Abstract
The association of Merkel cell polyomavirus (MCPyV) with Merkel cell carcinoma (MCC) in immunocompromised individuals has been revealed in a number of surveys. The study of MCPyV specific antibody titers and viral loads in such patients has a great attraction for research groups interested in viral reactivation. In this cross-sectional study to evaluate MCPyV antibody titer, DNA prevalence and viral load in peripheral blood mononuclear cells (PBMCs), we examined 205 HIV-1 infected patients and 100 un-infected controls. The HIV-1 infected patients divided into two groups (HIV/AIDS and non-AIDS) according to their CD4 status. Total IgG antibody titer against MCPyV was analyzed by virus like particle (VLP)-based enzyme linked immunosorbent assay (ELISA). Presence of MCPyV-DNA in subject’s PBMCs was examined by quantitative real-time PCR assay. Levels of anti-MCPyV IgG in HIV/AIDS patients were significantly higher than those in non-AIDS HIV-infected and control subjects (p value = <0.001). The prevalence rate of MCPyV-DNA in PBMCs of HIV/AIDS, non-AIDS HIV-infected and un-infected controls were 17%, 16%, and 14% respectively. The MCPyV viral load among the groups ranged between 0.15 to 2.9 copies/103cells (median, 1.9 copies/103cells), with no significant difference between the studied populations (p value = 0.3).
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The association of viral infections with the frequent malignancies in AIDS patients has been proven. Human herpes virus 8 (KSHV/HHV8)-associated Kaposi’s sarcoma (KSHV/HHV-8) or Epstein-Barr virus-associated non-Hodgkin lymphoma are examples demonstrated many years ago [1, 2]. HIV/AIDS patients also showed a 13.4-fold higher risk of Merkel cell carcinoma (MCC), an aggressive neuroendocrine cutaneous tumor, when compared with the general population [3, 4]. MCC is a rare skin cancer, but its incidence has been linked to aging, sun exposure, the use of immunosuppressive medication, and indeed AIDS [3, 5]. Surprisingly, the incidence of this rare skin cancer has tripled over the past 20 years [6].
In 2008, Feng et al. revealed a link between MCC and a novel human polyomavirus which had been identified in the majority of MCC tumor cases [7–9]. This newly discovered virus was named Merkel cell polyomavirus (MCPyV). They also showed that in more than 80% of MCC cases, the MCPyV DNA clonally-integrated into the genome of tumor cells [8]. Subsequent studies indicated persistent expression of the large tumor-associated antigen (LT-Ag) of MCPyV in a truncated form; this viral oncoprotein likely promotes cell division in the host by interacting with tumor suppressors [10–12].
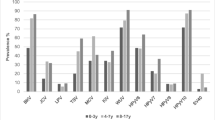
Serological studies indicate that the majority of adults are MCPyV IgG-positive and the seroprevalence in general population is around 45-85%, which increases with age [13], suggesting that exposure to MCPyV is very common. Our previous age-specific seroprevalence study in healthy individuals also revealed an increased level of MCPyV IgG antibody, correlating with age.
Prior studies showed that, while the majority of healthy adults are serologically positive to MCPyV, the magnitude of humoral immune responses to the viral major capsid (VP1) varies dramatically across a wide range [14]. Regarding these findings, patients with MCC displayed a very high-titer of specific MCPyV antibody when compared with control subjects [15–20]. So far, several research groups, including ours, have applied virus like particle based enzyme-linked immunosorbent assay (VLP-based ELISA) to identify/quantify specific antibody against MCPyV in different population groups [18–20].
