Abstract
Lithium’s inhibitory effect on enzymes involved in sulfation process, such as inhibition of 3’(2’)-phosphoadenosine 5’-phosphate (PAP) phosphatase, is a possible mechanism of its therapeutic effect for bipolar disorder (BD). 3’-Phosphoadenosine 5’-phosphosulfate (PAPS) is translocated from cytosol to Golgi lumen by PAPS transporter 1 (PAPST1/SLC35B2), where it acts as a sulfa donor. Since SLC35B2 was previously recognized as a molecule that facilitates the release of D-serine, a co-agonist of N-methyl-D-aspartate type glutamate receptor, altered function of SLC35B2 might be associated with the pathophysiology of BD and schizophrenia (SCZ). We performed genetic association analyses of the SLC35B2 gene using Japanese cohorts with 366 BD cases and 370 controls and 2012 SCZ cases and 2170 controls. We then investigated expression of SLC35B2 mRNA in postmortem brains by QPCR using a Caucasian cohort with 33 BD and 34 SCZ cases and 34 controls and by in situ hybridization using a Caucasian cohort with 37 SCZ and 29 controls. We found significant associations between three SNPs (rs575034, rs1875324, and rs3832441) and BD, and significantly reduced SLC35B2 mRNA expression in postmortem dorsolateral prefrontal cortex (DLPFC) of BD. Moreover, we observed normalized SLC35B2 mRNA expression in BD subgroups who were medicated with lithium. While there was a significant association of SLC35B2 with SCZ (SNP rs2233437), its expression was not changed in SCZ. These findings indicate that SLC35B2 might be differentially involved in the pathophysiology of BD and SCZ by influencing the sulfation process and/or glutamate system in the central nervous system.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Accumulating evidence has shown the importance of the glutamate system in the pathophysiology of bipolar disorder (BD) (Kim et al. 2017) and schizophrenia (SCZ) (O’Donovan et al. 2017). Genetic studies have demonstrated associations of genes involved in glutamatergic neurotransmission with BD (Benedetti et al. 2018) and SCZ (Uezato et al. 2012, 2017). Moreover, expression of genes involved in the glutamate system is altered in postmortem brains of subjects with BD (Uezato et al. 2009; McCullumsmith et al. 2007; Beneyto et al. 2007; Dean et al. 2016) and SCZ (Funk et al. 2017; Uezato et al. 2009; Uno and Coyle 2019) in brain regions of interest such as the dorsolateral prefrontal cortex (DLPFC), anterior cingulate cortex (ACC), and temporal lobe (hippocampus).
As a co-agonist, D-serine modulates the activity of the N-methyl-D-aspartate (NMDA) type glutamate receptor in the presence of glutamate (Mothet et al. 2000). In our previous rat study, we identified a molecule that facilitates the release of D-serine from cells and designated it as DSM-1 (D-serine modulator-1) (Shimazu et al. 2006). Dsm-1 is a rat ortholog of human 3’-phosphoadenosine 5’-phosphosulfate (PAPS) transporter 1 (PAPST1)/solute carrier family 35 member B2 (SLC35B2). SLC35B2 was originally identified as a molecule that translocates PAPS from cytosol into the Golgi lumen where PAPS acts as a sulfa donor (Kamiyama et al. 2003). Since PAPS is one of the substrates of PAP phosphatase (EC 3.1.3.7), which is inhibited by lithium (Li+), PAP phosphatase might be a novel target of Li+ therapy for BD (Yenush et al. 2000). Moreover, PAP phosphatase also has an inositol-polyphosphate 1-phosphatase activity (López-Coronado et al. 1999). Thus, elucidating the sulfation process in BD may extend an understanding of the pathophysiology of BD and connect it to the inositol depletion hypothesis (Berridge 1985; Yu and Greenberg 2016).
The SLC35B2 gene resides in the chromosomal position 6p21.1, where multiple genetic studies have demonstrated linkage or associations with BD (Doyle et al. 2010) and SCZ (Bamne et al. 2012; Yue et al. 2011; Zhang et al. 2013; Schizophrenia Psychiatric Genome-Wide Association Study 2011), suggesting genetic involvement of SLC35B2. We hypothesized that SLC35B2 has a role in the pathophysiology of BD and SCZ, through its involvement in sulfation process and glutamate system. Here, we report the findings of the multiple experiments performed in the separate settings. In these experiments, we performed case–control genetic association analyses for single-nucleotide polymorphisms (SNPs) in the SLC35B2 in BD and SCZ. We then investigated mRNA expression for SLC35B2 in postmortem brain from patients with BD and SCZ.
Methods
Case–control genetic association analysis
Subjects
Informed consent was obtained from all participants. They were all Japanese. All patients were diagnosed by experienced psychiatrists using semi-structured interviews according to the Diagnostic and Statistical Manual of Mental Disorders, Fourth Edition (DSM-IV) (American Psychiatric Association 1994). For controls (CTL), experienced psychiatrists performed clinical interviews and confirmed that they did not have major mental disorders. We utilized one cohort for BD and three for SCZ, and each of them was studied in separate settings for the current study. The characteristics of each cohort are described as follows.
BD cohort
The subjects were collected in the Tokyo Medical and Dental University, Hamamatsu University School of Medicine, Shiga University of Medical Science Hospital, University of Tokyo Hospital, Laboratory for Molecular Dynamics of Mental Disorders and Laboratory for Molecular Psychiatry. These include 366 patients with BD (50.1 ± 13.4 years old, 181 males and 185 females) and 370 controls (50.6 ± 12.6 years old, 185 males and 185 females). CTL were selected from students, nurses, office workers, and doctors in participants’ institutes, and their acquaintances.
SCZ cohort A
The cohort consists of 281 SCZ (51.3 ± 11.6 years old, 136 males and 145 females) and 289 CTL (48.4 ± 12.4 years old, 123 males and 166 females) collected by the Laboratory for Molecular Psychiatry of RIKEN Brain Science Institute. CTL were recruited from hospital staff and their acquaintances. Participants were residing in the vicinity of RIKEN which is in the western part of Tokyo. Therefore, population stratification should be negligible.
SCZ cohort B
The cohort was collected in the Tokyo Medical and Dental University. These include 237 SCZ (41.0 ± 10.8 years old, 138 males and 99 females) and 233 CTL (42.8 ± 12.2 years old, 119 males and 114 females). CTL were recruited from hospital staff and their acquaintances. Participants were residing in the vicinity of the Tokyo Medical and Dental University which is in the eastern part of Tokyo. Therefore, population stratification should be negligible.
