Abstract
Bacteriophage JBP901, isolated from fermented food, is specific for Bacillus cereus group species and exhibits a broad host spectrum among a large number of B. cereus isolates. Genome sequence analysis revealed a linear 159,492-bp genome with overall G+C content of 39.7 mol%, and 201 ORFs. The presence of a putative methylase, the large number of tRNAs, and the large number of nucleotide-metabolism- and replication-related genes in JBP901 reflects its broad lytic capacity. Most of the ORFs showed a high degree of similarity to Bcp1, Bc431v3 and BCP78, and various comparative genomics analyses also consistently clustered JBP901 with orphan (unclassified) Bacillus phages in the subfamily Spounavirinae of the family Myoviridae, supporting the presence of a distinguishable group in the subfamily.
Similar content being viewed by others

Avoid common mistakes on your manuscript.
Bacillus cereus, a member of B. cereus sensu lacto group [32], is an opportunistic foodborne pathogen that causes diarrhea, vomiting, and nausea [5, 8]. It has been isolated from many foods and food additives such as milk, cheese [1, 33], cereals, rice [23], dried red pepper [13], and fermented foods [22, 30, 31], with contamination levels of higher than 104 CFU/g [9], which is not acceptable by regulatory authorities [11, 17].
Bacteriophage JBP901, isolated from Korean fermented food, exhibited a lysis profile specific to B. cereus group species, such as B. cereus, B. thuringiensis, B. mycoides, B. weihenstephanensis [26] and even B. anthracis (unpublished data). Further studies showed that JBP901 exhibited a broad host spectrum among the B. cereus isolates, being able to lyse 279 of 344 (81 %) food and human isolates without affecting the growth of B. subtilis (50 food isolates) and B. licheniformis (12 food isolates), which are necessary for beneficial fermentation [4]. In this study, we sequenced and annotated the genome of JBP901 and provided evidence for its classification in the subfamily Spounavirinae.
Single-plaque isolation, large-scale preparation, PEG (polyethylene glycol) precipitation, CsCl gradient ultracentrifugation, and dialysis to yield a high titer of phage JBP901 have been reported previously [26]. The nucleic acid of JBP901 was manually extracted from a phage stock using a phenol-chloroform extraction protocol after DNase and SDS treatment [24]. Whole-genome sequencing of JBP901 was carried out at Macrogen Inc., South Korea, using a 454 GS-FLX system (Titanium GS70 Chemistry, Roche Life Science, Mannheim, Germany). Assembly of 97,624 reads using the Roche Newbler assembly software 2.9 yielded a single contig and an average coverage of 55× (minimum coverage of 17×) with a quality filter threshold of Q30. Geneious software v8.14 [12] was used to map total reads to the assembled genome of JBP901 to test for the presence of terminal repeats [15]. Coding sequences (CDSs) were predicted using Glimmer 3.02 [6], FgeneB [27], GeneMark.hmm 2.8 [20] and RAST 4.0 [3]. Only CDSs predicted by at least three of the above tools were selected for further analysis. The individual proteins were functionally annotated using the protein-protein BLAST (BLASTP) algorithm [2]. tRNA genes were identified using the tRNA-SE v.1.21 program [19] with default parameters. We also employed Easyfig [29] and CoreGenes 3.0 [21] to compare the genome with other similar Myoviridae phages at the nucleotide and protein level, respectively.
