Abstract
Previous molecular phylogenetic investigations of Apiaceae tribe Pimpinelleae have focused primarily on its largest genus Pimpinella and its closest allies. The monophyly and phylogenetic placements of five Iranian genera of the tribe have not been addressed sufficiently (Aphanopleura, Demavendia, Haussknechtia, Psammogeton, and Zeravschania). To examine relationships, a nrDNA ITS matrix including 169 accessions representing 49 genera of Apiaceae (including 10 Iranian taxa not analyzed previously) and a cpDNA rps16 matrix containing 37 accessions representing 24 genera of the family, representing the greatest sampling to date of the aforementioned genera, were subjected to phylogenetic analyses using Bayesian inference and maximum parsimony methods. The trees obtained showed a close affinity among the examined species of Aphanopleura, Psammogeton and several species of Trachyspermum. Neither Aphanopleura nor Psammogeton resolved as monophyletic, and A. leptoclada allied with Pimpinella. The genera Demavendia, Haussknechtia and Zeravschania also comprised a well-supported clade, with Demavendia and Haussknechtia (in the ITS trees) arising from within a paraphyletic Zeravschania. To recognize monophyletic genera, one new combination is proposed in Pimpinella and six new combinations are proposed in Psammogeton. A broader circumscription of Zeravschania to include Demavendia and Haussknechtia may also be warranted, but must await further study.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Tribe Pimpinelleae Spreng. includes the genera Aphanopleura Boiss., Arafoe Pimenov & Lavrova, Bubon L., Demavendia Pimenov, Frommia H.Wolff, Haussknechtia Boiss., Nothosmyrnium Miq., Phellolophium Baker, Pimpinella L., Psammogeton Edgew., and Zeravschania Korovin (Downie et al. 2010). It also includes segregates from other genera, such as Cryptotaenia africana Drude, Physospermopsis cuneata H.Wolff, and two species of Trachyspermum Link (T. scaberulum (Franch.) H.Wolff and T. triradiatum H.Wolff; Downie et al. 2010). Six genera of Pimpinelleae occur in Iran (Aphanopleura, Demavendia, Haussknechtia, Pimpinella, Psammogeton, and Zeravschania), of which only Demavendia and Haussknechtia are endemic (Hedge et al. 1987; Mozaffarian 2007; Mousavi et al. 2021). The members of Pimpinelleae in Iran are morphologically diverse and cannot be identified easily, although most species have small fruits (shorter than 5 mm) covered by glochidiate hairs (except in Demavendia and Zeravschania where the fruits are glabrous). Representatives of the tribe have been included in several molecular phylogenetic studies using nuclear ribosomal DNA internal transcribed spacer (nrDNA ITS) sequences (Downie et al. 2000a, b, 2001; Valiejo-Roman et al. 2006; Spalik and Downie 2007; Ajani et al. 2008; Zhou et al. 2008; Downie et al. 2010; Magee et al. 2010; Wang et al. 2014; Fernández Prieto et al. 2018; Mousavi et al. 2021). However, in all of these studies, several Iranian genera were incompletely sampled and the available results do not support the monophyly of four genera in their current circumscriptions: Aphanopleura (Fig. 1a–b), Pimpinella, Psammogeton (Fig. 1f–g), and Zeravschania (Fig. 1l–o).
Pimpinella is the largest genus of the tribe, includes about 150 species, and is taxonomically the most challenging. The genus is widely distributed in the Old World through Europe, Africa, and Asia, with the majority of species (ca. 125 spp.) occurring in Asia (Wolff 1921; Pimenov and Leonov 1993, 2004; Pu and Watson 2005). Members of the genus are characterized by cordate-ovoid or oblong-ovoid leaves and laterally compressed mericarps constricted at their commissures, each bearing five filiform ribs (Pu and Watson 2005). Pimpinella sensu lato (s.l.) is highly polyphyletic, with its members assigned to no less than seven tribes and other major clades within Apiaceae (Downie et al. 2010; Wang et al. 2014; Fernández Prieto et al. 2018). Nineteen species of Pimpinella occur in Iran, of which six (P. tragioides Boiss., P. deverroides Boiss., P. pastinacifolia H.Wolff, P. khorasanica Engstrand, P. anisactis Rech.f. and P. khayyamii Mozaff.) are endemic (Hedge et al. 1987; Mozaffarian 2007; Khajepiri et al. 2010). The relationships among many of these SW Asian Pimpinella species have been reported previously (Tabanca et al. 2005; Fereidounfar et al. 2016; Khajepiri et al. 2010). Fruit anatomical studies of Iranian Pimpinella confirmed differences between P. anthriscoides Boiss. and other species of Pimpinella and resulted in the placement of this species into the new genus Pseudopimpinella F.Ghahrem., Khajepiri & Mozaff. [as Pseudopimpinella anthriscoides (Boiss.) F.Ghahrem., Khajepiri & Mozaff.; Khajepiri et al. 2010]. Pimpinella anthriscoides has been transferred to the genus Aegopodium L. as A. tribracteolatum Schmalh. and P. anthriscoides var. cruciata (Bornm. & H.Wolff) V.A.Matthew has been transferred to Tamamschjanella Pimenov & Kljuykov as T. cruciata (Bornm. & H.Wolff) Pimenov & Zakharova (Zakharova et al. 2012). The monotypic Iranian genus Opsicarpium Mozaff. and the polytypic genus Reutera Boiss. have been merged recently with Pimpinella (Fereidounfar et al. 2016).
Aphanopleura and Psammogeton each include five species distributed mainly in the Iranian highlands (Hedge et al. 1987). They are morphologically similar, being small annual plants having white petals and laterally compressed mericarps covered by trichomes (Hedge et al. 1987). Demavendia is a monotypic genus [D. pastinacifolia (Boiss. & Hausskn.) Pimenov] endemic to Iran and Turkmenistan. Zeravschania includes nine species distributed in Iran, eastern Turkmenistan, Afghanistan, and Pakistan (Hedge et al. 1987; Mozaffarian 2007; Pimenov et al. 2007). Demavendia and Zeravschania possess similar membranous bracts and bracteoles and differ by their leaf shape (Hedge et al. 1987). Haussknechtia is a monotypic genus (H. elymaitica Boiss.) characterized by globose ultimate umbels (Hedge et al. 1987; Mozaffarian 2007). The Iranian genera Aphanopleura, Demavendia (Fig. 1c–e), Haussknechtia, Psammogeton, and Zeravschania have heretofore been poorly investigated phylogenetically. Previous phylogenetic studies suggested the non-monophyly of Aphanopleura, Psammogeton, and Zeravschania, and revealed the placement of these taxa along with Demavendia and Haussknechtia (and the Asian genus Nothosmyrnium Miq.) as basal branching lineages within the tribe (Spalik and Downie 2007; Wang et al. 2014; Fernández Prieto et al. 2018).
The main aim of this study is to investigate the monophyly and phylogenetic placements of the Iranian genera assigned to tribe Pimpinelleae, particularly those constituting the most basal branching lineages within the tribe that were underrepresented in previous molecular phylogenetic analyses (i.e., Aphanopleura, Demavendia, Haussknechtia, Psammogeton, and Zeravschania). We also highlight the morphological characters supporting each of the resolved clades containing these taxa.