SCZ cohort C
The cohort consists of 2,012 unrelated SCZ (48.1 ± 14.4 years old, 1,111 males and 901 females) and 2,170 CTL (42.4 ± 14.2 years old, 889 males and 1,281 females) collected by the Laboratory for Molecular Psychiatry of RIKEN Brain Science Institute. All participants were recruited from the Hondo area. A genome-wide study of the Japanese population has shown that the Hondo forms a distinct population cluster that is clearly separated from Han-Chinese and another Japanese cluster, Ryukyu (Yamaguchi-Kabata et al. 2008). In addition, recruitment was mostly restricted to the Kanto district of the Hondo area, which includes Tokyo and its surrounding areas. Therefore, population stratification should be negligible. CTL subjects were recruited from hospital staff and volunteers who showed no present or past evidence of psychoses and no family history of mental illness within the second degree of relationship during brief interviews by expert psychiatrists.
SNP selection
SNPs were obtained from Japanese genotype data of the HapMap database (release 27) (http://hapmap.ncbi.nlm.nih.gov/index.html). We used Paul de Bakker's Tagger tag SNP selection algorithm as implemented in Haploview v4.2 (http://www.broadinstitute.org/scientific-community/science/programs/medical-and-population-genetics/haploview/haploview /) (Barrett et al. 2005) [correlation coefficient (r2) > 0.80 and minor allele frequency (MAF) > 0.1] to select the most informative SNPs (tag SNPs) in terms of linkage disequilibrium (LD), from SNPs located within the 10 kb up- and down-stream regions of the SLC35B2 gene. In addition to these tag SNPs, SNPs that might affect gene function but are not covered by HapMap database were obtained from NCBI dbSNP (http://www.ncbi.nlm.nih.gov/SNP/). We prepared two sets of SNPs according to their range of distributions (Narrow and wide range SNP sets) (Supplementary Table 1, Supplementary Fig. 1). The narrow range SNP set (Narrow SNPs set) was studied for the BD cohort and SCZ cohort A and B, while the wide range SNP set (Wide SNPs set) was studied for the SCZ cohort C (Table 1).
Genotyping
Genomic DNAs from all subjects were prepared from peripheral whole blood cells by the phenol extraction method or by the DNA Extraction Kit (Stratagene). We used TaqMan assay (Applied Biosystems) to genotype SNPs. For the TaqMan assay, primer/probes were designed using Assays-by-Design™ SNP genotyping systems (Applied Biosystems) and fluorescence was determined using the ABI 7900 Sequence Detection System and SDS v2.0 software (Applied Biosystems, Foster City, California).
Postmortem brain mRNA expression analysis
Brain regions and mRNA measurements
The DLPFC and ACC of SCZ are investigated by various strands of scientific methods, because cognitive deficits, a fundamental symptom of this disorder, reflect dysfunction of these brain regions. For example, changes in gamma oscillatory activity reflecting cognitive dysfunction are speculated to be a consequence of alteration in the glutamate-GABA neural circuitry of the DLPFC (Schoonover et al. 2020). Because the DLPFC comprises a corticolimbic circuitry which is speculated to be disturbed and affecting emotion regulation in BD (Kebets et al. 2021), it is an attractive brain region to investigate for the pathophysiology of this disorder (Jun et al. 2014). For these reasons, in conjunction with availability of postmortem brain samples, the DLPFC (of BD and SCZ) and ACC (of SCZ only) were selected for the current postmortem study.
The current study investigated the mRNA expression of SLC35B2 by real-time quantitative reverse transcriptase polymerase chain reaction (qRT-PCR) and in situ hybridization (ISH) on postmortem brain tissues from different brain banks.
Brain RNA and qRT-PCR
The RNA from the postmortem DLPFC of subjects comprising 33 BD and 34 SCZ cases and 34 CTL was donated by the Stanley Medical Research Institute (SMRI; http://www.stanleyresearch.org) (Torrey et al. 2000). Left hemisphere or right hemisphere was randomly allocated for each sample by the brain bank.
qRT-PCR analysis was performed using the ABI PRISM 7900 Sequence Detection System. TaqMan primer/probes for SLC35B2 (GenBank accession number NM_178148) and for glyceraldehyde-3-phosphate dehydrogenase (GAPDH; NM_002046), which served as the endogenous reference, were purchased (Assay-on-Demand) from Applied Biosystems. All reactions were performed in duplicate, according to the manufacturer’s protocol. A comparative threshold cycle (CT) method validation experiment was done to check whether the efficiencies of target and reference amplifications were approximately equal (the slope of the log input amount vs. ∆CT < 0.1). One sample was randomly chosen as the calibrator, and was amplified in each plate, to correct for experimental differences among consecutive PCR runs. SLC35B2 mRNA was normalized to the endogenous reference, and expressed relative to the calibrator as 2–∆∆CT (comparative CT method). The diagnoses of the subjects were masked, while the assays were performed.
Brain sections and in situ hybridization
The postmortem brain tissues of ACC were donated from the Mount Sinai Medical Center/Bronx VA Medical Center Department of Psychiatry Brain Bank (MSMC/Bronx) (Davidson et al. 1995). The cohort comprised 37 patients with SCZ and 29 elderly control subjects. No evidence for neurodegenerative changes or Alzheimer disease was found in any of the subjects (Purohit et al. 1998). The methodology of tissue preparation has been described elsewhere in detail (Bauer et al. 2010).
mRNA expression was measured by ISH using subclones that were generated by amplifying unique segments of SLC35B2 (region used for probe: 594–1089) from a human cDNA brain library (Human Adult Brain Unamplified cDNA Library, Edge Biosystems; Gaithersburg, MD) and Polymerase Chain Reaction (PCR). The methodology of ISH has been described elsewhere in detail (Uezato et al. 2009). After obtaining the final images, film background values were subtracted from gray-scale values of either gray or underlying white matter regions of each section and converted to optical density. Values for two sections per subject were averaged and used for statistical analysis.
Statistical analysis
For statistical analysis in genetic association study, we used the PLINK v1.07 program (Purcell et al. 2007) (http://pngu.mgh.harvard.edu/~purcell/plink/). For genotype distribution of each polymorphism, deviation from Hardy–Weinberg equilibrium was examined by the Chi-square test. Differences between the patients and the controls with respect to the allele frequencies and genotype distributions were assessed using Fisher’s exact test.
