Abstract
The morphological variation and cytotype diversity were investigated among Central European populations traditionally recognized as Dactylorhiza fuchsii, recently incorporated in D. maculata s.l. Flow cytometry was employed to assess the ploidy levels of 738 individuals from 77 localities and multivariate morphometrics for a total of 531 individuals from 27 localities. Three ploidy levels were found: diploid (2n = 2x = 40), DNA-triploid and tetraploid (2n = 4x = 80). Whereas diploids and tetraploids often occurred as pure-cytotype populations, individuals of DNA-triploids always co-occurred with at least one of the other cytotypes. Qualitative morphological traits were inferred to be the most important drivers of morphological variation among the investigated plants, with the most striking differences in flower colouration and leaf spotting. The combination of morphological and cytological characters enabled to delimit two separate groups of populations. The first corresponded to D. maculata subsp. fuchsii with morphologically indistinguishable diploid, DNA-triploid and tetraploid individuals, sometimes occurring in mixed-ploidy populations. A complex geographical pattern of cytotype distributions was observed, with diploids scatteredly occurring throughout Central Europe except for Bohemian Massif, which was dominated by tetraploids. The other group of populations represented newly described in this study D. maculata subsp. sooana, subsp. nova, morphologically well-defined and strictly diploid taxon with a restricted geographical range, occurring in the Western Carpathians. A new combination for a hybrid taxon D. × dinglensis nothosubsp. smitakii, comb. nova (= D. maculata subsp. sooana × D. majalis subsp. majalis), was also proposed.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The genus Dactylorhiza Nevski belongs to the taxonomically most complicated groups of the orchid family in Europe (Heslop-Harrison 1968; Reinhard et al. 1991; Pedersen 1998; Delforge 2006; Pillon et al. 2006). Frequent polyploidization, hybridization, and gene introgression have resulted in reticulate evolution and multiple origins of some of its taxa (Lord and Richards 1977; Hedrén 1996; Hedrén et al. 2001; Pillon et al. 2007; Nordström and Hedrén 2009; De hert et al. 2012; Balao et al. 2016; Brandrud et al. 2020). High morphological variation, phenotypic plasticity (Meyer 1968) and putative epigenetic changes (Paun et al. 2010) further complicated the reconstruction of the phylogeny and taxonomic inferences within this group. The biosystematics and evolution of the genus has recently been a subject of many investigations, with the main focus on the D. incarnata/maculata polyploid complex, which consists of three groups of taxa: the diploid D. incarnata group, the diploid and autopolyploid D. maculata group, and the allopolyploid taxa of the D. majalis/traunsteineri group (Hedrén 2001; Devos et al. 2003; Pillon et al. 2007; Hedrén et al. 2008; Nordström and Hedrén 2009; Naczk et al. 2015; Bateman et al. 2018).
A number of taxa have been recognized within the D. maculata group across its distribution range from Europe to East Asia (cf. Vermeulen 1947; Senghas 1968; Soó 1980; Delforge 2006; Eccarius 2016), but no consensus on taxonomic treatment has been introduced to date, and the number of currently recognized species ranges from three to 15 (cf. Buttler 2000; Delforge 2006; Eccarius 2016). Two species are traditionally recognized in Central Europe within the D. maculata group: D. maculata (L.) Soó (s. str.) and D. fuchsii (Druce) Soó (e.g. Soó 1980; Reinhard et al. 1991; Delforge 2006; Danihelka et al. 2012; Eccarius 2016). Both taxa were lectotypified by Vermeulen (1947); the type specimen of D. maculata was selected from Linné’s material collected in the surroundings of Uppsala, while the name of D. fuchsii is based on Druce’s collection from Wantage in Oxfordshire. They were distinguished based on their morphology (Druce 1915; Vermeulen 1947; Heslop-Harrison 1951; Gathoye and Tyteca 1987; Dufrêne et al. 1991; Tyteca and Gathoye 2003; Ståhlberg and Hedrén 2008): plants with narrow, acute leaves and broad labellum with a small and thin middle lobe were assigned to D. maculata, while D. fuchsii was characterized by broad, obtuse leaves and deeply three-lobed labellum with the wide and long middle lobe. Later, some differences were stated also in ecology (Heslop-Harrison 1951; Jagiełło 1988; Dufrêne et al. 1991; Ståhlberg 2009) and, above all, in chromosome numbers: diploids with 2n = 40 were considered D. fuchsii, while tetraploids with 2n = 80 were assigned to D. maculata (Heslop-Harrison 1951; Vöth and Greilhuber 1980; Averyanov 1982, 1990).
However, subsequent research disproved the correlation between morphology and ploidy levels of D. maculata group, particularly in Central Europe. Whereas D. maculata has always been found to be tetraploid, plants morphologically corresponding to D. fuchsii were reported to be either diploid or tetraploid (cf. Vermeulen 1968; Májovský 1976, 1978; Vöth 1978; Jagiełło and Lankosz-Mróz 1988; Měsíček and Javůrková-Jarolímová 1992; Krahulcová 2003). Moreover, the morphological differences between both taxa in Central Europe seem to be rather weak based on sparsely published data (Jagiełło 1988; Ståhlberg and Hedrén 2010; Kaplan et al. 2017). Therefore, many authors prefer to merge both these taxa into a single species D. maculata s.l. and recognize them as subspecies (e.g. Cauwet-Marc and Balayer 1984; Reinhard et al. 1991; Buttler 2000; Baumann et al. 2002; Ströhle 2003; GIROS 2009; Ståhlberg and Hedrén 2008; Naczk et al. 2015; Kurtto et al. 2019). This treatment also better reflects the genetic structure of the D. maculata group (Ståhlberg and Hedrén 2010). On the other hand, a recent molecular study of Brandrud et al. (2020) recognized D. maculata and D. fuchsii as two well-separated evolutionary lineages; their sampling in Central Europe was however scarce and did not include polyploid individuals of the latter taxon. It follows that D. fuchsii (D. *fuchsii from hereafter) has still an undefined taxonomic position within the D. maculata group and requires more detailed studies.
Considering all previous findings, it is obvious that Central European populations of D. *fuchsii are considerably variable both concerning morphological traits and ploidy levels (e.g. Delforge 2006; Eccarius 2016). However, little is known about the correlation between morphological variation and ploidy level, as well as the distribution patterns of particular cytotypes. This also applies to the most peculiar morphotype of white-flowering populations clearly derived from D. *fuchsii and sometimes recognized as Dactylorhiza fuchsii subsp. sooana Borsos, which is however an invalid name. This taxon was first mentioned from Northern Hungary (Borsos 1959) and nowadays is considered endemic to the Czech Republic, Slovakia, and Hungary (Kliment 1999; Vlčko et al. 2003). A brief description of this taxon provided by Borsos (1959, 1961) was supplemented by Batoušek (1995), referring to D. fuchsii subsp. sooana as possessing white flowers (with or without markings), white anther caps, and spotted leaves. Nonetheless, the range of morphological variation of this taxon overlaps with D. fuchsii subsp. fuchsii according to some authors (Borsos 1961; Soó 1980; Vlčko et al. 2003), and the delimitation of these taxa is thus complicated, which also causes taxonomic ambiguities. Kreutz (2004) recognized these two taxa as varieties of D. fuchsii subsp. fuchsii, while Eccarius (2016) listed D. fuchsii subsp. sooana just among synonyms of D. fuchsii. A population of D. *fuchsii, labelled as ‘sooana’, was also included in the analysis by Ståhlberg and Hedrén (2010) as D. maculata subsp. fuchsii, with a note that it may be classified into a lower taxonomic unit because of possible morphological and/or geographical distinctions; nonetheless, the distinctions have not been scrutinized. Even the ploidy level of this putative taxon is unknown, and though both diploids and tetraploids have been mentioned in literature, reliable data are missing (Kubát 2010). Moreover, D. fuchsii subsp. sooana has never been validly described, as Borsos (1959) did not state the type specimen along with the protologue, and the epithet ‘sooana’ has never been validated.
Several more taxa in various taxonomic ranks were recognized within D. *fuchsii in Central Europe (e.g. ‘longibracteata’, ‘meyeri’), but they are usually not accepted in recent literature (cf. Kubát 2010, Eccarius 2016). Besides D. fuchsii subsp. sooana, the only widely accepted taxon is D. fuchsii subsp. sudetica (Rchb.) Verm., often synonymized with D. fuchsii subsp. psychrophila (Schlecht.) Holub. (e.g. Procházka 1979; Ponert 2019), resp. D. fuchsii subsp. fuchsii var. psychrophilla (Schlecht.) Soó (e.g. Kubát 2010; Danihelka et al. 2012). These names are applied to plants of subtle habitus and strikingly coloured flowers, occurring in mountain regions of Central Europe (Devillers & Devillers-Terschuren 2000). However, it was shown that the populations from the Sudeten Mts are rather transitional between D. maculata s. str. and D. *fuchsii in their morphology (Jagiełło 1988), and only tetraploid chromosome numbers have been found in these plants (Jagiełło 1988; Krahulcová 2003). Therefore, they are often incorporated into D. maculata s. str., under the name of D. maculata subsp. sudetica (Rchb.) Vöth (e.g. Vöth & Greilhuber 1980; Jagiełło 1988; Eccarius 2016). The taxonomic riddle of this taxon must be solved in a larger taxonomic and geographical context.
Flow cytometry provides a rapid estimate of the ploidy level of large populational samples and may be considered a useful non-invasive method (Doležel et al. 2007; Loureiro et al. 2010). This method was employed to assess the cytotype diversity of Central European populations of D. *fuchsii (including ‘sooana’). Simultaneous analyses of cytogenetic and morphological variation allowed us to address the following questions: (1) What is the extent and structure of morphological and genome size (cytotype) variation within this group in Central Europe? (2) What are the morphological characters diagnostic for the ploidy levels (cytotypes)? Revealed patterns of morphological and cytotype diversity allowed us to make some taxonomical inferences which follow here.
Materials and methods
Plant material
Only populations morphologically corresponding to D. *fuchsii according to literature (Soó 1980; Delforge 2006; Eccarius 2016) were studied. Each population was further classified as belonging to informal groups, either ‘fuchsii’ or ‘sooana’ (not italicized). Populations consisting of plants predominantly (with at least 95% individuals) with spotted leaves, white flowers (both with or without markings), and white anther caps were classified as ‘sooana’, while all others were considered ‘fuchsii’, comprising plants with spotted or unspotted leaves, white to purple flowers and mostly purple anther caps (Batoušek 1995). Flow cytometric estimation of ploidy levels enabled further assignment of the populations belonging to the fuchsii group as ‘fuchsii-2x’, ‘fuchsii-3x’ and ‘fuchsii-4x’. In mixed ploidy populations, each ploidy level was analysed as a separate subpopulation. The sooana group was uniform in ploidy level, and any further division of the group was thus not applicable.
