Abstract
Karyotype analyses in species of the family Marantaceae (550 species, 31 genera) were conducted to shed light on the reported strong variation of chromosome number and size and the occurrence of polyploidy. Special attention was paid to the alterations in basic chromosome numbers, karyotypes and ploidy levels. Taxon sampling covered the whole distribution area of Marantaceae in Africa, Asia and America. We applied mitotic chromosome counting using conventional rapid squash techniques in 43 accessions (39 species, 16 genera), evaluated literature records for 51 species and conducted karyotype analyses. Eleven different somatic chromosome numbers were found (2n = 20, 22, 24, 26, 28, 33, 44, 36, 52, 65, 72). Based on the presumed basic chromosome numbers of x = 9, 10, 11, 12, 13, 14, this may correspond to diploid, triploid, tetraploid, pentaploid and octoploid levels, respectively. Dysploid variation, polyploidy and, to a lesser extend, hybridization may be the main factors in chromosome number evolution of the family. Our results also point to a certain degree of association with species diversification and geographical patterns.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Changes in chromosome number and structure are important processes that often reflect speciation events as they can establish crossing barriers between populations (e.g. Lowry and Willis 2010; Ouyang and Zhang 2013; Winterfeld et al. 2014, 2016; Baack et al. 2015; Wölk et al. 2015). Chromosome variations such as polyploidy, hybridization or dysploid change may provide the cytological basis for ecological differentiation, adaptation and isolation. The phylogenetic effects in chromosome variations, their ecological significance and their impact on speciation and diversification are the subject of many studies in major angiosperm families (e.g. Knight et al. 2005; Peruzzi et al. 2012; Grabowska-Joachimiak et al. 2015; Carta et al. 2018 and references therein).
An outstanding system to study the role of chromosomal changes in species diversification is the pantropically distributed monocot family Marantaceae which, on the one hand, exhibits a geographically uneven distribution of high species diversity and, on the other hand, provides a remarkable variation in morphological traits and chromosome number. Marantaceae is the second largest family of Zingiberales (Kennedy 1978a, b, 2000; Dhetchuvi 1996; Andersson 1998; Suksathan and Borchsenius 2005; Jongkind 2008; Ley and Claßen-Bockhoff 2011) and comprises approximately 550 species in 31 genera (Stevens 2001 onwards, Prince and Kress 2006; Al-Gharaibeh 2017). The plants are perennial herbs and lianas from the understorey and gaps of tropical lowland rainforest and are distributed throughout tropical and warm temperate parts of the world (Andersson 1998). Their diversity centre is in the New World, where they are represented by ca. 450 species in 13 genera. Ca. 50 species in 11 genera are found in Africa including Madagascar and ca. 50 species in 6 genera in Asia (Prince and Kress 2006; Al-Gharaibeh 2017).
Available chromosome counts reveal a strong variation of chromosome number in Marantaceae. Some chromosome numbers were recurrently found, but in general there seems to be a swarm of possibly aneuploid forms as reviewed by Bisson et al. (1968) and Mahanty (1970). While the sister family Cannaceae is characterized by a uniform basic number of x = 9, Marantaceae seems to have monoploid numbers ranging from as low as 4 up to 13. Since the beginning of karyological studies in Marantaceae (Venkatasubban 1946; Mangenot and Mangenot 1957, 1958; Sharma and Bhattacharyya 1958; Sato 1960; Bisson et al. 1968; Mahanty 1970; Mukhopadhyay and Sharma 1987), there has been continuous discussion on the partly conflicting results and the ancestral states in the chromosomal evolution of the family.