The JBP901 genome is 159,492 bp long with an overall G+C content of 39.7 mol%. A higher read coverage was observed in the region between nucleotides 22,000 and 29,500 using Geneious software (data not shown), suggesting the existence of a terminal redundancy region (data not shown) [34]. A total of 201 open reading frames (ORFs) (172 on the reverse strand and 29 on the forward strand) and 19 tRNAs were identified (Supplementary Fig. 1). Among the total ORFs, 41 putative proteins exhibited significant similarity (higher than 50 % amino acid sequence identity) to the functionally annotated proteins of bacteriophages in the database, and these were then classified into six functional groups (Supplementary Table 1), including structure (tail protein, tail sheath protein, major capsid precursor, prohead protease, adsorption associated tail protein, baseplate protein I, baseplate protein II, minor tail protein, tail lysin and prohead protease), nucleotide metabolism (thymidylate synthase, methyltransferase, C-5 cytosine-specific DNA methylase I, II & III, nucleotidyltransferase, ribonucleoside-diphosphate reductase alpha & beta subunit, and deoxyuridine 5’-triphosphate) DNA replication and transcription (DNA primase, DNA-binding protein, exonuclease I & II, DNA helicase, RecA-like recombinase, ssDNA binding protein, and DNA polymerase I & II, transcriptional regulator I & II), DNA packaging (terminase large subunit and portal protein), host lysis (cell wall hydrolase/autolysin and holin), and others (PhoH family protein, metallo-beta-lactamase superfamily protein, FtsK_SpoIIIE-family protein, RNA polymerase sigma-70 factor and metallo-dependent phosphatase protein).
It is generally accepted that phages for biocontrol must possess a broad host range, be strictly lytic, and lack genes responsible for virulence, or those that may enhance the pathogenic properties of the host [10]. Genomic sequence analysis confirmed that JBP901 does not contain integrase or virulence-related proteins. However, ORF061 was annotated as a metallo-beta-lactamase superfamily protein that contains a conserved domain for the lactamase_B family. Considering the low amino acid sequence similarity (approximately 27 % identity with 62 % query coverage) to metal-dependent hydrolase in B. cereus (GenBank taxon: 1396), its functional identity remains unclear and needs to be studied further.
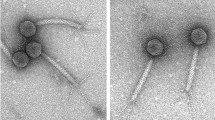
Previously, JBP901 was tentatively classified as a member of the SPO1-like group phages of the family Myoviridae, based on its morphological features [26]. However, genome analysis revealed that JBP901 lacks dUMP hydroxymethylase (deoxyuridylate hydroxymethyl-transferase), a signature gene (Gp 29 in SPO1) of SPO1 phage [28]. In addition, the top-scoring hits in BLAST analysis (BLASTP) of JBP901 ORFs are from Bcp1 [25], Bc431v3 [7], and BCP78 [18] genomes (Supplementary Table 1), not from SPO1-like or Twort-like phages. Likewise, when the whole nucleotide sequence of JBP901 was compared to the large genomes of Myoviridae phages using Easyfig [29], a shared synteny was found with the phages Bcp1, BCP78, Bc431v3 and JP901 (Fig. 1A) but not with SPO1 and Twort (Fig. 1B).
(A) Genome comparison of phage JBP901 and the related phages Bcp1 (NC_024137), Bc431v3 (JX094431), BCP78 (JN797797), phiAGATE (NC_020081), and Bastille (JF966203). The figure was constructed with Easyfig v 2.1. (B) Genome comparison of phage SPO1 (NC_011421), Twort (AY954970), and BCP8-2 using Easyfig v 2.1. The gray region shows sequence similarity
CoreGenes analysis using a BLASTP threshold score of 75 as described previously [16] showed that JBP901 shared a high proportion of its proteome with Bc431v3 (the percentage of proteins shared was 89.6 %), Bcp1 (84.1 %) and BCP78 (67.2 %) but a low proportion with Twort (33.3 %) or SPO1 (29.9 %). When pairwise comparison among the three most similar phages, Bc431v3, Bcp1 and JBP901, was performed using CoreGenes, 163 genes were similar for all three phages, while 34, 17 and 9 genes were shared only between Bcp1 and Bc431v3, between Bc431v3 and JBP901, and between Bcp1 and JBP901, respectively (Supplementary Fig. 2). Twenty-three, 24 and 12 genes were identified to be unique to Bcp1, Bc431v3 and JBP901, respectively (Supplementary Fig. 2).