Plant material and data were collected in 2011–2018 from 77 localities in Central Europe (Online Resource 1), including Austria (11), Czech Republic (29), Germany (3), Hungary (5), Poland (6), Romania (3), Slovakia (16) and Slovenia (4). In total, 738 individuals were investigated for their DNA-ploidy levels (Suda et al. 2006). Morphological data were collected for 531 individuals from 27 populations (Online Resource 2). Preferably, individuals with estimated DNA-ploidy level were used for morphometric analysis. In some cases also other plants were used, but DNA-ploidy level was estimated from a representative number of other plants in the same population, and the population must have shown to be pure-cytotype. Because of the conservation status of the studied taxa, herbarium vouchers were usually not collected; instead, a series of photographs was taken for most of the individuals used in the morphometric analysis and stored in archive of the first author.
Analyses of chromosome numbers, DNA-ploidy levels and genome sizes
Number of chromosomes was established from chromosomal spreads prepared from microspores (haplophasic chromosome number, n). Flower buds were collected in the field ca 10 days before blossoming, fixed in acetic acid: ethanol (1: 3) and stored at –20 °C until processed. Standard protocol of Feulgen staining was used to stain the tissue (Weiss et al. 2003). Briefly, flower buds were hydrolyzed in 5 N HCl for 30 min in 20 °C, washed with water and stained with Schiff’s reagent (Sigma, Vienna, Austria) for 1–2 h in darkness. The anthers were dissected and squashed in a drop of 60% acetic acid. Chromosome spreads were analysed under 1000 × magnification using Axioplan light microscope (Carl Zeiss, Vienna, Austria).
DNA-ploidy level was estimated by flow cytometry (FCM) following a standard protocol with internal standards (Doležel et al. 2007) and ploidy level was assessed based on calibration with plants for which chromosome numbers were counted. Pisum sativum cv. ‘Ctirad’ (2C = 9.09 pg; Doležel et al. 1998) was used as the internal standard for diploids and tetraploids, and Zea mays cv. ‘CE-777’ (2C = 5.43 pg; Lysák and Doležel 1998) for DNA-triploids. Fresh ovaries of Dactylorhiza were used for the analysis because the vegetative plant tissues (typically leaves) may provide erroneous results due to more prominent occurrence of progressively partial endoreplication (PPE; Trávníček et al. 2015), alternatively mentioned as strict partial endoreplication (Brown et al. 2017).This is a specific process of DNA endopolyploidization characteristic for the orchid family, leading to a disproportional increase in nuclear DNA content of somatic cells, including those of ovaries, which however contain sufficiency of non-replicated nuclei, yielding to 2C peaks in FCM analysis (Trávníček et al. 2015; Hřibová et al. 2016). Ovaries were collected in the field and stored in wet paper tissue in 4 °C until processed, typically up to 5 days, but no more than 10 days. In the laboratory, one or two ovaries and 0.5 × 0.5 cm of internal standard tissue were co-chopped using a razor blade (Galbraith et al. 1983) in a Petri dish in LB01 buffer with PVP (Doležel et al. 2007). The nuclei solution was filtered through the 40 µm nylon mesh and stained with 30 µl of either 4,6-diamidino-2-phenylindole (DAPI; 4 µg/ml) or propidium iodide (PI, 50 µg/ml). In the analysis with PI, 30 µl of RNase was added to the sample to digest the RNAs.
The analysis was conducted with the following instruments: BD Accuri C6 (BD Biosciences, San Jose, CA, USA); Partec Cy Flow ML (Partec GmbH, Münster, Germany), both at the Department of Botany, Palacký University Olomouc; Partec Cy Flow ML at the Department of Botany and Biodiversity Research, University of Vienna; and Partec Cy Flow ML at the Institute of Experimental Botany, Olomouc. Each individual was analysed separately and the fluorescence of at least 3,000 particles was recorded. Only results with peak CV ≤ 5.0 were accepted. Several diploid and tetraploid individuals were analysed with both PI and DAPI to calibrate the position of the peaks for the different dyes used.
BD Accuri software and Partec FloMax software were used to evaluate the histograms with two or more (because of frequent endoreplication) peaks. The G0/G1 peak of the standard and G0/G1 peak (2C-peak after Trávníček et al. 2015) of the analysed plant were identified. For every plant, an index (relative genome size) was calculated as the ratio of the mean G0/G1 peak of the Dactylorhiza / mean G0/G1 peak of the internal standard. The ratios obtained from the analysis using Z. mays as the standard were recalculated to the values expected from the measurement with P. sativum.
Absolute genome size was measured for several plants, using a similar protocol as for DNA-ploidy level estimation with the following settings: suspension was stained with PI, each plant was measured three times, and at least 3,333 nuclei were analysed in each measurement with a maximum peak CV = 3.5%. The peak ratios obtained for each plant were averaged and the genome size was calculated as the average peak ratio multiplied with the genome size of the internal standard.
Morphological data recording and analyses
Twenty-four morphological characters were measured (16 characters), numbered (four characters) or scored (three binary characters and one multistate character) (Tables 1 and 2; Online Resource 3). Characters studied included morphological characters traditionally used in the determination keys and special taxonomic literature for delimitation of various Dactylorhiza taxa as well as characters found useful in our preliminary screening of Central European populations of D. maculata group. Vegetative traits were measured with an adjusted ruler on living plants directly in the field, to minimize the damage of the individuals. Floral traits were measured from a digital picture. For each individual, one flower from the middle-low part of the inflorescence (typically the 4th flower from the bottom) was removed. The lip was separated, put on the scanner glass, and weighted down with a microscope slide; this led to flattening of the lip, which was subsequently digitized by a scanner with high resolution (1200 dpi). ImageJ software (Schneider et al. 2012) was used for the size measurement of the traits. Besides the primary traits, 15 additional ratios and indices were derived from primary traits for further analyses.
In total, 531 individuals from 27 populations of D. *fuchsii were included in the morphometric analyses. Several datasets were used: (1) matrix 1 – complete dataset including all 531 individuals as OTU and all primary and derived characters; (2) matrix 2 – complete dataset including all 531 individuals as OTU and reduced set of characters. Specifically, two primary characters (in1, in2) and 5 ratios derived from these characters (plH_in1, plH_in2, in2_in1, lL1_in1, lL2_in2) were excluded from the dataset due to the absence of their records for some populations. Problem of multicollinearity was assessed by variance inflation factor (VIF) for quantitative traits using the library usdm (Naimi 2017) in R (R Foundation for Statistical Computing, Vienna, Austria). Only those variables were retained in the analyses whose VIF was lower than 15, which is slightly higher than the recommended VIF ≤ 10 (O'Brien 2007). Consequently, six primary quantitative characters were excluded (plH, wL1, lL2, A, C, E). The potential problem of multicollinearity in categorical characters was accessed by Cramer’s V (Legendre and Legendre 2012). Only one variable (pPe) had Cramer’s V higher than 0.9 in two paired analyses (with pAc and cLaW) and therefore it was excluded from the dataset; (3) matrix 3 – complete dataset including all 531 individuals as OTU and reduced set of characters. Only quantitative characters and their ratios identical to those in matrix 2 were considered. All nominal variables, including those considered as diagnostic for the sooana group, were excluded from the matrix; (4) matrix 4 – a dataset with 27 population samples as OTU characterized by the population’s median values of quantitative characters and their ratios and proportional representation of each category for each studied categorical variable per each population. After excluding the collinear variables with VIF ≥ 15, just 13 variables remained as follows: plH/in1, in2/in1, lL1/in1, lL2/in2, pLeP, pLeB, pLaA, pLaP, pAx, cLaB, cLaP, LAS1, LAS2.
To compare groups (as defined above), the matrix 1 was firstly analysed using univariate statistics. Nested ANOVA with populations nested within groups and Tukey multiple comparison test were used for quantitative characters and their ratios using NCSS 9 (NCSS, LLC., Kaysville, Utah, USA, ncss.com/software/ncss). Bonferroni correction of P-values of ANOVAs was applied additionally. Before statistical tests, some quantitative characters were log-transformed to improve their normality. Descriptive statistics based on the original (untransformed) values are presented in tables and visualized in plots.
Generalized linear mixed model (GLMM) with the logit link function and binomial distribution was used for the analyses of binary characters. In the GLMM, a group was considered a fixed factor and a population a random factor nested within groups. For GLMM, the lme4 library (Bates et al. 2019) and afex library (Singmann et al. 2016) in R were used. LRT test was used for the estimation of significance level and emmeans library (Lenth et al. 2020) was used for multiple comparisons between groups using Tukey method with P value adjustment. Due to convergence problems when using GLMM with multinomial distribution of multistate categorical characters in Statistica 10 software (StatSoft, Inc., Tulsa, USA) using population as a nested random factor, log-linear models were calculated instead, using likelihood ratio χ2 test on pooled data (i.e. ignoring population identity within each group) in NCSS 9. After significant overall χ2 test, separate χ2 tests were done for each pair of groups and P-values were adjusted using Bonferroni correction. Small value (0.2; i.e. delta value) was added to each cell count when 0's were present in the table.
Principal coordinate analysis (PCoA), using a Gower’s dissimilarity coefficient for mixed data consisting a mixture of quantitative, count and qualitative characters (Legendre and Legendre 2012), was used to obtain insight into the phenetic relationships among all studied individuals (matrix 2). Principal component analysis (PCA) was done based on the correlation matrix of the quantitative characters (matrix 3) to observe the structuring of individuals in the ordination space based on the quantitative characters. A third analysis (PCA) was performed on matrix 4 containing populations as OTU. Before multivariate analyses, some quantitative characters were log-transformed. PCoA and PCA were run using the software Canoco 5.12 (ter Braak and Šmilauer 2012).
Results
Cytotype diversity and population composition: chromosome numbers and flow cytometry
Chromosome numbers were obtained for six individuals from three populations (Online Resource 1). Three plants were diploids (n = x = 20), with one individual representing the fuchsii-2x group (pop. 28, Furth an der Triesting) and two individuals representing the sooana group (pop. 4, Hluboče; Fig. 1a). The other three plants were tetraploids (n = 2x = 40), belonging to a single population (27, Alland; Fig. 1b) and assigned to the fuchsii-4x group. Peak ratios for all of these reference individuals are shown in Online Resource 4.
