Abstract
The northwest of Iran shares long borders with three neighboring countries; therefore, it is considered one of the main entry portals of Newcastle disease virus (NDV) into the country. Ten virulent NDVs were recovered from 19 poultry farms of various prefectures in northwestern Iran during Newcastle disease outbreaks in 2010. The isolates were genotypically analyzed using an F-gene-specific reverse transcription polymerase chain reaction (RT-PCR) assay. The amplified F gene (nucleotides 189-1666) sequences of the NDV isolates were compared phylogenetically with those of previously published strains in GenBank. All of the NDV isolates belonged to genotype VIIb and were closely related to some isolates from Iran, Russia, and Sweden. Therefore, it can be postulated that these isolates evolved from previously reported strains. The velogenic viruses carried the motif 112R-R-Q-K-R/F117 at the F0 cleavage site and a unique substitution of 190L→F which had never been reported in any NDV genotype VIIb isolate. They shared high sequence similarity with each other but were distinct from current NDV vaccines and NDV strains reported from other countries. This information is fundamental for improving the efficacy of controlling strategies and vaccine development for NDV.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Newcastle disease (ND) is one of the most serious and devastating diseases in the poultry industry [35]. The etiological agent, Newcastle disease virus (NDV), also known as avian paramyxovirus 1 (APMV-1), belongs to the genus Avulavirus within the family Paramyxoviridae [30]. This enveloped virus has a negative-sense, single-stranded RNA genome of approximately 15.2 kb that encodes six proteins, including nucleoprotein (NP), phosphoprotein (P), matrix protein (M), fusion protein (F), hemagglutinin-neuraminidase (HN), and a large RNA-directed RNA polymerase (L) [6].
Based on conventional in vivo pathogenicity indices for chickens, NDV strains can be categorized into three main pathotypes: highly virulent (velogenic), intermediate (mesogenic) and low-virulent (lentogenic) [5]. This difference in pathogenicity is due to differences in the amino acid sequence of the fusion protein cleavage site (FPCS). This protein is synthesized as an inactive precursor (F0) in non-functional state, which has to be cleaved into active F1 and F2 subunits for the virus to be infectious [16]. The virulent NDV (vNDV) has the motif 112R/K-R-Q-R/K-R↓F117 in its FPCS, whereas 112G/E-K/R-Q-G/E-R↓L117 is the consensus sequence for the cleavage site of low-virulent viruses. The presence of multiple basic amino acids in vNDV permits the virus to be cleaved by ubiquitous host proteases found in most tissues, resulting in a fatal systemic infection [38]. Conversely, the F0 protein of low-virulence APMV-1 strains is cleaved only in cells containing unique trypsin-like enzymes, limiting infection to mucosal tissues of the respiratory or intestinal tract of the host. Therefore, the FPCS is used as a virulence criterion along with biological virulence determinations [8].
All NDV isolates characterized to date belong to a single serotype, but significant genetic diversity among NDV isolates has been recognized [4]. Comparison of the nucleotide sequences of different strains of NDV has revealed two distinct NDV genotype groups, class Ι and ΙΙ, consisting of at least nine genotypes, designed 1-9 and I-IX, respectively [32, 46]. Class I viruses are distributed worldwide and have been isolated mainly from aquatic birds and live-bird markets, and most of them are avirulent [23]. Only class II viruses are responsible for fatal diseases in poultry [33]. The genotypes VI and VII are further divided into seven (VIa-g) and five (VIIa-e) subgenotypes, respectively [9, 27]. Genotypes V, VI, and VII of virulent viruses are the predominant genotypes circulating worldwide [32, 33]. Of these, genotype VII is particularly important, given that it has been associated with many of the most recent outbreaks in Asia, Africa, and the Middle East [22, 23, 27]. Thus, phylogenetic analysis of NDV is a powerful tool for investigating epidemiological relationships among NDV isolates present in various parts of the world.