Materials and methods
Sampling strategy and marker selection
We included all genera but one previously assigned to tribe Pimpinelleae and many species from throughout the entire range of each genus. The genus Bubon, reinstated with the single species B. macedonicum L. and separated from Athamanta L. in tribe Scandiceae by Spalik et al. (2001), could not be included because material was unavailable. We included representatives from most tribes and other major clades of Apiaceae subfamily Apioideae (Downie et al. 2010) and rooted all trees with Sanicula epipactis E.H.L. Krause (subfamily Saniculoideae). The nrDNA ITS region, comprising ITS1, 5.8S rDNA, and ITS2, and the plastid rps16 intron were selected as markers for phylogenetic analyses. These markers were suggested as effective and informative for tribal- and generic-level assignments in Apiaceae (Downie et al. 2010). We newly generated the sequences of 10 taxa for ITS and nine taxa for rps16 from the following Iranian species: Aphanopleura breviseta (Boiss.) Heywood & Jury, A. leptoclada Lipsky, A. trachysperma Boiss., Demavendia pastinacifolia (Boiss. & Hausskn.) Pimenov, Opopanax armeniacus Bordz., Psammogeton stocksii (Boiss.) Nasir, Trachyspermum ammi (L.) Sprague, T. copticum (L.) Link, Zeravschania membranacea (Boiss.) Pimenov, and Z. pauciradiata (Tamamsch.) Pimenov. We did not have access to material of Aphanopleura zangelanica Gogina & Matsenko because it is not distributed in Iran. We downloaded all available ITS and rps16 intron sequences of Pimpinelleae from GenBank and created a data matrix for each marker. The final nrDNA ITS data set included 169 accessions representing 49 genera (including the aforementioned 10 species not analyzed previously) and the cpDNA rps16 data set included 37 accessions representing 24 genera. The number of included taxa in the rps16 data set was considerably smaller than that of ITS due to failed PCR or sequencing efforts for the rps16 region and the preponderance of ITS sequences available in GenBank. The accessions included in this study, including voucher information for the newly generated sequences, are presented in Appendix. A list of all new combinations proposed in this study is presented in Table 2.
DNA extractions, PCR amplifications, and sequencing
Total genomic DNA was extracted from dried leaves removed from herbarium sheets held in the herbarium of the Research Institute of Forests and Rangelands, Tehran (TARI) using a NucleoSpin Plant DNA Extraction kit (Macherey–Nagel, Düren, Germany) according to the manufacturer’s protocol. For PCR-amplification of the nrDNA ITS region, we used primer pairs ITS4 and ITS5 (White et al. 1990). The cpDNA rps16 intron region was amplified using primer pairs rpsF and rpsR3R (Oxelman et al. 1997; Petri and Oxelman 2011; Kool et al. 2012). PCR amplifications were performed in T-Personal 48 (Biometra, Göttingen, Germany), Primus 96 plus (MWG: Biotech, Ebersberg, Germany), or 2720 (Applied Biosystems, Carlsbad, California, U.S.A.) thermocyclers. Cycle sequencing was performed using the BigDye Terminator v.3.1 Cycle Sequencing kit (Applied Biosystems). PCR products were sequenced bi-directionally using an ABI 3730 DNA Analyzer 48-well capillary sequencer (Applied Biosystems).
Alignment and phylogenetic analyses
The sequences were aligned using default parameters in Mafft v.7 (Katoh and Standley 2013), and alignment errors were corrected using Mesquite 2.75 (Maddison and Maddison 2011). Bayesian inference (BI) and maximum parsimony (MP) analyses were done for each data set on the CIPRES server (Miller et al. 2010). Before running the BI analysis, the optimal substitution models were estimated using the Akaike information criterion (AIC) in jModelTest v.0.1.1 (Posada 2008). The general time reversible model of nucleotide substitution with gamma-shaped rate variation and a proportion of invariable sites (GTR + I + G) was the estimated best-fit model for ITS and GTR + G was the estimated best-fit model for rps16. For BI, we used MrBayes v.3.2.6 (Ronquist and Huelsenbeck 2003) with 20 million generations for each data set. Trees were sampled every 1000 generations with the default of three “heated” and one “cold” chain. According to the results obtained from Tracer v.1.7.1 (Rambaut et al. 2018), the burn-in values were 10% and 13% for ITS and rps16 datasets, respectively. The effective sample sizes (ESS) were 438.3 and 271.8 for ITS and rps16 datasets, respectively. The obtained trees from each data set were summarized using a 50% majority-rule consensus tree. Bayesian posterior probability (PP) values were presented on the trees. For MP, we used PAUP* v.4.0b10 (Swofford 2003). Characters were equally weighted, the heuristic search was set with random sequence addition, tree-bisection-reconnection branch swapping, 50 random addition sequence replicates, and the maxtree option set to 20,000. The obtained trees were summarized using a strict consensus tree. Bootstrap (BS) values were calculated using heuristic searches with the bootstrap nreps set at 5,000. Additional details describing the alignment and phylogenetic strategies and analyses used are presented in Mousavi et al. (2021).
Results
A summary of alignment characteristics and parsimony statistics for the ITS and rps16 data sets is presented in Table 1. The ITS matrix had over 2.5 times as many parsimony-informative characters as did the rps16 intron matrix. We present only the trees obtained from BI analyses of these data sets. Though the number of accessions differed considerably between these two data sets, their results were largely congruent in revealing the groupings of the Iranian genera under study, their non-monophyly, and the placement of A. leptoclada in a clade within Pimpinella saxifraga L., the nomenclatural type of Pimpinella. The ITS and rps16 alignments are available in the Online Resources 1, 2.
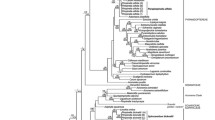
Although the tribe Pimpinelleae comprised a highly supported, monophyletic group in the ITS phylogeny (PP = 1, BS = 98, Clades A-F, Fig. 2), it was not resolved as monophyletic in the rps16 trees (Fig. 3). In both sets of trees, several clades can be recognized in Pimpinelleae. The Iranian species of Pimpinella fell within the Pimpinella core group (i.e., Clades E and F). In the ITS trees, Clade A had strong support (PP = 1, BS = 98) and included members of Aphanopleura, Nothosmyrnium, Psammogeton, and Trachyspermum. Neither Aphanopleura, Psammogeton, nor Trachyspermum resolved as monophyletic. Clade B contained representatives of the Iranian genera Demavendia, Haussknechtia, and Zeravschania. Demavendia and Haussknechtia comprised monophyletic sister groups, with this clade arising from within a paraphyletic Zeravschania. The single accession of A. leptoclada allied weakly with Pimpinella procumbens H.Wolff in Clade F. Clades C-F comprised Pimpinella species already attributable to tribe Pimpinelleae in previous studies, plus all examined accessions of Physospermopsis H.Wolff, the Afro-Malagasian members of the genera Frommia, Phellolophium and Cryptotaenia DC., and Arafoe. Pimpinella saxifraga, the type of the genus, occurred in Clade F. Twelve additional species of Pimpinella (14 accessions) fell outside of tribe Pimpinelleae, none of which occur in Iran.
