Abstract
Twenty-five years have passed since the causative gene for familial Parkinson's disease (PD), Parkin (now PRKN), was identified in 1998; PRKN is the most common causative gene in young-onset PD. Parkin encodes a ubiquitin-protein ligase, and Parkin is involved in mitophagy, a type of macroautophagy, in concert with PTEN-induced kinase 1 (PINK1). Both gene products are also involved in mitochondrial quality control. Among the many genetic PD-causing genes discovered, discovering PRKN as a cause of juvenile-onset PD has significantly impacted other neurodegenerative disorders. This is because the involvement of proteolytic systems has been suggested as a common mechanism in neurodegenerative diseases in which inclusion body formation is observed. The discovery of the participation of PRKN in PD has brought attention to the involvement of the proteolytic system in neurodegenerative diseases. Our research group has successfully isolated and identified CHCHD2, which is involved in the mitochondrial electron transfer system, and prosaposin (PSAP), which is involved in the lysosomal system, in this Parkin mechanism. Hereditary PD is undoubtedly an essential clue to solitary PD, and at least 25 or so genes and loci have been reported so far. This number of genes indicates that PD is a very diverse group of diseases. Currently, the diagnosis of PD is based on clinical symptoms and imaging studies. Although highly accurate diagnostic criteria have been published, early diagnosis is becoming increasingly important in treatment strategies for neurodegenerative diseases. Here, we also describe biomarkers that our group is working on.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The importance of hereditary Parkinson's disease (PD), a form of PD first reported in 1997, has attracted much attention. The α-synuclein was found to be a significant component of Lewy bodies, and it is believed that research on hereditary PD will lead to a better understanding of the pathogenesis of sporadic forms of PD, which account for the majority of the disease. In 1995, we were studying mitochondrial abnormalities and oxidative stress in the pathogenesis of PD. Our research found a family in which four siblings had familial PD with an autosomal recessive inheritance. The family had a polymorphism in the manganese superoxide dismutase gene. All affected individuals were homozygous for the alanine allele at position -9 (valine is wild-type), while the unaffected individuals were heterozygous for this polymorphism (Shimoda-Matsubayashi et al. 1997; Oji et al. 2020). Manganese-SOD is expressed in the mitochondrial matrix and is pivotal in controlling oxidative stress and mitochondrial abnormalities. We surmised that the causative gene in this form of familial PD might be located close to the Mn-SOD region, which had been mapped to the long arm of chromosome 6.
As we had only two families with autosomal recessive PD, we decided to collaborate with Prof. Shoji Tsuji, the chairman of the Department of Neurology at Niigata University at that time. He and his colleagues were studying 11 families with autosomal recessive familial PD with his colleagues. We performed linkage analyses on these families and found that the genetic locus for these families was located at 6q25.2–27 (Matsumine et al. 1997). Soon after, we found an autosomal recessive PD patient who lacked the D6S301 marker (Matsumine et al. 1998) among the highest logarithm of odds (LOD) scores in our linkage analysis. Subsequently, we identified the causative gene, PRKN, in 1998 based on our collaboration with Prof. Nobuyoshi Shimizu of Keio University. Since identifying the first causative gene, a-synuclein (SNCA), in 1997, the study of hereditary PD has accelerated dramatically. PRKN was the first causative gene for young-onset PD and is considered the most frequent causative gene so far (Table 1).
On the other hand, as the study of hereditary PD has progressed, the diversity of the disease has become apparent, and the need for early diagnosis and stratification has been advocated.
Discovery of PRKN that is responsible for young-onset PD (YOPD)
Clinical features of patients with PRKN mutations
The clinical features of autosomal recessive early-onset PD were first reported by Yamamura et al. (1973). They coined the term “early-onset parkinsonism with diurnal fluctuation” and described the clinical features as average age at onset of 26.1 years, parkinsonism with marked diurnal fluctuation, a remarkable effect of levodopa, dystonia, hyperreflexia, absence of dementia, and a benign course (Yamamura et al. 1998). Autonomic symptoms are only mild, if present at all. The patients developed dyskinesia soon after the initiation of levodopa therapy. Thus, since the discovery of PRKN, mutations within PRKN have been known to cause YOPD, particularly in patients who develop symptoms before age 40 (Schrag and Schott 2006). In addition, mutations in the parkin gene are a common cause of familial YOPD and sporadic cases with an age of onset < 40 years. Clinical features are very similar to those of dopa-responsive dystonia (DYT5) (Khan et al. 2003). An autopsy study of one of their patients by Takahashi et al. revealed neuronal loss without Lewy bodies and the presence of melanin-poor neurons in the substantia nigra (SN). They found a loss of neurons in the SN and the locus coeruleus, with the nigral loss much more significant than that in the locus coeruleus (Takahashi et al. 1994). We confirmed their autopsy findings in our patient with a linkage to 6q25.2–27 (Mori et al. 1998). We also confirmed that Yamamura’s cases mapped to the same locus (Matsumine et al. 1998).
Molecular cloning of PRKN and its mutations
We screened the candidate gene using the only complete human genomic DNA library that had been constructed by Asakawa et al. (Asakawa et al. 1997) at the Department of Molecular Biology at Keio University in Japan. We collaborated with Dr. Nobuyoshi Shimizu, the chairman of that department. We cloned a gene from Asakawa’s complete human bacterial artificial clones (BAC) DNA library using the marker D6S301. We isolated a complementary DNA clone of 2960 base pairs (bp) with a 1395-bp open reading frame, encoding a protein of 465 amino acids over 12 exons (Kitada et al. 1998). Dr. Shimizu named it ‘‘parkin’’ and is now known as PRKN. The gene product has a ubiquitin-like structure in the amino-terminal region, and two RING-finger motives in the carboxyl side are present. This was the second familial PD gene discovered, the first being PARK1/4, now known as SNCA (Polymeropoulos et al. 1997).