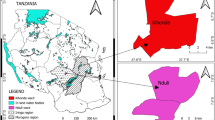
Despite the extensive and strict vaccination policy for prevention and control of NDV infections, ND is regarded as an endemic disease in some parts of the world, posing a constant threat to the poultry industry. In 2010, infection by vNDV was confirmed in domestic poultry in countries of Asia, Africa, Europe and North and South America [35]. However, very little data is available on virulent NDVs responsible for recent outbreaks in Iran. As the northwestern part of the country shares borders with two post-Soviet states, Armenia and Azerbaijan, besides Turkey and Iraq, it has been speculated that the diversity of vNDVs is high in that region. In order to evaluate the degree of genetic diversity of NDV strains circulating in commercial poultry in this part of Iran and to estimate their relationship to other NDV isolates, a part of the F gene from NDV isolates was characterized phylogenetically.
Materials and methods
Sampling and virus isolation
Tissue samples from four clinically diseased chickens suspected to have ND were collected from each of 19 different farms in various parts of northwestern Iran in 2010. Clinical signs observed on farms affected by ND in 2010 outbreaks included torticollis, paralysis of the legs and wings, tremors, diarrhea, lack of muscular coordination, and lethargy. Specimens including brain, spleen, and intestines of chickens with suspected infection were transferred to Razi Vaccine and Serum Research Institute, Iran. Filtrates of pooled homogenized tissues were used to inoculate fertilized specific-pathogen-free (SPF) chicken eggs (Lohmann, Australia) using standard procedures [27, 36]. The second egg passage of virus stocks was used for serological analysis. Also, allantoic fluid samples were harvested, divided into aliquots, stored at -70 °C and used as a working stock for molecular analysis. The identity of the virus was confirmed by haemagglutination inhibition assay with anti-NDV La Sota hyperimmune serum [27, 35] and F-gene-targeted reverse transcription polymerase chain reaction (RT-PCR) [21].
Virus pathotyping
The virulence of each NDV isolate was evaluated by mean death time (MDT) index [5, 18] and F-gene-based pathotype-specific primers as described previously [21]. All manipulations of live viruses were performed in a biosafety level 3 laboratory containment facility.
RNA extraction and F gene amplification
Genomic RNA was extracted directly from infected allantoic fluid using a commercial RNA extraction kit (Roche, Germany) according to the manufacturer’s instructions. Afterwards, a Titan One Tube RT-PCR System (Roche, Germany) was used to prime synthesis of first-strand cDNA and to perform PCR in one step. The reaction mixture (7 μL of purified template RNA, 4 μL of dNTPs mix [2.5 mM each], 1 μL of each forward and reverse primer [10 pM], 2.5 μL of DTT [100 mM], 0.5 μL of protector RNase inhibitor, 10 μL of 5X RT-PCR buffer, 1.5 mM MgCl2, and 1 μL of titan enzyme mix) were mixed in a final volume of 50 μL Using the previously published primers 5′-TTGATGGCAGGCCTCTTGC-3′ [21] and 5′-TTTGTAGTGGCTCTCATCTG-3′ [48], a 1561-bp product of the F gene was amplified. The thermal cycle profile was 45 °C for 45 min and 94 °C for 2 min, followed by 35 cycles of amplification (consisting of denaturation at 94 °C for 1 min, annealing at 60 °C for 75 s, and extension at 68 °C for 135 s) and a final extension step at 68 °C for 10 min. The 1561-bp products were subjected to electrophoresis on a 1 % agarose gel, and the DNA bands were excised from the gel and purified using a gel extraction kit (Roche, Germany).
Nucleotide sequencing and phylogenetic analysis
The purified products were submitted to the MWG-Biotech Company (Germany) for sequencing with the same primers from both ends. A 1478-bp sequence of the F gene was deposited in the GenBank database under accession numbers listed in Table 1. Nucleotide sequence editing and alignment and prediction of deduced amino acid sequences were conducted using BioEdit software version 7.0.9 [17]. Phylogenetic analysis was performed using Molecular Evolutionary Genetics Analysis version 6.0 (MEGA 6.0) [49]. Evolutionary distances were inferred using the neighbor-joining algorithm [43] with 1000 bootstrap replicates (Fig. 1). For comparison, the corresponding sequences of representative NDV strains retrieved from the GenBank database were also included (Table 2).