Up to now very few studies have been published examining molecular or seroprevalence of MCPyV in HIV-infected patients. In one report, MCPyV DNA was detected in 31% of anal, 35% of penile, and 36% of oral swabs of HIV-infected individuals although no correlation between oral or anogenital dysplasia and the presence of MCPyV DNA was seen [21]. Since it is possible that the elevation of MCPyV-DNA levels might raise the chance of its integration into the host cell genome, the prevalence or load of this viral genome in different samples drew the attention of more researches. In 2011, Wieland et al. showed that MCPyV-genome detected in skin swabs of HIV-positive men had a higher load than in HIV-uninfected controls [22]. Although, there are several reports of a lymphatic tropism for MCPyV, the prevalence and load of MCPyV-genomes in PBMC or lymphoid tissues, and a pathogenic outcome for this infection, in HIV/AIDS patients still remains controversial. In the current study, we applied VLP-based enzyme-linked immunosorbent assay (ELISA) in order to measure MCPyV IgG titers among 3 groups which included HIV-infected patients with CD4 cell counts higher than 350, HIV-infected patients with CD4 cell counts less than 200, and un-infected control individuals. We also examined the presence of MCPyV DNA in peripheral blood mononuclear cells (PBMC) in these study groups using real-time PCR assay. The prevalence of HIV infection in Iranian populations is relatively low but recent studies have shown that about a quarter of these identified individuals have progressed to develop AIDS [23, 24].
Materials and methods
Human subjects and clinical examination
In this study, 205 HIV-infected patients (115 males, 90 females) were included, from November 2013 to May 2015. According to classification system for HIV/AIDS provided by the US Centers for Disease Control and Prevention (CDC), the severity of disease in these patients is characterized by CD4 cell counts and other AIDS related conditions. All included patients were referred to the three Tehran general hospitals during their follow-up periods and their CD4 cell counts were determined at the time of blood sampling. In our study, progression of a HIV-infected patient to AIDS phase was determined by a CD4 cell count less than 200 cells/µl (29). Regarding the aforementioned categories our 3 study groups were; Group 1: 105 HIV-infected patients with CD4 cell counts exceeding 350 cells/µl, named as the non-AIDS HIV-infected group; Group 2: 100 HIV-infected patients with CD4 cell counts below 200 cells/µl, named as AIDS stage patients; and Group3: 100 HIV-negative healthy individuals used as control patients. The age-range and gender of the control group were chosen to match with the HIV-infected patients.
The study protocol was approved by the ethical committee of the Iran University of Medical Sciences (Approval No. 21091). All patients and controls agreed to participate in our study by giving their written informed consent. Blood samples were collected in tubes containing anti-coagulant agent. Plasma samples were separated and stored at -20 °C until used.
Production and purification of MCPyV-like particles
Generation of MCPyV-like particles was performed by expression of the major capsid protein (VP1) of the virus in a Spodoptera Frugiperda insect cell line (sf9 cells) using the Bac-to-BacR Baculovirus Expression System (Life technology), as previously described (20). Briefly, the MCPyV VP1 coding sequence that had been commercially synthesized (GenScript, USA), was sub-cloned into a pFastBacTM HT A plasmid. After transformation of these plasmids into DH10 BacTM Escherichia coli cells, the recombinant bacmid was selected according to the manufacturer’s instructions. Then, the purified recombinant bacmid vector was transfected into sf9 cells using CellfectinR II Reagent (Life technology) according to the manufacturer’s instructions. Recombinant baculovirus was harvested six days’ post-transfection and stored at 4 °C. To produce MCPyV-like particles, sf9 cells were infected with a recombinant baculovirus. 3 days after infection we harvested cells and the VLPs were released by 3 freeze-thaw cycles. VLPs were purified by using the protocol previously described by Chen et al. [18] using cesium chloride (CsCl) density-gradient centrifugation in a swing SW41Ti rotor (Beckman) at 24,000 rpm. The expression of the VP1 protein was confirmed using western blot analysis. Self-assembly of the MCPyV-like particles was showed by transmission electron microscopy (TEM), according to the staining protocol described before [20].