One to 35 plants per (sub)population (mean ± SD; 8.2 ± 6.8) were analysed by FCM, accounting for a total of 738 plants representing 90 (sub)populations from 77 localities. Three DNA-ploidy levels were identified, corresponding to diploids, tetraploids, and a cytotype with a relative genome size intermediate between that of diploids and tetraploids, referred to as DNA-triploid (Suda et al. 2006). PPE was frequently observed. In leaf tissue, the non-replicated (2C) nuclei of tetraploids were detected, but endoreplicated (2C + P) nuclei prevailed in diploids, for which 2C peaks were not detectable on the FCM histograms. To avoid erroneous results, ovaries were used for all FCM analyses.
Within the analysed 140 individuals of the sooana group, only diploid plants were found. All three cytotypes were found in the fuchsii group. A majority of these plants corresponded to tetraploids (373; 62.4%), followed by diploids (196; 32.8%) and DNA-triploids (29 individuals; 4.8%). Most of the analysed populations of the fuchsii group (85.9%) were uniform in terms of ploidy level: 23 populations were exclusively diploid (37.7%) and 34 populations were tetraploid (55.7%). Only four mixed-ploidy populations were found in which diploids and tetraploids co-occurred with DNA-triploid individuals (30, Nasswald; 37, Weissenbach; 55, Zajačkova lúka; 74, Kramplje). DNA-triploids were also sporadically found as a minority cytotype in four predominantly diploid (8, Ransko; 14, Zakopane; 31, Fronbach; 65, Pârâul Rece) and one tetraploid (36, Postalm) populations. A higher proportion of DNA-triploids (5 out of 8 individuals) was found in only one population (74, Kramplje) comprising all three cytotypes.
Significant differences in relative genome size were found between all pairs of groups (Welch's test of means allowing for unequal variances; DAPI: F3, 47.8 = 8829.0, P < 0.001; PI: F3, 58.0 = 6677.5, P < 0.001), except for the sooana and fuchsii-2x groups with nearly the same genome size (Table 3). The genome size of polyploids was not additive compared to their diploid relatives. The average monoploid relative genome size of tetraploids corresponded to 88% of that of diploids, and that of DNA-triploids was exactly intermediate between the average monoploid relative genome sizes of diploids and tetraploids. PI and DAPI measurements yielded consistent results (Table 3).
Absolute genome size was measured for five plants from two populations. Two individuals were diploids classified as fuchsii-2x (28, Furth an der Triesting), and three individuals were tetraploids classified as fuchsii-4x (27, Alland). The absolute genome size of diploids was estimated to be 2C = 6.55 and 6.64 pg, while the absolute genome size of tetraploids ranged from 2C = 11.89 to 12.22 pg (Online Resource 5). Chromosome number of n = x = 20 was counted for the diploid plant with 2C = 6.55 pg.
Morphological variation of Dactylorhiza *fuchsii populations
Only 13 out of 31 quantitative characters (42%) were significantly different at least between some of the groups (Table 1 and Online Resources 6). The majority of characters differing between groups were those recorded on flowers (A, B, C, E) or represented ratios (HH, A/D) derived from floral traits. The second set of characters differing among groups were related to plant habit, i.e. the length of internodes (in1, in2) and their ratios with plant height and length of leaf (e.g. plH/in1, plH/in2). However, just two characters (in2 and IL2/dBW) remained significant after the application of Bonferroni correction (Table 1 and Online Resource 6).
Every binary character studied showed significantly different patterns at least between some groups (Fig. 2 and Table 2). Most plants of all three groups were without pigmentation on the inflorescence axis. Only fuchsii-4x plants had more frequently dark anthocyanin pigmentation on the inflorescence axis compared to the sooana group. The majority (94%) of sooana plants did not have pigmentation on the perianth (excluding labellum), while the majority (90%) of fuchsii-2x and fuchsii-4x individuals had. Similarly, almost all fuchsii-2x and fuchsii-4x plants had anthocyanin pigmentation on the anther cap, while most of the sooana plants had anther caps without pigmentation.
Stacked bar charts of eight qualitative characters in studied groups. Vertical axes represent proportions. The abbreviations of the characters see in Table 2
Frequency distributions of the categories of every multistate categorical variable differed significantly among groups (Fig. 2 and Table 2). Intensity of spots on leaves increased in the direction fuchsii-2x → fuchsii-4x → sooana. While approximately half of the plants (47%) of the fuchsii-2x group were without spots on the leaves, 71% of sooana plants had bold spots on leaves. More than 75% of both fuchsii-2x and fuchsii-4x plants had bold labellum markings, while the sooana group had almost equal frequencies of plants with either bold or pale labellum markings. The sooana group also differed from both fuchsii groups in labellum colour, having a white labellum in most plants (97%), while both fuchsii-2x and fuchsii-4x groups had similar proportions of plants of three colour categories, with only predominantly purple labellum plants. All groups also differed in the shape of leaf apexes. Just a minority of plants in all groups possessed an acute leaf apex, with the highest proportion of such plants found in the fuchsii-4x and lowest in the sooana group.
The PCoA based on quantitative and qualitative characters (matrix 2; Fig. 3a, b) revealed a near complete separation of the fuchsii-4x and sooana groups along the first ordination axis, with just some fuchsii-4x individuals situated within the sooana cluster; most of these individuals belonged to one population (32, Giesshübl). On the other hand, the clump of fuchsii-2x individuals overlapped with the fuchsii-4x clump on the left part of the ordination diagram. Some fuchsii-2x individuals from two populations (1, Smutné udolí; 14, Zakopane) occurred in the right part of the ordination diagram where they overlapped with the sooana group (Fig. 3a). The observed pattern in the distribution of the groups along the first axis was almost completely caused by several qualitative characters related to labellum and anther cap colour and labellum marking. All these characters are tightly correlated with the first axis (Fig. 3b): cLaW (point biserial correlation coefficient; r = 0.67***), pLaP (0.39***), pAc (-0.75***), pLaB (-0.47***), and cLaP (-0.56***). It follows that the resemblance of some individuals of the fuchsii-2x, fuchsii-4x and sooana groups was due to sharing some of the diagnostic traits of the sooana group, particularly white flowers. Other characters, including all quantitative ones, did not significantly correlate with the first ordination axis; only some characters were related to the second ordination axis, suggesting phenotypic variation in size regardless of group identity (Fig. 3b).
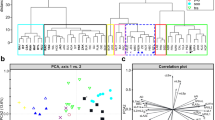
Results of multivariate analyses of morphological characters of Dactylorhiza *fuchsii plants. a, b Principal coordinate analysis based on 32 quantitative and qualitative characters (matrix 2) with individual plants as OTUs. The first and second ordination axes explained 17.3% and 11.3% of the total variation, respectively. Characters, of which the larger absolute value of the two correlations with the ordination axes exceed 0.3, were shown in the diagram. c, d Principal component analysis based on 22 quantitative characters (matrix 3) with individual plants as OTUs. The first and second ordination axes explained 23.4% and 17.2% of the total variation, respectively. Characters, whose individual fit on both displayed axes exceed 10%, were shown in the diagram. e, f Principal component analysis based on 15 characters with populations as OTUs (matrix 4). The first and second ordination axes explained 30.4% and 14.0% of the total variation, respectively. Characters, whose individual fit on both displayed axes exceed 10%, were shown in the diagram. Symbols: fuchsii-2x – empty circle, fuchsii-4x – black circle, sooana – cross. The abbreviations of the characters see in Table 1 and Online Resource 3, the codes of populations see in Online Resource 1
After the removal of qualitative characters, incl. diagnostic traits of the sooana group, from the dataset (matrix 3), the PCA based on 22 quantitative characters (incl. their ratios) revealed no morphological differentiation among groups (Fig. 3c). Main gradient along the first axis was correlated with the size dimensions of the labellum and leaf width, irrespective of group identity (Fig. 3d).
The PCA based on a reduced set of 13 characters representing populations as OTUs (matrix 4) revealed a pattern of group distribution in the ordination space (Fig. 3e, f) similar to that in the PCoA analysis of matrix 2. The sooana group was nearly completely separated from the remaining groups; only two populations of fuchsii-2x (1, Smutné údolí; 14, Zakopane) were situated in an intermediate position between the sooana clump and fuchsii-2x clump. Both the fuchsii-4x and fuchsii-2x groups partly overlapped in the centre of the ordination diagram, but fuchsii-4x group also showed considerably higher variability in the multivariate space than the fuchsii-2x group. Scores of populations along the first axis were significantly correlated with the following variables: pLaP (Pearson r = 0.73***), pLeB (0.58***), cLaP (-0.57**), plH/in1 (-0.78***), lL1/in1 (-0.71***) (Fig. 3f). Population 32, Giesshübl together with population 35, Sittersdorf were situated in the upper left part of the ordination diagram, in rather isolated positions from all remaining populations (Fig. 3e).
Distribution and cytogeography of the groups
Populations of the fuchsii group exhibited a clear geographical pattern in the distribution of their cytotypes throughout Central Europe (Fig. 4). Tetraploid populations (fuchsii-4x) prevailed in the Bohemian Massif, with only a single diploid population (58, Ranský brook) and one mixed-ploidy population with diploids and DNA-triploids (8, Ransko) found in this region (the Žďárské vrchy Mts). Solely three purely diploid populations (15, Tanew; 64, Cisnădioara; 66, Cheia) and one mixed-ploidy population of diploids with a single DNA-triploid plant (65, Pârâul Rece) were found in the Carpathians and peri-Carpathian region east and southeast of the Tatra Mts in Slovakia. The western half of the Western Carpathians, the Eastern Alps and Dinarides proved to be a transitional zone, where pure diploid, pure tetraploid, and mixed-ploidy populations containing all three cytotypes (30, Nasswald; 37, Weissenbach; 55, Zajačkova lúka; 74. Kramplje) were found. Several uniformly diploid populations (4, 5, 33, 47, 48, 49, 50, 51, 52, 53, 54, 59, 60; see Online Resource 2) corresponding to the sooana group were found in the Carpathian area of Northern Hungary, Southern Slovakia and Southeastern Czechia.
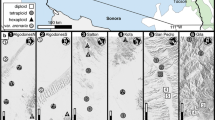
Groups and cytotypes of Dactylorhiza *fuchsii as recognized in this study. a Map of populations analysed by flow cytometry and proportions of diverse groups occurring at common localities: green = sooana, yellow = fuchsii-2x, red = fuchsii-3x, blue = fuchsii-4x. Symbol size is proportional to the sample size. Examples of plants belonging to different groups: b sooana (49, Mátraszentimre); c fuchsii-2x (58, Ranský brook); d fuchsii-3x (55, Zajačkova lúka); e fuchsii-4x (7, Adamova rokle). Photographs: V. Taraška
Discussion
A considerable cytotype diversity and morphological variation were found among populations of D. *fuchsii in Central Europe. Three cytotypes were identified, diploids, DNA-triploids and tetraploids. The distribution of these cytotypes was not even throughout Central Europe and regional cytotype diversity differed. The most important variation of phenotype concerned flower colouration and leaf spotting. Combination of morphological and karyological data allowed reliable delimitation of the fuchsii and sooana groups as two well-defined taxa.