Alignment of the predicted amino acid sequences of the fusion proteins of NDV isolates. Partial sequences for 10 Iranian NDV isolates and 11 representative NDV strains from GenBank are shown. The sequences were derived by translating primary nucleotide sequences obtained by direct sequencing of RT-PCR products
Results
Identification and pathogenicity assessment of the NDV strains
Ten vNDVs were isolated from 19 ND outbreaks occurring on different farms. The isolates were confirmed using an HI test and by amplification of the F gene by PCR (data not shown) [2]. The MDT varied from 43.6 to 57.9 (Table 1). Additionally, a 254-bp amplicon produced from all of the isolates using primers specific for virulent strains, indicated that these viruses were probably highly pathogenic for chickens (data not shown) [2]. A 1561-bp amplicon corresponding to nucleotide positions 141-1701 of the F gene of NDV encompassing the FPCS was generated from all 10 virulent NDV isolates using oligonucleotide primers. Following nucleotide sequencing, the predicted amino acid sequence of the F protein was determined for each isolate.
Comparison of F gene nucleotide sequences of NDVs and their corresponding amino acid sequences
Comparison of nucleotide sequences of the 10 NDV isolates showed high sequence similarity (98.3 to 99.6 % identity at the nucleotide level and 97.3 to 98.9 % at the amino acid level) among them. Sequence analysis of the 10 NDVs based on partial F gene sequences showed 98.5 to 99 % nucleotide sequence identity and 86.4 to 87.3 % amino acid sequence identity to previously characterized MK13/75 NDV isolate from Iran. Comparison of the F gene sequences of NDVs isolated in the present study to those of reference strains from GenBank revealed a high level of similarity of the isolated NDVs to members of genotype VII of NDV. A high level of nucleotide and amino acid sequence identity was found among the isolates of the present study and some NDV strains (98.3 to 98.9 % nucleotide identity to strains chicken/Sweden/97, chicken/Itali/3286/00 and VOL95/Russia and 97.8 to 98.1 % nucleotide sequence identity to strain Sterna/Astr/2755/2001/Russia]. The isolated NDVs had low nucleotide and amino acid sequence similarity to the NDV vaccines commonly used on Iranian poultry farms (84 to 85.1 % identity to the La Sota vaccine and 84.4 to 85.5 % identity to the B1 vaccine at the nucleotide sequence level).
Proteolytic cleavage site of the F0 protein and virulence
The sequences of nucleotides 330–347 of the F0 gene corresponding to amino acid residues 112–117 at the F0 protein cleavage site of 10 NDV isolates are shown in Table 1. The MDT values were in accordance with those predicted based on the sequence of the FPCS. The FPCS sequence of the 10 NDV isolates had four basic amino acids [112R-R-Q-R-R116] at the C-terminus of the protein and a phenylalanine (F) at residue 117, representing a virulence motif according to the definition of the OIE (Table 1) [35].
Analysis of the nucleotide and deduced amino acid sequences of the F protein gene
The partial predicted amino acid sequences of the F proteins of 10 NDV isolates were 470 residues in length (aa 70 to 540) (Fig. 1). Cysteine residues in the partial F protein were conserved in all NDV isolates. There were 11 cysteine residues located at positions 76, 199, 338, 347, 362, 370, 394, 399, 401, 424, and 523 in the F protein. The predicted N-glycosylation sites of 10 NDV isolates were conserved. All of the isolates had five potential N-glycosylation sites, Asn-X-Ser/Thr (N-X-S/T), located at positions 85NRT87, 191NNT193, 366NTS368, 447NIS449, and 471NNS473. At least 11 neutralizing epitopes, positioned at residues 75, 79, 158, 159, 160, 164, 165, 168, 171, 343, and 378 in the partial F protein, which are believed to be critical for the structure and function of the F protein, have been identified in all examined isolates [34, 51, 52]. The GC content of the partial sequenced F gene was estimated to be 43.57 to 44.05 %.