Anti-MCPyV antibody analysis using VLP-based ELISA
Analysis of anti-VP1 antibody levels by ELISA was determined by using MCPyV strain 339 VLPs generated in Spodoptera Frugiperda insect cells as previously described [20]. For the quantitation of anti- MCPyV IgG, we used a modified protocol that had been described by Viscidi et al. [13]. Briefly, the 2-fold dilutions of sera and triplicate dilutions of a reference serum were incubated on VLP-coated plates. Following serial dilution of a reference serum, we applied a four parameter equation using SoftMax Pro7 software (Sunnyvale, CA 94089 USA) to draw a standard curve. Once we reached the highest dilution point of the reference sample (an OD value two times higher than the mean OD of the PBS controls) this was set as 1 Enzyme Immuno Assay (EIA) unit. The standard curve was constructed with the operational range from 1 to 2048 units. Accordingly, the EIA units of every test subject at each dilution were obtained by interpolation into the given standard curve, before multiplication by the dilution factor. VLP-competition assays were done to identify the possible occurrence of cross-reactivity among other human polyomaviruses, as described previously by Tolstov et al [17]. We used JCPyV VLP (kindly gifted from Keyvan Lab) as a competitor in this assay.
Detection of MCPyV DNA in PBMC
Isolation of PBMCs from blood samples was performed by Ficol-Hypaque gradient centrifugation (Lymphocyte®-H, CEDARLANE®) and according to the manufacturer’s instructions, PBMCs were resuspended in sterile phosphate buffer saline (PBS) and stored at -20 °C until use. In the next step, DNA was extracted from 1–2 × 106 PBMCs of each sample, using the QIAamp DNA Mini Kit (Qiagen, Hilden, Germany) according to the manufacturer’s instructions. Detection of MCPyV-DNA was carried out by using the real-time PCR assay (Rotor-Gene 6000) and specific MCPyV large T antigen primer sets along with a TaqMan probe. In addition, a primers/probe specific human RNase P gene was used as an internal control (26). Standard curves were generated by applying the 10-fold diluted series of the purified plasmid containing MCPyV TAg gene (ranging from 2 × 101 to 2 × 106 copies/ml) to the aforementioned real-time PCR experiment, as previously described. Viral gene copy numbers per cell were calculated by dividing the virus copy number into half of the RNase P copy number, since each diploid cell contains two copies of RNase P [25].
Statistical analysis
All statistical analysis was carried out with the IBM SPSS v20.0 software. A comparison between anti-MCPyV IgG levels in uninfected, non-AIDS HIV-infected, and HIV/AIDS populations was performed using one-way ANOVA analysis with Tukey’s post hoc test. 1000 bias-corrected and accelerated bootstrap samples were used since log transformation of the data did not yield a normal distribution and a significance criterion less than 0.05 has been adopted. To adjust the effect of gender, multinomial logistic regression was used with status and gender and the level of serum antibody considered as dependent variables. The copy number data of MCPyV genome in different groups were statistically analyzed using Mann-Whitney test.
χ 2 test was used for proportional analysis between study groups and association between the anti-MCPyV antibody titers/ quantity of the MCPyV genome and HIV/AIDS stages or other characteristics of patients. Kruskal-Wallis test was used to compare geometric mean titers (GMT) of different groups.
Results
Generation of MCPyV VLP
The major capsid protein coding sequence of MCPyV (VP1) strain 375804) with EcoRI/BamHI restriction sites at each end was artificially generated. The VP1 coding sequence was successfully sub-cloned into a transfer vector (pFastBac HTA) and transformed into DH10 BacTM Escherichia coli cells. The recombinant bacmid containing the MCPyV VP1 gene downstream of the polyhedrin promoter was extracted and the accuracy of the insertion was verified by colony-PCR using bacmid specific primers (data not shown). Recombinant baculovirus was generated by transfecting insect cells with the recombinant bacmid and lysates from the virus-infected sf9 cells were subjected to a purification protocol using cesium chloride density gradient centrifugation.
Through western blot analysis with Anti-6X His tag® antibody (HRP) (Abcam, US), a protein band of 46 KDa was observed on nitrocellulose paper (Fig.1A). In the electron microscope, purified VLPs were revealed with a spherical shape and a diameter of 20- 45 nm (Fig.1B).