To determine the prevalence and genotype–phenotype correlation of the PRKN gene variants in PD, we screened 2,322 patients for enrollment, including 1,204 familial PD patients and 1,118 sporadic PD patients. We identified 137 familial PD patients and 105 sporadic PD patients, 242 of whom had PRKN mutations as possible susceptibility factors. We divided the cohort into two groups: monoallelic and biallelic mutation. Patients with the biallelic mutation had a significantly younger age of onset than those with the monoallelic mutation. The presence of the biallelic mutation reflected a higher frequency of normal [123I]metaiodobenzylguanidine myocardial scintigraphy. The log-rank test revealed exacerbations associated with the biallelic mutation over a 15-year course. Patients with the PRKN variant reported specific symptoms dependent on the number of mutant alleles (Yoshino et al. 2022) (Fig. 1). The allele frequencies of deletion and insertion mutations were significantly higher than those of point mutations (Yoshino et al. 2022; Hattori et al. 1998).
The Parkin and PINK1 genes are both responsible for autosomal recessive PD, typically caused by mutations in two alleles, though some cases have mutations in only one allele. The number of mutated alleles can influence the age of onset, with single allele mutations in either gene leading to onset at ages 45 or 48, slightly younger than those without the mutation. For mutations in two alleles of either gene, the age of onset is 29 for Parkin and 31 for PINK1. Cases with two mutated Parkin alleles and one mutated PINK1 allele have an even younger onset at 26. Genetic studies in Drosophila have shown that Parkin and PINK1 functionally cooperate in mitochondrial clearance
Recently, we aimed to identify complex structural variants in PRKN using long-read sequencing. Our research involved investigating the genetic cause of young onset dystonia-parkinsonism in monozygotic twins. A heterozygous exon 3 deletion in PRKN and a large novel inversion spanning over 7 Mb were identified. The study also analyzed whole genome sequencing data from the UK-Biobank and AMP-PD datasets, identifying several inversions likely to affect PRKN isoforms. The findings underscore the importance of using long-read whole genome sequencing for structural variant analysis in unresolved young-onset PD cases (Daida et al. 2023) (Fig. 2). This suggests the possibility that some cases of young-onset PD may be caused by inversions, as in this case. Therefore, it is believed that long-read sequencing is necessary for PRKN mutations, which have a high frequency of structural variants including copy number variants.
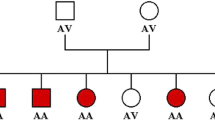
A Family pedigree of the family with PRKN inv/exon 3 deletion. Squares, men; circles, women; oblique lines, deceased; black squares, clinically diagnosed with PD. B MLPA result for PRKN exons and screenshot from IGV presenting with Exon 3 deletion in PRKN. C Screenshot from IGV presenting breakpoints of 7 M bp inversion including PRKN. Left figure represents the 5ʹ break point. The breakpoint is located on the intron 11 of PRKN. Right figure shows the 3ʹ breakpoint. Schematic representation of the inversion. Upper panel shows the overall picture of the inversion. Lower panel shows the sequence around both breakpoints. 5ʹ breakpoint Chr6:161,351,957, 3ʹ breakpoint Chr6:168,777,862. REF reference, INV inversion
Based on this structure, a new domain, RING0, consisting of two RING boxes (RING1 and 2) plus two different conserved cysteine-rich clusters between Cys150-Cys169 and Cys196-His215 consisting of CX2-3CX11CX2C and CX4-6CX10-16-CX2(H/C) motifs, was identified. Several mutations, such as K161N, S167N, M192V/L, S193I, K211R/N, and C212G/Y, have been found in the newly discovered RING0 domain (Hristova et al. 2009).
Hedrich et al. (2002) introduced a new technique to detect biallelic mutations in PRKN that used quantitative duplex PCR to detect alterations in gene dosage. They found that 14% of the 50 probands had biallelic mutations of PRKN, and 12% had monoallelic mutations. Mitsui et al. authors reported a mutation in the aphidicolin-induced common fragile sites (CFSs), like that seen in genes responsible for Duchenne and Becker muscular dystrophy (DMD/BMD) (Mitsui et al. 2010).
Biallelic variants in PRKN can cause PD, while monoallelic variants may increase PD risk through altered mitochondrial function. Castelo Rueda et al. studied cells from non-manifesting monoallelic PRKN variant carriers, finding hyperactive mitochondrial respiration and other signs of altered mitochondrial function. These findings could help identify individuals at risk of PD and test potential therapies before neurodegeneration progresses (Castelo Rueda et al. 2023).
Function of parkin
The product of this gene, E3 ubiquitin-protein ligase parkin, is involved in protein degradation as a ubiquitin-protein ligase, working with the enzyme UbcH7. Mutations in PRKN from PD patients result in a loss of this activity, leading to an accumulation of unidentified proteins and selective neural cell death. This discovery could aid in understanding neurodegeneration in not only PD and other diseases related to abnormal protein ubiquitination, such as AD, ALS, CAG triplet diseases, and tauopathies (Shimura et al. 2000). Moreover, the gene PRKN, which codes for the ubiquitin ligase parkin, is linked to early onset PD due to loss-of-function mutations. While the role of parkin in neuron maintenance is unclear, it has been associated with regulating mitochondria. Its absence leads to swollen mitochondria, muscle degeneration in Drosophila melanogaster, and mitochondrial dysfunction in other species. Narendra et al. (2008) show that in mammalian cells, parkin is selectively recruited to dysfunctional mitochondria with low membrane potential. It then facilitates autophagosomes' engulfment of these mitochondria, leading to the selective elimination of impaired mitochondria. This suggests that parkin promotes the autophagy of damaged mitochondria and that the inability to eliminate dysfunctional mitochondria may contribute to the development of PD. In addition, Matsuda et al. reveal that the genes PRKN and PINK1, linked to familial forms of PD, play a crucial role in disease pathogenesis through ubiquitylation and maintaining mitochondrial integrity (Matsuda et al. 2010). The exact mechanism of their interaction remains unclear. Our group shows that PINK1 is rapidly degraded in a mitochondrial membrane potential-dependent manner under steady conditions. Loss of mitochondrial membrane potential stabilizes PINK1. PINK1 phosphorylates and recruits parkin to mitochondria with low membrane potential to initiate autophagic degradation of damaged mitochondria (Shiba-Fukushima et al. 2012, 2014). Ubiquitin ligase activity of parkin is usually repressed but is activated upon PINK1-dependent mitochondrial localization. Some pathogenic mutations of PINK1 and PRKN disrupt these processes, indicating their importance in PD etiology. This provides significant insight into PD pathogenic mechanisms (Matsuda et al. 2010).