Majority-rule consensus tree inferred from Bayesian Inference analysis of nrDNA ITS sequences. Numbers above the branches indicate posterior probability values; those below branches indicate MP bootstrap values for similar clades occurring in the MP strict consensus tree. The major clades recognized in tribe Pimpinelleae (Clades A-F) comprise a monophyletic group. The relationships of those accessions indicated in red are those highlighted in the present study
Majority-rule consensus tree inferred from Bayesian Inference analysis of plastid rps16 sequences. Numbers above the branches indicate posterior probability values; those below branches indicate MP bootstrap values for similar clades occurring in the MP strict consensus tree. Tribe Pimpinelleae (Clades A-B and D-F; the same as those presented in Fig. 2) does not comprise a monophyletic group, as members of tribe Pyramidoptereae (incl. Trachyspermum copticum) and several other members of tribes Apieae, Careae, and Selineae and the Cachrys and Conium clades occur within
In the rps16 trees, the placements of the Iranian genera were similar, with Psammogeton stocksii arising from within a paraphyletic Aphanopleura and this clade sister group to Nothosmyrnium (Clade A) and Demavendia arising from within a paraphyletic Zeravschania (Clade B). Aphanopleura leptoclada was sister to the clade of Pimpinella anisum L. + P. saxifraga.
Discussion
Though Pimpinella is a polyphyletic genus, with its species occurring in at least seven different tribes and other major clades of subfamily Apioideae (Downie et al. 2010; Wang et al. 2014; Fernández Prieto et al. 2018), the Iranian species of Pimpinella examined herein all fall in the same well-supported group as P. saxifraga, the type of the genus. This group reflects the core Pimpinella group (Fernández Prieto et al. 2018) and is reflected in the current study by Clades E and F. The Iranian genera Aphanopleura, Demavendia, Haussknechtia, Psammogeton, and Zeravschania, along with Nothosmyrnium and several species of Trachyspermum, constitute basally branching lineages of tribe Pimpinelleae in the ITS trees (Clades A and B). Based on the phylogenetic results obtained herein, we recircumscribe Psammogeton to include six new combinations and transfer Aphanopleura leptoclada to Pimpinella (Table 2). We also highlight the morphological characters supporting each of the resolved clades containing these taxa.
Aphanopleura and Psammogeton
Aphanopleura is a genus of five species (A. breviseta, A. capillifolia Lipsky, A. leptoclada Lipsky, A. trachysperma, and A. zangelanica). Aphanopleura zangelanica was not included in our study. These species are distributed in Central and SW Asia, sometimes reaching China (2 spp.), but mostly they occur in Iran, Afghanistan, Turkmenistan, and Caucasus (Hedge et al. 1987). The plants are characterized by an annual life form, small and thin stems, ternate (3-segmented) leaves, the presence of bracts and bracteoles, a clearly dentate calyx, white to red petals, and mericarps ovate or almost circular, laterally compressed, and covered by papillate, clavate or capitellate trichomes. Previous molecular phylogenetic investigations of Aphanopleura only clarified its placement in Pimpinelleae, not its interspecific relationships (Spalik and Downie 2007; Zhou et al. 2008; Downie et al. 2010; Wang et al. 2014; Fernández Prieto et al. 2018). According to our results, Aphanopleura is a polyphyletic genus, with most of its species nested within Clade A. The type of genus, A. trachysperma, presents a sister group relationship with Nothosmyrnium japonicum Miq., while the other species of Aphanopleura within this clade assemble with Trachyspermum or Psammogeton. Aphanopleura leptoclada is nested within Clade F, supporting the hypothesis that its annual habit and other correlated morphological features have evolved independently in tribe Pimpinelleae. We transfer A. leptoclada into Pimpinella (see “Taxonomic Treatment” section), while acknowledging that the annual habit of this species is rare in Pimpinella.
Psammogeton includes five species distributed in Central and SW Asia, mainly in the Iranian Highlands (Hedge et al. 1987), and all species were included herein. Its members are characterized by an annual life form, bi-ternate leaves with lanceolate or cuneate ultimate segments, the presence of bracts and bracteoles, white to red petals, and oblong, slightly laterally compressed mericarps densely covered by a glochidiate indumentum. The geographical distribution and morphological features of Aphanopleura and Psammogeton are very similar, with only the mericarp indumentum used for their separation (Boissier 1872; Hedge et al. 1987). In this study, Psammogeton forms a robust clade in Clade A, along with Aphanopleura breviseta and A. capillifolia. These two Aphanopleura species show a closer relationship with Psammogeton than they do to the type species of Aphanopleura (A. trachysperma). For this reason, we transfer A. capillifolia into Psammogeton and resurrect P. brevisetum Boiss. Aphanopleura breviseta had been previously treated as a synonym of P. brevisetum Boiss. (Hedge et al. 1987; Mozaffarian 2007).
Psammogeton, as circumscribed herein, includes species of Trachyspermum and Aphanopleura. This group is characterized by an annual or occasionally perennial habit, ultimate leaf segments mostly linear, bracts and bracteoles absent or small (shorter than 3 mm), and ovoid-globose mericarps covered by a diverse array of trichrome types (i.e., clavate, capitellate, tubular papillose, T-shaped, glochidiate, inflated vesicles, tubercles, or scales). Aphanopleura trachysperma, the type of the genus, is separate from the Psammogeton clade and is sister to Nothosmyrnium. It is distinct from members of the Psammogeton clade primarily by its much longer and conspicuous involucral bracts.
Trachyspermum
Trachyspermum includes some 12 to 15 species (Pimenov and Leonov 1993; Downie et al. 2000c; Wang et al. 2014). Trachyspermum copticum (L.) Link (a synonym of T. ammi (L.) Sprague) is the type of the genus (Fig. 1h–k). Trachyspermum is another taxonomically challenging genus (Pimenov and Leonov 1993). The genus was originally described based on a few general characters (Link 1821), such as pinnate involucral leaves, a striate, finely muricate mericarp having a short rough indumentum, a 5-merous and rudimentary calyx, a withering style, and a conical stylopodium, resulting in a vague circumscription. The genus is distributed worldwide, but predominantly occurs in Asia. The Iranian Highlands is a diversification center of the genus and harbors ten species including the newly described T. reginei Ajani & Mozzaff. (Hedge et al. 1987; Ajani and Mozaffarian 2019). The description of the genus was revised in Flora Iranica (Hedge et al. 1987) to include plants having the following features: annual or perennial habit, 1 to 3-pinnate leaves with linear ultimate segments, distinct calyx teeth, white petals incurved at apex, ovate or elliptic mericarps with prominent obtuse ribs and narrow at apex, and densely covered by white vesicular, papillose, T-shape or setulose indumentum (each type of indumentum characterizes a separate species), and a depressed or conical stylopodium. The members of Trachyspermum are only obscurely distinguishable from Aphanopleura, Pimpinella, and Psammogeton, with the main differences among them associated with the mericarp indumentum. Pimpinella possesses pilose, papillose or tuberculate trichomes, Aphanopleura (in its traditional circumscription, sensu Hedge et al. 1987) has clavate trichomes, Psammogeton bears glochidiate or martelliform trichomes at its apex, and Trachyspermum has papillose mericarps (Hedge et al. 1987). In addition, the perennial species of Trachyspermum are morphologically similar to Pimpinella, differing only in their number of vallecular vittae (which are 2‒4 in Pimpinella and one in Trachyspermum; Hedge et al. 1987).
Very few species of Trachyspermum were included in previous molecular phylogenetic studies, but not the type species T. copticum (Downie et al. 2000c, 2010; Zhou et al. 2008). These previous studies, though based on a limited sampling of species, rendered Trachyspermum as polyphyletic, with its members placed in three different tribes: T. aethusifolium Chiov. in Echinophoreae; T. ammi in Pyramidoptereae; and T. scaberulum and T. triradiatum H.Wolff in Pimpinelleae (Downie et al. 2010). The latter two species were recently united under Pimpinella scaberula (Franch.) H.Boissieu (Pimenov 2017) and in our study are placed in Clade E alongside members of Pimpinella. This placement is in accordance with previous studies (Magee et al. 2010; Wang et al. 2014; Pimenov 2017; Fernández Prieto et al. 2018).