Detection of expressed MCPyV VP1 and formation of VLP: A) Western blot analysis were performed to identify VP1 protein expression using a specific monoclonal antibody anti-HN and secondary antibody corresponding to horseradish peroxidase (HRP)-labeled anti-mouse IgG; Lane1: first control sample containing un-infected sf9 cell lysate; Lane 2: second control sample containing cell lysate that infected by baculovirus without insert (mock); Lane 3: lysate supernatant of sf9 cells which have been infected with recombinant baculovirus expressing MCPyV VP1 B) Electron micrograph of MCPyV-like particles at ×50,000 nominal magnification. Scale bar = 100 nm
MCPyV DNA detection
The prevalence of MCPyV DNA in PBMC samples of the HIV/AIDS patients was 17% (17/100) which was not significantly different compared to those of HIV-infected patients (15%, 16/105) and un-infected individuals (14%, 14/100) (P-value = 0.06, OD 3.9, 95% CI 3.332–10.30) (Table 1).
MCPyV viral loads in HIV/AIDS patients was slightly higher (mean load of 1.45 copies/103 cells, a range of 0.15–2.9 copies/103cells) than in HIV-infected patients (mean load of 0.95 copies/103 cells, range 0.3–2.7 copies/103 cells) and in uninfected individuals (mean load of 1.01 copies/103 cells, range 0.2–2.5 copies/103 cells); however, the difference was not statistically significant (P value = 0.3) (data not shown).
Anti-MCPyV total IgG measurement
The anti- MCPyV antibody titers was significantly higher in HIV/AIDS patients than those in HIV-infected or uninfected individuals (p value < 0.001). Yet no significant difference was seen between MCPyV IgG titers in HIV-infected and un-infected (p value = 0.34) (Fig 2). The logistic regression test results revealed that the anti- MCPyV antibody titers in these study groups was not associated with gender (p value = 0.7). To determine if this VLP-based EIA system recognizes specific antibodies against MCPyV, VLP-competition assays using MCPyV and JCPyV VLPs were carried out on 5 separate high reactive sera. It was observed that pre-incubation of these reactive sera with 100 ng of purified MCPyV VLP are adequate to block EIA reactivity. However, pre-incubation of the reactive sera with up to 500 ng of purified JCPyV VLP failed to compete with reactivity of MCPyV specific antibodies in this developed EIA assay.
Discussion
Since the discovery of MCPyV in 2008, several studies have examined an association of the viral infection with different malignancies and its role in immuno-compromised hosts [8]. Here, we demonstrated that higher MCPyV IgG levels among HIV-infected individuals associated with immuno-competence or progression to AIDS, while the presence of MCPyV DNA in PBMCs in the studied groups did not significantly correlate. Our results showed that MCPyV IgG titer in HIV-infected patients with CD4 cell count lower than 350, is significantly higher than those in HIV-infected patients with normal CD4 count as well as when compared to uninfected individuals as well (p value < 0.001).
To our knowledge seroprevalence studies examining MCPyV in these populations have not yet been published. In our previous study we detected MCPyV specific antibody in 57% of Tehran’s general population and the IgG antibody levels tended to increase along with age but we found no significant difference between males and females [20].
In order to determine MCPyV lymphotropism as has been seen previously in JC and BK polyomavirus infections, PBMC samples from all subjects were collected and tested by quantitative real-time PCR assay. The MCPyV DNA positivity in PBMCs among our non-MCC HIV/AIDS patients (17%) was slightly higher than the prevalence rate reported by Shuda et al. in a similar population [12]. In 2010, Leude et al. detected MCPyV genome in PBMC samples from about 50% of MCC patients and they revealed a link between the MCC’s stages or evolution and the frequency of MCPyV DNA in patients’ PBMCs [26]. They also suggested that the presence of MCPyV DNA in PBMCs may correspond to viral replication followed by reactivated infection in lymphoid tissue, as observed with other polyomaviruses [27, 28]. Notably, our data showed no significant difference in prevalence or load of MCPyV DNA in the PBMC samples of our study groups. Therefore, we assumed that the possible reactivation of MCPyV caused an increase of anti- MCPyV IgG titers in HIV/AIDS group that may correlate with the active replication of virus in a site other than lymphoid tissues. It’s notable that a very recent study has suggested dermal fibroblasts near hair follicles as the target of productive MCPyV infection [29], accordingly it is possible to assume a role for such cells as a reactivation site in the context of immunodeficiency.