The discovery of ubiquitin enrichment in neurodegenerative disorders like PD and AD suggested the importance of the involvement of the proteolytic system. Ubiquitin marks proteins for elimination, leading to the belief that the ubiquitin–proteasome system (UPS) is inactive in these disorders, causing protein accumulation, aggregation into inclusion bodies, and neuronal death. The gene PRKN, linked to young-onset PD, was found to function as a ubiquitin ligase, further supporting this theory. Moreover, recent studies, including ours, show ubiquitylation relevance to the autophagy system, with parkin promoting autophagy of dysfunctional mitochondria after mitochondrial membrane potential loss.
Parkin is crucial for mitochondrial quality control in cells. PRKN mutations, which cause parkin dysfunction, lead to dopaminergic cell loss and decreased mitochondrial function due to impaired mitochondrial clearance. This dysfunction also causes mitochondrial injury and astrocytic dysfunction. Studies using immunohistochemical methods and patient-specific induced pluripotent stem cells showed fewer astrocytes in PRKN-mutated subjects and organoids (Kano et al. 2020). This suggests that PRKN mutations may cause astrocytic alterations and potentially non-autonomous cell death in dopaminergic neurons (Kano et al. 2020). PINK1, a mitochondrial kinase, phosphorylates ubiquitin and Parkin, another gene linked to parkinsonism (Shiba-Fukushima et al. 2012). PINK1 protein levels are typically low due to degradation after mitochondrial import but can accumulate on the mitochondrial membrane when the membrane potential is disrupted (Narendra et al. 2010). This leads to increased ubiquitin phosphorylation and parkin recruitment, targeting dysfunctional mitochondria for degradation. Despite the critical role of PINK1 in neuron maintenance, its activity in primary neurons is hard to detect. Evidence suggests that non-neuronal cells, like astrocytes and microglia, play a role in PD pathogenesis. Barodia et al. found that PINK1-dependent ubiquitin phosphorylation mainly in astrocytes, highlighting the need for further research on the function of PINK1 in astrocytes and their contribution to PD (Barodia et al. 2019).
Moreover, Cossu et al. investigated the role of the mitophagy-related gene PRKN in neuroinflammation using a mouse model of experimental autoimmune encephalomyelitis (EAE) (Cossu et al. 2021). Compared to wild-type mice, PRKN-/- mice showed earlier onset and greater severity of EAE, with increased T-cell activity, monocyte/macrophage recruitment, and microglia activation. In persistent disease, these mice also exhibited reduced glial cell numbers and abnormal mitochondrial morphology. The findings highlight the importance of parkin modulating immune responses during EAE and its potential in developing new neuroprotective therapies. The differences observed in the EAE model are presumed to indicate impairments in the immune system. Given the previously noted dysfunction in glial cells, the pathogenesis of hereditary PD by parkin may involve non-autonomous cell death. Neuroinflammation is a significant cause of developing PD.
The six factors potentially contributing to immune system dysfunction in PD include: (1) PD risk-associated genetic variants possibly affecting immune function; (2) Mitochondrial dysfunction linked to both immune system activation and neurodegeneration in PD; (3) The Braak hypothesis suggesting Lewy body pathology may originate in peripheral areas like the nose and gut; (4) Epidemiological evidence indicating a mix of factors, including infection, may increase PD risk during the prodromal phase; (5) Increased leakage of the blood–brain barrier and cerebrospinal fluid barrier in PD patients and animal models, suggesting altered peripheral immune responses and drug effects may influence disease progression; 6) Age-related immune system changes potentially increasing susceptibility to infection and age-related autoimmunity.
Discovery of CHCHD2 responsible for autosomal dominant PD
Our linkage analysis and whole genome sequencing of Japanese families with an autosomal dominant form of PD led to the discovery of mutations in coiled-coil-helix-coiled-coil-helix domain containing 2 (CHCHD2) (Funayama et al. 2015). The patients exhibited unique symptoms such as typical parkinsonism, middle-aged onset, good response to levodopa, largely benign course, and fewer complications of dysautonomia, psychosis, and cognitive decline. Subsequent studies reported the association of P2L and R8H in sporadic PD and V66M in a patient with multiple system atrophy (MSA) (Ikeda et al. 2017; Yang et al. 2016; Nicoletti et al. 2018). Additionally, several N-terminal variants were identified as a risk for PD and dementia with Lewy bodies (DLB) (Ogaki et al. 2015). At the same time, other studies found no evidence linking CHCHD2 to PD in both Asian and Caucasian populations, suggesting that CHCHD2 mutations are relatively rare in PD.
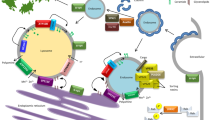
CHCHD2, found in the intermembrane spaces of the mitochondria, is a multifunctional protein that regulates mitochondrial respiration, transcriptional regulation of complex IV, and mitochondria-associated apoptosis. The loss of CHCHD2 destabilizes cytochrome c and complex IV, reducing oxygen consumption and complex IV activity in cultured mammalian cells (Fig. 3) (Meng et al. 2017). In Drosophila, the loss of CG5010, an ortholog of CHCHD2 and CHCHD10, led to high production of reactive oxygen species (ROS), degeneration of mitochondrial crista structure, and impairment of oxygen respiration, resulting in PD-like phenotypes such as DA neuronal loss and motor dysfunction with age.
Heterozygous mutations in PSAP saposin D domain in PD. A Genetic mutation screening identified three independent probands with autosomal dominant inherited PD with mutations in PSAP saposin D domain. B Mutations in PSAP saposin D domain can impair intracellular transport of PSAP and cause lysosomal dysfunction. Impairment of the delivery of saposins to lysosomes might be involved in the lysosomal dysfunction. Autophagy-lysosomal dysfunction can be related to α-synuclein aggregation
CHCHD10, which shares 54% amino acid sequence identity with CHCHD2, has been linked to symptoms consistent with amyotrophic lateral sclerosis (ALS) and frontotemporal dementia in individuals with the CHCHD10 S59L mutation (Bannwarth et al. 2014). This mutation causes mitochondrial respiratory chain deficiency and partial disassembly of complex VI and complex V. CHCHD2 forms a heterodimer with CHCHD10, contributing to efficient mitochondrial respiration and stress response, suggesting that these two proteins may play multiple roles in the pathogenesis of neurodegenerative diseases (Ikeda et al. 2022).