For statistical analysis in postmortem study, we used the IBM SPSS Statistics version 23. One-way analysis of variance (ANOVA) or chi-squared test was used to examine the difference in the distribution of demographic variables between the groups. Linear regression analysis was performed to test for associations between mRNA expression and age, brain pH, and postmortem interval (PMI). When significant associations with age, pH, or PMI were found, ANCOVA was utilized; otherwise, ANOVA was utilized. Post hoc analyses were performed with Dunnett’s test for qRT-PCR data and Tukey's HSD test for ISH data. α = 0.05 was used for significance. We did not apply correction for multiple testing for the current study.
Results
The summary of the findings of the genetic association and postmortem studies is shown in Table 1.
Association between SLC35B2 and BD
All SNP markers were within the Hardy–Weinberg Equilibrium (HWE) in control subjects, while SNP8 in BD subjects showed deviation from HWE. Genotype and allele frequencies of the SNP5 (rs575034), SNP6 (rs1875324), and SNP8 (rs3832441) were significantly different between BD and CTL (Table 2). The genotype frequency of the SNP6 remained significant even after Bonferroni correction (α = 0.0071).
Since our analyses showed associations in several regions of SLC35B2 gene, we performed further haplotype analyses. LD block structure was assessed for the combined data of BD and CTL (Supplementary Fig. 2). Due to SNP4 to SNP8 being in the same LD block, we performed haplotypic association analysis for these 5 SNPs. Haplotypic distributions were significantly different between BD and CTL (P = 0.008, Supplementary Table 2).
Association between SLC35B2 and SCZ
In the SCZ cohort A, all SNP markers were within the HWE in CTL and SCZ subjects. The case–control analysis of SLC35B2 showed no evidence of association with SCZ for allelic or genotypic distributions of the 7 SNPs (Supplementary Table 3a).
In the SCZ cohort B, all SNP markers were within the HWE in CTL, while SNP7 in SCZ subjects showed deviation from HWE. Allele frequencies of SNP4 (rs9394996) were significantly different between SCZ and CTL (P = 0.045, odds ratio (OR) [95% confidence interval (95% CI) = 0.71 (0.51–1.00)]), although not surviving after Bonferroni correction (α = 0.0071) (Supplementary Table 3b).
In the SCZ cohort C, all SNP markers were within the HWE in CTL and SCZ subjects. In genotypic and allelic test, SNP2 (rs2233437) showed a significant association (C allele is over-represented in the disease subjects; genotypic P = 0.005 and allelic P = 0.035; OR = 1.10 (1.01–1.20)) (Table 3). The genotypic P value survived the multiple testing using false discovery rate (P = 0.030).
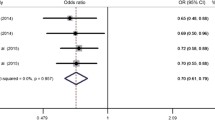
Because SNP4 and SNP9 are the only SNPs which are commonly examined for the SCZ cohorts A and B (Narrow SNPs set) and SCZ cohort C (Wide SNPs set) (Supplementary Table 1), we performed meta-analysis on these 2 SNPs. The meta-analysis yielded a significant odds ratio (Mantel–Haenszel OR = 0.89, 95% CI (0.81, 0.98)) for SNP4 (Table 1).
SLC35B2 mRNA expression in BD and SCZ measured by qRT-PCR
The demographic details of the subjects are shown in Supplementary Table 4. Among the BD group, 8 patients were medicated with Li+.
No significant difference in age distribution was observed between the three groups. However, there was a significant difference in gender distribution (P = 0.022, BD was different from other groups) and brain pH (P = 0.015, CTL < BD). Linear regression analysis revealed no significant association between SLC35B2 mRNA expression and age, PMI, or brain pH.
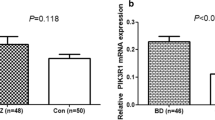
Using ANOVA, we detected a main effect for diagnosis (F(2, 98) = 3.10, P = 0.049). Dunnett’s post hoc test revealed that SLC35B2 mRNA expression is significantly decreased in DLPFC of BD (0.90 ± 0.35 (mean ± SD)) compared to CTL (1.12 ± 0.36) (P = 0.030) (Fig. 1a).
SLC35B2 mRNA expression measured by qRT-PCR. Box and whiskers plots showing the distribution of SLC35B2 mRNA expression measured by qRT-PCR in the three diagnostic groups (a) and BD subgroups divided according to the Li+ use. The boxes delineate the second and third quartiles, the horizontal lines in the boxes represent the medians, and the vertical bars (whiskers) show the extent of the data spread. The crosses indicate the means. SLC35B2, 3’-phosphoadenosine 5’-phosphosulfate transporter 1 (PAPST1); CTL control, BD bipolar disorder, SCZ schizophrenia, Li+ lithium. *P < 0.05
Because SLC35B2 mRNA expression could be influenced by Li+ use, we divided BD group into two subgroups according to Li+ status and re-analyzed the data BD vs. CTL. BD-Li+( +) (subjects who were on Li+ therapy) consisted of 8 patients and BD-Li+(–) (subjects who were not) consisted of 25 patients. There was no difference between BD-Li+(–) and BD-Li+( +) in age, PMI, or brain pH. ANOVA showed a main effect for Li+ group (F(2, 64) = 3.37, P = 0.040). Post hoc analysis revealed a significant reduction of SLC35B2 mRNA expression in BD-Li+(–) (0.89 ± 0.32) compared to CTL (1.12 ± 0.36) (P = 0.027). There was no significant difference in SLC35B2 mRNA expression between CTL and BD-Li( +) (0.95 ± 0.43) (p = 0.37) (Fig. 1b). For SCZ, three cases were on Li+ (SCZ-Li+( +)) and their mean SLC35B2 mRNA expression was 1.62 ± 0.28, while the one of SCZ cases without Li+ use (SCZ-Li+(–)) was 0.99 ± 0.37. Pearson correlation analysis revealed no correlation between SLC35B2 mRNA expression and lifetime antipsychotics dose (chlorpromazine equivalent) in BD or SCZ.