Although there were similarities among NDV isolates in the present study and their worldwide counterparts, as shown in the phylogenetic tree, the newly characterized NDVs still contained several amino acid substitutions. For instance, substitutions of 190F to L in all 10 newly characterized NDV isolates, 265G to S in NR_2, NR_3, NR_9, NR_10, and NR_14, 479D to N in NR_6, NR_7, NR_11, NR_13, and NR_15, 514F to S in all isolates except NR_2 and NR_3, 489N to G in NR_2 and NR_3, 226T to N, 235I to F, and 269I to N in NR_11 and NR_15 were unique to these isolates. Moreover, analysis of unique residue substitution revealed that the 190th amino acid position of NDVs analyzed from northwestern Iran have leucine (L) replaced by phenylalanine (F), which has never seen in other strains of genotype VIIb.
Phylogenetic analysis
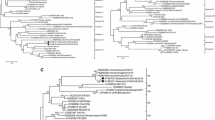
The evolutionary distances among the 2010 NDV isolates from northwestern Iran and other NDV isolates were assessed by phylogenetic analysis. A 1478-bp region corresponding to nucleotides 189-1666 of the F gene was compared to those of other strains belonging to genotypes I to VIII of class II, together with sequences of isolates representing subtypes VIIa, VIIb, VIIc, VIId, and VIIe. The resulting phylogenetic tree is shown in Fig. 2. From the topology of the tree, it was apparent that all NDV isolates under study belonged to genotype VIIb, or lineage 5b, the prevalent pathogen involved in ND outbreaks in northwest of Iran in 2010 to 2011. These viruses were related to chicken/Sweden/97 and Sterna/Astr/2755/2001/Russia as possible progenitor-type viruses. Analysis of the deduced amino acid sequences of the collected samples and data obtained from GenBank did not show any difference in the phylogenetic grouping of samples (results not shown).
Phylogenetic comparison of NDV isolates based on their partial F gene sequences (nt189 to 1666) constructed using the neighbor-joining method with 1000 bootstrap replicates (numbers at branching points are bootstrap values given as a percentage). Northwestern Iran isolates from this study are indicated by a solid circle. Provisional designations of the genotypes are shown at the right
Discussion
From 2009 to the middle of 2012, outbreaks of ND on poultry farms throughout Iran were frequently reported to World Organization for Animal Health [35]. Because it shares long borders with several countries, northwestern Iran is considered one of the main entry portals of NDV.
In the present study, the molecular characteristics of 10 vNDV isolates recovered from northwestern Iran during 2010 outbreaks were investigated. Sequence analysis of a 1487-bp fragment of the F gene from each of the NDV isolates, showed that they clustered with virulent NDVs belong to genotype VIIb and were closely related to some Iranian (AY928933), Swedish (GU585905) and Russian isolates (AY865652). Moreover, the predominant sub-genotype VIIb existed in Iran and Indian subcontinent countries during 2008-2011 [14]. The close nucleotide and amino acid sequence similarity among the newly isolates and the representative virulent isolates from Iran supports the idea of a local evolutionary trend with regional viral dissemination [14, 44]. Therefore, it could be hypothesized that genotype VIIb NDV was present later and is still circulating in the region. A possibility for the maintenance of the virus in the northwestern region of Iran is the existence of a large bird population as the virus reservoir. In contrast, vNDVs obtained from the 2010 to 2011 outbreaks in Shiraz, a city in the south of the country in Iran’s Region 2, were phylogenetically different from prior isolates, and consequently, from the isolates from northwestern Iran. NDV isolates in the present study possessed the velogenic motif 112R-R-Q-K-R/F117 at the FPCS and clustered in genotype VIId with some Chinese strains, possibly as a result of the import of agricultural products from China to Iran [10]. Conversely, the vNDVs isolated from Industrial poultry throughout the country in the 2011 and 2012 outbreaks, including Ardabil (in the northwest of the country in Iran’s Region 3) and Isfahan (in the center of the country in Iran’s Region 2) as well as five other provinces, were related to some Chinese and Palestinian isolates based on the motif 112R-R-Q-K-R/F117 at the FPCS [42]. In addition, phylogenetic analysis revealed a close association among the prevailing vNDVs isolated from industrial poultry of Isfahan province almost at the end of 2012 with isolates from India (AY339401.1) and Kazakhstan (HQ445947.1). They carried a 112R-R-Q-R-R/F117 motif at their cleavage sites [41].
