Abstract
The 9573-nucleotide genome of a potyvirus was sequenced from a Coriandrum sativum plant from India with viral symptoms. On analysis, this virus was shown to have greater than 85 % nucleotide sequence identity to vanilla distortion mosaic virus (VDMV). Analysis of the putative coat protein sequence confirmed that this virus was in fact VDMV, with greater than 91 % amino acid sequence identity. The genome appears to encode a 3083-amino-acid polyprotein potentially cleaved into the 10 mature proteins expected in potyviruses. Phylogenetic analysis confirmed that VDMV is a distinct but ungrouped member of the genus Potyvirus.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
During a routine import inspection, unusual symptoms of chlorosis, bronzing and necrosis were observed spreading from affected petioles and stems on a consignment of Coriandrum sativum from India. In the laboratory, the sample tested positive by ELISA for potyvirus (antibody from AS-0573/1, DSMZ, Braunschweig, Germany) and negative for strawberry latent ringspot virus (antibody from Bioreba, Reinach, Switzerland), alfalfa mosaic virus (antibody from Bioreba, Switzerland) and cucumber mosaic virus (antibody from Agdia, Indiana, USA).
Total RNA was extracted from the sample using an RNeasy kit (QIAGEN, UK), and an indexed sequencing library was constructed using a ScriptSeq library preparation kit (EpoBio, USA) following the manufacturer’s instructions. The resulting library was sequenced on an Illumina MiSeq using a 500-cycle V2 sequencing kit (Illumina, USA). The resulting 5,056,797,250 nucleotide-paired end sequences were quality-trimmed to a phred score of 20 using SolexaQA [5] and then assembled using Trinity [7]. The resultant contigs were compared to the NCBI database using the BLASTx algorithm [3], and viral sequences were identified using MEGAN [8]. One large contig of 9573 nucleotides was identified as the genome of a potyvirus (accession number KF906523) with >85 % nucleotide sequence identity and >90 % protein sequence identity to sequences of vanilla distortion mosaic virus (VDMV) present in the GenBank database (accession numbers AY948436-37, AY943944-6 [host Vanilla planifolia] and AM261869 [host Stevia sp.]). Based on the species demarcation criteria for potyviruses [10], this suggests that the potyvirus identified in Indian coriander was in fact VDMV and confirms that it is a member of a distinct species within the genus Potyvirus. No published details are available on VDMV, with the exception of six partial sequences deposited in the GenBank database in 2006. Details within the GenBank submissions suggest that the virus was isolated in India on two separate occasions, initially from Vanilla planifolia and subsequently from Stevia sp. This is therefore the first published report describing details of the genome of VDMV, and the first description of the symptoms caused by this virus in coriander.
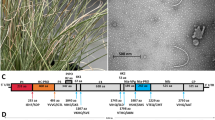
The VDMV genome investigated in this study consists of a 121-nucleotide 5′ UTR, a 181-nucleotide 3′ UTR ending in a polyadenosine tail, and a single putative open reading frame of 3083 aa encoding a 349-kDa polyprotein. The polyprotein contains the predicted cleavage sites typical of a potyvirus [1], yielding ten putative mature proteins (PI-Pro, HC-Pro, P3, 6k1, CI, 6k2, Vpg, NIa, Nib, CP), as shown in Fig. 1. The predicted mature proteins contain the expected potyvirus motifs: 210Hx8Dx33S253 and 274FIVRG278 in PI, 354KITC357 and 483FRNK486 in HC-Pro, 1247GSGKSx3P1255, 1264VLLxEPTRPL1273 and 1333DExH1336 in CI, 2090 Hx34Dx67GxCGx14H2211 in NIa, 2533CDADGS2538 and 2639GDD2641 in NIb, and DAGx in CP [12]. KITC and DAGx are known aphid transmission motifs [2], suggesting that this virus may be transmissible by aphids. As expected in a typical potyvirus, an 8-aa, 9.1-kDa Pipo coding region was found in the P3 region of the genome, starting with the expected G2A7 motif [4].