SLC35B2 mRNA expression in SCZ measured by ISH
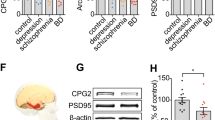
The pattern of expression for the probe was relatively uniform within the gray and underlying white matter, and no lamination was observed (Fig. 2a). Therefore, we analyzed the entire profile of either the gray or the white matter of each section for the transcript. We detected higher levels of SLC35B2 mRNA expression in the gray matter of the ACC, compared to those in the white matter.
SLC35B2 mRNA expression measured by ISH. Box and whiskers plots showing the distribution of SLC35B2 mRNA expression measured by ISH in the three diagnostic groups (a) and BD subgroups divided according to the Li+ use. The boxes delineate the second and third quartiles, the horizontal lines in the boxes represent the medians, and the vertical bars (whiskers) show the extent of the data spread. The crosses indicate the means. SLC35B2, 3’-phosphoadenosine 5’-phosphosulfate transporter 1 (PAPST1); ISH in situ hybridization, CTL control, SCZ schizophrenia, O.D. optical density
The demographic details of the subjects are shown in Supplementary Table 5. No significant difference in age was observed between the groups. However, there was a significant group difference in gender distribution (P = 0.004) and PMI (P = 0.003). Regression analysis revealed a significant association between SLC35B2 mRNA expression and pH in the gray matter of the ACC. ANCOVA with pH as a covariate revealed no significant difference in SLC35B2 mRNA expression in SCZ either in the gray matter (F(1, 63) = 0.336, P = 0.564) or in the white matter (F(1, 63) = 0.208, P = 0.650) (Fig. 2b, c).
Discussion
The remarkable findings of the current study were the significant allelic and genotypic associations of SLC35B2 gene with BD and the decreased SLC35B2 mRNA expression in the DLPFC of patients with BD. For SCZ, although there was a significant genotypic association with SLC35B2 gene, no change in SLC35B2 mRNA expression was observed in DLPFC or ACC (Table 1).
SNP5 resides in the intron 1. SNP6 is a silent mutation and SNP8 is located in 3’-UTR, both residing in the exon 4 (Supplementary Fig. 1, Supplementary Table 1). Although neither of these SNP changes the amino acid sequence of the SLC35B2 protein, there is increasing evidence that 3’-UTRs of mRNAs contain regulatory elements with important roles in post-transcriptional control of gene expression (Hesketh 2004). It is also reported that the polymorphisms in the intron may affect transcriptional activity (Tsukada et al. 2006). Although determining the role of the polymorphisms in these regions in SLC35B2 gene will require further biological experiments, these BD associated SNPs may have important roles in gene regulation. Otherwise, the functional SNPs in a regulatory region of the gene, which is in linkage disequilibrium with these BD associated SNPs, may contribute to the reduction of SLC35B2 mRNA expression as demonstrated in the present study. We used GTEx Portal (https://gtexportal.org/home/) to test if the SNPs in this study are expression quantitative trait locus (eQTL) by referencing the data source GTEx Analysis Release V8 (dbGaP Accession phs000424.v8.p2). No data were available for SNP4, 7, or 8. None of the other SNPs demonstrated significant effect on SLC35B2 expression in ACC or DLPFC. However, the SNP13 demonstrated a significant effect (p = 0.045, normalized effect size = − 0.14) in the tissue ‘Cortex’, suggesting possible involvement of this SNP in gene expression.
Our findings are not consistent with the recent genome-wide association study (GWAS) on European descent by the Bipolar Disorder Working Group of the Psychiatric Genomics Consortium which did not identify our gene locus as genome-wide significant (Stahl et al. 2019). This study also used MAGMA (Multi-marker Analysis of GenoMic Annotation) to conduct gene-wise GWAS and reported data for SNPs in SLC35B2. The gene-wide GWAS for SLC35B using 136 SNPs which included 8 SNPs of our study (SNP1, 2, 5, 6, 8, 10, 11, and 13) was non-significant (the joint p = 0.73, the highest p = 0.67). The discrepancy between the two studies might be attributed to ethnic difference.
SLC35B2 translocates PAPS from cytosol into the Golgi lumen where PAPS acts as a sulfa donor for sulfation (Fig. 3) (Kamiyama et al. 2003). Decreased SLC35B2 may result in the reduction of PAPS availability, leading to the interference of sulfation process. It follows that important physiological processes that might be impaired include deactivation and bioactivation of xenobiotics, inactivation of hormones and catecholamines, structure and function of macromolecules, and elimination of end products of catabolism (Klaassen and Boles 1997), leading to dysfunction of neuronal systems in BD. Consistent with this hypothesis, SLC35B2 is essential for sulfation-dependent cell growth during neural development and stem cell maintenance/differentiation (Bhattacharya et al. 2009; Sasaki et al. 2009).
Conceptual diagram of sulfation process involving SLC35B2. Li+, lithium; APS, adenosine 5’-phosphosulfate; PAPS, 3’-phosphoadenosine 5’-phosphosulfate; SULT, sulfotransferase; AMP, adenosine 5’-phosphate; PAPP, 3’-phosphoadenosine 5’-phosphate phosphatase; BPNT-1, bisphosphate 3’-nucleotidase; SLC35B2, 3’-phosphoadenosine 5’-phosphosulfate transporter 1 (PAPST1); IMPAD1, inositol monophosphatase domain containing 1. In cytosol, APS is phosphorylated by PAPSS to form PAPS. PAPS is degraded by two different pathways. In the first pathway, PAPS is desulfated by SULT, forming PAP, which is then dephosphorylated by PAPP to yield AMP. In the second pathway, PAPS is dephosphorylated by PAPP to yield APS. PAPS is translocated into Golgi lumen by SLC35B2. In Golgi lumen, PAPS is desulfated by gSULT, forming PAP, which is then dephosphorylated by gPAPP to yield AMP. The inhibition of PAPP by Li+ raises intracellular concentration of PAPS by reducing the dephosphorylation of PAPS to APS, or by the reduction of SULTs activity as a consequence of accumulated PAP. The diagram is
Despite the limitation due to the small sample size and lack of protein measurement, the finding that SLC35B2 mRNA expression in BD-Li+(-) is reduced (Fig. 1b) might represent a novel therapeutic effect of Li+ for BD. In the PAPS metabolism pathway conceptualized from studies in yeast (Klaassen and Boles 1997) and C. elegans (Meisel and Kim 2016) (Fig. 3), inhibition of PAP phosphatase by Li+ raises intracellular concentration of PAPS by reducing the dephosphorylation of PAPS to adenosine 5’-phosphosulfate (APS), or by the reduction of PAPS reductase (sulfotransferase, SULT) activity as a consequence of accumulated PAP (Murguía et al. 1996). Thus, PAPS availability would be ‘recovered’ to the normal level. Increased cytosolic PAPS then upregulates SLC35B2 mRNA, as is demonstrated in the present study in which the SLC35B2 mRNA level in BD-Li+( +) is apparently restored to the level of controls. Consistently, SCZ-Li+( +) demonstrated higher SLC35B2 mRNA level than SCZ-Li+(−) in our study, while evidence of beneficial clinical effects of Li+ and the role of sulfation in SCZ is limited (Luo et al. 2020).