Some species assigned previously to Trachyspermum, viz. T. roxburghianum (DC.) H.Wolff (= T. involucratum (Roxb.) H.Wolff), T. anethifolium H.Wolff, T. microcarpum Hedge, Lamond & Rech.f., T. papillare (Boiss.) Hedge, Lamond & Rech.f., and T. paktianum Hedge, Lamond & Rech.f., are nested within Clade A and show close affinity to Psammogeton. As a consequence, we transfer these species to Psammogeton. Trachyspermum copticum, a synonym of T. ammi (Menglan et al. 2005) and the type of the genus, is placed among the members of tribe Pyramidoptereae. Accordingly, the genus Trachyspermum is circumscribed narrowly.
Demavendia, Haussknechtia, and Zeravschania
Zeravschania, as circumscribed currently, includes nine species in SW Asia (Pimenov et al. 2007). It was originally described as a monotypic genus from the western Pamir-Alai mountains in Central Asia (Korovin 1948). The type of the genus, Z. regeliana Korovin, was not sufficiently described originally due to its lack of mature fruits. Subsequent morphological studies have shown that its mericarps are dorsally compressed and possess filiform dorsal ribs beside marginal ones that develop into wings (Hedge et al. 1987; Mozaffarian 2007). The most important characteristics of the genus are its white membranous and broadly ovate or lanceolate bracts and bracteoles. The phylogenies presented herein show Zeravschania as paraphyletic (Clade B), with all examined accessions of Demavendia and Haussknechtia nested within. To recognize a monophyletic Zeravschania, one treatment would be to include members of Demavendia and Haussknechtia in a broadly circumscribed Zeravschania. The monotypic genus Demavendia [D. pastinacifolia (Boiss. & Hausskn.) Pimenov] is endemic to Iran and also bears white, membranous bracts. The species currently placed in Demavendia and Zeravschania were previously treated as members of Peucedanum [unranked] Membranacea Boiss. which was characterized by such membranous bracts and bracteoles (Boissier 1872). Leaf shape and margin provide important differences between these two genera. Demavendia possesses large oblong leaf segments with irregularly dentate margins, whereas Zeravschania has ovate, ovate-oblong or lanceolate leaf segments which are 2‒4-ternate at its ultimate segments. Sister group to Demavendia and Zeravchania in the ITS trees is the monotypic genus Haussknechtia Boiss. (H. elymaitica Boiss.), a species characterized by tall plants up to 2 m or more, having sessile, very dense, and globose ultimate umbels (Hedge et al. 1987). We refrain herein from presenting a new formal taxonomic treatment of this group, in which Zeravschania is circumscribed much wider to include members of both Haussknechtia and Demavendia, due to the low resolution of relationships within the corresponding clade. Demavendia and Zeravschania are obviously closely related, especially upon considering their phylogenetic placement and morphological similarities. Haussknechtia also shows a close phylogenetic relationship to these genera, although its morphology is quite distinct. Further molecular phylogenetic investigations involving more informative markers should shed light on the relationships among Demavendia, Haussknechtia, and Zeravschania.
Taxonomic treatment
Psammogeton is circumscribed broadly to include species previously assigned to Aphanopleura and Trachyspermum. We increase the number of species under Psammogeton to 12 and reduce the number of species under Aphanopleura to two. In total, we recognize seven new combinations and resurrect one name.
New combinations
Pimpinella leptoclada (Aitch. & Hemsl.) Mousavi, Mozaff. & Zarre, comb. nov. ≡ Carum leptocladum Aitch. & Hemsl., Trans. Linn. Soc. Bot. Ser. 2, 3: 66. 1888–1889. ≡ Aphanopleura leptoclada (Aitch. & Hemsl.) Lipsky, Izv. Imp. Akad. Nauk V, 4: 377. 1898. ≡ Carum aphanopleura Koso-Pol., Not. Syst. Herb. Hort. Bot. Petrop. 3, 18: 70. 1922. —LECTOTYPE (designated here): Afghanistan, Herat Prov., Hari-rud valley, 5 Jun 1885, J.E.T. Aitchison 603 (G barcode G00359737; isolectotypes: FI barcode FI014839, GH barcode GH00075662, W barcode W1963-0000628). —SYNTYPES: Afghanistan, 5 Jun 1885, J.E.T. Aitchison 603 (P barcode P00834708); Afghanistan, Herat, Hari-rud valley, 1884, J.E.T. Aitchison 603 (C barcode C10008329).
Diagnosis: Annual plants, stem puberulent or glabrous, 10–40 cm tall; leaves ternate with linear or oblong-linear ultimate segments; ultimate umbels long pedunculated; bracts 2‒5, linear-lanceolate, membranous, scabrous at margins; bracteoles present; umbellules numerous with filiform rays; petals white or red, dorsally pilose; mericarps ovoid-globose or ovoid, covered by dense claviform or capitate indumentum.
Distribution: Iran (mostly in the northeast), Afghanistan, Tajikistan, Turkmenistan, and Uzbekistan.
Psammogeton anethifolium (D.Don) Mousavi, Mozaff. & Zarre, comb. nov. ≡ Pimpinella anethifolia D.Don, Prodr. Fl. Nepal. 184. 1825. ≡ Athamanta anethifolia (D.Don) Wall., Numer. List no. 569. 1829. ≡ Ptychotis anethifolia (D.Don) DC., Prodr. 4: 108. 1830. ≡ Carum anethifolium (D.Don) C.B.Clarke in Benth. & Hook.f., Fl. Brit. India, 2(6): 683. 1879. ≡ Trachyspermum anethifolium (D.Don) H.Wolff, Pflanzenr. IV, 228(90): 90. 1927. —LECTOTYPE (designated here): Nepalia, 1821, N. Wallich 569 (K barcode K000685646; isolectotypes: G barcode G00367346, M barcode M0172780).
Diagnosis: Annual plants, pubescent, up to 40–50 cm tall; leaves small, 1–4 cm long, ternate with linear ultimate segments; ultimate umbels small, with unequal, and filiform rays; bracts and bracteoles numerous, linear, ± as long as the peduncles; petals white; mericarps rounded-elliptic, pubescent.
Distribution: Nepal (endemic).
Psammogeton capillifolium (Regel & Schmalh.) Mousavi, Mozaff. & Zarre, comb. nov. ≡ Pimpinella capillifolia Regel & Schmalh., Izv. Imp. Obshch. Lyubit. Estestv. Moskovsk. Univ. 34(2): 29. 1881. ≡ Aphanopleura capillifolia (Regel & Schmalh.) Lipsky, Izv. Imp. Akad. Nauk. 4: 379. 1896. = Carum capillifolium (Regel & Schmalh.) Koso-Pol., Bull. Soc. Imp. Naturalistes Moscou n.s., 29: 199. 1915 (publ. 1916). —TYPE: Kazakhstan, Karatan of fluv. Tschygan [Tschilak, am Nordabhang der Berge Karatan], 1880, A. Regel s.n. SYNTYPEYNTYPE; Kazakhstan, Turkestan: Suleimanfels bei Osch., 25 May 1880, A. Regel s.n. (P barcode P03214061; isosyntype: MPU barcode MPU019088).
Diagnosis: Annual plants; glabrous, sometimes purplish-red in lower parts of stem; leaves petiolate, 2-pinnate or 2-ternate, ultimate segments filiform or linear; umbels long, usually with more than 2 flowers and unequal rays; bracts absent or one; bracteoles lanceolate or linear; mericarps broadly ovate.