A maximum-likelihood tree (ML) (Supplementary Fig. 3) drawn with either the putative major capsid proteins or tail sheath proteins of JBP901 and other phages (including all ICTV classified Spounavirinae phages) supported that JBP901 is not a member of the SP01-like phages. Taken together, our data support the presence of a distinguishable group in the subfamily Spounavirinae, different from the SPO1-like phages under the current ICTV [14] classification scheme for the family Myoviridae.
Nucleotide sequence accession number
The whole genome sequence of phage JBP901 was deposited in the GenBank database with the accession number KJ676859.
References
Ahmed A (1983) Incidence of Bacillus cereus in milk and some milk products. J Food Prot 46:126–128
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ (1990) Basic local alignment search tool. J Mol Biol 215:403–410
Aziz RK, Bartels D, Best AA, DeJongh M, Disz T, Edwards RA, Formsma K, Gerdes S, Glass EM, Kubal M (2008) The RAST Server: rapid annotations using subsystems technology. BMC Genomics 9:75
Bandara N, Kim CH, Kim K-P (2014) Extensive host range determination and improved efficacy of the bacteriophage JBP901 in the presence of divalent cations for control of Bacillus cereus in Cheonggukjang. Food Sci Biotechnol 23:499–504
Bottone EJ (2010) Bacillus cereus, a volatile human pathogen. Clin Microbiol Rev 23:382–398
Delcher AL, Bratke KA, Powers EC, Salzberg SL (2007) Identifying bacterial genes and endosymbiont DNA with Glimmer. Bioinformatics 23:673–679
El-Arabi TF, Griffiths MW, She Y-M, Villegas A, Lingohr EJ, Kropinski AM (2013) Genome sequence and analysis of a broad-host range lytic bacteriophage that infects the Bacillus cereus group. Virol J 10:48
Granum PE, Lund T (1997) Bacillus cereus and its food poisoning toxins. FEMS Microbiol Lett 157:223–228
Guinebretière M-H, Broussolle V (2002) Enterotoxigenic profiles of food-poisoning and food-borne Bacillus cereus strains. J Clin Microbiol 40:3053–3056
Hagens S, Loessner MJ (2010) Bacteriophage for biocontrol of foodborne pathogens: calculations and considerations. Curr Pharm Biotechnol 11:58–68
Hugas M, Tsigarida E, Robinson T, Calistri P (2007) Risk assessment of biological hazards in the European Union. Int J Food Microbiol 120:131–135
Kearse M, Moir R, Wilson A, Stones-Havas S, Cheung M, Sturrock S, Buxton S, Cooper A, Markowitz S, Duran C (2012) Geneious Basic: an integrated and extendable desktop software platform for the organization and analysis of sequence data. Bioinformatics 28:1647–1649
Ki Kim S, Kim K-P, Jang SS, Mi Shin E, Kim M-J, Oh S, Ryu S (2009) Prevalence and toxigenic profiles of Bacillus cereus isolated from dried red peppers, rice, and Sunsik in Korea. J Food Prot 72:578–582
King AMQ, Adams MJ, Lefkowitz EJ, Carstens EB (2012) Virus taxonomy: classification and nomenclature of viruses: ninth report of the International Committee on Taxonomy of Viruses. Elsevier Academic Press
Klumpp J, Lavigne R, Loessner MJ, Ackermann H-W (2010) The SPO1-related bacteriophages. Arch Virol 155:1547–1561
Lavigne R, Darius P, Summer EJ, Seto D, Mahadevan P, Nilsson AS, Ackermann HW, Kropinski AM (2009) Classification of Myoviridae bacteriophages using protein sequence similarity. BMC Microbiol 9:224
Lee G-I, Lee H-M, Lee C-H (2012) Food safety issues in industrialization of traditional Korean foods. Food Cont 24:1–5
Lee J-H, Shin H, Son B, Ryu S (2012) Complete genome sequence of Bacillus cereus bacteriophage BCP78. J Virol 86:637–638
Lowe TM, Eddy SR (1997) tRNAscan-SE: a program for improved detection of transfer RNA genes in genomic sequence. Nucleic Acids Res 25:0955–0964