An alternative view of the effect of Li+ on PAPS metabolism pathway is based on evidence that PAP accumulation has a toxic effect on RNA processing and PAPS-utilizing enzymes (López-Coronado et al. 1999). mRNA stabilization and enzyme inhibition might be related to the normalization of overactive neurons in BD as therapeutic effects. This hypothesis is inconsistent with the findings in the present study in which SLC35B2 mRNA expression is reduced in BD, which should reduce PAPS availability and further decelerate sulfation process. However, it is consistent with the other studies which demonstrated that several Li+-related biochemical measures in BD were altered in the direction of their response to Li+ treatment. For example, PAP phosphatase protein was reduced in the frontal cortex of patients with BD, although Li+ inhibits PAP phosphatase (Shaltiel et al. 2002). Likewise, inositol was reduced in frontal cortex of patients with BD, although Li+ inhibits IMPase (Shimon et al. 1997).
Regardless of its specific direction of change, it is likely that the effect of Li+ on PAP phosphatase plays an important role in sulfation process in the nervous system, as a recent study demonstrated that loss of the Li+-sensitive PAP phosphatase, bisphosphate 3’-nucleotidase (BPNT-1), or inhibition of BPNT-1 by Li+ causes selective toxicity to specific neurons, resulting in corresponding effects on behavior in the simple animal C. elegans (Meisel and Kim 2016).
Accumulating evidence suggests the involvement of the glutamatergic system in the etiology and treatment of BD (Henter et al. 2021). The glutamatergic system also appears to be associated with treatment response to Li+ in BD (Vecera et al. 2021). In addition to the role of SLC35B2 as a PAPS transporter, SLC35B2 is involved in the transport of D-serine presumably from the Golgi apparatus to a certain cytosolic site for extracellular release in glia or neurons (Shimazu et al. 2006). The decrease in the expression of SLC35B2 mRNA in BD demonstrated in the current study may result in the reduction of D-serine release leading to dysfunction of NMDAR in this disorder. In this context, unaltered SLC35B2 mRNA expression in SCZ was somewhat unexpected, since the NMDAR dysfunction is well established pathophysiology of SCZ (O'Donovan et al. 2017; McCullumsmith et al. 2004). This might be due to the methodological limitation of the current study which measured mRNA at the region level, while cell-level alterations of expression have been reported in SCZ (McCullumsmith et al. 2016). Alternatively, the findings that there was a significant genotypic association between SLC35B2 gene and SCZ, while its mRNA expression was not changed which may suggest that the function, not expression itself, of SLC35B2 is altered in SCZ.
In summary, while the current study is an exploratory study and no final conclusions can be drawn, we demonstrated genetic and expressional changes in the SLC35B2 gene, which indicate the involvement of sulfation process and/or glutamate system in the pathophysiology of BD and SCZ. Particularly, the upregulation of SLC35B2 gene by Li+ might be associated with therapeutic effect of Li+ on PAP phosphatase, which is involved in sulfation and inositol pathways. Because of the low number of lithium-treated patients in the postmortem study, results should be replicated in an independent sample.
Availability of data and materials
The data that support the findings of this study are available from the corresponding author upon reasonable request.
Code availability
Not applicable.
References
Bamne M, Wood J, Chowdari K, Watson AM, Celik C, Mansour H, Klei L, Gur RC, Bradford LD, Calkins ME, Santos AB, Edwards N, Kwentus J, McEvoy JP, Allen TB, Savage RM, Nasrallah HA, Gur RE, Perry RT, Go RC, Devlin B, Yolken R, Nimgaonkar VL (2012) Evaluation of HLA polymorphisms in relation to schizophrenia risk and infectious exposure. Schizophr Bull 38(6):1149–1154. https://doi.org/10.1093/schbul/sbs087
Barrett JC, Fry B, Maller J, Daly MJ (2005) Haploview: analysis and visualization of LD and haplotype maps. Bioinformatics 21(2):263–265. https://doi.org/10.1093/bioinformatics/bth457
Bauer D, Haroutunian V, Meador-Woodruff JH, McCullumsmith RE (2010) Abnormal glycosylation of EAAT1 and EAAT2 in prefrontal cortex of elderly patients with schizophrenia. Schizophr Res 117(1):92–98. https://doi.org/10.1016/j.schres.2009.07.025
Benedetti F, Poletti S, Locatelli C, Mazza E, Lorenzi C, Vitali A, Riberto M, Brioschi S, Vai B, Bollettini I, Melloni E, Aggio V, Falini A, De Bartolomeis A, Colombo C (2018) A homer 1 gene variant influences brain structure and function, lithium effects on white matter, and antidepressant response in bipolar disorder: a multimodal genetic imaging study. Prog Neuropsychopharmacol Biol Psychiatry 81:88–95. https://doi.org/10.1016/j.pnpbp.2017.10.011
Beneyto M, Kristiansen LV, Oni-Orisan A, McCullumsmith RE, Meador-Woodruff JH (2007) Abnormal glutamate receptor expression in the medial temporal lobe in schizophrenia and mood disorders. Neuropsychopharmacology 32(9):1888–1902. https://doi.org/10.1038/sj.npp.1301312