Distribution: Kazakhstan, Kyrgyzstan, Tajikistan, Turkmenistan, Uzbekistan, and China.
Psammogeton microcarpum (Hedge, Lamond & Rech.f.) Mousavi, Mozaff. & Zarre, comb. nov. ≡ Trachyspermum microcarpum Hedge, Lamond & Rech.f., Fl. Iran. 162: 342. 1987. —TYPE: Afghanistan, Kabul, in valle Logar 12 km S Kabul prope “Stupa Guldara”, 1970 ma. s. l., in rupibus verticalibus, 24 Sep 1967, H. Freitag 1991 (holotype M Hb.Freitag; isotypes: E, W barcode W0054255).
Diagnosis: Perennial plants, stem up to 40 cm; leaves trisected or pinnate with cuneate ultimate segments; umbels with 2–5 unequal rays; bracts and bracteoles linear or setaceous; mericarps ovate-elliptic, covered with T-shaped trichomes; stylopodium conical.
Distribution: Afghanistan (endemic).
Psammogeton paktianum (Hedge, Lamond & Rech.f.) Mousavi, Mozaff. & Zarre, comb. nov. ≡ Trachyspermum paktianum Hedge, Lamond & Rech.f., Fl. Iran. 162: 343. 1987. —TYPE: Afghanistan, Jaji, slopes 1 km below Alikhe, 20 Aug 1907, H.Freitag 1829 (holotype M Hb. Freitag; isotypes: E barcode E00000356, MSB barcode MSB003195, MSB barcode MSB003196).
Diagnosis: Perennial plants; leaves long petiolate, subtripinnate with cuneate-trisected or acute ultimate segment; umbels long, thin, pedunculated with 4–6 rays; bracts subulate; bracteoles lanceolate; petals white and short papillous; mericarps ovoid or subpyriform, covered with short trichomes.
Distribution: Afghanistan (endemic).
Psammogeton papillare (Boiss.) Mousavi, Mozaff. & Zarre, comb. nov. ≡ Reutera papillaris Boiss., Diagn. Pl. Orient. Ser. 2, 2: 76. 1856. ≡ Pimpinella papillaris (Boiss.) Benth. & Hook.f. ex Drude, in Engler & Prantl, Natürl. Pflanzenfam. III, 8: 196. 1898. ≡ Carum papillare (Boiss.) Koso-Pol., Bull. Soc. Imp. Naturalistes Moscou n.s., 29: 198. 1915 (publ. 1916). ≡ Trachyspermum papillare (Boiss.) Hedge, Lamond & Rech.f., Fl. Iran. 162: 342. 1987. —LECTOTYPE (designated here): Afghanistan, Jumra and Karabagh, W.Griffith s.n. (K, barcode K000685651; isolectotype: E, barcode E00000357).
Diagnosis: Perennial plants; leaves pinnate with linear, acuminate ultimate segments, cauline leaves linear-filiform, apiculate; umbels with few, unequal, thin rays; bracts and bracteoles lanceolate, acute, unequal, densely hirsute, hyaline at margin; petals white, short hirsute; mericarps ovoid, covered with dense T-shape trichomes; stylopodium depressed or conical.
Distribution: Afghanistan (endemic).
Psammogeton involucratum Mousavi, Mozaff. & Zarre, comb. nov. ≡ Apium involucratum Roxb. in Fleming, Asiat. Res. 11: 157. 1810. ≡ Athamanta roxburghiana Wall., Cat. no. 571. 1829, nom. superfl. ≡ Ptychotis roxburghiana DC., Prodr. [A.P. de Candolle] 4: 109. 1830, nom. superfl. ≡ Pimpinella involucrata (Roxb.) Wight & Arn., Prodr. Fl. Ind. Orient. 1: 369. 1834. ≡ Carum roxburghianum (DC.) Benth. & Hook.f., Gen. Pl. 1: 891. 1867, nom. superfl. ≡ Carum involucratum (Roxb.) Baill., Hist. Pl. 7: 179. 1879. ≡ Carum involucratum (Roxb.) Kuntze, Rev. Gen. 265. 1891, nom. illeg., non Baill. ≡ Trachyspermum roxburghianum Craib, Fl. Siam. Enum. 1: 788. 1931. ≡ Trachyspermum matthewii M.R.Almeida, Fl. Maharashtra 2: 363. 1998, nom. superfl. —LECTOTYPE (selected by I.M.Turner, 2014 in Kew Bulletin 69: 9489): Roxburgh drawing no. 1388 Icones Roxburghianae no. 1388 (K!, available online at http://apps.kew.org/floraindica/home.do).
= Trachyspermum stictocarpum (C.B.Clarke) H.Wolff, Pflanzenr. (Engler) Umbellif.-Apioid.-Ammin. 89. 1927. ≡ Carum stictocarpum C.B.Clarke, Fl. Brit. India 2: 681. 1879. —LECTOTYPE (designated here): India, Maharashtra, Concan, J.E. Stocks and J.S. Law s.n. (K barcode K000685630; isolectotypes: K barcode. K000685631, K000685629). —SYNTYPE: J.S. Law s.n. (K barcode K000685632).
= Carum stictocarpum var. hebecarpum C.B.Clarke, Fl. Brit. India 2: 682. 1879. —LECTOTYPE (designated here): India, Maharashtra, Concan, J.E. Stocks s.n. (K barcode K000685626; isolectotype: K barcode K000685627). —SYNTYPE: India, Maharashtra, Concan, J.S. Law s.n. (K barcode K000685628).
Diagnosis: Annual plants; stem 20–100 cm; leaves petiolate, blade ovate, 2-pinnate or ternate-pinnate with narrowly oblong ultimate segments; umbels 2–4, pedunculated, with 4–12 filiform rays; bracts and bracteoles few, linear-subulate or ciliate; mericarps ovoid, densely hirsutulous or glabrescent.
Distribution: Java, India, Malaysia, Borneo, Philippines, Vietnam, Laos, New Guinea, Thailand, Andaman Islands, Myanmar, China, and Bangladesh.
Resurrected name
Psammogeton brevisetum Boiss., Fl. Or. 2: 1079. 1872. ≡ Cuminum brevisetum (Boiss.) Kos-Pol., Bull. Soc. Imp. Nat. Mosc. Ser. Nov. 29: 209. 1916. ≡ Aphanopleura breviseta (Boiss.) Heywood & Jury, Ombell. Contrib. Pluridisc. Syst. 2: 733. 1977 (publ. 1978). —LECTOTYPE (designated here): Iran, Inter Kerman et Jesd, May.1859, A. von Bunge (P barcode P00834707).
= Athamantha grisea Stapf & Wettst., Denkschr. Akad. Wiss. Wien. Math. Nat. Kl. 51, 2: 319. 1886. —Lectotype (designated here): Iran, Kuschkek inter Hamadan & Teheran, 19 Jul 1882, T.Pichler s.n. (JE barcode JE00003614).
= Psammogeton flabellatus Bornm. & Gauba, Repert. Spec. Nov. Regni Veg. 36: 341. 1934. —TYPE: Iran, Keredj: Salzberge bei Mardabad (SW Keredj in der Salzsteppe), 1934, D.E. Gauba s.n. (Holotype: B barcode B100366020).