We found widespread α-synuclein pathology in an autopsy case with a CHCHD2 T61I mutation, a member of a large family with PD. DA neuron cultures prepared from CHCHD2 T61I-induced pluripotent stem cells (iPSCs) showed increased accumulation of detergent-resistant α-synuclein. Furthermore, Drosophila expressing CHCHD2 T61I in DA neurons replicated the acceleration of α-synuclein aggregation. Our results provide direct genetic evidence that mitochondrial dysfunction due to the CHCHD2 mutation could be a risk factor for α-synuclein aggregation and propagation in the pathogenesis of PD (Ikeda et al. 2019).
Previous reports suggest that mitochondrial dysfunction and accumulation of α-synuclein may play a vital role in the development of PD. However, the sequence of these events leading to neurodegeneration remains unclear. In this review, we review data supporting that mitochondrial dysfunction is preceded and indirectly induced by α-synuclein accumulation or that mitochondrial dysfunction initiates neuronal dysfunction and α-synuclein accumulation (Zaltieri et al. 2015). The exact temporal sequence of these events and their contribution to PD remains challenging.
Discovery of PSAP responsible for autosomal dominant PD
The importance of lysosomal storage disorders in the pathogenesis of PD arose from the discovery of the involvement of Gaucher disease in PD. Gaucher disease is an autosomal recessive lysosomal storage disorder caused by homozygous mutations in GBA1, whereas heterozygous mutations in GBA1 were reported to be the most common genetic risk factor for PD (Aflaki et al. 2017; Sidransky et al. 2009). The depletion of glucocerebrosidase, a product of GBA1, leads to the accumulation of α-synuclein, causing neurotoxicity through aggregation. For several lysosomal hydrolases, including glucocerebrosidase, to be fully active, the presence of sphingolipid activator proteins (saposins), which are degradation products of prosaposin (PSAP), is necessary.
The genetic variability in PSAP has been linked to PD. Three pathogenic mutations in the PSAP saposin D domain were found in families with autosomal dominant PD (Fig. 4A). Two variants in the intronic regions of the PSAP saposin D domain were found in sporadic PD in Japan and Taiwan. Abnormal accumulation of autophagic vacuoles, impaired autophagic flux, altered intracellular localization of prosaposin, and aggregation of α-synuclein were observed in patient-derived cells such as skin fibroblasts or induced pluripotent stem cell-derived dopaminergic neurons (Fig. 4B). In mice, a Psap saposin D mutation caused motor decline and dopaminergic neurodegeneration, providing new genetic evidence for the involvement of the PSAP saposin D domain in PD (Oji et al. 2020).
Altered PSAP levels were observed in the plasma, cerebrospinal fluid, and post-mortem brain of PD patients, and these changes correlated with motor impairments. PSAP-deficient mice showed symptoms like PD, including reduced movement and depression/anxiety-like symptoms. Lipidomic analysis revealed an accumulation of highly unsaturated and shortened lipids and a reduction of sphingolipids in the brains of these mice. Overexpression of PSAP protected against toxicity in wild-type rodents, suggesting that PSAP may maintain dopaminergic lipid homeostasis, which is disrupted in PD, and counteract experimental parkinsonism (He et al. 2023). PSAP's role in maintaining the homeostasis of dopaminergic neurons, particularly in lipid metabolism, is revealed through experiments with PSAP deletion. This deletion increases the susceptibility of mice to α-synuclein overexpression. However, Parkinsonism induced by α-synuclein can be mitigated by PSAP injection. PSAP also protects against oxidative stress and neurotoxicity induced by 6-OHDA. Furthermore, PSAP implantation shows therapeutic effects against Parkinsonism induced by α-synuclein. Therefore, PSAP could potentially modify the pathogenesis of PD, and its replenishment could help slow the progression of the disease (He et al. 2023).
Toward the era of biological biomarkers for PD
Disease staging, originating from cancer research, is crucial for categorizing individuals based on shared biomedical characteristics along a disease trajectory. This approach, defined by anatomical and biological features, has been practical in establishing prognosis and directing treatment protocols. There is growing interest in applying a similar system for PD. However, operationalizing a biological definition into a diagnosis or staging system requires measurability. This poses a challenge for assays involving α-synuclein, a protein whose function isn't fully understood and can aggregate in the brain. Current methods to measure α-synuclein require a cerebrospinal fluid (CSF) sample, which is challenging for most PD patients. However, the field is rapidly evolving with the potential development of blood-based α-synuclein seed amplification assays (SAA) and possibilities to document α-synuclein presence in other body fluids or tissues (Cardoso et al. 2023).
The SAA has finally laid the foundation for the biological diagnosis of PD, and it has made it possible to detect the disease even before any clinical or physical changes are detected. However, the collection of CSFs is more burdensome than conventional testing methods, and the development of blood tests and other less-than-burdensome methods may be necessary to spread the use of α-synuclein as a clue for diagnosing PD (Siderowf et al. 2023).
PD, DLB, and MSA comprise a group of neurodegenerative diseases called synucleinopathies. α-Syn seeds have been identified in various tissues in patients with synucleinopathies. The diseases are caused by abnormal aggregation of proteins called alpha-synuclein. These aggregates gradually appear in the patient's brain, causing neuronal cell death and producing different neurological symptoms depending on the part of the brain affected. Furthermore, differentiation of these diseases is difficult in the early stages. Therefore, early diagnosis is desirable because early treatment of these conditions can improve patients' quality of life. In synucleinopathies, it has been observed that α-synuclein aggregates are widely distributed in the central nervous system and spread to peripheral nerves. However, the mechanism of α-synuclein spread is still unclear. The spread of α-synuclein seeds cannot be explained by a single focus of nerve-mediated spread alone, as the evidence suggests that α-synuclein deposited in the skin may spread via the blood. Therefore, the multifocal theory assumes that the seed spreads by other pathways, including blood circulation.