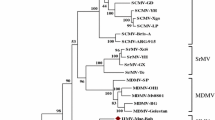
Multiple sequence alignment was carried out using MAFFT7 [9] on the VDMV genome sequence and 96 other complete potyvirus genome sequences. Based on this genome analysis, the most closely related viruses to VDMV are yam mosaic virus (accession no. NC_004752, 59 % nucleotide sequence identity) and lettuce mosaic virus (accession no. NC_003605, 55 % nucleotide sequence identity). These viruses have previously been shown to be related to each other but not grouped with other potyviruses [6]. Figure 2 shows a neighbour-joining tree constructed from coat protein amino acid sequences of VDMV and related potyviruses using Mega 5 [11] (note that the sequence of VDMV in GenBank isolated from Stevia sp. was not used, as it was too short). The tree shows that, based on CP amino acid sequence comparisons, the isolate of VDMV from coriander is indeed closely related to VDMV isolates from vanilla (>91 % amino acid sequence identity) and distinct from other viruses. The other most closely related viruses are ornamental onion stripe virus (accession no. ABU41870, 70 % amino acid sequence identity) and endive necrotic mosaic virus (accession no. CAA11564, 69 % amino acid sequence identity).
In summary, these results confirm that the virus sequenced from coriander was vanilla distortion mosaic virus and that this virus is a distinct but ungrouped potyvirus infecting coriander and vanilla in India.
References
Adams MJ, Antoniw JF, Beaudoin F (2005) Overview and analysis of the polyprotein cleavage sites in the family Potyviridae. Mol Plant Pathol 6:471–487
Atreya PL, Lopez-Moya JJ, Chu M, Atreya CD, Pirone TP (1995) Mutational analysis of the coat protein N-terminal amino acids involved in potyvirus transmission by aphids. J Gen Virol 76:265–270
Camacho C, Coulouris G, Avagyan V, Ma N, Papadopoulos J, Bealer K, Madden TL (2009) BLAST + : architecture and applications. BMC Bioinform 10:421
Chung BY-W, Miller WA, Atkins JF, Firth AE (2008) An overlapping essential gene in the Potyviridae. Proc Nat Acad Sci 105:5897–5902
Cox MP, Peterson DA, Biggs PJ (2010) SolexaQA: at-a-glance quality assessment of illumina second-generation sequencing data. BMC Bioinform 11:485
Gibbs A, Ohshima K (2010) Potyviruses and the digital revolution. Annu Rev Phytopathol 48:205–223
Grabherr MG, Haas BJ, Yassour M, Levin JZ, Thompson DA, Amit I, Adiconis X, Fan L, Raychowdhury R, Zeng Q, Chen Z, Mauceli E, Hacohen N, Gnirke A, Rhind N, di Palma F, Birren BW, Nusbaum C, Lindblad-Toh K, Friedman N, Regev A (2011) Full-length transcriptome assembly from RNA-Seq data without a reference genome. Nat Biotechnol 29:644–652
Huson DH, Auch AF, Qi J, Schuster SC (2007) MEGAN analysis of metagenomic data. Genome Res 17:377–386
Katoh K, Standley DM (2013) MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol Biol Evol 30:772–780
Adams MJ, Zerbini FM, French R, Rabenstein F, Stenger DC, Valkonen JPT (2011) Family Potyviridae. In: King E, Kowitz E, Adams M, Carstens (eds) Virus taxonomy: ninth report of the international committee on taxonomy of viruses. Elsevier, London
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S (2011) MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28:2731–2739
Urcuqui-Inchima S, Haenni A-L, Bernardi F (2001) Potyvirus proteins: a wealth of functions. Virus Res 74:157–175
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Adams, I.P., Rai, S., Deka, M. et al. Genome sequence of vanilla distortion mosaic virus infecting Coriandrum sativum . Arch Virol 159, 3463–3465 (2014). https://doi.org/10.1007/s00705-014-2215-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00705-014-2215-z