Diagnosis: Annual plants; stem 3–30 cm; leaves ternate or bi-ternate, petiolate, densely covered by short indumentum; umbels pedunculate with 5–10 rays; bracts 5, short, membranous at margin, subulate; bracteoles 5–7, lanceolate; petals white, curved at apex; mericarps oblong-ovoid covered by clavate, or vesiculate trichomes.
Distribution: Central Iran (endemic).
References
Ajani Y, Mozaffarian V (2019) Trachyspermum reginei sp. nov. (Apiaceae) from southwest Iran. Nordic J Bot 2019:e02238. https://doi.org/10.1111/njb.02238
Ajani Y, Ajani A, Cordes JM, Watson MF, Downie SR (2008) Phylogenetic analysis of nrDNA ITS sequences reveals relationships within five groups of Iranian Apiaceae subfamily Apioideae. Taxon 57:383–401. https://doi.org/10.2307/25066011
Boissier E (1872) Umbelliferae. In: Boissier E (ed) Flora Orientalis, vol 2. H Georg, Geneva & Basileae, pp 819–1090
Downie SR, Katz-Downie DS, Spalik K (2000a) A phylogeny of Apiaceae tribe Scandicinae: evidence from nuclear ribosomal DNA internal transcribed spacer sequences. Amer J Bot 87:76–95. https://doi.org/10.2307/2656687
Downie SR, Watson MF, Spalik K, Katz-Downie DS (2000b) Molecular systematics of Old World Apioideae (Apiaceae): relationships among some members of tribe Peucedaneae sensu lato, the placement of several island-endemic species, and resolution within the apioid superclade. Canad J Bot 78:506–528. https://doi.org/10.1139/cjb-78-4-506
Downie SR, Katz-Downie DS, Watson MF (2000c) A phylogeny of the flowering plant family Apiaceae based on chloroplast DNA rpl16 and rpoC1 intron sequences: towards a suprageneric classification of subfamily Apioideae. Amer J Bot 87:273–292. https://doi.org/10.2307/2656915
Downie SR, Plunkett GM, Watson MF, Spalik K, Katz-Downie DS, Valiejo-Roman CM, Terentieva EI, Troitsky AV, Lee B-Y, Lahham J, El-Oqlah A (2001) Tribes and clades within Apiaceae subfamily Apioideae: the contribution of molecular data. Edinburgh J Bot 58:301–330. https://doi.org/10.1017/S0960428601000658
Downie SR, Spalik K, Katz-Downie DS, Reduron JP (2010) Major clades within Apiaceae subfamily Apioideae as inferred by phylogenetic analysis of nrDNA ITS sequences. Pl Diversity Evol 128:111–136. https://doi.org/10.1127/1869-6155/2010/0128-0005
Fereidounfar S, Ghahremaninejad F, Khajehpiri M (2016) Phylogeny of the Southwest Asian Pimpinella and related genera based on nuclear and plastid sequences. Genet Molec Res 15:gmr15048767. https://doi.org/10.4238/gmr15048767
Fernández Prieto JA, Sanna M, Sánchez ÁB, Molero-Mesa J, García LL, Cires E (2018) Polyphyletic origin in Pimpinella (Apiaceae): evidence in Western Europe. J Pl Res 131:747–758. https://doi.org/10.1007/s10265-018-1046-5
Hedge IC, Lamond JM, Rechinger KH (1987) Umbelliferae. In: Rechinger KH (ed) Flora Iranica, vol. 162. Akademische Druck und Verlagsanstalt, Graz, pp 1–555
Katoh K, Standley DM (2013) MAFFT Multiple Sequence Alignment Software Version 7: Improvements in Performance and Usability. Molec Bio Evol 30:772–780. https://doi.org/10.1093/molbev/mst010
Khajepiri M, Ghahremaninejad F, Mozaffarian V (2010) Fruit anatomy of the genus Pimpinella L. (Apiaceae) in Iran. Flora 205:344–356. https://doi.org/10.1016/j.flora.2009.12.030
Kool A, Perrigo A, Thulin M (2012) Bristly versus juicy: Phylogenetic position and taxonomy of Sphaerocoma (Caryophyllaceae). Taxon 61:67–75. https://doi.org/10.1002/tax.611005
Korovin EP (1948) Species novae Umbelliferarum Florae Uzbekistanicae. Bot Mater Gerb Inst Bot Zool Akad Nauk Uzbeksk SSR 12:28
Link HF (1821) Enumeratio Plantarum Horti Regii Berolinensis Altera. Parts 1. G. Reimer, Berolini
Maddison WP, Maddison DR (2011) Mesquite: A modular system for evolutionary analysis version 2.75. Available at: http://mesquiteproject.org
Magee AR, van Wyk B-E, Tilney PM, Downie SR (2010) Phylogenetic position of African and Malagasy Pimpinella species and related genera (Apiaceae, Pimpinelleae). Pl Syst Evol 288:201–211. https://doi.org/10.1007/s00606-010-0325-y
Menglan S, Fading P, Zehui P, Watson MF, Cannon JFM, Holmes-Smith I, Kljuykov EV, Phillippe LR, Pimenov MG (2005) Apiaceae (Umbelliferae). In: Flora of China Editorial Committee (eds) Flora of China, vol. 14. Science Press & Missouri Botanical Garden, Beijing & St Louis, pp 1–205
Miller MA, Pfeiffer W, Schwartz T (2010) Creating the CIPRES Science Gateway for inference of large phylogenetic trees. pp: 45–52 in: Proceedings of the Gateway Computing Environments Workshop (GCE) New Orleans Louisiana 14 Nov 2010 Piscataway: IEEE. https://doi.org/10.1109/GCE.2010.5676129
Mousavi S, Mozaffarian V, Mummenhoff K, Downie SR, Zarre S (2021) An updated lineage-based tribal classification of Apiaceae subfamily Apioideae with special focus on Iranian genera. Syst Biodivers 19:89–109. https://doi.org/10.1080/14772000.2020.1834002
Mozaffarian V (2007) Flora of Iran, vol. 54. Umbelliferae. Research Institute of Forests and Rangelands, Tehran
Oxelman B, Lidén M, Berglund D (1997) Chloroplast rps16 intron phylogeny of the tribe Sileneae (Caryophyllaceae). Pl Syst Evol 206:393–410. https://doi.org/10.1007/BF00987959
Petri A, Oxelman B (2011) Phylogenetic relationships within Silene (Caryophyllaceae) section Physolychnis. Taxon 60:953–968. https://doi.org/10.1002/tax.604002
Pimenov MG (2017) Updated checklist of Chinese Umbelliferae: nomenclature synonymy typification distribution. Turczaninowia 20:106–239. https://doi.org/10.14258/turczaninowia.20.2.9
Pimenov MG, Leonov MV (1993) The genera of the Umbelliferae. Royal Botanic Gardens, Kew
Pimenov MG, Leonov MV (2004) The Asian Umbelliferae Biodiversity Database (ASIUM) with particular references to southwest Asian taxa. Turkish J Bot 28:139–145
Pimenov MG, Kljuykov EV, Ostroumova TA (2007) Critical taxonomic analysis of Dichoropetalum, Johrenia, Zeravschania related genera of Umbelliferae-Apioideae-Peucedaneae. Willdenowia 37:465–502. https://doi.org/10.3372/wi.37.37208
Posada D (2008) jModelTest: Phylogenetic model averaging. Molec Biol Evol 25:1253–1256. https://doi.org/10.1093/molbev/msn083
Pu FD, Watson MF (2005) Pimpinella. In: Wu ZY, Raven PH, Hong DY (eds) Flora of China, vol 14. Missouri Botanical Garden Press, St Louis, pp 93–104