Based on the findings, we have developed a method combining RT-QuIC and immunoprecipitation (IP/RT-QuIC) to enrich α-synuclein seeds (Okuzumi et al. 2023). This approach allows us to detect microscopic α-synuclein seeds in the serum of individuals with synucleinopathies with high sensitivity. Previous studies have shown that the structure of α-synuclein fibrils can vary between PD and MSA. Therefore, we examined the morphology of α-synuclein fibrils produced by serum seeds using transmission electron microscopy and a cell-based assay. Both techniques revealed morphological differences between α-synuclein fibrils from individuals with synucleinopathies (PD, DLB, or MSA), providing diagnostic solid power. We also found that the transmission of serum α-synuclein seeds from patients with PD or MSA differs. When injected directly into the striatum of mice, PD and MSA α-synuclein seeds spread at different rates, correlating with the progression of clinical symptoms in these diseases (Okuzumi et al. 2023). These results suggest that the amplified α-synuclein seeds retain their disease-specific morphological and transmission characteristics.
Looking ahead, detecting serum α-synuclein seeds by IP/RT-QuIC could be a valuable biomarker for diagnosing synucleinopathy. However, most participants in our study were clinically diagnosed without neuropathological confirmation, so the accuracy of our method needs further validation. Additionally, it remains unclear whether the presence of α-synuclein seeds in serum influences disease progression and pathogenic mechanisms. To address this, longitudinal monitoring of patients with REM sleep behavior disorder (RBD), early PD, DLB, and MSA is necessary. Notably, RBD can be a prodromal state to PD, DLB, or, less commonly, MSA. The fact that some RBD patients tested positive with IP/RT-QuIC suggests that our method could potentially detect the disease before clinical or physical changes become apparent. Therefore, our IP/RT-QuIC method could be beneficial for diagnosing synucleinopathies from blood samples (an easily accessible source of pathological α-synuclein seeds) and for understanding the pathogenic mechanisms of synucleinopathies. Furthermore, serum α-synuclein seeds could serve as a surrogate marker for disease-modifying therapies.
The landscape of PD diagnosis is being significantly transformed by the recent development of SAA and new sensitive assays for biomarker detection in biofluids. However, most studies are conducted post-diagnosis and are cross-sectional during the earliest stages of the disease. The reliability and stability of results, particularly for SAA assays, need to be verified through inter-laboratory exchange and cross-validation, as most studies are based on single laboratory experiences. The diverse clinical presentation of Alpha-synucleinopathy should be considered for at-risk populations and marker distribution in different tissues, which may highlight different pathophysiological mechanisms and disease subtypes. The combination of various techniques and markers is crucial to detect and differentiate markers of state and progression, which are particularly interesting as target engagement for disease-modifying treatments and strategies. The use of biological markers for a biological diagnosis of PD and synucleinopathies even before symptom development is a significant challenge for the research and clinical community, but it will undoubtedly transform the landscape in the coming decades.
Conclusions
The causative genes for hereditary PD have been isolated up to PARK25. Our group has successfully isolated and identified three of these causative genes. Among them, parkin is the most frequent causative gene in young onset PD, and there is a consensus that its function is mitophagy. It is necessary to consider the involvement of mitophagy not only in neurons but also in glial cells. The second gene we isolated and identified, CHCHD2, is localized in mitochondria and is known to be directly involved in the electron transport system. The binding of CHCHD2 with ALS-associated CHCHD10 suggests a common involvement of mitochondrial dysfunction in neurodegenerative diseases. The third gene is a cofactor of the lysosomal system, and its relevance to Gaucher disease is profound. The fact that these single genes can express the phenotype of PD implies that dopamine neuron loss occurs for multiple etiologies, and even a single cause can be multifaceted.
In the diversity, neuropathology can be divided mainly into two by the presence or absence of Lewy bodies. In our IP/RT-QuIC, all parkin mutation-positive cases showed negative of IP/RT-QuIC, and we believe that precision medicine will be realized by diagnosing with blood in the future. Undoubtedly, aiming to establish effective biomarkers and elucidate the cause of hereditary PD will be a breakthrough. The most important task is to develop from lab-based biomarkers to tests that can be included in insurance coverage.
Data availability
Not applicable.