Some authors (Venkatasubban 1946; Sato 1948, 1960; Bisson et al. 1968) suggested that basic numbers of x = 9, 10, 11, 12, 13 were the derivatives of the original x = 4 (in the following: p = 4 according to Peruzzi 2013). However, 2n = 8 was reported in only two species (Table 1), for which higher numbers were recorded by other authors (e.g. Venkatasubban 1946) that makes this assumption questionable. Further, there was a strong geographical and taxonomic bias since the early chromosome surveys were restricted to a couple of genera, especially Goeppertia (under Calathea) and Maranta, which are mainly American, while Old World taxa had not been considered so far. Subsequently p = 11, which is found in several species of the American genus Goeppertia (Venkatasubban 1946; Sato 1948; Sharma and Bhattacharyya 1958), as suggested as ancestral in Marantaceae (Sharma and Bhattacharyya 1958). The other numbers should have been derived from p = 11 by ascending or descending dysploidy. Mahanty (1970) worked on the Malayan and primarily African genera Stachyphryium, Marantochloa, and Thalia and emphasized that geographical distribution may have played a role in the evolution of Marantaceae and their different basic numbers. Based on the assumption that the origin of the family was tropical America (Holttum 1951), migration to the Far East and the African tropics might have been linked with the acquisition of ‘derived’ chromosomal character states. Mahanty (1970) suggested that x = 13 as frequently found in African Marantaceae is a secondary basic number, which became established in Old World genera. Recent molecular phylogenetic and morphological investigations (Andersson and Chase 2001; Prince and Kress 2006), however, point to an origin of Marantaceae in the Old World. This means that the presumed cytogeographical pattern of ‘original’ monoploid chromosome numbers in the New and ‘derived’ ones in the Old World must be called into question. In addition to the varying basic numbers, also high chromosome numbers being based on polyploidy were reported in the American genera Maranta and Stromanthe (Venkatasubban 1946; Bisson et al. 1968; Sharma and Mukhopadhyay 1984; Hanson et al. 1999; see Table 1). Polyploidy was suggested to be related to the vegetative mode of propagation, which is important in the whole family Marantaceae. In this way, chromosomal ‘biotypes’, which otherwise would become extinct, would be preserved (Mahanty 1970).
Some chromosome numbers reported in the literature for the Marantaceae should be treated with caution, due to shortcomings in some of the chromosome techniques employed and also with regard to the fact that the plant specimens were partly unreliably identified. Up to now, chromosome counts are available from only a relatively small number of species. Therefore, reliable conclusions cannot be drawn on the actual range of chromosomal variation in Marantaceae, not to speak of chromosomal and genome evolution in this family.
The present study aimed at providing a detailed survey of chromosome variation in Marantaceae. In particular, (1) we determined chromosome numbers and made karyotype analyses of 43 accessions, focussing on taxa that were not examined before, in which available information seemed to be doubtful or several different chromosome numbers have been reported; (2) we critically examined the numerical variation in monoploid chromosome sets, checked for different ploidy levels and possible aneuploid forms to explain the enormous variation in somatic chromosome numbers; and (3) we chose taxa from all major tropical regions (Africa, America, Asia) to find out whether there are trends in karyotype evolution that can be associated with the diversification of evolutionary lineages in Marantaceae and/or biogeography.
Materials and methods
Plant material
In total, 43 plant accessions belonging to 16 genera were included in the chromosome survey, of which 37 were identified to species level. The species of Hypselodelphys (M69) could not have been identified. One accession is cf. Sarcophrynium (M63). For Ctenanthe burle-marxii, Goeppertia bella and G. zebrina two different provenances could be investigated. Root tips were excised from Marantaceae plants of the living collection of the Botanical Garden of the Martin Luther University of Halle-Wittenberg. Voucher specimens are deposited in the herbarium HAL. For 51 further species, data from the literature were evaluated. The taxa studied, collection and voucher details and references are listed in Table 1.
Chromosome counting, karyotyping and regression analysis
Chromosome numbers were counted in actively growing root tips. They were washed with tap water a few times, treated in 8-hydroxyquinoline (0.002 M aqueous solution) at 21 °C for 3 h to accumulate metaphases, fixed in absolute ethanol/glacial acetic acid (3:1) at 21 °C for 3 h and stored in absolute ethanol at -20 °C until preparation. Enzyme-treated root tips (Schwarzacher et al. 1980) were squashed on slides in a drop of 45% propionic acid with 2% carmine and covered with a coverslip. Photographs of metaphase chromosomes were taken on a Zeiss Axiophot microscope using a computer-assisted cooled CCD camera (Zeiss Axiocam HRC) employing Zeiss Axiovision software. A total of 4–20 metaphase plates were used for chromosome counts.