Chromosome numbers and genome size
Three cytotypes were detected among populations of D. *fuchsii in Central Europe, diploid, DNA-triploid, and tetraploid. Chromosomal spreads confirmed the previously reported chromosome numbers, i.e. 2n = 2x = 40 for diploids and 2n = 4x = 80 for tetraploids (Heslop-Harrison 1951; Vöth and Greilhuber 1980; Jagiełło and Lankosz-Mróz 1988; Averyanov 1990; Amich et al. 2007). Progressively partial endoreplication (Bory et al. 2008; Trávníček et al. 2015) as well as genome downsizing in polyploids (Leitch and Bennett 2004; Parisod et al. 2010) occurred frequently. Relative genome size of some DNA-triploids was quite similar to the lowest values measured for plants considered to be tetraploids. However, DNA-triploids always co-occurred with plants of other cytotypes and their average relative genome size corresponded to the presumptive triploid genome size from the respective locality (with both PI and DAPI). Therefore, the three cytotypes were clearly distinguishable even despite some intracytotype variation of relative genome sizes.
This paper presents the first extensive ploidy level screening using FCM in the D. maculata s.l. taking into consideration the methodological task of PPE. Genome size, either absolute or relative, of the D. maculata group in Northern Europe has previously been investigated by using Feulgen-densitometry (Aagaard et al. 2005) and FCM (Ståhlberg and Hedrén 2008). However, FCM analyses relied on leaf tissue which could potentially obscure the results due to the incidence of PPE and should be regarded with caution (cf. Trávníček et al. 2015). The FCM estimate of the genome size of the D. maculata s.l. using ovaries was first presented by Šmarda et al. (2019), who analysed a single plant designated as D. fuchsii from the Hrubý Jeseník Mts (Bohemian Massif, Czech Republic), which was considered diploid, although its chromosomes were not counted. The genome size of this plant was estimated to be 2C = 10.83 pg, which is just a slightly lower value than the lowest estimates for tetraploids in the present study (2C = 11.89 pg), as well as the genome size of the tetraploid D. maculata investigated by Aagaard et al. (2005; 2C = 11.32 pg). Considering the lower estimates for internal standards by Šmarda et al. (2019), compared to Doležel et al. (1998) followed in this study, it may be suggested that the plant used in their analyses was rather tetraploid.
Morphological variability
Morphology may be strongly influenced by environmental factors, ontogenetic developmental stages, or interspecific interactions in orchids (Bateman and Denholm 1988, 1989). Similarly, the major part of the morphological variation among analysed groups recognized within D. *fuchsii is likely to be connected to environmental factors and the impact of local selection pressures, particularly concerning several quantitative traits of the flowers. High variation in floral traits in many orchid species is a consequence of a deceptive pollination system (Ackerman et al. 2011), where spatially and temporally variable selection pressures related to different pollinators or negative frequency-dependent selection (Gigord et al. 2001) or even non-adaptive processes (Vereecken and Schiestl 2009) might promote the persistence of phenotypic variance in floral traits (Ackerman et al. 2011). Flower characters were hypothesized not to be correlated with the phylogeny of the genus Orchis s.l. (Aceto et al. 1999), and these traits alone are probably unsuitable for taxonomic conclusions even in the genus Dactylorhiza. They may, however, be considered if they are correlated with other characters, ecological preferences, and/or patterns of geographical distribution (Pedersen 2009).
The most striking morphological differences were found between the sooana and fuchsii groups, the latter comprising both fuchsii-2x and fuchsii-4x individuals. These differences were connected to several qualitative traits related to leaf spotting and flower colouration, characters that were used for the classification of groups in this study. Importantly, these characters were also drivers of the main gradient of morphological variability among the analysed individuals as well as populations. There was just a slight overlap between the sooana group and the cluster formed by fuchsii-2x and fuchsii-4x plants in the PCA diagram based on individuals. This was caused by the presence of several albinotic individuals within both the fuchsii-2x and fuchsii-4x groups, which were similar to the sooana group in the flower colouration. Unlike sooana, such albinotic plants, however, lacked bold spots on their leaves. Furthermore, the sooana group was well-separated from the other groups in the PCA diagram based on populations. Therefore, the sooana group represented the most distinct, morphologically well-defined group within D. *fuchsii and it showed considerable dissimilarity from diploid as well as tetraploid fuchsii groups.
The fuchsii-2x and fuchsii-4x groups were similar to each other in their morphology. It was shown that autopolyploids in general may differ from their diploid progenitors in quantitative morphological traits; being more robust, possessing larger flowers, leaves, and stems (Parisod et al. 2010; Spoelhof et al. 2017). Only a few quantitative differences were detected between the diploid and tetraploid fuchsii groups, and significance was proved for just a single quantitative trait (length of the 2nd internode) after application of Bonferroni correction. Instead, the most apparent differences between these two groups were found in qualitative traits, i.e. leaf and labellum pigmentations. Diploid plants often lack spots on the leaves and their flowers are pale, with less conspicuous or even absent markings. Individuals with bold leaf spots and striking anthocyanin pigmentation of flowers are much more frequent among tetraploids. Notably, the intensity of leaf spotting is clearly correlated with the intensity of flower pigmentations in individuals of both fuchsii-2x and fuchsii-4x groups.
Populations of the fuchsii-4x group comprise larger morphological variability than those of the fuchsii-2x group. Some of the morphological differences observed between diploids and tetraploids may be also caused by putative gene introgression among tetraploid D. *fuchsii and other tetraploid taxa of the D. maculata group, as it was suggested by Jagiełło (1988) and later indicated by molecular markers (Ståhlberg and Hedrén 2010; Brandrud et al. 2020). Gene admixture could occur to various extents in tetraploid populations of D. *fuchsii, which may verge to D. maculata s. str. in some morphological traits. Such a process may have affected the Heslop-Harrison index, which is slightly lower in the fuchsii-4x group, or the shape of the leaf apex, which is more frequently acute in the fuchsii-4x group compared to fuchsii-2x. Genetic structure of these tetraploids therefore requires further investigation.
Cytotype diversity and cytogeography
Diploid populations were found mainly in the Carpathians, Alps, and Dinarides, which is in congruence with previous karyological reports (e.g. Skalińska et al. 1957; Groll 1966; Vaucher 1966; Löve 1971; Májovský 1978; Vöth and Greilhuber 1980; Jagiełło and Lankosz-Mróz 1988; Uhríková 2007). Diploids have also been mentioned from Bohemian Massif (Potůček 1969; Kubát 2010), but most populations of D. *fuchsii from this region analysed in the current study were tetraploid. Žďárské vrchy Mts are the only region within the Bohemian Massif where a diploid population (58, Ranský brook) has been confirmed to date. Diploids were also reported from the vicinity of Jagniątków in the Karkonosze Mts (Poland; Jagiełło and Lankosz-Mróz 1988), but this population (9, Jagniątków) was shown to be tetraploid in the current analysis.
Diploid populations were found in both the fuchsii and the sooana groups. Unlike fuchsii, the sooana group was exclusively diploid. Both groups were also largely geographically separated: the sooana group was found in the southern part of Western Carpathians (i.e. Northern Hungary, Southern Slovakia, and the White Carpathians in the Czech Republic), while populations in other regions corresponded to the fuchsii group. The distribution areas of both groups slightly overlapped in Northwestern Slovakia. On the other hand, at least some literature records of D. fuchsii from Hungary may represent the sooana group, depicted under this name in the Atlas of Hungarian Orchids (Molnár et al. 2011). The sooana group is also the only one found in Hungary during our field survey.
Tetraploid populations were widespread in Bohemian Massif, as well as in the Alps and Western Carpathians, where they reached Tatra Mts as the easternmost region. Despite D. *fuchsii has been considered exclusively diploid by many authors (Heslop-Harrison 1951; Vöth and Greilhuber 1980; Kubát 2010), tetraploids were reported repeatedly (e.g. Jagiełło and Lankosz-Mróz 1988, Měsíček and Javůrková-Jarolímová 1992, Bertolini et al. 2000) from this area. Ståhlberg and Hedrén (2010) suggested that tetraploid populations of D. *fuchsii were geographically limited to Central Europe, which may be explained by the relatively recent origin of this evolutionary lineage, dated to Holocene. Nevertheless, sporadic records of tetraploid individuals were also published from Pyrenees (Cauwet-Marc and Balayer 1984) and Apennines (Bertolini et al. 2000), which points to ongoing recurrent polyploidization.
DNA-triploids together with diploid and/or tetraploid individuals, were found in the Western Carpathians (i.e. Northwestern Slovakia), the Eastern Alps (Austria) and the Northern Dinarides (Slovenia), which are putative contact zones between the diploid and tetraploid lineages of D. *fuchsii. They were also rarely found in the Žďárské vrchy Mts, where diploids and tetraploids also co-occur. Furthermore, DNA-triploids were found within a diploid population (65, Pârâul Rece) in Southern Carpathians, where tetraploids were not recorded. DNA-triploids always co-occurred with other cytotype(s) and never formed a uniformly DNA-triploid population. Two different processes may have led to the establishment of ploidy-heterogeneous populations: (1) triploid formation within diploid populations via unreduced gamete formation in diploid individuals or (2) secondary contact of individuals of different ploidy levels (diploids and tetraploids) resulting in occasional hybridization giving rise to triploids (cf. Ramsey and Schemske 1998; Kolář et al. 2017; Popelka et al. 2019a, 2019b), which was observed in D. maculata s.l. in Scandinavia (Ståhlberg 2009). The DNA-triploids in the current study may have originated by either of these ways.
Taxonomic consequences
Using various approaches (morphological traits and ploidy level estimation) allows to delimit two groups of populations, representing two different taxa. The first consists of morphologically indistinguishable populations of fuchsii-2x and fuchsii-4x, but the fuchsii-3x group may be obviously included too, although its morphology was not evaluated. The other group comprises the populations here classified as the sooana group. These groups differ from each other in phenotypic variation, cytotype diversity and distribution patterns, but probably also in ecology, as populations of the sooana group are able to occupy more mesic habitats and avoid acidic substrates (V. Taraška et al., pers. observ.). Regarding all distinctions between these taxa, the rank of subspecies seems to be the most appropriate for them.
