, Brettle RP, McMenamin J, Prins M, Cochrane A, Simmonds P, Coutinho RA, Bruisten S (2004) European and Italian Seroconverter Studies. Spread of hepatitis C virus among European injection drug users infected with HIV: a phylogenetic analysis. J Infect Dis 189:292–302
Tallo T, Norder H, Tefanova V, Krispin T, Schmidt J, Ilmoja M, Orgulas K, Pruunsild K, Priimägi L, Magnius LO (2007) Genetic characterization of hepatitis C virus strains in Estonia: fluctuations in the predominating subtype with time. J Med Virol 79:374–382
Markov PV, van de Laar TJ, Thomas XV, Aronson SJ, Weegink CJ, van den Berk GE, Prins M, Pybus OG, Schinkel J (2012) Colonial history and contemporary transmission shape the genetic diversity of hepatitis C virus genotype 2 in Amsterdam. J Virol 86:7677–7687
Harris KA, Gilham C, Mortimer PP, Teo CG (1999) The most prevalent hepatitis C virus genotypes in England and Wales are 3a and 1a. J Med Virol 58:127–131
Ranjith-Kumar CT, Kao CC (2006) Biochemical activities of the HCV NS5B RNA-dependent RNA polymerase. In: Tan SL (ed) Hepatitis C viruses: genomes and molecular biology, Chapter 10. Horizon Bioscience, Norfolk
Lesburg CA, Cable MB, Ferrari E, Hong Z, Mannarino AF, Weber PC (1999) Crystal structure of the RNA-dependent RNA polymerase from hepatitis C virus reveals a fully encircled active site. Nat Struct Biol 6:937–943
Poch O, Sauvaget I, Delarue M, Tordo N (1989) Identification of four conserved motifs among the RNA-dependent polymerase encoding elements. EMBO J 8:3867–3874
Lohmann V, Korner F, Herian U, Bartenschlager R (1997) Biochemical properties of hepatitis C virus NS5B RNA-dependent RNA polymerase and identification of amino acid sequence motifs essential for enzymatic activity. J Virol 71:8416–8428
Cheney IW, Naim S, Lai VC, Dempsey S, Bellows D, Walker MP, Shim JH, Horscroft N, Hong Z, Zhong W (2002) Mutations in NS5B polymerase of hepatitis C virus: impacts on in vitro enzymatic activity and viral RNA replication in the subgenomic replicon cell culture. Virology 297:298–306
Behrens SE, Tomei L, De Francesco R (1996) Identification and properties of the RNA-dependent RNA polymerase of hepatitis C virus. EMBO J 15:12–22
Di Maio VC, Cento V, Mirabelli C, Artese A, Costa G, Alcaro S, Perno CF, Ceccherini-Silberstein F (2014) Hepatitis C virus genetic variability and the presence of NS5B resistance-associated mutations as natural polymorphisms in selected genotypes could affect the response to NS5B inhibitors. Antimicrob Agents Chemother 58:2781–2797
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
All authors declare that they have no conflict of interest.
Ethical approval
All procedures performed in studies involving human participants were in accordance with the ethical standards of the institutional and/or national research committee and with the 1964 Helsinki declaration and its later amendments or comparable ethical standard.
Informed consent
Informed consent was obtained from all individual participants included in the study.
Additional information
B. Mugosa, E. Cella and A. Lai contributed equally.
Electronic supplementary material
Below is the link to the electronic supplementary material.
705_2017_3224_MOESM1_ESM.tif
Supplementary material 1 (TIFF 59 kb) Fig. S1 Likelihood mapping of the HCV-3a dataset of nucleotide sequences. The three corners represent fully resolved tree topologies, i.e., the presence of a tree-like phylogenetic signal, in the given dataset
705_2017_3224_MOESM2_ESM.tif
Supplementary material 2 (TIFF 367 kb) Fig. S2 Detail of the variant positions 250, 330 and 334. The protein is represented as a salmon- colored cartoon, while the side chain of the variable amino acids are shown by sticks. In A, reference amino acid Arg250 is shown, while the replacing Lys is represented in B. H-bonds between side chains of the residues in positions 250 and Gln241 are depicted by yellow dashed lines, and distances are shown. In C and D reference residues are shown with salmon-colored stick models, while the variant residues are represented by grey sticks
705_2017_3224_MOESM3_ESM.tif
Supplementary material 3 (TIFF 444 kb) Fig. S3 Detail of the variant positions 304, 305 and 307. The protein is represented as a cartoon, while the side chains of the variable amino acids are shown by sticks. In A, reference amino acids Lys304, Ala305, Asn307 are shown, while the variant amino acids are represented by grey sticks in B, C and D. In B, the interactions between residue side chains in position 304 and the residues Thr66 and Glu70 are indicated by yellow dashed lines, labeled with atomic distances
705_2017_3224_MOESM4_ESM.tif
Supplementary material 4 (TIFF 240 kb) Fig. S4 Residue conservation analysis. The residue conservation was analyzed using Consurf through the comparison of 250 NS5B protein sequences of different HCV genotypes obtained from the UniProt database. The reference sequence of the HCV-3a NS5B is displayed with the residue conservation score at each site color-coded into it. The conservation scale was defined from the most variable amino acid position (grade 1, colored turquoise) to the most conserves amino acid position (grade 9, colored maroon). Positions for which the inferred conservation level was assigned with low confidence are marked with light yellow. The first row below the sequence lists the predicted burial status of the site (i.e., ‘‘b’’– buried versus ‘‘e’’ – exposed). The second row indicates residues predicted to be structurally and functionally important: ‘‘s’’ and ‘‘f’’, respectively. Positions 219 and 221, marked by black arrows, are discussed in the text
705_2017_3224_MOESM5_ESM.tif
Supplementary material 5 (TIFF 678 kb) Fig. S5 Sequence alignment of HCV 3a NS5B protein sequences. Multiple sequence alignment of the NS5B sequences. Alignment was obtained with the program Clustal Omega and displayed with Jalview. Sequences are reported with the amino acid one-letter code; the last sequence is the HCV 3a NS5B reference sequence labeled with its RefSeq code. The black arrows indicate the 219, 221 and 307 positions (numbers are referred to the reference sequence), which have an high mutation frequency with respect to reference sequence. Residues are colored according to the ClustalX scheme, which for each alignment column, takes into account conservation and amino acid type
Rights and permissions
About this article
Cite this article
Mugosa, B., Cella, E., Lai, A. et al. Hepatitis C virus genotype 3A in a population of injecting drug users in Montenegro: Bayesian and evolutionary analysis. Arch Virol 162, 1549–1561 (2017). https://doi.org/10.1007/s00705-017-3224-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00705-017-3224-5