Rambaut A, Drummond AJ, Xie D, Baele G, Suchard MA (2018) Posterior summarization in bayesian phylogenetics using tracer 1.7. Syst Biol 67:901–904. https://doi.org/10.1093/sysbio/syy032
Ronquist F, Huelsenbeck JP (2003) MrBayes: Bayesian Phylogenetic Inference under Mixed Models. Bioinformatics 19:1572–1574. https://doi.org/10.1093/bioinformatics/btg180
Spalik K, Downie SR (2007) Intercontinental disjunctions in Cryptotaenia (Apiaceae Oenantheae): an appraisal using molecular data. J Biogeogr 34:2039–2054. https://doi.org/10.1111/j.1365-2699.2007.01752.x
Spalik K, Wojewódzka A, Downie SR (2001) The evolution of fruit in Scandiceae subtribe Scandicinae (Apiaceae). Canad J Bot 79:1358–1374
Swofford DL (2003) PAUP: Phylogenetic analysis using parsimony version 40b10 for 32-bit Microsoft Windows Sunderland. Sinauer, Massachusetts
Tabanca N, Douglas AW, Bedir E, Dayan FE, Kirimer N, Can Baser KH, Aytac Z, Khan IA, Scheffler BE (2005) Patterns of essential oil relationships in Pimpinella (Umbelliferae) based on phylogenetic relationships using nuclear and chloroplast sequences. Pl Genet Resources 3:149–169. https://doi.org/10.1079/PGR200573
Valiejo-Roman CM, Terentieva EI, Samigullin TH, Pimenov MG, Ghahremani-Nejad F, Mozaffarian V (2006) Molecular data (nrITS-sequencing) reveal relationships among Iranian endemic taxa of the Umbelliferae. Feddes Repert 177:367–388. https://doi.org/10.1002/fedr.200611106
Wang Z-X, Downie SR, Tan J-B, Liao C-Y, Yan Y, He X-J (2014) Molecular phylogenetics of Pimpinella and allied genera (Apiaceae), with emphasis on Chinese native species, inferred from nrDNA ITS and cpDNA intron sequence data. Nordic J Bot 32:642–657. https://doi.org/10.1111/j.1756-1051.2013.00343.x
White TJ, Bruns T, Lee S, Taylor J (1990) Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In: Innis MA, Gelfand DH, Sninsky JJ, White TJ (eds) PCR protocols: A guide to methods and applications. Academic Press, San Diego, pp 315–322. https://doi.org/10.1016/B978-0-12-372180-850042-1
Wolff H (1921) Spiroceratium bicknellii nov. gen. Umbelliferarum balearicum. Repert Spec Nov Regni Veg 17:45
Zakharova EA, Degtjareva GV, Pimenov MG (2012) Redefined generic limits of Carum (Umbelliferae, Apioideae) and new systematic placement of some of its taxa. Willdenowia 42:149–168. https://doi.org/10.3372/wi4242201
Zhou J, Peng H, Downie SR, Liu Z-W, Gong X (2008) A molecular phylogeny of Chinese Apiaceae subfamily Apioideae inferred from nuclear ribosomal DNA internal transcribed spacer sequences. Taxon 57:402–416. https://doi.org/10.2307/25066012
Acknowledgements
S.S.M. is grateful to the University of Tehran and the Research Institute of Forests and Rangelands in Tehran for supporting her work. We are grateful to the directors and curators of the herbaria TARI and TUH for their generous assistance in providing plant materials and photographs for this study. S.Z. also appreciates the financial support provided by Alexander Humboldt Stiftung (Germany) and the University of Tehran. Lastly, we thank three anonymous reviewers and Deborah S. Katz-Downie for their valuable constructive comments on early drafts of the manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Handling editor: Christoph Oberprieler.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Appendices
Appendix
Accessions of Pimpinelleae and other taxa of Apiaceae subfamilies Apioideae and Saniculoideae included in the study. Previously published nrDNA ITS and rps16 sequences are represented herein by GenBank accession numbers only. The first GenBank number following a species name refers to ITS and the second for rps16, except for those instances where there are three numbers and these refer to separate ITS1, ITS2, and rps16 sequences. A single GenBank number often refers to ITS but in a few instances it refers to rps16. The number “1” or “2” added to the species name coincides with their placement in Figs. 2 and 3. For those species with newly generated ITS and rps16 sequences, voucher information including species name, geographical origin, collector(s), vouchers (herbarium: TARI, Tehran Ariamehr Research Institute) is provided. Species names in brackets follow the taxonomic treatment proposed in the present study.
Aciphylla glacialis, KF160671; Aegopodium tribracteolatum, AY581783; Aethusa cynapium, AF110539; Aletes macdougalii, KF619604; Ammi trifoliatum, KJ473904; Ammodaucus leucotrichus, KT347642, KT347831; Annesorhiza macrocarpa, DQ368835; Anthriscus sylvestris, KT347715, KT347879; Aphanopleura capillifolia [Psammogeton capillifolium], DQ516368; Aphanopleura breviseta [Psammogeton brevisetum], Iran, Prov. Azarbaijan, ca 2 km from Tabriz to Bababaghi,1400 m a. s. l., Mozaffarian 71,290 (TARI), MZ687311, MZ706974; Aphanopleura leptoclada [Pimpinella leptoclada], Iran, Prov. Yazd, Mehriz to Tezerjan, Sand dunes around the roads, 1500 m a. s. l., Mozaffarian 77,439 (TARI), MZ687308, MZ706975; Aphanopleura trachysperma1, AF008629; Aphanopleura trachysperma2, Iran, Prov. West Azarbaijan, Maku, Shut, SE of Sufi village, 1070 m a. s. l., Mozaffarian 94,396 (TARI), MZ687310, MZ706976; Apium graveolens, AH003471; Arafoe aromatica1, AF077874; Arafoe aromatica2, U78383, U78443; Astrodaucus orientalis, MG098986, AF123748; Bunium allioides, JX312805; Bunium setaceum, EF544477; Bupleurum microcephalum, GU269882; Carum carvi, KF454471, FJ385182; Chamaesium paradoxum, EU236161; Choritaenia capensis, DQ368842; Conium maculatum, AF110546; Coriandrum sativum, HQ377205; Cryptotaenia africana1, DQ516371; Cryptotaenia africana2, DQ516370; Cymbocarpum wiedemannii, GU291352; Demavendia pastinacifolia, AY911857, AY911863; Demavendia pastinacifolia, Iran, Prov. Tehran, W Tehran, Suleghun valley, 1500–2000 m a. s. l., Assadi & Mozaffarian 32,621 (TARI), MZ687312, MZ706977; Ducrosia flabellifolia, DQ427051; Echinophora chrysantha, AF077883; Erigenia bulbosa, MG218514; Eryngium proteiflorum, EU070531; Ferula grigoriewii, KJ660784, KJ698392; Frommia ceratophylloides, DQ647630; Galagania neglecta, HM229391; Haussknechtia elymaitica1, EU169273; Haussknechtia elymaitica2, AY911859, AY911865; Heteromorpha involucrata, DQ368853; Komarovia