In the traditional view, they should be recognized as two subspecies of D. fuchsii. However, the taxonomic concept used by Scandinavian authors (Hedrén et al. 2001; Ståhlberg and Hedrén 2010) seems to be more appropriate, incorporating D. *fuchsii into the broadly interpreted species D. maculata s.l. Unlike the concept of two separate species, D. maculata s. str. and D. fuchsii, this approach is rather conservative and is applicable in the whole distribution area of both taxa, including Central Europe where they tend to merge secondarily. Consequently, the correct name for the subspecies represented by the fuchsii-2x, -3x and -4x groups is D. maculata subsp. fuchsii (Druce) Hyl. The other taxon, comprising populations of the sooana group, is being mentioned under various names based on the basionym Dactylorhiza fuchsii subsp. sooana Borsos (e.g. Vlčko et al. 2003; Kreutz 2004; Kubát 2010) and its taxonomic reassessment is discussed below.
Dactylorhiza maculata subsp. fuchsii is widely distributed throughout Europe and it includes diploids, DNA-triploids, and tetraploids. Nevertheless, these could be hardly classified as separate taxa, as they do not differ in morphology nor ecology, and they often co-occur. Furthermore, DNA-triploids may be involved in bidirectional gene exchange between diploids and tetraploids (Thórsson et al. 2001; Ståhlberg 2009). Relatively frequent occurrence of DNA-triploid individuals within diploid populations also indicates a recent polyploidization. Coexistence of multiple cytotypes should be regarded as a hidden intrapopulation diversity, with serious evolutionary potential and conservation importance (Soltis et al. 2007). Cytotype variation should be considered besides the population size when setting conservation priorities, as it was stated also for the closely related genus Gymnadenia (Trávníček et al. 2011). High cytotype diversity of D. maculata subsp. fuchsii was detected mainly in the Western Carpathians, Eastern Alps, and Northern Dinarides. These regions are situated in the contact zone of diploid and tetraploid lineages (Ståhlberg and Hedrén 2010; Eccarius 2016). The Žďárské vrchy Mts must be regarded as one of the diversity hotspots of D. maculata subsp. fuchsii in the Bohemian Massif, because it is the only known place in that area where all three cytotypes co-occur. High morphological variability of Central European populations may be partly a consequence of recent or former hybridization and gene introgression between D. maculata subsp. maculata and D. maculata subsp. fuchsii at the tetraploid level (Ståhberg and Hedrén 2010). Genetic structure of tetraploid populations of D. maculata subsp. fuchsii therefore needs further investigation.
Several taxa are often mentioned to be derived from D. *fuchsii in Central Europe. Their taxonomic value as well as position within D. maculata s.l. however, remains unclear. Tetraploid plants from the population 12, Velká kotlina, are usually assigned to D. fuchsii var. psychrophila (Schltr.) Soó (cf. Kubát 2010; Bureš 2013; Kaplan et al. 2017). This name, however, relates to diploid taxon described from Northern Europe (Vermeulen 1947; Eccarius 2016). Taxonomic evaluation of this population thus requires a wider geographical and taxonomical context. Another noteworthy tetraploid population was that of the locality Giesshübl (32), which is locus classicus of the unclear taxon D. maculata subsp. austriaca Vöth. Although it was subordinated to D. maculata s. str. because of its tetraploid chromosome number, even the protologue admits that this taxon is morphologically close rather to D. *fuchsii (Vöth 1978). The most striking morphological characteristic of this population is a high proportion of individuals with low pigmentation of both flowers and leaves. Hypochromic individuals can be often found in populations of D. maculata subsp. fuchsii, although usually not in such a high proportion (Bateman and Denholm 1988; Pikner 2012). Locality Giesshübl consists of two small meadow enclaves in the forest, and the population is probably reproductively isolated. Various evolutionary processes, including stochastic events, could lead to increase in the number of the hypochromatic plants (Narbona et al. 2017). Recently, this taxon is usually not accepted (cf. Redl 2003; Fischer et al. 2008). Giesshübl is also probably the only locality from where D. maculata subsp. austriaca has been reliably reported. Ståhlberg and Hedrén (2010) mention this taxon also from the surroundings of the town of Furth an der Triesting, Lower Austria. The exact location is however not known (M. Hedrén, in litt.) and only diploid D. maculata subsp. fuchsii (28, Furth an der Triesting) was found in this area within our field work. Thus, D. maculata subsp. austriaca should be rather considered only a colour morph, which should not be recognized taxonomically (cf. Pedersen 1998).
Populations corresponding to the sooana group were found in several localities in hilly regions of the Western Carpathians, and they are usually mentioned under the name of D. fuchsii subsp. sooana. Some authors (Borsos 1961; Potůček 1969; Soó 1980; Vlčko et al. 2003) circumscribe this taxon solely based on the white colour of flowers; the flower colouration alone, however, cannot be used for its delimitation. These plants may be almost invariably characterized by white flowers with white anther caps and pale to bold spots on the leaves, and they are always diploid. In analogy to D. maculata subsp. fuchsii, the sooana group should be subordinated to D. maculata in the rank of subspecies. The oldest epithet related to this taxon at the subspecific level must be thus found.
The name D. fuchsii subsp. sooana commonly appears in the literature (Procházka 1979; Soó 1980; Batoušek 1995; Kubát 2010; Vlačiha 2013; Ponert 2019), but it is not valid, as no type specimen was stated for it in its protologue (cf. Borsos 1959), nor later. Thus, other names must be considered. In British Isles, plants with similar morphological characteristics, i.e. white flowers with markings and spotted leaves, are recognized as D. fuchsii subsp. okellyi (Druce) Soó (e.g. Eccarius 2016). Bateman and Denholm (1988) stated that there are no differences between ‘okellyi’ (recognized at variety level) and ‘sooana’ that could justify their separation. Nevertheless, their description of D. fuchsii var. okellyi (Druce) Bateman et Denholm implies that British plants are considerably subtler than those from Central Europe. In addition, Harrap and Harrap (2009) mention that white-flowered individuals in British Isles represent only part of a population of plants which are more variable in flower colour. Even the distribution pattern suggests that the Carpathian populations and the populations from the British Isles represent separate evolutionary units of independent origin. Their similarity in some morphological traits is likely to be just a result of convergence, which is quite common in Dactylorhiza (Averyanov 1982; Delforge 2006; Efimov et al. 2016).
The high proportion of white-flowering individuals within the sooana group could indicate some relation with D. maculata subsp. austriaca; this name should also be applied if both taxa were found to be identical. The distribution areas of these taxa border on each other, as D. maculata subsp. austriaca is known from the Northeastern Alps (Vöth 1978). A considerable morphological overlap between D. maculata subsp. austriaca and the sooana group is also apparent in our data. However, unlike the sooana group, D. maculata subsp. austriaca is tetraploid. It is also improbable that D. maculata subsp. austriaca is a polyploid derivate of the sooana group, because its flower colouration is positively correlated with leaf pigmentation: white-flowered individuals typically lack spots on the leaves. This is not the case of the sooana group, and D. maculata subsp. austriaca seems to be derived rather from the tetraploid cytotype of D. maculata subsp. fuchsii.
According to our knowledge, there is no valid name available for the taxon represented by the sooana group at the subspecies level. With no doubt, the invalid name ‘Dactylorhiza fuchsii subsp. sooana’ used by Borsos (1959) is related to this taxon. The epithet ‘sooana’ (originally “soóiana”, which is a typographical error) is thus adopted here, and a valid name of the subspecies is introduced, providing a diagnosis and stating the holotype.
Conclusions
Populations of D. *fuchsii in Central Europe are considerably variable both in morphology and ploidy level. Despite the commonly shared conviction that they are strictly diploid, a number of tetraploid populations was detected, as well as several DNA-triploids representing a minority cytotype within diploid or tetraploid, or even mixed ploidy populations. Tetraploid populations utterly prevail in the Bohemian Massif, while diploids are more common in the Carpathians, but all three cytotypes occur throughout Central Europe. This is the first large-scale screening of ploidy levels in D. maculata s.l. based on FCM considering PPE.
Based on the combination of phenotypic traits, ploidy level variation, and geographical distribution patterns, it is justifiable to separate a group of West Carpathian populations, which typically possess white flowers with white anther caps, pale to bold spots on the leaves, and strictly diploid chromosome numbers. In contrast, the other group of populations, widespread in Central Europe, is more variable, characterized by white to purple flowers, spotted or unspotted leaves; but importantly, with a positive correlation between the intensity of leaves and flower pigmentation, and with purple anther caps even in plants with completely white flowers. All three cytotypes were found in this group, but they were morphologically indistinguishable. Following the more appropriate taxonomic concept, the latter of the groups should be recognized as D. maculata susbp. fuchsii, while the first is here described as D. maculata subsp. sooana, subsp. nova. A new combination of its hybrid with D. majalis subsp. majalis is also suggested, which is D. × dinglensis nothosubsp. smitakii, comb. nova.
Taxonomic treatment
Dactylorhiza maculata subsp. sooana Borsos ex Batoušek, Taraška & Trávn., subsp. nova. [Dactylorhiza fuchsii subsp. sooana Borsos, nom. inval., Acta Bot. Acad. Sci. Hung. 5: 324. 1959 (‘soóiana’)].—TYPE: Slovakia, Štiavnické vrchy Hills, Banský Studenec Village, meadow in the valley of the Bystrý potok brook east of the village, 655 m a. s. l., 48°26′31"N, 19°00′49"E, 13 Jun 2017, leg. excursion group (holotype: OL 37871!; isotypes: OL 37872!, OL 37873!, BRNM 826419!) (photographs of the live holotype plant see Fig. 5, photograph of its herbarium specimen see Online Resource 7).
Etymology: The epithet ‘sooana’ was adopted from Borsos (1959) and it refers to Károly Rezsö Soó (1903–1980), a Hungarian botanist and taxonomist with interest in genus Dactylorhiza.
Description: Herbaceous perennial plant with palmate tubers. Stem (26)37–61(67) cm high, with (4)5–9(13) leaves, often with brownish stripes. Lower 3–6 leaves with sheaths, upper leaves bract-like; at least lower leaves with bold or pale spots. Lowermost leaf obovate or oblong, usually obtuse at the apex, (46)74–141(180) × (14)18–35(52) mm, (2.3)3.1–5.6(7.5) times longer than wide. The 2nd lowermost leaf obovate, oblong or lanceolate, usually obtuse or subacute at the apex, (82)103–165(200) × (11)17–34(52) mm, (2.4)4.0–7.3(11.6) times longer than wide. Inflorescence a dense-flowered spike. Tepals white, sometimes with markings. Lip three-lobed, the Heslop-Harrison index (1.1)1.2–1.5(1.8), white with or without purple marking and white anther caps. Capsules cylindrical, seeds dust-like.