Based on available data, different vNDVs strains have circulated in Iran [11, 31, 44]. Emergence of new strains of vNDVs might be attributed either to the increased trade of agricultural products including poultry or to the movement of migratory birds from the north to the east of the world, involving Europe, Asia, and the Middle East. The presence of multiple velogenic strains of NDV at different times and in various regions of the country emphasizes the need for continued isolation and epidemiological investigation of NDVs circulating across Iran to find the dominant patterns of vNDVs. In a study in Pakistan, a neighboring country to the southeast of Iran, NDVs isolated from commercial poultry were clustered within sub-lineage 5b and were closely related to the viruses isolated in the present study [47]. Therefore, it could be estimated that if all suspicious cases of ND throughout Iran had been evaluated, more diversity of vNDV strains, including the isolates prevailing before 2010, might have been identified.
Although the vNDVs involved outbreaks of the northwest of Iran in 2010 originated from identical or quite similar velogenic strains of the virus, they contained several point mutations, among which the unique residue substitution 190L→F was recognized as a specified marker in these isolates and was reported for the first time in VIIb NDVs. Based on the published phylogenetic data, the only other L-to-F substitution in the vNDV fusion protein was at amino acid 23 in genotype VIId reported by Lien et al. [26], which differs in position and genotype from our findings. Therefore, the importance of the190L→F substitution in virus pathogenicity and vaccine escape could not be determined in the present study due to the lack of information. Furthermore, amino acid sequence analysis revealed that the fusion-inducing hydrophobic stretch at the N-terminus of the F1 protein (amino acid residues 117 to 142) and cysteine at position 76 are important for disulphide bond formation between F1 and F2. The results also showed that amino acids at potential glycosylation sites were conserved in all 10 ND viruses, as previously reported by other researchers [45, 50]. All of these conserved regions have a considerable role in the precise conformation of the fusion protein vis-à-vis survival of the virus in nature. The FPCS sequence is considered the molecular determinant of NDV pathogenicity in chickens [3, 40]. A typical cleavage site sequence (112R-R-Q-R-R/F117) of virulent NDV present in all 10 isolates in the present study remained conserved. Furthermore, the presence of a phenylalanine (F) residue at position 117 is also potentially associated with neurological signs [25].
In northwestern Iran and the rest of the country as well, a rigorous vaccination program using live (during the rearing and laying period) and killed vaccines (only in the rearing period) are employed in order to induce high levels of maternal antibodies to protect breeders against egg production losses and clinical disease [11, 20, 29]. Nonetheless, ND outbreaks in vaccinated broilers reared under field conditions has increased, and these outbreaks are responsible for high mortality rate and production losses in the poultry industry [15, 20]. As all NDV isolates belong to the same serotype, ND vaccines prepared with any NDV should protect poultry from clinical disease and mortality in the event of a virulent challenge with heterologous strains of different genotypes [35, 39]. La Sota and Clone 30, the most extensively used live vaccines in Iran, belong to genotype II and are different from the prevalent genotype VII. The inability of live vaccines to elicit a protective immune response might be due to several reasons, such as an improper cold chain supply, an inappropriate route of vaccination, or uneven vaccination schedules [47].
In summary, the presence of velogenic NDVs belonging to genotype VIIb has been confirmed in northwestern Iran. Genotype VII of NDV is now regarded as the major pathogen responsible for the fourth panzootic of ND [1, 7, 9, 12, 13, 19, 24, 28, 36, 37]. Therefore, the development and administration of new NDV vaccines that are closely related to predominant VII viruses may confer better protection than conventional vaccines. Further investigation of the molecular epidemiology of NDV and ND vaccinal strains is recommend to explore circulating NDVs in the region in order to develop the most effective and protective methods.