anisosperma, AF077897; Lecokia cretica, EU169294; Lichtensteinia trifida, EU434683; Muretia lutea, DQ516359; Nothosmyrnium japonicum1, DQ516367, FJ385211; Nothosmyrnium japonicum2, EU236179; Oedibasis tamerlanii, HM229403; Opopanax armeniacus, Iran, Prov. East Azarbayejan: Mianeh, Bozgoush Mount. Neshagh village, 1880 m a. s. l., 2007, Mozaffarian 93,480 (TARI), MZ687309; Orlaya kochii, AH003483, AF123733; Ormopterum turcomanicum, GQ379313; Petroselinum crispum, GQ148800, AF110544; Phellolophium madagascariense1, DQ647627, KJ173509; Phellolophium madagascariense2, KJ173483, KJ173508; Physospermopsis cuneata, FJ385055, FJ385221; Physospermopsis shaniana, EU236192, FJ385225; Pimpinella acuminata, EU236193, FJ385226; Pimpinella affinis, AY581780; Pimpinella alismatifolia, FM986448; Pimpinella anisum1, KR150177, KJ173510; Pimpinella anisum2, KR150177; Pimpinella armena, KX982510; Pimpinella arguta, JF831512; Pimpinella aromatica, AY581784; Pimpinella aurea, AY581785; Pimpinella austriaca, KX982512; Pimpinella betsileensis1, DQ647626, KJ173511; Pimpinella betsileensis2, KJ173488; Pimpinella brachycarpa, AY548230; Pimpinella brachystyla, GQ379270; Pimpinella buchananii, FM986455; Pimpinella caffra, FM986447; Pimpinella candolleana1, MH117649, FJ385227; Pimpinella candolleana2, MH117649; Pimpinella cappadocica, AY581786; Pimpinella caudata, JF831514; Pimpinella chungdienensis, JF831515; Pimpinella confuse, KX982508; Pimpinella coriacea, JF831516; Pimpinella corymbosa, AY581787; Pimpinella cretica, AY581788; Pimpinella diversifolia1, JF831517; Pimpinella diversifolia2, KF806585; Pimpinella eriocarpa, AY581790; Pimpinella espanensis, MH377861; Pimpinella fargesii, JF831518; Pimpinella favifolia1, FM986458; Pimpinella favifolia2, FM986453; Pimpinella flabellifolia, AY581791; Pimpinella flaccida, JF831519; Pimpinella henryi, EU236195; Pimpinella heyneana, GQ379276; Pimpinella huillensis1, FM986454; Pimpinella huillensis2, FM986443; Pimpinella insignis, AY941280, AY941308; Pimpinella isaurica, AY581792; Pimpinella khayyamii, KX982523; Pimpinella kingdon-wardii, JF831520; Pimpinella kotschyana, DQ516373; Pimpinella krookii, FM986445; Pimpinella kyimbilaensis, FM986452; Pimpinella ledermannii, FM986457; Pimpinella lutea, DQ516374; Pimpinella major1, MH377862; Pimpinella major2, KX982513; Pimpinella nigra, KX982509; Pimpinella niitakayamensis, DQ516375; Pimpinella nudicaulis, AY581794; Pimpinella olivieri1, KX982506; Pimpinella olivieri2, KX922690; Pimpinella oliverioides, AY581795; Pimpinella oreophila, FM986450; Pimpinella peregrina1, AY581797; Pimpinella peregrina2, AY581797; Pimpinella peucedanifolia1, AY581798; Pimpinella peucedanifolia2, KX982524; Pimpinella procumbens, MH377863; Pimpinella puberula, AY581799; Pimpinella purpurea1, JF831521; Pimpinella purpurea2, EU236197; Pimpinella rhodantha1, KX982511; Pimpinella rhodantha2, KX982511; Pimpinella rhomboidea, JF831522; Pimpinella rigidistyla, FM986459; Pimpinella rockii1, JF831523; Pimpinella rockii2, FJ385057; Pimpinella rubescens, JF831524; Pimpinella saxifraga1, MH377864, DQ133876; Pimpinella saxifraga2, AY581801; Pimpinella serbica, KX982507; Pimpinella siifolia1, MH377868; Pimpinella siifolia2, MH377867; Pimpinella sintenisii, AY581802; Pimpinella smithii1, GQ379272, FJ385230; Pimpinella smithii2, JF831526; Pimpinella thellungiana, JF831527; Pimpinella tibetanica, JF831528; Pimpinella tragium1, MH377869; Pimpinella tragium2, MH377869; Pimpinella transvaalensis, FM986449; Pimpinella trifurcata, FM986446; Pimpinella valleculosa, JF831529; Pimpinella villosa, MH377871; Pimpinella yunnanensis1, JF831530, FJ385231; Pimpinella yunnanensis2, JF831530; Pleurospermum rivulorum, HQ824798; Postiella capillifolia, DQ422829, DQ422848; Prangos eriantha, MT250532; Prangos ferulacea, KX982519; Prangos lophoptera, KU168369; Psammogeton biternatum, AF164839, AF164864; Psammogeton canescens, MG827069; Psammogeton lamondiae, MG827073; Psammogeton ranunculifolius, MG920277; Psammogeton sp., MG827076; Psammogeton stocksii, Iran, Prov. Baluhestan, Between Iranshahr and Bam, Bazman, 1200 m a. s. l., Assadi 23,009 (TARI), MZ687313, MZ706978; Pyramidoptera cabulica, AF008631; Sanicula epipactis, EU169013, EU168959; Selinum broteri, AY179029; Smyrnium olusatrum, MK050086; Trachydium simplicifolium, FJ385067; Trachyspermum ammi, Iran, Prov. Baluchistan: Iranshahr to Chahbahar, Pole Jaligur over the Sarbay river, 300 m a. s. l., Mozaffarian 74,383 (TARI), MZ687314, MZ706979; Trachyspermum anethifolium [Psammogeton anethifolium], MG827079; Trachyspermum copticum, Iran, Prov. Baluchistan: Khash to Iranshahr, Dashte Abkhan, 1400 m a. s. l., Mozaffarian 72,711 (TARI), MZ687315, MZ706980; Trachyspermum microcarpum [Psammogeton microcarpum], MG920276; Trachyspermum paktianum [Psammogeton paktianum], MG827078; Trachyspermum papillare [Psammogeton papillare], MG920275; Trachyspermum roxburghianum [Psammogeton involucratum], MG827080; Trachyspermum scaberulum1, JF831531, FJ385258; Trachyspermum scaberulum2, EU236215; Trachyspermum triradiatum, EU236216, FJ385259; Zeravschania aucheri, AY911860, AY911866; Zeravschania membranacea1, AY911862, AY911868; Zeravschania membranacea2, Iran, Prov. Khorassan: Mashhad, Karde. All village, 1460 m a. s. l., Mozaffarian 87,007 (TARI), MZ687317, MZ706982; Zeravschania pauciradiata, Iran, Prov. East Azarbaijan, Siahrud to Kakaleh, near to Kalaleh, 390 m a. s. l., Ajani 1584 (TARI), MZ687316, MZ706981; Zeravschania regeliana, AY911861, AY911867.
Information on Electronic Supplementary Material
Online Resource 1. The alignment of nrDNA ITS sequences which has been used to reconstruct the ITS trees obtained in this study.
Online Resource 2. The alignment of plastid rps16 intron sequences which has been used to reconstruct the rps16 trees obtained in this study.
Rights and permissions
About this article
Cite this article
Mousavi, S.S., Mozaffarian, V., Mummenhoff, K. et al. Systematics of the Iranian genera Aphanopleura, Demavendia, Haussknechtia, Psammogeton, and Zeravschania (Apiaceae tribe Pimpinelleae). Plant Syst Evol 308, 2 (2022). https://doi.org/10.1007/s00606-021-01792-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00606-021-01792-x