Berridge MJ (1985) Phosphoinositides and signal transductions. Rev Clin Basic Pharm 5(Suppl):5S-13S
Bhattacharya R, Townley RA, Berry KL, Bülow HE (2009) The PAPS transporter PST-1 is required for heparan sulfation and is essential for viability and neural development in C. elegans. J Cell Sci 122(Pt 24):4492–4504. https://doi.org/10.1242/jcs.050732
Davidson M, Harvey PD, Powchik P, Parrella M, White L, Knobler HY, Losonczy MF, Keefe RS, Katz S, Frecska E (1995) Severity of symptoms in chronically institutionalized geriatric schizophrenic patients. Am J Psychiatry 152(2):197–207
Dean B, Gibbons AS, Boer S, Uezato A, Meador-Woodruff J, Scarr E, McCullumsmith RE (2016) Changes in cortical N-methyl-D-aspartate receptors and post-synaptic density protein 95 in schizophrenia, mood disorders and suicide. Aust N Z J Psychiatry 50(3):275–283. https://doi.org/10.1177/0004867415586601
Doyle AE, Biederman J, Ferreira MA, Wong P, Smoller JW, Faraone SV (2010) Suggestive linkage of the child behavior checklist juvenile bipolar disorder phenotype to 1p21, 6p21, and 8q21. J Am Acad Child Adolesc Psychiatry 49(4):378–387
Funk AJ, Mielnik CA, Koene R, Newburn E, Ramsey AJ, Lipska BK, McCullumsmith RE (2017) Postsynaptic density-95 isoform abnormalities in schizophrenia. Schizophr Bull. https://doi.org/10.1093/schbul/sbw173
Henter ID, Park LT, Zarate CA Jr (2021) Novel glutamatergic modulators for the treatment of mood disorders: current status. CNS Drugs. https://doi.org/10.1007/s40263-021-00816-x
Hesketh J (2004) 3’-Untranslated regions are important in mRNA localization and translation: lessons from selenium and metallothionein. Biochem Soc Trans 32(Pt 6):990–993. https://doi.org/10.1042/BST0320990
Jun C, Choi Y, Lim SM, Bae S, Hong YS, Kim JE, Lyoo IK (2014) Disturbance of the glutamatergic system in mood disorders. Exp Neurobiol 23(1):28–35. https://doi.org/10.5607/en.2014.23.1.28
Kamiyama S, Suda T, Ueda R, Suzuki M, Okubo R, Kikuchi N, Chiba Y, Goto S, Toyoda H, Saigo K, Watanabe M, Narimatsu H, Jigami Y, Nishihara S (2003) Molecular cloning and identification of 3’-phosphoadenosine 5’-phosphosulfate transporter. J Biol Chem 278(28):25958–25963. https://doi.org/10.1074/jbc.M302439200
Kebets V, Favre P, Houenou J, Polosan M, Perroud N, Aubry JM, Van De Ville D, Piguet C (2021) Fronto-limbic neural variability as a transdiagnostic correlate of emotion dysregulation. Transl Psychiatry 11(1):545. https://doi.org/10.1038/s41398-021-01666-3
Kim Y, Santos R, Gage FH, Marchetto MC (2017) Molecular mechanisms of bipolar disorder: progress made and future challenges. Front Cell Neurosci 11:30. https://doi.org/10.3389/fncel.2017.00030
Klaassen CD, Boles JW (1997) Sulfation and sulfotransferases 5: the importance of 3’-phosphoadenosine 5’-phosphosulfate (PAPS) in the regulation of sulfation. FASEB J 11(6):404–418
López-Coronado JM, Bellés JM, Lesage F, Serrano R, Rodríguez PL (1999) A novel mammalian lithium-sensitive enzyme with a dual enzymatic activity, 3’-phosphoadenosine 5’-phosphate phosphatase and inositol-polyphosphate 1-phosphatase. J Biol Chem 274(23):16034–16039
Luo DZ, Chang CY, Huang TR, Studer V, Wang TW, Lai WS (2020) Lithium for schizophrenia: supporting evidence from a 12-year, nationwide health insurance database and from Akt1-deficient mouse and cellular models. Sci Rep 10(1):647. https://doi.org/10.1038/s41598-019-57340-8
McCullumsmith RE, Clinton SM, Meador-Woodruff JH (2004) Schizophrenia as a disorder of neuroplasticity. Int Rev Neurobiol 59:19–45. https://doi.org/10.1016/S0074-7742(04)59002-5
McCullumsmith RE, Kristiansen LV, Beneyto M, Scarr E, Dean B, Meador-Woodruff JH (2007) Decreased NR1, NR2A, and SAP102 transcript expression in the hippocampus in bipolar disorder. Brain Res 1127(1):108–118. https://doi.org/10.1016/j.brainres.2006.09.011
McCullumsmith RE, O’Donovan SM, Drummond JB, Benesh FS, Simmons M, Roberts R, Lauriat T, Haroutunian V, Meador-Woodruff JH (2016) Cell-specific abnormalities of glutamate transporters in schizophrenia: sick astrocytes and compensating relay neurons? Mol Psychiatry 21(6):823–830. https://doi.org/10.1038/mp.2015.148
Meisel JD, Kim DH (2016) Inhibition of lithium-sensitive phosphatase BPNT-1 causes selective neuronal dysfunction in C. elegans. Curr Biol 26(14):1922–1928. https://doi.org/10.1016/j.cub.2016.05.050
Mothet JP, Parent AT, Wolosker H, Brady RO Jr, Linden DJ, Ferris CD, Rogawski MA, Snyder SH (2000) D-serine is an endogenous ligand for the glycine site of the N-methyl-D-aspartate receptor. Proc Natl Acad Sci U S A 97(9):4926–4931
Murguía JR, Bellés JM, Serrano R (1996) The yeast HAL2 nucleotidase is an in vivo target of salt toxicity. J Biol Chem 271(46):29029–29033
O’Donovan SM, Sullivan CR, McCullumsmith RE (2017) The role of glutamate transporters in the pathophysiology of neuropsychiatric disorders. NPJ Schizophr 3(1):32. https://doi.org/10.1038/s41537-017-0037-1
Purcell S, Neale B, Todd-Brown K, Thomas L, Ferreira MA, Bender D, Maller J, Sklar P, de Bakker PI, Daly MJ, Sham PC (2007) PLINK: a tool set for whole-genome association and population-based linkage analyses. Am J Hum Genet 81(3):559–575. https://doi.org/10.1086/519795