References
Abolnik C, Horner RF, Bisschop SP, Parker ME, Romito M, Viljoen GJ (2004) A phylogenetic study of South African Newcastle disease virus strains isolated between 1990 and 2002 suggests epidemiological origins in the Far East. Arch Virol 149:603–619
Ahmadi E, Pourbakhsh SA, Ahmadi M, Talebi A (2014) Pathotypic characterization of the Newcastle disease virus isolated from commercial poultry in northwest Iran. Turk J Vet Anim Sci 38:383–387
Aldous EW, Alexander DJ (2001) Detection and differentiation of Newcastle disease virus (avian paramyxovirus type 1). Avian Pathol 30:117–128
Aldous EW, Mynn JK, Banks J, Alexander DJ (2003) A molecular epidemiological study of avian paramyxovirus type 1 (Newcastle disease virus) isolates by phylogenetic analysis of a partial nucleotide sequence of the fusion protein gene. Avian Pathol 32:239–256
Alexander DJ (2000) Newcastle disease and other avian paramyxoviruses. Rev Sci Tech 19:443–462
Alexander DJ (2011) Newcastle disease in the European Union 2000 to 2009. Avian Pathol 40:547–558
Ballagi-Pordany A, Wehmann E, Herczeg J, Belak S, Lomniczi B (1996) Identification and grouping of Newcastle disease virus strains by restriction site analysis of a region from the F gene. Arch Virol 141:243–261
Berinstein A, Sellers HS, King DJ, Seal BS (2001) Use of a heteroduplex mobility assay to detect differences in the fusion protein cleavage site coding sequence among Newcastle disease virus isolates. J Clin Microbiol 39:3171–3178
Bogoyavlenskiy A, Berezin V, Prilipov A, Usachev E, Lyapina O, Levandovskaya S, Korotetskiy I, Tolmacheva V, Makhmudova N, Khudyakova S, Tustikbaeva G, Zaitseva I, Omirtaeva E, Ermakova O, Daulbaeva K, Asanova S, Kydyrmanov A, Sayatov M, King D (2005) Molecular characterization of virulent Newcastle disease virus isolates from chickens during the 1998 NDV outbreak in Kazakhstan. Virus Gen 31:13–20
Boostani A, Pourbakhsh S, Momayez R, Charkhkar S (2013) Molecular characterization and phylogenetic study of Newcastle disease virus isolates from the 2010 to 2011 outbreaks in Shiraz, Iran. Afr J Microbiol Res 7:657–660
Boroomand Z, Jafari RA, Mayahi M (2016) Molecular characterization and phylogenetic study of the fusion genes of Newcastle disease virus from the recent outbreaks in Ahvaz, Iran. Virusdisease 27:102–105
Chen JP, Wang CH (2002) Phylogenetic analysis of Newcastle disease virus in Taiwan. J Microbiol Immunol Infect 35:223–228
Czegledi A, Herczeg J, Hadjiev G, Doumanova L, Wehmann E, Lomniczi B (2002) The occurrence of five major Newcastle disease virus genotypes (II, IV, V, VI and VIIb) in Bulgaria between 1959 and 1996. Epidemiol Infect 129:679–688
Ebrahimi MM, Shahsavandi S, Moazenijula G, Shamsara M (2012) Phylogeny and evolution of Newcastle disease virus genotypes isolated in Asia during 2008–2011. Virus Gen 45:63–68
Ghahramani B, Alipour R, Mehrani K, Mehrvarz M, Gerayili Moghaddam S (2014) Evaluation of two different Newcastle disease vaccination programs in broiler breeder chickens by HI tests. Eur J Exp Biol 4:133–136
Gould AR, Kattenbelt JA, Selleck P, Hansson E, Della-Porta A, Westbury HA (2001) Virulent Newcastle disease in Australia: molecular epidemiological analysis of viruses isolated prior to and during the outbreaks of 1998–2000. Virus Res 77:51–60
Hall TA (1999) BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic Acids Symp Ser 41:95–98
Hanson RP, Brandly CA (1955) Identification of vaccine strains of Newcastle disease virus. Science 122:156–157
Herczeg J, Wehmann E, Bragg RR, Travassos Dias PM, Hadjiev G, Werner O, Lomniczi B (1999) Two novel genetic groups (VIIb and VIII) responsible for recent Newcastle disease outbreaks in Southern Africa, one (VIIb) of which reached Southern Europe. Arch Virol 144:2087–2099