References
Aflaki E, Westbroek W, Sidransky E (2017) The complicated relationship between Gaucher disease and parkinsonism: insights from a rare disease. Neuron 93(4):737–746. https://doi.org/10.1016/j.neuron.2017.01.018
Asakawa S, Abe I, Kudoh Y, Kishi N, Wang Y, Kubota R, Kudoh J, Kawasaki K, Minoshima S, Shimizu N (1997) Human BAC library: construction and rapid screening. Gene 191(1):69–79. https://doi.org/10.1016/s0378-1119(97)00044-9
Bannwarth S, Ait-El-Mkadem S, Chaussenot A, Genin EC, Lacas-Gervais S, Fragaki K, Berg-Alonso L, Kageyama Y, Serre V, Moore DG, Verschueren A, Rouzier C, Le Ber I, Auge G, Cochaud C, Lespinasse F, N’Guyen K, de Septenville A, Brice A, Yu-Wai-Man P, Sesaki H, Pouget J, Paquis-Flucklinger V (2014) A mitochondrial origin for frontotemporal dementia and amyotrophic lateral sclerosis through CHCHD10 involvement. Brain 137(Pt 8):2329–2345. https://doi.org/10.1093/brain/awu138
Barodia SK, McMeekin LJ, Creed RB, Quinones EK, Cowell RM, Goldberg MS (2019) PINK1 phosphorylates ubiquitin predominantly in astrocytes. NPJ Parkinsons Dis 5:29. https://doi.org/10.1038/s41531-019-0101-9
Cardoso F, Goetz CG, Mestre TA, Sampaio C, Adler CH, Berg D, Bloem BR, Burn DJ, Fitts MS, Gasser T, Klein C, de Tijssen MAJ, Lang AE, Lim SY, Litvan I, Meissner WG, Mollenhauer B, Okubadejo N, Okun MS, Postuma RB, Svenningsson P, Tan LCS, Tsunemi T, Wahlstrom-Helgren S, Gershanik OS, Fung VSC, Trenkwalder C (2023) A statement of the MDS on biological definition, staging, and classification of Parkinson’s disease. Mov Disord. https://doi.org/10.1002/mds.29683
Castelo Rueda MP, Zanon A, Gilmozzi V, Lavdas AA, Raftopoulou A, Delcambre S, Del Greco MF, Klein C, Grunewald A, Pramstaller PP, Hicks AA, Pichler I (2023) Molecular phenotypes of mitochondrial dysfunction in clinically non-manifesting heterozygous PRKN variant carriers. NPJ Parkinsons Dis 9(1):65. https://doi.org/10.1038/s41531-023-00499-9
Cossu D, Yokoyama K, Sato S, Noda S, Sechi LA, Hattori N (2021) PARKIN modifies peripheral immune response and increases neuroinflammation in active experimental autoimmune encephalomyelitis (EAE). J Neuroimmunol 359:577694. https://doi.org/10.1016/j.jneuroim.2021.577694
Daida K, Funayama M, Billingsley KJ, Malik L, Miano-Burkhardt A, Leonard HL, Makarious MB, Iwaki H, Ding J, Gibbs JR, Ishiguro M, Yoshino H, Ogaki K, Oyama G, Nishioka K, Nonaka R, Akamatsu W, Blauwendraat C, Hattori N (2023) Long-read sequencing resolves a complex structural variant in PRKN Parkinson’s disease. Mov Disord 38(12):2249–2257. https://doi.org/10.1002/mds.29610
Funayama M, Ohe K, Amo T, Furuya N, Yamaguchi J, Saiki S, Li Y, Ogaki K, Ando M, Yoshino H, Tomiyama H, Nishioka K, Hasegawa K, Saiki H, Satake W, Mogushi K, Sasaki R, Kokubo Y, Kuzuhara S, Toda T, Mizuno Y, Uchiyama Y, Ohno K, Hattori N (2015) CHCHD2 mutations in autosomal dominant late-onset Parkinson’s disease: a genome-wide linkage and sequencing study. Lancet Neurol 14(3):274–282. https://doi.org/10.1016/S1474-4422(14)70266-2
Hattori N, Kitada T, Matsumine H, Asakawa S, Yamamura Y, Yoshino H, Kobayashi T, Yokochi M, Wang M, Yoritaka A, Kondo T, Kuzuhara S, Nakamura S, Shimizu N, Mizuno Y (1998) Molecular genetic analysis of a novel Parkin gene in Japanese families with autosomal recessive juvenile parkinsonism: evidence for variable homozygous deletions in the Parkin gene in affected individuals. Ann Neurol 44(6):935–941. https://doi.org/10.1002/ana.410440612
He Y, Kaya I, Shariatgorji R, Lundkvist J, Wahlberg LU, Nilsson A, Mamula D, Kehr J, Zareba-Paslawska J, Biverstal H, Chergui K, Zhang X, Andren PE, Svenningsson P (2023) Prosaposin maintains lipid homeostasis in dopamine neurons and counteracts experimental parkinsonism in rodents. Nat Commun 14(1):5804. https://doi.org/10.1038/s41467-023-41539-5
Hedrich K, Marder K, Harris J, Kann M, Lynch T, Meija-Santana H, Pramstaller PP, Schwinger E, Bressman SB, Fahn S, Klein C (2002) Evaluation of 50 probands with early-onset Parkinson’s disease for Parkin mutations. Neurology 58(8):1239–1246. https://doi.org/10.1212/wnl.58.8.1239
Hristova VA, Beasley SA, Rylett RJ, Shaw GS (2009) Identification of a novel Zn2+-binding domain in the autosomal recessive juvenile Parkinson-related E3 ligase parkin. J Biol Chem 284(22):14978–14986. https://doi.org/10.1074/jbc.M808700200
Ikeda A, Matsushima T, Daida K, Nakajima S, Conedera S, Li Y, Yoshino H, Oyama G, Funayama M, Nishioka K, Hattori N (2017) A novel mutation of CHCHD2 p. R8H in a sporadic case of Parkinson’s disease. Parkinsonism Relat Disord 34:66–68. https://doi.org/10.1016/j.parkreldis.2016.10.018
Ikeda A, Nishioka K, Meng H, Takanashi M, Hasegawa I, Inoshita T, Shiba-Fukushima K, Li Y, Yoshino H, Mori A, Okuzumi A, Yamaguchi A, Nonaka R, Izawa N, Ishikawa KI, Saiki H, Morita M, Hasegawa M, Hasegawa K, Elahi M, Funayama M, Okano H, Akamatsu W, Imai Y, Hattori N (2019) Mutations in CHCHD2 cause alpha-synuclein aggregation. Hum Mol Genet 28(23):3895–3911. https://doi.org/10.1093/hmg/ddz241