Several potential limitations of this study should also be considered, including the absence of serological data for other AIDS-related viruses (e.g. CMV, EBV, and HHV8) and the cross-sectional design of sampling. As expected, all groups (HIV-infected and HIV-uninfected and control individuals) had detectable MCPyV IgG titers, so the comparison between MCPyV-exposed and un-exposed groups was not possible.
In the current study we observed an association between increased MCPyV-IgG titer and the level of immunocompetency in HIV/AIDS patients that probably suggests a reactivation tumorigenesis in these patients. Prior studies clearly revealed the fact that the geometric mean titer of specific IgG antibody against MCPyV in a MCC patient was around 10 to 14-fold higher than controls [14]. Although it may seem irrational that this strong humoral response in MCC patients could not abrogate MCPyV infection, it has been reported that these IgG antibodies are neutralizing [16]. Nevertheless, the explanation for this paradoxical situation can be found through deductions based on studies of another human polyomaviruses. For instance, in BK virus it has been observed that viral replication persists for a long time despite an efficient immune response, and that the level of BK IgG antibody reported can be even higher in individuals with viral replication than in those with no active replication [30, 31]. Additionally it has been shown that when immunosuppression with any cause occurred, BK polyomavirus reactivates from a latent state, following by a simultaneous increase in IgG antibody levels against the viral capsid [30, 31]. This phenomenon also explains the increased level of MCPyV IgG titer in HIV/AIDS patients and suggests a reactivation of MCPyV infection in the case of virally induced-immunosuppression.
References
Li M, Saghafi N, Freymiller E, Basile JR, Lin YL (2013) Metastatic Merkel cell carcinoma of the oral cavity in a human immunodeficiency virus-positive patient and the detection of Merkel cell polyomavirus. Oral Surg Oral Med Oral Pathol Oral Radiol 115(5):e66–e71. doi:10.1016/j.oooo.2012.09.002
Tedeschi R, Bortolin MT, Bidoli E, Zanussi S, Pratesi C, Vaccher E, Tirelli U, De Paoli P (2012) Assessment of immunovirological features in HIV related non-Hodgkin lymphoma patients and their impact on outcome. J Clin Virol 53(4):297–301. doi:10.1016/j.jcv.2011.12.021
Engels EA, Frisch M, Goedert JJ, Biggar RJ, Miller RW (2002) Merkel cell carcinoma and HIV infection. Lancet 359(9305):497–498. doi:10.1016/S0140-6736(02)07668-7
Lanoy E, Dores GM, Madeleine MM, Toro JR, Fraumeni JF Jr, Engels EA (2009) Epidemiology of nonkeratinocytic skin cancers among persons with AIDS in the United States. Aids 23(3):385–393. doi:10.1097/QAD.0b013e3283213046
Clarke CA, Robbins HA, Tatalovich Z, Lynch CF, Pawlish KS, Finch JL, Hernandez BY, Fraumeni JF Jr, Madeleine MM, Engels EA (2015) Risk of merkel cell carcinoma after solid organ transplantation. J Natl Cancer Inst. doi:10.1093/jnci/dju382
Hodgson NC (2005) Merkel cell carcinoma: changing incidence trends. J Surg Oncol 89(1):1–4. doi:10.1002/jso.20167
Becker JC, Houben R, Ugurel S, Trefzer U, Pfohler C, Schrama D (2009) MC polyomavirus is frequently present in Merkel cell carcinoma of European patients. J Invest Dermatol 129(1):248–250. doi:10.1038/jid.2008.198
Feng H, Shuda M, Chang Y, Moore PS (2008) Clonal integration of a polyomavirus in human Merkel cell carcinoma. Science 319(5866):1096–1100. doi:10.1126/science.1152586