Purohit DP, Perl DP, Haroutunian V, Powchik P, Davidson M, Davis KL (1998) Alzheimer disease and related neurodegenerative diseases in elderly patients with schizophrenia: a postmortem neuropathologic study of 100 cases. Arch Gen Psychiatry 55(3):205–211
Sasaki N, Hirano T, Ichimiya T, Wakao M, Hirano K, Kinoshita-Toyoda A, Toyoda H, Suda Y, Nishihara S (2009) The 3’-phosphoadenosine 5’-phosphosulfate transporters, PAPST1 and 2, contribute to the maintenance and differentiation of mouse embryonic stem cells. PLoS One 4(12):e8262. https://doi.org/10.1371/journal.pone.0008262
Schizophrenia Psychiatric Genome-Wide Association Study C (2011) Genome-wide association study identifies five new schizophrenia loci. Nat Genet 43(10):969–976. https://doi.org/10.1038/ng.940
Schoonover KE, Dienel SJ, Lewis DA (2020) Prefrontal cortical alterations of glutamate and GABA neurotransmission in schizophrenia: insights for rational biomarker development. Biomark Neuropsychiatr. https://doi.org/10.1016/j.bionps.2020.100015
Shaltiel G, Kozlovsky N, Belmaker RH, Agam G (2002) 3’(2’)-phosphoadenosine 5’-phosphate phosphatase is reduced in postmortem frontal cortex of bipolar patients. Bipolar Disord 4(5):302–306
Shimazu D, Yamamoto N, Umino A, Ishii S, Sakurai S, Nishikawa T (2006) Inhibition of D-serine accumulation in the Xenopus oocyte by expression of the rat ortholog of human 3’-phosphoadenosine 5’-phosphosulfate transporter gene isolated from the neocortex as D-serine modulator-1. J Neurochem 96(1):30–42
Shimon H, Agam G, Belmaker RH, Hyde TM, Kleinman JE (1997) Reduced frontal cortex inositol levels in postmortem brain of suicide victims and patients with bipolar disorder. Am J Psychiatry 154(8):1148–1150
Stahl EA, Breen G, Forstner AJ, McQuillin A, Ripke S, Trubetskoy V, Mattheisen M, Wang Y, Coleman JRI, Gaspar HA, de Leeuw CA, Steinberg S, Pavlides JMW, Trzaskowski M, Byrne EM, Pers TH, Holmans PA, Richards AL, Abbott L, Agerbo E, Akil H, Albani D, Alliey-Rodriguez N, Als TD, Anjorin A, Antilla V, Awasthi S, Badner JA, Baekvad-Hansen M, Barchas JD, Bass N, Bauer M, Belliveau R, Bergen SE, Pedersen CB, Boen E, Boks MP, Boocock J, Budde M, Bunney W, Burmeister M, Bybjerg-Grauholm J, Byerley W, Casas M, Cerrato F, Cervantes P, Chambert K, Charney AW, Chen D, Churchhouse C, Clarke TK, Coryell W, Craig DW, Cruceanu C, Curtis D, Czerski PM, Dale AM, de Jong S, Degenhardt F, Del-Favero J, DePaulo JR, Djurovic S, Dobbyn AL, Dumont A, Elvsashagen T, Escott-Price V, Fan CC, Fischer SB, Flickinger M, Foroud TM, Forty L, Frank J, Fraser C, Freimer NB, Frisen L, Gade K, Gage D, Garnham J, Giambartolomei C, Pedersen MG, Goldstein J, Gordon SD, Gordon-Smith K, Green EK, Green MJ, Greenwood TA, Grove J, Guan W, Guzman-Parra J, Hamshere ML, Hautzinger M, Heilbronner U, Herms S, Hipolito M, Hoffmann P, Holland D, Huckins L, Jamain S, Johnson JS, Jureus A, Kandaswamy R, Karlsson R, Kennedy JL, Kittel-Schneider S, Knowles JA, Kogevinas M, Koller AC, Kupka R, Lavebratt C, Lawrence J, Lawson WB, Leber M, Lee PH, Levy SE, Li JZ, Liu C, Lucae S, Maaser A, MacIntyre DJ, Mahon PB, Maier W, Martinsson L, McCarroll S, McGuffin P, McInnis MG, McKay JD, Medeiros H, Medland SE, Meng F, Milani L, Montgomery GW, Morris DW, Muhleisen TW, Mullins N, Nguyen H, Nievergelt CM, Adolfsson AN, Nwulia EA, O’Donovan C, Loohuis LMO, Ori APS, Oruc L, Osby U, Perlis RH, Perry A, Pfennig A, Potash JB, Purcell SM, Regeer EJ, Reif A, Reinbold CS, Rice JP, Rivas F, Rivera M, Roussos P, Ruderfer DM, Ryu E, Sanchez-Mora C, Schatzberg AF, Scheftner WA, Schork NJ, Shannon Weickert C, Shehktman T, Shilling PD, Sigurdsson E, Slaney C, Smeland OB, Sobell JL, Soholm Hansen C, Spijker AT, St Clair D, Steffens M, Strauss JS, Streit F, Strohmaier J, Szelinger S, Thompson RC, Thorgeirsson TE, Treutlein J, Vedder H, Wang W, Watson SJ, Weickert TW, Witt SH, Xi S, Xu W, Young AH, Zandi P, Zhang P, Zollner S, Adolfsson R, Agartz I, Alda M, Backlund L, Baune BT, Bellivier F, Berrettini WH, Biernacka JM, Blackwood DHR, Boehnke M, Borglum AD, Corvin A, Craddock N, Daly MJ, Dannlowski U, Esko T, Etain B, Frye M, Fullerton JM, Gershon ES, Gill M, Goes F, Grigoroiu-Serbanescu M, Hauser J, Hougaard DM, Hultman CM, Jones I, Jones LA, Kahn RS, Kirov G, Landen M, Leboyer M, Lewis CM, Li QS, Lissowska J, Martin NG, Mayoral F, McElroy SL, McIntosh AM, McMahon FJ, Melle I, Metspalu A, Mitchell PB, Morken G, Mors O, Mortensen PB, Muller-Myhsok B, Myers RM, Neale BM, Nimgaonkar V, Nordentoft M, Nothen MM, O’Donovan MC, Oedegaard KJ, Owen MJ, Paciga SA, Pato C, Pato MT, Posthuma D, Ramos-Quiroga JA, Ribases M, Rietschel M, Rouleau GA, Schalling M, Schofield PR, Schulze TG, Serretti A, Smoller JW, Stefansson H, Stefansson K, Stordal E, Sullivan PF, Turecki G, Vaaler AE, Vieta E, Vincent JB, Werge T, Nurnberger JI, Wray NR, Di Florio A, Edenberg HJ, Cichon S, Ophoff RA, Scott LJ, Andreassen OA, Kelsoe J, Sklar P, Bipolar Disorder Working Group of the Psychiatric Genomics C (2019) Genome-wide association study identifies 30 loci associated with bipolar disorder. Nat Genet 51(5):793–803. https://doi.org/10.1038/s41588-019-0397-8