Hosseini H, Langeroudi AG, Torabi R (2014) Molecular characterization and phylogenetic study of Newcastle disease viruses isolated in Iran, 2010–2012. Avian Dis 58:373–376
Kant A, Koch G, Van Roozelaar DJ, Balk F, Huurne AT (1997) Differentiation of virulent and non-virulent strains of Newcastle disease virus within 24 hours by polymerase chain reaction. Avian Pathol 26:837–849
Khan TA, Rue CA, Rehmani SF, Ahmed A, Wasilenko JL, Miller PJ, Afonso CL (2010) Phylogenetic and biological characterization of Newcastle disease virus isolates from Pakistan. J Clin Microbiol 48:1892–1894
Kim LM, King DJ, Suarez DL, Wong CW, Afonso CL (2007) Characterization of class I Newcastle disease virus isolates from Hong Kong live bird markets and detection using real-time reverse transcription-PCR. J Clin Microbiol 45:1310–1314
Kwon HJ, Cho SH, Ahn YJ, Seo SH, Choi KS, Kim SJ (2003) Molecular epidemiology of Newcastle disease in Republic of Korea. Vet Microbiol 95:39–48
Lamb RA, Parks G (2007) Paramyxoviridae: the viruses and their replication. In: Knipe DM, Howley PM, Griffin DE, Lamb RA, Martin MA, Roizman B, Straus SE (eds) Fields virology. Lippincott Williams, Philadelphia, pp 1449–1496
Lien YY, Lee JW, Su HY, Tsai HJ, Tsai MC, Hsieh CY, Tsai SS (2007) Phylogenetic characterization of Newcastle disease viruses isolated in Taiwan during 2003–2006. Vet Microbiol 123:194–202
Liu XF, Wan HQ, Ni XX, Wu YT, Liu WB (2003) Pathotypical and genotypical characterization of strains of Newcastle disease virus isolated from outbreaks in chicken and goose flocks in some regions of China during 1985–2001. Arch Virol 148:1387–1403
Liu YL, Hu SL, Zhang YM, Sun SJ, Romer-Oberdorfer A, Veits J, Wu YT, Wan HQ, Liu XF (2007) Generation of a velogenic Newcastle disease virus from cDNA and expression of the green fluorescent protein. Arch Virol 152:1241–1249
Madadi MS, Ghaniei A, Shojaei H (2014) Evaluation of B1 and Lasota vaccines for hyperimmunization of breeder flocks in production phase. Int J Basic Sci Appl Res 3:946–950
Mayo MA (2002) Virus taxonomy—Houston 2002. Arch Virol 147:1071–1076
Mehrabanpour MJ, Khoobyar S, Rahimian A, Nazari MB, Keshtkar MR (2014) Phylogenetic characterization of the fusion genes of the Newcastle disease viruses isolated in Fars province poultry farms during 2009–2011. Vet Res Forum 5:187–191
Miller PJ, Kim LM, Ip HS, Afonso CL (2009) Evolutionary dynamics of Newcastle disease virus. Virology 391:64–72
Miller PJ, Decanini EL, Afonso CL (2010) Newcastle disease: evolution of genotypes and the related diagnostic challenges. Infect Genet Evol 10:26–35
Neyt C, Geliebter J, Slaoui M, Morales D, Meulemans G, Burny A (1989) Mutations located on both F1 and F2 subunits of the Newcastle disease virus fusion protein confer resistance to neutralization with monoclonal antibodies. J Virol 63:952–954
OIE (2012) Newcastle disease. In: OIE (ed) World Organization for Animal Health (OIE). OIE, Paris, pp 1–19
Otim MO, Christensen H, Jorgensen PH, Handberg KJ, Bisgaard M (2004) Molecular characterization and phylogenetic study of newcastle disease virus isolates from recent outbreaks in eastern Uganda. J Clin Microbiol 42:2802–2805
Pedersen JC, Senne DA, Woolcock PR, Kinde H, King DJ, Wise MG, Panigrahy B, Seal BS (2004) Phylogenetic relationships among virulent Newcastle disease virus isolates from the 2002–2003 outbreak in California and other recent outbreaks in North America. J Clin Microbiol 42:2329–2334
Peeters BP, de Leeuw OS, Koch G, Gielkens AL (1999) Rescue of Newcastle disease virus from cloned cDNA: evidence that cleavability of the fusion protein is a major determinant for virulence. J Virol 73:5001–5009