Diagnosis: Dactylorhiza maculata subsp. sooana differs from the type D. maculata subsp. maculata by broader, obtuse lower leaves, and deeply three-lobed lips of flowers (Heslop-Harrison index ≥ 1.3), as well as diploid chromosome number (2n = 2x = 40). These characteristics are mostly shared with D. maculata subsp. fuchsii, from which D. maculata subsp. sooana differs by a combination of several qualitative traits: white flowers, sometimes with purple markings and always with white anther caps, and spotted leaves, even in individuals with completely white flowers. Both taxa also differ in cytotype diversity, as D. maculata subsp. sooana is always diploid, while D. maculata subsp. fuchsii may be di-, tri- or tetraploid.
Chromosome numbers: 2n = 2x = 40.
Habitats: Mesophilous to wet meadows, open broad-leaved (beech) forests.
Distribution area: Czech Republic, Slovakia and Hungary. Endemic to Western Carpathians.
Dactylorhiza × dinglensis nothosubsp. smitakii (Batoušek) Batoušek, Taraška & Trávn., comb. nova. [D. maculata subsp. sooana × D. majalis (Rchb.) P.F.Hunt & Summerh. subsp. majalis]. ≡ Dactylorhiza × braunii nothosubsp. smitakii Batoušek, J. Eur. Orch. 29: 643. 1997.—HOLOTYPE: Moravia meridioorientalis, montes Bílé Karpaty, distr. Zlín: Nedašov, pratum clivis septentrionalis montis Cigán (744 m), 550 m a. s. l., 15 Jun 1980, P. Batoušek (GM 29845!).
Note: A hybrid of D. maculata subsp. sooana and D. majalis subsp. majalis was described by Batoušek (1997) as D. × braunii nothosubsp. smitakii Batoušek from Eastern Moravia (Czech Republic). The name D. × braunii (Halácsy) Soó is however applied to interspecific hybrids of D. fuchsii and D. majalis, where the first is recognized at the species level. Following the here accepted taxonomic concept, in which D. fuchsii is considered as an infraspecific taxon of D. maculata, a new combination is required for the hybrid. The interspecific hybrids of D. maculata and D. majalis are recognized as D. × dinglensis (Wilmott) Soó, Nom. Nov. Gen. Dactylorhiza 10, 1962 based on the name of Orchis × dinglensis Wilmott, Proc. Linn. Soc. London 148: 128, 1936. This hybrid taxon was noted by us on the locus clasicus of D. maculata subsp. sooana (near Banský Studenec Village in Štiavnické vrchy Hills, Slovakia; photographs in Online Resource 8), as well as in further localities in Slovakia (54, Rudno nad Hronom), Czech Republic (5, Bylničky) and Hungary (48, Bohó-hegy). From Slovakian territory, this hybrid was reported by Vlčko et al. 2003: 97 (from the Biele Karpaty Mts).
References
Aagaard SMD, Såstad SM, Greilhuber J, Moen A (2005) A secondary hybrid zone between diploid Dactylorhiza incarnata ssp. cruenta and allotetraploid D. lapponica (Orchidaceae). Heredity 94:488–496. https://doi.org/10.1038/sj.hdy.6800643
Aceto S, Caputo P, Cozzolino S, Gaudio L, Moretti A (1999) Phylogeny and evolution of Orchis and allied genera based on ITS DNA variation: morphological gaps and molecular continuity. Molec Phylogen Evol 13:67–76. https://doi.org/10.1006/mpev.1999.0628
Ackerman JD, Cuevas AA, Hof D (2011) Are deception-pollinated species more variable than those offering a reward? Pl Syst Evol 293:91–99. https://doi.org/10.1007/s00606-011-0430-6
Amich F, García-Barriuso M, Bernardos S (2007) Polyploidy and speciation in the orchid flora of the Iberian Peninsula. Bot Helv 117:143–157. https://doi.org/10.1007/s00035-007-0804-0
Averyanov LV (1982) Dactylorhiza maculata s. l. (Orchidaceae) na territorii SSSR. Bot Zhur 67:303–312 (in Russian)
Averyanov LV (1990) A review of the genus Dactylorhiza. In: Arditti J (ed) Orchid biology Reviews and perspectives, vol 5. Timber Press, Portland, pp 159–206
Balao F, Tannhäuser M, Lorenzo MT, Hedrén M, Paun O (2016) Genetic differentiation and admixture between sibling allopolyploids in the Dactylorhiza majalis complex. Heredity 116:351–361. https://doi.org/10.1038/hdy.2015.98
Bateman RM, Denholm I (1988) A reappraisal of the British and Irish dactylorchids, 3. The Spotted-Orchids. Watsonia 17:319–349
Bateman RM, Denholm I (1989) On measuring marsh-orchids. Morphometric procedure, taxonomic objectivity and marsh-orchid systematics. Watsonia 17:449–462
Bateman RM, Murphy ARM, Hollingsworth PM, Hart ML, Denholm I, Rudall PJ (2018) Molecular and morphological phylogenetics of the digitate-tubered clade within subtribe Orchidinae s.s. (Orchidaceae: Orchideae). Kew Bull 73:54. https://doi.org/10.1007/s12225-018-9782-1
Bates D, Maechler M, Bolker BM, Walker S, Bojesen RH, Singmann H, Dai B, Grothendieck G, Green P (2019) lme4: Linear Mixed effects Models Using S4 Classes. Available at: http://lme4.r-forge.r-project.org/. Accessed 1 Feb 2019
Batoušek P (1995) Zur Kenntnis von Dactylorhiza fuchsii (Druce) Soó ssp. sooana (Borsos) Borsos. J Eur Orchid 27:51–74
Batoušek P (1997) Zwei neue Hybriden aus der Tschechischen Republik. J Eur Orchid 29:641–652
Baumann H, Künkele S, Lorenz R (2002) Taxonomische Liste der Orchideen Deutschlands. J Eur Orchid 34:129–206
Bertolini V, del Prete C, Garbari F (2000) Karyological and biometrical studies on some species of the genus Dactylorhiza Necker ex Nevski, sect. Dactylorhiza (Orchidaceae) of Central-Northern Italy. Port Acta Biol 19:249–265
Borsos O (1959) Dactylorchis fuchsii Druce et son affinité dans les flores Hongroise et Carpatique. Acta Bot Acad Sci Hung 5:321–325
Borsos O (1961) Geobotanische Monographie der Orchideen der pannonischen und karpatischen Flora V. Ann Univ Sci Budapest Rolando Eotvos, Sect Biol 4:51–82
Bory S, Catrice O, Brown S, Leitch IJ, Gigant R, Chiroleu F, Grisoni M, Duval MF, Besse P (2008) Natural polyploidy in Vanilla planifolia (Orchidaceae). Genome 51:816–826. https://doi.org/10.1139/G08-068
Brandrud MK, Baar J, Lorenzo MT, Athanasiadis A, Bateman RM, Chase MW, Hedrén M, Paun O (2020) Phylogenomic relationships of diploids and the origins of allotetraploids in Dactylorhiza (Orchidaceae). Syst Biol 69:91–109. https://doi.org/10.1093/sysbio/syz035
Brown SC, Bourge M, Maunoury N, Wong M, Wolfe Bianchi M, Lepers-Andrzejewski S, Besse P, Siljak-Yakovlev S, Dron M, Satiat-Jeunemaître B (2017) DNA remodeling by Strict Partial Endoreplication in Orchids, an original process in the Plant Kingdom. Genome Biol Evol 9:1051–1071. https://doi.org/10.1093/gbe/evx063
Bureš L (2013) Chráněné a ohrožené rostliny Chráněné krajinné oblasti Jeseníky. Rubico, Olomouc
Buttler KP (2000) Orchideje. Planě rostoucí druhy a poddruhy Evropy, Přední Asie a severní Afriky. Ikar, Praha
Cauwet-Marc AM, Balayer M (1984) Les genres Orchis L., Dactylorhiza Necker ex Nevski, Neotinea Reichb. et Traunsteinera Reichb.: caryologie et proposition de phylogénie et d’évolution. Bot Helv 94:391–406
Danihelka J, Chrtek J Jr, Kaplan Z (2012) Checklist of vascular plants of the Czech Republic. Preslia 84:647–811
De hert K, Jacquemyn H, Van Glabeke S, Roldán-Ruiz I, Vandepitte K, Leus L, Honnay O (2012) Reproductive isolation and hybridization in sympatric populations of three Dactylorhiza species (Orchidaceae) with different ploidy levels. Ann Bot (Oxford) 109:709–720. https://doi.org/10.1093/aob/mcr305
Delforge P (2006) Orchids of Europe. North Africa and the Middle East. A&C Black, London
Devillers P, Devillers-Terschuren J (2000) Dactylorhiza sudetica (Pöch ex Rchb. fil. 1851) Averyanov 1982 dans les monts des Géants. Naturalistes Belges 81:331–338
Devos N, Tyteca D, Raspé O, Wesselingh RA, Jacquemart A-L (2003) Patterns of chloroplast diversity among western European Dactylorhiza species (Orchidaceae). Pl Sys Evol 243:85–97. https://doi.org/10.1007/s00606-003-0068-0
Doležel J, Greilhuber J, Suda J (2007) Estimation of nuclear DNA content in plants using flow cytometry. Nat Protoc 2:2233–2244. https://doi.org/10.1038/nprot.2007.310
Doležel J, Greilhuber J, Lucretti S, Meister A, Lysák MA, Nardi L, Obermayer R (1998) Plant genome size estimation by flow cytometry: Inter-laboratory comparison. Ann Bot (Oxford) 82(Suppl. A):17–26
Druce GC (1915) Orchis maculata L. and O. fuchsii. Rep Bot Exch Club Soc Brit Isles 4:99–108
Dufrêne M, Gathoye J-L, Tyteca D (1991) Biostatistical studies on western European Dactylorhiza (Orchidaceae) – the D. maculata group. Pl Syst Evol 175:55–72
Eccarius W (2016) Die Orchideengattung Dactylorhiza. Phylogenie, Taxonomie, Morphologie, Biologie, Verbreitung, Ökologie und Hybridisation. W. Eccarius, Eisenach
Efimov PG, Philippov EG, Krivenko DA (2016) Allopolyploid speciation in Siberian Dactylorhiza Orchidaceae Orchidoideae. Phytotaxa 258:101–120. https://doi.org/10.11646/phytotaxa.258.2.1
Fischer MA, Oswald K, Adler W (2008) Exkursionsflora für Österreich, Liechtenstein und Südtirol, 3rd edn. Biologiezentrum der Oberösterreichischen Landesmuseen, Linz
Galbraith DW, Harkins KR, Maddox JM, Ayres NM, Sharma DP, Firoozabady E (1983) Rapid flow cytometric analysis of the cell cycle in intact plant tissues. Science 220:1049–1051