Ikeda A, Imai Y, Hattori N (2022) Neurodegeneration-associated mitochondrial proteins, CHCHD2 and CHCHD10-what distinguishes the two? Front Cell Dev Biol 10:996061. https://doi.org/10.3389/fcell.2022.996061
Kano M, Takanashi M, Oyama G, Yoritaka A, Hatano T, Shiba-Fukushima K, Nagai M, Nishiyama K, Hasegawa K, Inoshita T, Ishikawa KI, Akamatsu W, Imai Y, Bolognin S, Schwamborn JC, Hattori N (2020) Reduced astrocytic reactivity in human brains and midbrain organoids with PRKN mutations. NPJ Parkinsons Dis 6(1):33. https://doi.org/10.1038/s41531-020-00137-8
Khan NL, Graham E, Critchley P, Schrag AE, Wood NW, Lees AJ, Bhatia KP, Quinn N (2003) Parkin disease: a phenotypic study of a large case series. Brain 126(Pt 6):1279–1292. https://doi.org/10.1093/brain/awg142
Kitada T, Asakawa S, Hattori N, Matsumine H, Yamamura Y, Minoshima S, Yokochi M, Mizuno Y, Shimizu N (1998) Mutations in the parkin gene cause autosomal recessive juvenile parkinsonism. Nature 392(6676):605–608. https://doi.org/10.1038/33416
Matsuda N, Sato S, Shiba K, Okatsu K, Saisho K, Gautier CA, Sou YS, Saiki S, Kawajiri S, Sato F, Kimura M, Komatsu M, Hattori N, Tanaka K (2010) PINK1 stabilized by mitochondrial depolarization recruits Parkin to damaged mitochondria and activates latent Parkin for mitophagy. J Cell Biol 189(2):211–221. https://doi.org/10.1083/jcb.200910140
Matsumine H, Saito M, Shimoda-Matsubayashi S, Tanaka H, Ishikawa A, Nakagawa-Hattori Y, Yokochi M, Kobayashi T, Igarashi S, Takano H, Sanpei K, Koike R, Mori H, Kondo T, Mizutani Y, Schaffer AA, Yamamura Y, Nakamura S, Kuzuhara S, Tsuji S, Mizuno Y (1997) Localization of a gene for an autosomal recessive form of juvenile Parkinsonism to chromosome 6q25.2–27. Am J Hum Genet 60(3):588–596
Matsumine H, Yamamura Y, Hattori N, Kobayashi T, Kitada T, Yoritaka A, Mizuno Y (1998) A microdeletion of D6S305 in a family of autosomal recessive juvenile parkinsonism (PARK2). Genomics 49(1):143–146. https://doi.org/10.1006/geno.1997.5196
Meng H, Yamashita C, Shiba-Fukushima K, Inoshita T, Funayama M, Sato S, Hatta T, Natsume T, Umitsu M, Takagi J, Imai Y, Hattori N (2017) Loss of Parkinson’s disease-associated protein CHCHD2 affects mitochondrial crista structure and destabilizes cytochrome c. Nat Commun 8:15500. https://doi.org/10.1038/ncomms15500
Mitsui J, Takahashi Y, Goto J, Tomiyama H, Ishikawa S, Yoshino H, Minami N, Smith DI, Lesage S, Aburatani H, Nishino I, Brice A, Hattori N, Tsuji S (2010) Mechanisms of genomic instabilities underlying two common fragile-site-associated loci, PARK2 and DMD, in germ cell and cancer cell lines. Am J Hum Genet 87(1):75–89. https://doi.org/10.1016/j.ajhg.2010.06.006
Mori H, Kondo T, Yokochi M, Matsumine H, Nakagawa-Hattori Y, Miyake T, Suda K, Mizuno Y (1998) Pathologic and biochemical studies of juvenile parkinsonism linked to chromosome 6q. Neurology 51(3):890–892. https://doi.org/10.1212/wnl.51.3.890
Narendra D, Tanaka A, Suen DF, Youle RJ (2008) Parkin is recruited selectively to impaired mitochondria and promotes their autophagy. J Cell Biol 183(5):795–803. https://doi.org/10.1083/jcb.200809125
Narendra DP, Jin SM, Tanaka A, Suen DF, Gautier CA, Shen J, Cookson MR, Youle RJ (2010) PINK1 is selectively stabilized on impaired mitochondria to activate Parkin. PLoS Biol 8(1):e1000298. https://doi.org/10.1371/journal.pbio.1000298
Nicoletti G, Gagliardi M, Procopio R, Iannello G, Morelli M, Annesi G, Quattrone A (2018) A new CHCHD2 mutation identified in a southern Italy patient with multiple system atrophy. Parkinsonism Relat Disord 47:91–93. https://doi.org/10.1016/j.parkreldis.2017.12.005
Ogaki K, Koga S, Heckman MG, Fiesel FC, Ando M, Labbe C, Lorenzo-Betancor O, Moussaud-Lamodiere EL, Soto-Ortolaza AI, Walton RL, Strongosky AJ, Uitti RJ, McCarthy A, Lynch T, Siuda J, Opala G, Rudzinska M, Krygowska-Wajs A, Barcikowska M, Czyzewski K, Puschmann A, Nishioka K, Funayama M, Hattori N, Parisi JE, Petersen RC, Graff-Radford NR, Boeve BF, Springer W, Wszolek ZK, Dickson DW, Ross OA (2015) Mitochondrial targeting sequence variants of the CHCHD2 gene are a risk for Lewy body disorders. Neurology 85(23):2016–2025. https://doi.org/10.1212/WNL.0000000000002170
Oji Y, Hatano T, Ueno SI, Funayama M, Ishikawa KI, Okuzumi A, Noda S, Sato S, Satake W, Toda T, Li Y, Hino-Takai T, Kakuta S, Tsunemi T, Yoshino H, Nishioka K, Hattori T, Mizutani Y, Mutoh T, Yokochi F, Ichinose Y, Koh K, Shindo K, Takiyama Y, Hamaguchi T, Yamada M, Farrer MJ, Uchiyama Y, Akamatsu W, Wu YR, Matsuda J, Hattori N (2020) Variants in saposin D domain of prosaposin gene linked to Parkinson’s disease. Brain 143(4):1190–1205. https://doi.org/10.1093/brain/awaa064
Okuzumi A, Hatano T, Matsumoto G, Nojiri S, Ueno SI, Imamichi-Tatano Y, Kimura H, Kakuta S, Kondo A, Fukuhara T, Li Y, Funayama M, Saiki S, Taniguchi D, Tsunemi T, McIntyre D, Gerardy JJ, Mittelbronn M, Kruger R, Uchiyama Y, Nukina N, Hattori N (2023) Propagative alpha-synuclein seeds as serum biomarkers for synucleinopathies. Nat Med 29(6):1448–1455. https://doi.org/10.1038/s41591-023-02358-9