Lukashin AV, Borodovsky M (1998) GeneMark. hmm: new solutions for gene finding. Nucleic Acids Res 26:1107–1115
Mahadevan P, King JF, Seto D (2009) CGUG: in silico proteome and genome parsing tool for the determination of “core” and unique genes in the analysis of genomes up to ca. 1.9 Mb. BMC Res Notes 2:168
Oguntoyinbo FA, Mathew Oni O (2004) Incidence and characterization of Bacillus cereus isolated from traditional fermented meals in Nigeria. J Food Prot 67:2805–2808
Park Y-B, Kim J-B, Shin S-W, Kim J-C, Cho S-H, Lee B-K, Ahn J, Kim J-M, Oh D-H (2009) Prevalence, genetic diversity, and antibiotic susceptibility of Bacillus cereus strains isolated from rice and cereals collected in Korea. J Food Prot 72:612–617
Sambrook J, Russell DW, Russell DW (2001) Molecular cloning: a laboratory manual (3-volume set). Cold Spring Harbor Laboratory Press, Cold Spring Harbor
Schuch R, Pelzek AJ, Fazzini MM, Nelson DC, Fischetti VA (2014) Complete genome sequence of Bacillus cereus sensu lato bacteriophage Bcp1. Genome Announc 2:e00334-00314
Shin H, Bandara N, Shin E, Ryu S, Kim KP (2011) Prevalence of Bacillus cereus bacteriophages in fermented foods and characterization of phage JBP901. Res Microbiol 162:791–797
Solovyev V, Salamov A (2011) Automatic annotation of microbial genomes and metagenomic sequences. Metagenomics and its applications in agriculture, biomedicine and environmental studies. Nova Science Publishers, Hauppauge, pp 61–78
Stewart CR, Casjens SR, Cresawn SG, Houtz JM, Smith AL, Ford ME, Peebles CL, Hatfull GF, Hendrix RW, Huang WM (2009) The genome of Bacillus subtilis bacteriophage SPO1. J Mol Biol 388:48–70
Sullivan MJ, Petty NK, Beatson SA (2011) Easyfig: a genome comparison visualizer. Bioinformatics 27:1009–1010
Thorsen L, Azokpota P, Hansen BM, Hounhouigan DJ, Jakobsen M (2010) Identification, genetic diversity and cereulide producing ability of Bacillus cereus group strains isolated from Beninese traditional fermented food condiments. Int J Food Microbiol 142:247–250
Thorsen L, Azokpota P, Munk Hansen B, Rønsbo MH, Nielsen KF, Hounhouigan DJ, Jakobsen M (2011) Formation of cereulide and enterotoxins by Bacillus cereus in fermented African locust beans. Food Microbiol 28:1441–1447
Van der Auwera GA, Feldgarden M, Kolter R, Mahillon J (2013) Whole-genome sequences of 94 environmental isolates of Bacillus cereus sensu lato. Genome Announc 1:e00380-00313
Wong HC, Chang M, Fan J (1988) Incidence and characterization of Bacillus cereus isolates contaminating dairy products. Appl Environ Microbiol 54:699–702
Yee LM, Matsumoto T, Yano K, Matsuoka S, Sadaie Y, Yoshikawa H, Asai K (2011) The genome of Bacillus subtilis phage SP10: a comparative analysis with phage. Biosci Biotechnol Biochem 75:944–952
Acknowledgments
This research was supported by High Value-Added Food Technology Development Program (project No. 112108-2), Korea Institute of Planning and Evaluation for Technology in Food, Agriculture, Forestry and Fisheries (IPET) and the Ministry for Food, Agriculture, Forestry, and Fisheries of Republic of Korea, and by the “Cooperative Research Program for Agriculture & Technology Development (Project No. PJ009842)”, Rural Development Administration, Republic of Korea.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Asare, P.T., Ryu, S. & Kim, KP. Complete genome sequence and phylogenetic position of the Bacillus cereus group phage JBP901. Arch Virol 160, 2381–2384 (2015). https://doi.org/10.1007/s00705-015-2485-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00705-015-2485-0