Kassem A, Schopflin A, Diaz C, Weyers W, Stickeler E, Werner M, Zur Hausen A (2008) Frequent detection of Merkel cell polyomavirus in human Merkel cell carcinomas and identification of a unique deletion in the VP1 gene. Cancer Res 68(13):5009–5013. doi:10.1158/0008-5472.CAN-08-0949
Bhatia K, Goedert JJ, Modali R, Preiss L, Ayers LW (2010) Merkel cell carcinoma subgroups by Merkel cell polyomavirus DNA relative abundance and oncogene expression. Int J Cancer 126(9):2240–2246. doi:10.1002/ijc.24676
Houben R, Shuda M, Weinkam R, Schrama D, Feng H, Chang Y, Moore PS, Becker JC (2010) Merkel cell polyomavirus-infected Merkel cell carcinoma cells require expression of viral T antigens. J Virol 84(14):7064–7072. doi:10.1128/JVI.02400-09
Shuda M, Arora R, Kwun HJ, Feng H, Sarid R, Fernandez-Figueras MT, Tolstov Y, Gjoerup O, Mansukhani MM, Swerdlow SH, Chaudhary PM, Kirkwood JM, Nalesnik MA, Kant JA, Weiss LM, Moore PS, Chang Y (2009) Human Merkel cell polyomavirus infection I. MCV T antigen expression in Merkel cell carcinoma, lymphoid tissues and lymphoid tumors. Int J Cancer 125(6):1243–1249. doi:10.1002/ijc.24510
Viscidi RP, Rollison DE, Sondak VK, Silver B, Messina JL, Giuliano AR, Fulp W, Ajidahun A, Rivanera D (2011) Age-specific seroprevalence of Merkel cell polyomavirus, BK virus, and JC virus. Clin Vaccine Immunol 18(10):1737–1743. doi:10.1128/CVI.05175-11
Touze A, Le Bidre E, Laude H, Fleury MJ, Cazal R, Arnold F, Carlotti A, Maubec E, Aubin F, Avril MF, Rozenberg F, Tognon M, Maruani A, Guyetant S, Lorette G, Coursaget P (2011) High levels of antibodies against merkel cell polyomavirus identify a subset of patients with merkel cell carcinoma with better clinical outcome. J Clin Oncol 29(12):1612–1619. doi:10.1200/JCO.2010.31.1704
Carter JJ, Paulson KG, Wipf GC, Miranda D, Madeleine MM, Johnson LG, Lemos BD, Lee S, Warcola AH, Iyer JG, Nghiem P, Galloway DA (2009) Association of Merkel cell polyomavirus-specific antibodies with Merkel cell carcinoma. J Natl Cancer Inst 101(21):1510–1522. doi:10.1093/jnci/djp332
Pastrana DV, Tolstov YL, Becker JC, Moore PS, Chang Y, Buck CB (2009) Quantitation of human seroresponsiveness to Merkel cell polyomavirus. PLoS Pathog 5(9):e1000578. doi:10.1371/journal.ppat.1000578
Tolstov YL, Pastrana DV, Feng H, Becker JC, Jenkins FJ, Moschos S, Chang Y, Buck CB, Moore PS (2009) Human Merkel cell polyomavirus infection II. MCV is a common human infection that can be detected by conformational capsid epitope immunoassays. Int J Cancer 125(6):1250–1256. doi:10.1002/ijc.24509
Chen T, Hedman L, Mattila PS, Jartti T, Ruuskanen O, Soderlund-Venermo M, Hedman K (2011) Serological evidence of Merkel cell polyomavirus primary infections in childhood. J Clin Virol 50(2):125–129. doi:10.1016/j.jcv.2010.10.015
Touze A, Gaitan J, Arnold F, Cazal R, Fleury MJ, Combelas N, Sizaret PY, Guyetant S, Maruani A, Baay M, Tognon M, Coursaget P (2010) Generation of Merkel cell polyomavirus (MCV)-like particles and their application to detection of MCV antibodies. J Clin Microbiol 48(5):1767–1770. doi:10.1128/JCM.01691-09
Vahabpour R, Aghasadeghi MR, Salehi-Vaziri M, Mohajel N, Keyvani H, Nasimi M, Esghaei M, Monavari SH (2016) Prevalence of Merkel Cell polyomavirus in tehran: an age-specific serological study. Iran Red Crescent Med J 18(5):e26097. doi:10.5812/ircmj.26097
Wieland U, Mauch C, Kreuter A, Krieg T, Pfister H (2009) Merkel cell polyomavirus DNA in persons without merkel cell carcinoma. Emerg Infect Dis 15(9):1496–1498. doi:10.3201/eid1509.081575