Polymeropoulos MH, Lavedan C, Leroy E, Ide SE, Dehejia A, Dutra A, Pike B, Root H, Rubenstein J, Boyer R, Stenroos ES, Chandrasekharappa S, Athanassiadou A, Papapetropoulos T, Johnson WG, Lazzarini AM, Duvoisin RC, Di Iorio G, Golbe LI, Nussbaum RL (1997) Mutation in the alpha-synuclein gene identified in families with Parkinson’s disease. Science 276(5321):2045–2047. https://doi.org/10.1126/science.276.5321.2045
Schrag A, Schott JM (2006) Epidemiological, clinical, and genetic characteristics of early-onset Parkinsonism. Lancet Neurol 5(4):355–363. https://doi.org/10.1016/S1474-4422(06)70411-2
Shiba-Fukushima K, Imai Y, Yoshida S, Ishihama Y, Kanao T, Sato S, Hattori N (2012) PINK1-mediated phosphorylation of the Parkin ubiquitin-like domain primes mitochondrial translocation of Parkin and regulates mitophagy. Sci Rep 2:1002. https://doi.org/10.1038/srep01002
Shiba-Fukushima K, Arano T, Matsumoto G, Inoshita T, Yoshida S, Ishihama Y, Ryu KY, Nukina N, Hattori N, Imai Y (2014) Phosphorylation of mitochondrial polyubiquitin by PINK1 promotes Parkin mitochondrial tethering. PLoS Genet 10(12):e1004861. https://doi.org/10.1371/journal.pgen.1004861
Shimoda-Matsubayashi S, Hattori T, Matsumine H, Shinohara A, Yoritaka A, Mori H, Kondo T, Chiba M, Mizuno Y (1997) Mn SOD activity and protein in a patient with chromosome 6-linked autosomal recessive parkinsonism in comparison with Parkinson’s disease and control. Neurology 49(5):1257–1262. https://doi.org/10.1212/wnl.49.5.1257
Shimura H, Hattori N, Kubo S, Mizuno Y, Asakawa S, Minoshima S, Shimizu N, Iwai K, Chiba T, Tanaka K, Suzuki T (2000) Familial Parkinson disease gene product, parkin, is a ubiquitin-protein ligase. Nat Genet 25(3):302–305. https://doi.org/10.1038/77060
Siderowf A, Concha-Marambio L, Lafontant DE, Farris CM, Ma Y, Urenia PA, Nguyen H, Alcalay RN, Chahine LM, Foroud T, Galasko D, Kieburtz K, Merchant K, Mollenhauer B, Poston KL, Seibyl J, Simuni T, Tanner CM, Weintraub D, Videnovic A, Choi SH, Kurth R, Caspell-Garcia C, Coffey CS, Frasier M, Oliveira LMA, Hutten SJ, Sherer T, Marek K, Soto C, Parkinson’s Progression Markers I (2023) Assessment of heterogeneity among participants in the Parkinson’s Progression Markers Initiative cohort using alpha-synuclein seed amplification: a cross-sectional study. Lancet Neurol 22(5):407–417. https://doi.org/10.1016/S1474-4422(23)00109-6
Sidransky E, Nalls MA, Aasly JO, Aharon-Peretz J, Annesi G, Barbosa ER, Bar-Shira A, Berg D, Bras J, Brice A, Chen CM, Clark LN, Condroyer C, De Marco EV, Durr A, Eblan MJ, Fahn S, Farrer MJ, Fung HC, Gan-Or Z, Gasser T, Gershoni-Baruch R, Giladi N, Griffith A, Gurevich T, Januario C, Kropp P, Lang AE, Lee-Chen GJ, Lesage S, Marder K, Mata IF, Mirelman A, Mitsui J, Mizuta I, Nicoletti G, Oliveira C, Ottman R, Orr-Urtreger A, Pereira LV, Quattrone A, Rogaeva E, Rolfs A, Rosenbaum H, Rozenberg R, Samii A, Samaddar T, Schulte C, Sharma M, Singleton A, Spitz M, Tan EK, Tayebi N, Toda T, Troiano AR, Tsuji S, Wittstock M, Wolfsberg TG, Wu YR, Zabetian CP, Zhao Y, Ziegler SG (2009) Multicenter analysis of glucocerebrosidase mutations in Parkinson’s disease. N Engl J Med 361(17):1651–1661. https://doi.org/10.1056/NEJMoa0901281
Takahashi H, Ohama E, Suzuki S, Horikawa Y, Ishikawa A, Morita T, Tsuji S, Ikuta F (1994) Familial juvenile parkinsonism: clinical and pathologic study in a family. Neurology 44(3 Pt 1):437–441. https://doi.org/10.1212/wnl.44.3_part_1.437
Yamamura Y, Sobue I, Ando K, Iida M, Yanagi T (1973) Paralysis agitans of early onset with marked diurnal fluctuation of symptoms. Neurology 23(3):239–244
Yamamura Y, Kuzuhara S, Kondo K, Yanagi T, Uchida M, Matsumine H, Mizuno Y (1998) Clinical, pathologic and genetic studies on autosomal recessive early-onset Parkinsonism with diurnal fluctuation. Parkinsonism Relat Disord 4(2):65–72. https://doi.org/10.1016/s1353-8020(98)00015-7
Yang X, Zhao Q, An R, Zheng J, Tian S, Chen Y, Xu Y (2016) Mutational scanning of the CHCHD2 gene in Han Chinese patients with Parkinson’s disease and meta-analysis of the literature. Parkinsonism Relat Disord 29:42–46. https://doi.org/10.1016/j.parkreldis.2016.05.032
Yoshino H, Li Y, Nishioka K, Daida K, Hayashida A, Ishiguro Y, Yamada D, Izawa N, Nishi K, Nishikawa N, Oyama G, Hatano T, Nakamura S, Yoritaka A, Motoi Y, Funayama M, Hattori N, investigators of Japan Parkinson disease genetic s (2022) Genotype-phenotype correlation of Parkinson’s disease with PRKN variants. Neurobiol Aging 114:117–128. https://doi.org/10.1016/j.neurobiolaging.2021.12.014
Zaltieri M, Longhena F, Pizzi M, Missale C, Spano P, Bellucci A (2015) Mitochondrial dysfunction and alpha-synuclein synaptic pathology in Parkinson’s disease: who’s on first? Parkinsons Dis 2015:108029. https://doi.org/10.1155/2015/108029
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Hattori, N., Funayama, M., Imai, Y. et al. Pathogenesis of Parkinson’s disease: from hints from monogenic familial PD to biomarkers. J Neural Transm 131, 709–719 (2024). https://doi.org/10.1007/s00702-024-02747-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00702-024-02747-5