Gathoye J-L, Tyteca D (1987) Étude biostatistique des Dactylorhiza (Orchidaceae) de Belgique et des territoires voisins. Bull Jard Bot Natl Belg 57:389–424
Gigord LDB, Macnair MR, Smithson A (2001) Negative frequency-dependent selection maintains a dramatic flower color polymorphism in the rewardless orchid Dactylorhiza sambucina (L.) Soó. Proc Natl Acad Sci USA 98:6253–6255. https://doi.org/10.1073/pnas.111162598
GIROS (2009) Orchidee d’Italia. Guida alle orchidee spontanee. Il Castello, Cornaredo
Groll M (1966) Fruchtansatz, Bestäubung und Merkmalsanalyse bei diploiden und polyploiden Sippen von Dactylorchis (Orchis) maculata und Gymnadenia conopsea. Österr Bot Z 112:657–700
Harrap A, Harrap S (2009) Orchids of Britain and Ireland. A field and site guide. A&C Black, London
Hedrén M (1996) Genetic differentiation, polyploidization and hybridization in northern European Dactylorhiza (Orchidaceae): evidence from allozyme markers. Pl Syst Evol 201:31–55. https://doi.org/10.1007/BF00989050
Hedrén M (2001) Systematics of the Dactylorhiza euxina/incarnata/maculata polyploid complex (Orchidaceae) in Turkey: evidence from allozyme data. Pl Syst Evol 229:23–44. https://doi.org/10.1007/s006060170016
Hedrén M, Fay MF, Chase MW (2001) Amplified fragment length polymorphisms (AFLP) reveal details of polyploid evolution in Dactylorhiza (Orchidaceae). Amer J Bot 88:1868–1880. https://doi.org/10.2307/3558363
Hedrén M, Nordström S, Ståhlberg D (2008) Polyploid evolution and plastid DNA variation in the Dactylorhiza incarnata/maculata complex (Orchidaceae) in Scandinavia. Molec Ecol 17:5075–5091. https://doi.org/10.1111/j.1365-294X.2008.03965.x
Heslop-Harrison J (1951) A comparison of some Swedish and British forms of Orchis maculata L. sens. lat. Svensk Bot Tidskr 45:608–635
Heslop-Harrison J (1968) Genetic system and ecological habit as factors in Dactylorchid variation. Jahresber Naturwiss Vereins Wuppertal 21(22):20–27
Hřibová E, Holušová K, Trávníček P, Petrovská B, Ponert J, Šimková H, Kubátová B, Jersáková J, Čurn V, Suda J, Doležel J, Vrána J (2016) The enigma of progressively partial endoreplication: new insights provided by flow cytometry and next-generation sequencing. Genome Biol Evol 8:1996–2005. https://doi.org/10.1093/gbe/evw141
Jagiełło M, Lankosz-Mróz M (1988) Cytotaxonomic studies in the Dactylorhiza maculata (L.) Soó group in Poland (Orchidaceae). – Fragm Florist Geobot 31–32:385–394
Jagiełło M (1988) Analysis of population variability and distribution of species from the Dactylorhiza maculata group (Orchidaceae) in Poland. Fragm Florist Geobot 31–32:333–383
Kaplan Z, Danihelka J, Šumberová K, Chrtek J Jr, Rotreklová O, Ekrt L, Štěpánková J, Taraška V, Trávníček B, Prančl J, Ducháček M, Hroneš M, Kobrlová L, Horák D, Wild J (2017) Distributions of vascular plants in the Czech Republic. Preslia 89:333–439. https://doi.org/10.23855/preslia.2017.333
Kliment J (1999) Komentovaný prehľad vyšších rastlín flóry Slovenska, uvádzaných v literatúre ako endemické taxóny. Bull Slov Bot Spoločn 21(Suppl. 4):1–434
Kolář F, Čertner M, Suda J, Schönswetter P, Husband BC (2017) Mixed-ploidy species: progress and opportunities in polyploid research. Trends Pl Sci 22:1041–1055. https://doi.org/10.1016/j.tplants.2017.09.011
Krahulcová A (2003) Chromosome numbers in selected monocotyledons (Czech Republic, Hungary and Slovakia). Preslia 75:97–113
Kreutz CAJ (2004) Kompendium der Europäischen Orchideen. Kreutz Publishers, Landgraaf
Kubát K (2010) Dactylorhiza Nevski – prstnatec. In: Štěpánková J, Chrtek J, Kaplan Z (eds) Květena České republiky 8. Academia, Praha, pp 502–523
Kurtto A, Lampinen R, Piirainen M, Uotila P (2019) Checklist of the vascular plants of Finland. Suom Putkilokasvien Luettelo Norrlinia 34:1–206
Legendre P, Legendre L (2012) Numerical ecology. Elsevier, Amsterdam
Leitch IJ, Bennett MD (2004) Genome downsizing in polyploid plants. Biol J Linn Soc 82:651–663. https://doi.org/10.1111/j.1095-8312.2004.00349.x
Lenth R, Singmann H, Love J, Buerkner P, Herve M (2020) emmeans: Estimated Marginal Means, aka Least-Squares Means. Available at: https://cran.r-project.org/web/packages/emmeans/index.html. Accessed 1 Feb 2019
Lord RM, Richards AJ (1977) A hybrid swarm between the diploid Dactylorhiza fuchsii (Druce) Soó and the tetraploid D. purpurella (T. & T. A. Steph.) Soó in Durham. Watsonia 11:205–210
Loureiro J, Trávníček P, Rauchová J, Urfus T, Vít P, Štech M, Castro S, Suda J (2010) The use of flow cytometry in the biosystematics, ecology and population biology of homoploid plants. Preslia 82:3–21
Löve Á (1971) IOPB Chromosome number reports XXXIV. Taxon 20:785–797
Lysák MA, Doležel J (1998) Estimation of nuclear DNA content in Sesleria (Poaceae). Caryologia 52:123–132. https://doi.org/10.1080/00087114.1998.10589127
Májovský J (ed.) (1976) Index of chromosome numbers of Slovakian flora (Part 5). Acta Fac Rerum Nat Univ Comen Bot 25:1–18
Májovský J (ed) (1978) Index of chromosome numbers of Slovakian flora (Part 6). Acta Fac Rerum Nat Univ Comen Bot 26:1–42
Měsíček J, Javůrková-Jarolímová V (1992) List of chromosome numbers of the Czech vascular plants. Academia, Praha
Meyer FJ (1968) Genetisches System und ökologisches Verhalten als Faktoren der Variantenbildung bei Dactylorchis. Jahresber Naturwiss Vereins Wuppertal 21(22):28–31
Molnár VA, Sulyok J, Magos G, Bódis J (2011) Erdei ujjaskosbor – Dactylorhiza fuchsii (Druce) Soó 1962. In: Molnár VA (ed.) Magyarország orchideáinak atlasza. Kossuth Kiadó, Budapest, pp 353–354
Naczk AM, Górniak M, Szlachetko DL, Ziętara MS (2015) Plastid DNA haplotype diversity and morphological variation in the Dactylorhiza incarnata/maculata complex (Orchidaceae) in northern Poland. Bot J Linn Soc 178:121–137. https://doi.org/10.1111/boj.12267
Naimi B (2017) Uncertainty analysis for species distribution models. Available at: https://cran.r-project.org/web/packages/usdm/index.html. Accessed 31 Jul 2019
Narbona E, Wang H, Ortiz PL, Arista M, Imbert E (2017) Flower colour polymorphism in the Mediterranean Basin: occurrence, maintenance and implications for speciation. Pl Biol (Stuttgart) 20:8–20. https://doi.org/10.1111/plb.12575
Nordström S, Hedrén M (2009) Evolution, phylogeography and taxonomy of allopolyploid Dactylorhiza (Orchidaceae) and its implications for conservation. Nordic J Bot 27:548–556. https://doi.org/10.1111/j.1756-1051.2009.00548.x
O’Brien RM (2007) A caution regarding rules of thumb for variance inflation factors. Qual Quant 41:673–690. https://doi.org/10.1007/s11135-006-9018-6
Parisod C, Holderegger R, Brochmann C (2010) Evolutionary consequences of autopolyploidy. New Phytol 186:5–17. https://doi.org/10.1111/j.1469-8137.2009.03142.x
Paun O, Bateman RM, Fay MF, Hedrén M, Civeyrel L, Chase MW (2010) Stable epigenetic effects impact adaptation in allopolyploid orchids (Dactylorhiza: Orchidaceae). Molec Biol Evol 27:2465–2473. https://doi.org/10.1093/molbev/msq150
Pedersen HÆ (1998) Species concept and guidelines for infraspecific taxonomic ranking in Dactylorhiza (Orchidaceae). Nordic J Bot 18:289–310. https://doi.org/10.1111/j.1756-1051.1998.tb01881.x
Pedersen HÆ (2009) Apochromic populations of Dactylorhiza incarnata s. l. (Orchidaceae): diversity and systematic significance as revealed by allozyme markers and morphology. Bot J Linn Soc 159:396–407. https://doi.org/10.1111/j.1095-8339.2009.00954.x
Pikner T (2012) Taxonomic diversity of Dactylorhiza on Saaremaa. J Hardy Orchid Soc 9:128–143
Pillon Y, Fay MF, Hedrén M, Bateman RM, Devey DS, Shipunov AB, van der Bank M, Chase MW (2007) Evolution and temporal diversification of western European polyploid species complexes in Dactylorhiza (Orchidaceae). Taxon 56:1185–1208. https://doi.org/10.2307/25065911
Pillon Y, Fay MF, Shipunov AB, Chase MW (2006) Species diversity versus phylogenetic diversity: a practical study in the taxonomically difficult genus Dactylorhiza (Orchidaceae). Biol Conservation 129:4–13. https://doi.org/10.1016/j.biocon.2005.06.036
Ponert J (2019) Dactylorhiza Nevski – prstnatec (vstavač). In: Kaplan Z, Danihelka J, Chrtek J Jr, Kirschner J, Kubát K, Štech M, Štěpánek J (eds) Klíč ke květeně České republiky. Adademia, Praha, pp 183–187
Popelka O, Trávníček B, Šiková P, Jandová M, Duchoslav M (2019) Natural hybridization between diploid Ficaria calthifolia and tetraploid Ficaria verna subsp. verna in central Europe: evidence from morphology, ecology and life-history traits. Preslia 91:179–212. https://doi.org/10.23855/preslia.2019.179
Popelka O, Sochor M, Duchoslav M (2019b) Reciprocal hybridization between diploid Ficaria calthifolia and tetraploid Ficaria verna subsp. verna: evidence from experimental crossing, genome size and molecular markers. Bot J Linn Soc 189:293–310. https://doi.org/10.1093/botlinnean/boy085
Potůček O (1969) Klíč k určování československých druhů čeledi Orchidaceae. Východočeské muzeum v Pardubicích, Pardubice
Procházka F (1979) Okruh prstnatce plamatého (Dactylorhiza maculata agg.) v Československu. Zprávy Českoslov Bot Společ 14:9–12