Perozo F, Villegas P, Dolz R, Afonso CL, Purvis LB (2008) The VG/GA strain of Newcastle disease virus: mucosal immunity, protection against lethal challenge and molecular analysis. Avian Pathol 37:237–245
Pritzer E, Kuroda K, Garten W, Nagai Y, Klenk HD (1990) A host range mutant of Newcastle disease virus with an altered cleavage site for proteolytic activation of the F protein. Virus Res 15:237–242
Rahimian MD, Moghaddam AZ, Momtaz H, Niazi MH (2012) Detection and phylogenetic Analysis of Newsactle disease virus based on molecular techniques in broiler in Isfahan province. J Vet Microbiol 7:25–36
Rostamali T, Shushtari A, Charkhkar S, Bozorgmehri fard MH (2012) Phylogenetic analysis of Newcastle disease viruses Isolated in Iran between 2011 to 2012 based on F gene. Comp Pathobiol J 10:1065–1070
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
Samadi S, Kianizadeh M, Najafi MF, Nasab SD, Davatgar AM, Royaee A, Pilvar P (2014) Molecular characterization and phylogenetic study of velogenic Newcastle disease virus isolates in Iran. Virus Gen 48:290–295
Seal BS, King DJ, Bennett JD (1995) Characterization of Newcastle disease virus isolates by reverse transcription PCR coupled to direct nucleotide sequencing and development of sequence database for pathotype prediction and molecular epidemiological analysis. J Clin Microbiol 33:2624–2630
Seal BS, King DJ, Bennett JD (1996) Characterization of Newcastle disease virus vaccines by biological properties and sequence analysis of the hemagglutinin–neuraminidase protein gene. Vaccine 14:761–766
Shabbir MZ, Zohari S, Yaqub T, Nazir J, Shabbir MA, Mukhtar N, Shafee M, Sajid M, Anees M, Abbas M, Khan MT, Ali AA, Ghafoor A, Ahad A, Channa AA, Anjum AA, Hussain N, Ahmad A, Goraya MU, Iqbal Z, Khan SA, Aslam HB, Zehra K, Sohail MU, Yaqub W, Ahmad N, Berg M, Munir M (2013) Genetic diversity of Newcastle disease virus in Pakistan: a countrywide perspective. Virol J 10:170
Tahmasian R, Foruzandeh M, Madani R, Esmaeelzadeh M, Dehghani MR (2007) Cloning and sequencing of HR1 and HR2 parts of F gene from NR43 Newcastle disease virus isolate. Med Daneshvar 62:43–50
Tamura K, Stecher G, Peterson D, Filipski A, Kumar S (2013) MEGA6: molecular evolutionary genetics analysis version 6.0. Mol Biol Evol 30:2725–2729
Toyoda T, Sakaguchi T, Imai K, Inocencio NM, Gotoh B, Hamaguchi M, Nagai Y (1987) Structural comparison of the cleavage-activation site of the fusion glycoprotein between virulent and avirulent strains of Newcastle disease virus. Virology 158:242–247
Toyoda T, Gotoh B, Sakaguchi T, Kida H, Nagai Y (1988) Identification of amino acids relevant to three antigenic determinants on the fusion protein of Newcastle disease virus that are involved in fusion inhibition and neutralization. J Virol 62:4427–4430
Yusoff K, Nesbit M, McCartney H, Meulemans G, Alexander DJ, Collins MS, Emmerson PT, Samson AC (1989) Location of neutralizing epitopes on the fusion protein of Newcastle disease virus strain Beaudette C. J Gen Virol 70(Pt 11):3105–3109
Acknowledgments
This research was funded by Razi Vaccine and Serum Research Institute (RVSRI) (Grant No. A543). The authors would like thank RVSRI and Urmia University for providing technical and financial support for this research.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
All authors declare no conflict of interest.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Rights and permissions
About this article
Cite this article
Ahmadi, E., Pourbakhsh, S.A., Ahmadi, M. et al. Phylogenetic characterization of virulent Newcastle disease viruses isolated during outbreaks in northwestern Iran in 2010. Arch Virol 161, 3151–3160 (2016). https://doi.org/10.1007/s00705-016-3021-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00705-016-3021-6