Wieland U, Silling S, Scola N, Potthoff A, Gambichler T, Brockmeyer NH, Pfister H, Kreuter A (2011) Merkel cell polyomavirus infection in HIV-positive men. Arch Dermatol 147(4):401–406. doi:10.1001/archdermatol.2011.42
Jahanbakhsh F, Hattori J, Matsuda M, Ibe S, Monavari SH, Memarnejadian A, Aghasadeghi MR, Mostafavi E, Mohraz M, Jabbari H, Kamali K, Keyvani H, Azadmanesh K, Sugiura W (2013) Prevalence of transmitted HIV drug resistance in Iran between 2010 and 2011. PLoS One 8(4):e61864. doi:10.1371/journal.pone.0061864
Jahanbakhsh F, Ibe S, Hattori J, Monavari SH, Matsuda M, Maejima M, Iwatani Y, Memarnejadian A, Keyvani H, Azadmanesh K, Sugiura W (2013) Molecular epidemiology of HIV type 1 infection in Iran: genomic evidence of CRF35_AD predominance and CRF01_AE infection among individuals associated with injection drug use. AIDS Res Hum Retroviruses 29(1):198–203. doi:10.1089/AID.2012.0186
McNees AL, White ZS, Zanwar P, Vilchez RA, Butel JS (2005) Specific and quantitative detection of human polyomaviruses BKV, JCV, and SV40 by real time PCR. J Clin Virol 34(1):52–62. doi:10.1016/j.jcv.2004.12.018
Laude HC, Jonchere B, Maubec E, Carlotti A, Marinho E, Couturaud B, Peter M, Sastre-Garau X, Avril MF, Dupin N, Rozenberg F (2010) Distinct merkel cell polyomavirus molecular features in tumour and non tumour specimens from patients with merkel cell carcinoma. PLoS Pathog 6(8):e1001076. doi:10.1371/journal.ppat.1001076
Egli A, Infanti L, Dumoulin A, Buser A, Samaridis J, Stebler C, Gosert R, Hirsch HH (2009) Prevalence of polyomavirus BK and JC infection and replication in 400 healthy blood donors. J Infect Dis 199(6):837–846
White MK, Khalili K (2004) Polyomaviruses and human cancer: molecular mechanisms underlying patterns of tumorigenesis. Virology 324(1):1–16. doi:10.1016/j.virol.2004.03.025
Liu W, Yang R, Payne AS, Schowalter RM, Spurgeon ME, Lambert PF, Xu X, Buck CB, You J (2016) Identifying the target cells and mechanisms of Merkel cell polyomavirus infection. Cell Host Microbe 19(6):775–787. doi:10.1016/j.chom.2016.04.024
Bohl DL, Brennan DC, Ryschkewitsch C, Gaudreault-Keener M, Major EO, Storch GA (2008) BK virus antibody titers and intensity of infections after renal transplantation. J Clin Virol 43(2):184–189. doi:10.1016/j.jcv.2008.06.009
Randhawa P, Bohl D, Brennan D, Ruppert K, Ramaswami B, Storch G, March J, Shapiro R, Viscidi R (2008) longitudinal analysis of levels of immunoglobulins against BK virus capsid proteins in kidney transplant recipients. Clin Vaccine Immunol 15(10):1564–1571. doi:10.1128/CVI.00206-08
Acknowledgements
The authors would like to acknowledge the supervisor and staff of Keyvan virology laboratory. Funding was provided by Iran University of Medical Sciences (Grant No. 26059).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors report no conflicts of interest.
Ethical approval
This study followed the principles of the Declaration of Helsinki and was approved by the local ethics committee of the Iran University of Medical Sciences, Tehran, Iran.
Informed consent
Informed consent was obtained from all individual participants included in the study.
Rights and permissions
About this article
Cite this article
Vahabpour, R., Nasimi, M., Naderi, N. et al. Merkel cell polyomavirus IgG antibody levels are associated with progression to AIDS among HIV-infected individuals. Arch Virol 162, 963–969 (2017). https://doi.org/10.1007/s00705-016-3186-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00705-016-3186-z