Ramsey J, Schemske DW (1998) Pathways, mechanisms and rates of polyploid formation in flowering plants. Annual Rev Ecol Evol Syst 29:467–501. https://doi.org/10.1146/annurev.ecolsys.29.1.467
Redl K (2003) Wildwachsende Orchideen in Österreich – faszinierend und schützenswert, 3rd edn. K. Redl Eigenverlag, Altenmarkt
Reinhard HR, Gölz P, Peter R, Wildermuth H (1991) Die Orchideen der Schweiz und angrenzender Gebiete. Fotorotar AG, Egg
Schneider CA, Rasband WS, Eliceiri KW (2012) NIH Image to ImageJ: 25 years of image analysis. Nature Meth 9:671–675. https://doi.org/10.1038/nmeth.2089
Senghas K (1968) Taxonomische Übersicht der Gattung Dactylorhiza Necker ex Nevski. Jahresber Naturwiss Vereins Wuppertal 21(22):32–67
Singmann H, Bolker B, Westfall J, Aust F (2016) afex: Analysis of Factorial Experiments. R package version 0.16–1. Available at: https://CRAN.R-project.org/package=afex. Accessed 1 Feb 2019
Skalińska M, Banach-Pogan E, Wcisło H et al (1957) Further studies in chromosome numbers of Polish Angiosperms. Acta Soc Bot Poloniae 26:215–246
Soltis DE, Soltis PS, Schemske DW, Hancock JF, Thompson JN, Husband BC, Judd WS (2007) Autopolyploidy in angiosperms: have we grossly underestimated the number of species? Taxon 56:13–30. https://doi.org/10.2307/25065732
Soó R (1980) Dactylorhiza Necker ex Nevski. In: Tutin TG, Heywood VH, Burges NA, Moore DM, Valentine DH, Walters SM, Webb DA (eds) Flora Europaea, vol 5. Cambridge University Press, Cambridge, pp 335–337
Spoelhof JP, Soltis PS, Soltis DE (2017) Pure polyploidy: closing the gaps in autopolyploid research. J Syst Evol 55:340–352. https://doi.org/10.1111/jse.12253
Ståhlberg D, Hedrén M (2008) Systematics and phylogeography of the Dactylorhiza maculata complex (Orchidaceae) in Scandinavia: insights from cytological, morphological and molecular data. Pl Syst Evol 273:107–132. https://doi.org/10.1007/s00606-008-0035-x
Ståhlberg D, Hedrén M (2010) Evolutionary history of the Dactylorhiza maculata polyploid complex (Orchidaceae). Biol J Linn Soc 101:503–525. https://doi.org/10.1111/j.1095-8312.2010.01505.x
Ståhlberg D (2009) Habitat differentiation, hybridization and gene flow patterns in mixed populations of diploid and autotetraploid Dactylorhiza maculata s. l. (Orchidaceae). Evol Ecol 23:295–328. https://doi.org/10.1007/s10682-007-9228-y
Ströhle W (2003) Nomenklatorische Liste der europäischen Orchideentaxa – Arten und Unterarten. J Eur Orchid 35:771–860
Suda J, Krahulcová A, Trávníček P, Krahulec F (2006) Ploidy level versus DNA ploidy level: an appeal for consistent terminology. Taxon 55:447–450. https://doi.org/10.2307/25065591
Šmarda P, Knápek O, Březinová A, Horová L, Grulich V, Danihelka J, Veselý P, Šmerda J, Rotreklová O, Bureš P (2019) Genome sizes and genomic guanine cytosine (GC) contents of the Czech vascular flora with new estimates for 1700 species. Preslia 91:117–142. https://doi.org/10.23855/preslia.2019.117
ter Braak C, Šmilauer P (2012) Canoco reference manual and user’s guide: software for ordination (version 5.0). Microcomputer Power, Ithaca
Thórsson ÆT, Salmela E, Anamthawat-Jónsson K (2001) Morphological, cytogenetic, and molecular evidence for introgressive hybridization in birch. J Heredity 92:404–408. https://doi.org/10.1093/jhered/92.5.404
Trávníček P, Kubátová B, Čurn V, Rauchová J, Krajníková E, Jersáková J, Suda J (2011) Remarkable coexistence of multiple cytotypes of the Gymnadenia conopsea aggregate (the fragrant orchid): evidence from flow cytometry. Ann Bot (Oxford) 107:77–87. https://doi.org/10.1093/aob/mcq217
Trávníček P, Ponert J, Urfus T, Jersáková J, Vrána J, Hřibová E, Doležel J, Suda J (2015) Challenges of flow-cytometric estimation of nuclear genome size in Orchids, a plant group with both whole-genome and progressively partial endoreplication. Cytom Part A 87:958–966. https://doi.org/10.1002/cyto.a.22681
Tyteca D, Gathoye J-L (2003) Morphometric analyses of the Dactylorhiza maculata (L.) Soó group in western Europe. Ber Arbeitskreis Heimische Orchid 20:1–32
Uhríková A (2007) New records in Dactylorhiza. In: Marhold K, Mártonfi P, Mereďa P Jr, Mráz P (eds) Chromosome number survey of the ferns and flowering plants of Slovakia. Veda, Bratislava, p 221
Vaucher C (1966) Contribution à l’étude cytologique du genre Dactylorchis (Klinge) Vermeulen. Bull Soc Neuchâtel Sci Nat 89:75–85
Vereecken NJ, Schiestl FP (2009) On the roles of colour and scent in a specialized floral mimicry system. Ann Bot (Oxford) 104:1077–1084. https://doi.org/10.1093/aob/mcp208
Vermeulen P (1947) Studies on Dactylorchids. Drukkerij Fa. Schotanus & Jens, Utrecht
Vermeulen P (1968) Dactylorchis maculata und ihre Formen. Jahresber Naturwiss Vereins Wuppertal 21(22):68–76
Vlačiha V (2013) Prstnatce České republiky. ČSOP Launensia, Ústí nad Labem
Vlčko J, Dítě D, Kolník M (2003) Vstavačovité Slovenska. ZO SZOPK Orchidea, Zvolen
Vöth W, Greilhuber J (1980) Zur Karyosystematik von Dactylorhiza maculata s.l. und ihrer Verbreitung, insbesondere in Niederösterreich. Linz Biol Beitr 12:415–468
Vöth W (1978) Biometrische Untersuchungen an Dactylorhiza maculata s. l. – Sippen in Niederösterreich (Orchidaceae). Linz Biol Beitr 10:179–215
Weiss H, Stuessy TF, Grau J, Baeza CM (2003) Chromosome reports from South American Hypochaeris (Asteraceae). Ann Missouri Bot Gard 90:56–63.
Acknowledgements
We are thankful to many botanists who guided us in the field, helped us with searching localities or samplings, namely, A. Čejková, I. Jongepierová, A. Kantor, N. Kantorová, J. Košnar, E. Pietorová, J. Plekanec, M. Popelářová, M. & R. Puscarciuc, P. Salenka, A. Schmotzer, G. Schneeweiss, R. Štencl, Z. Václavová, V. Vlačiha, M. & M. Wolanin and many others. We are grateful to M. Hroneš and M. Jandová for service of the FCM laboratory at the Palacký University and their methodological support, as well as J. Doležel and J. Vrána, who enabled us to work in the laboratory of the Institute of Experimental Botany in Olomouc. We furthermore appreciate M. Oulehlová, who scanned the type specimen, and P. Trávníček for valuable consultations. We are grateful to both reviewers for critical reading and improvement of the manuscript. The authorities in nature conservation are acknowledged for granting us with the necessary permits. Research was supported by internal grant of the Palacký University IGA_PrF_2021_001 and by Austrian Federal Ministry of Education, Science and Research (BMBWF) within program Aktion CZ-AT, administrated by OeAD-GmbH.
Funding
VT was supported by internal grant of the Palacký University IGA_PrF_2021_001 and by Austrian Federal Ministry of Education, Science and Research within program Aktion CZ-AT, administrated by OeAD-GmbH.
Author information
Authors and Affiliations
Contributions
VT, PB, and BT conceived the project and conducted the field work. VT, EMT and HWS performed the analysis of genome size and chromosome numbers. MD designed and conducted the statistical analysis. VT drafted the manuscript with significant contributions of MD and BT. All authors commented on and approved the manuscript. BT supervised the project.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Handling editor: Martin A. Lysak.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Information on Electronic Supplementary Material
Information on Electronic Supplementary Material
Online Resource 1. Locality details of Dactylorhiza *fuchsii.
Online Resource 2. Details to (sub)populations of Dactylorhiza *fuchsii. Averaged relative DNA content (= fluorescence ratios between the positions of the sample and internal reference standard G0/G1 peaks) of investigated populations.
Online Resource 3. Explanations to quantitative characters used in the morphometrics.
Online Resource 4. Relative DNA content (= fluorescence ratios between the positions of the sample and internal reference standard G0/G1 peaks) of six plants with counted chromosome numbers; the stain was either DAPI or PI. All values are calculated relative to the Pisum sativum cv. ‘Ctirad’ as internal reference standard.
Online Resource 5. Absolute genome sizes (GS) of five individuals of Dactylorhiza *fuchsii estimated by flow cytometry.
Online Resource 6. Box plots of characters analysed for the fuchsii-2x, fuchsii-4x and sooana groups.
Online Resource 7. Holotype of Dactylorhiza maculata subsp. sooana Batoušek, Taraška & Trávn.
Online Resource 8. Images of Dactylorhiza maculata subsp. sooana, D. maculata subsp. fuchsii and D. ×dinglensis nothosubsp. smitakii.
Rights and permissions
About this article
Cite this article
Taraška, V., Batoušek, P., Duchoslav, M. et al. Morphological variability, cytotype diversity, and cytogeography of populations traditionally called Dactylorhiza fuchsii in Central Europe. Plant Syst Evol 307, 51 (2021). https://doi.org/10.1007/s00606-021-01770-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00606-021-01770-3