Torrey EF, Webster M, Knable M, Johnston N, Yolken RH (2000) The stanley foundation brain collection and neuropathology consortium. Schizophr Res 44(2):151–155
Tsukada S, Tanaka Y, Maegawa H, Kashiwagi A, Kawamori R, Maeda S (2006) Intronic polymorphisms within TFAP2B regulate transcriptional activity and affect adipocytokine gene expression in differentiated adipocytes. Mol Endocrinol 20(5):1104–1111. https://doi.org/10.1210/me.2005-0311
Uezato A, Meador-Woodruff JH, McCullumsmith RE (2009) Vesicular glutamate transporter mRNA expression in the medial temporal lobe in major depressive disorder, bipolar disorder, and schizophrenia. Bipolar Disord 11(7):711–725. https://doi.org/10.1111/j.1399-5618.2009.00752.x
Uezato A, Kimura-Sato J, Yamamoto N, Iijima Y, Kunugi H, Nishikawa T (2012) Further evidence for a male-selective genetic association of synapse-associated protein 97 (SAP97) gene with schizophrenia. Behav Brain Funct 8(1):2. https://doi.org/10.1186/1744-9081-8-2
Uezato A, Yamamoto N, Jitoku D, Haramo E, Hiraaki E, Iwayama Y, Toyota T, Umino M, Umino A, Iwata Y, Suzuki K, Kikuchi M, Hashimoto T, Kanahara N, Kurumaji A, Yoshikawa T, Nishikawa T (2017) Genetic and molecular risk factors within the newly identified primate-specific exon of the SAP97/DLG1 gene in the 3q29 schizophrenia-associated locus. Am J Med Genet B Neuropsychiatr Genet. https://doi.org/10.1002/ajmg.b.32595
Uno Y, Coyle JT (2019) Glutamate hypothesis in schizophrenia. Psychiatry Clin Neurosci 73(5):204–215. https://doi.org/10.1111/pcn.12823
Vecera CM, Fries GR, Shahani LR, Soares JC, Machado-Vieira R (2021) Pharmacogenomics of lithium response in bipolar disorder. Pharmaceuticals (Basel). https://doi.org/10.3390/ph14040287
Yamaguchi-Kabata Y, Nakazono K, Takahashi A, Saito S, Hosono N, Kubo M, Nakamura Y, Kamatani N (2008) Japanese population structure, based on SNP genotypes from 7003 individuals compared to other ethnic groups: effects on population-based association studies. Am J Hum Genet 83(4):445–456. https://doi.org/10.1016/j.ajhg.2008.08.019
Yenush L, Bellés J, López-Coronado J, Gil-Mascarell R, Serrano R, Rodríguez P (2000) A novel target of lithium therapy. FEBS Lett 467(2–3):321–325
Yu W, Greenberg ML (2016) Inositol depletion, GSK3 inhibition and bipolar disorder. Future Neurol 11(2):135–148. https://doi.org/10.2217/fnl-2016-0003
Yue WH, Wang HF, Sun LD, Tang FL, Liu ZH, Zhang HX, Li WQ, Zhang YL, Zhang Y, Ma CC, Du B, Wang LF, Ren YQ, Yang YF, Hu XF, Wang Y, Deng W, Tan LW, Tan YL, Chen Q, Xu GM, Yang GG, Zuo XB, Yan H, Ruan YY, Lu TL, Han X, Ma XH, Wang Y, Cai LW, Jin C, Zhang HY, Yan J, Mi WF, Yin XY, Ma WB, Liu Q, Kang L, Sun W, Pan CY, Shuang M, Yang FD, Wang CY, Yang JL, Li KQ, Ma X, Li LJ, Yu X, Li QZ, Huang X, Lv LX, Li T, Zhao GP, Huang W, Zhang XJ, Zhang D (2011) Genome-wide association study identifies a susceptibility locus for schizophrenia in Han Chinese at 11p11.2. Nat Genet 43(12):1228–1231. https://doi.org/10.1038/ng.979
Zhang Y, Lu T, Yan H, Ruan Y, Wang L, Zhang D, Yue W, Lu L (2013) Replication of association between schizophrenia and chromosome 6p21–6p221 polymorphisms in Chinese Han population. PLoS One 8(2):e56732. https://doi.org/10.1371/journal.pone.0056732
Acknowledgements
We thank Dr. Kazuo Yamada for his support in the genetic association studies.
Funding
This research was supported by research grants from the Ministry of Education, Culture, Sports, Science and Technology of Japan and the Ministry of Health, Labour and Welfare of Japan; AMED JP18dm0107083 (TY), AMED JP18H05435 (TY), CREST (TN), Grant-in-Aid for Scientific Research (B) No.24390280 (TN), Grant-in-Aid for Scientific Research (C) No. 26461713 (NY) and Grant-in-Aid for Scientific Research (C) No. 17K10267 (AU). In addition, a part of this study was supported by Tokyo Medical and Dental University funds. REM was supported by MH107487, AG057598, and MH121102.
Author information
Authors and Affiliations
Contributions
AU performed ISH and had major roles in overall data analysis, data interpretation, and writing the paper. DJ, DS, NY and AK performed the genetic association study. YI, TT and TY performed the qRT-PCR study. VH and JMW supervised the postmortem studies. RM supervised ISH. TN conceived, designed and directed this project and wrote the final version of the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors have no conflict of interest to declare.
Ethical approval
The ethics committees of Tokyo Medical and Dental University, RIKEN Center for Brain Science, and University of Alabama at Birmingham approved the present study. All experiments were performed in accordance with the Declaration of Helsinki.
Consent to participate
Informed consent to participate in this study was obtained from all participants.
Consent for publication
Consent for publication was included in the informed consent.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Uezato, A., Jitoku, D., Shimazu, D. et al. Differential genetic associations and expression of PAPST1/SLC35B2 in bipolar disorder and schizophrenia. J Neural Transm 129, 913–924 (2022). https://doi.org/10.1007/s00702-022-02503-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00702-022-02503-7