For karyotyping, one metaphase plate with sharp contours of individual chromosomes was chosen. Karyotypes were reconstructed in all species except for Goeppertia bella (M73) and G. lancifolia (M4), in which only chromosome numbers could be retrieved. Chromosomes were arranged and measured in CorelDraw Graphics Suite—X8 by hand. They were grouped as pairs or groups of putative homologues or homoeologues according to their similarities in chromosome length and position of centromeres and/or secondary constrictions. Selected metaphase plates and karyograms of each taxon studied are shown in Figs. 1, 2, 3 and 4, respectively. Calculations of karyological parameters, done with Excel, are as follows Paszko (2006) and Peruzzi and Eroğlu (2013): TML = total length of monoploid chromosome set [= (L + S)/ploidy level, L = total length of long arms, S = total length of short arms]; MCA = mean centromeric asymmetry [= (L − S)/(L + S) × 100]; CVCL = interchromosomal asymmetry [= (sCL/MCL) × 100, sCL = standard deviation of chromosome length in a chromosome complement, MCL = mean chromosome length].
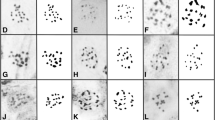
Somatic metaphase plates. aPhrynium terminale (2n = 22); bThalia geniculata M51 (2n = 33); cMegaphrynium macrostachyum M63 (2n = 36); dTrachyphrynium brauneanum M62 (2n = 22); eGoeppertia bachemiana M75 (2n = 26); fGoeppertia picturata M13 (2n = 28); gThaumatococcus danielii M53 (2n = 20); hStromanthe sanguinea M50 (2n = 44); iCtenanthe dasycarpa M79 (2n = 72); jMaranta arundinacea M29 (2n = 52); kMarantochloa conferta M54 (2n = 26); lMarantochloa congensis M58 (2n = 65); mMarantochloa cordifolia M56 (2n = 36); nStachyphrynium latifolium M86 (2n = 26) and oStachyphrynium placentarium M85. Scale bar 10 µm
For regression analysis, linear models were calculated between the four dependent variables MCl, TML, MCA and CVCL and the three predictor variables chromosome number (2n), ploidy level (pl) and basic chromosome number (x) using the software R (R Core Team 2014). Because a phylogenetic hypothesis including the species studied for their chromosomes is still lacking, we are using ordinary regression analysis instead of regressions on phylogenetic independent contrasts. It has been shown that results from the two types of analysis produce very similar results (Ricklefs and Starck 1996).
Results
Chromosome counts, basic numbers and ploidy levels
Chromosome counts in 43 accessions are listed in Table 1, along with information on previous reports for 51 taxa. For 24 species and for genus Hypselodelphys, chromosome numbers are reported for the first time. In 32 accessions, all cells studied consistently had the same chromosome number. Variable chromosome numbers, the most frequent being underlined in the following, occurred in eleven accessions, i.e. Goeppertia ornata (2n = 24, 26, 27), Hypselodelphys sp. (2n = 21, 22, 24), Maranta arundinacea (2n = 49, 50, 52; Fig. 1j), M. leuconeura (2n = 49, 52), Marantochloa conferta (2n = 25, 26; Fig. 1k), M. purpurea (2n = 21, 22, 24, 26), Megaphrynium macrostachyum (2n = 34, 36, 38; Fig. 1c), Thalia geniculata (2n = 29, 32, 33, 35; Fig. 1b) and Trachyphrynium brauneanum (2n = 22, 24; Fig. 1d). Numerical variation within the same individual was infrequently observed in Maranta noctiflora (2n = 52 counted in three cells, 2n = 56 in one cell) and Marantochloa cordifolia (2n = 36 in four cells, 2n = 34 in one cell).
The most likely basic chromosome numbers and ploidy levels were as follows: x = 9 in Ctenanthe setosa, Marantochloa cordifolia, Megaphrynium macrostachyum, all of which had 2n = 4x = 36, and Ctenanthe dasycarpa with 2n = 8x = 72 (Fig. 1c, i, m). x = 10 occurred in Ctenanthe burle-marxii (accessions M22 and M77) and Thaumatococcus danielii, both 2n = 2x = 20 (Fig. 1g). x = 11 was found in Donax canniformis, Hypselodelphys sp., Phacelophrynium interruptum, Phrynium imbricatum, P. pedunculiferum, P. pubinerve, P. terminale and Trachyphrynium brauneanum, all of which had 2n = 2x = 22, Thalia geniculata with 2n = 3x = 33 and Stromanthe sanguinea with 2n = 4x = 44 (Fig. 1a, b, d, h). x = 12 with 2n = 2x = 24 had only Pleiostachya pruinosa. The most frequent basic number of x = 13 was consistently found in nine of ten accessions of genus Goeppertia, in Halopegia azurea, Sarcophrynium brachystachyum, S. schweinfurthianum, Stachyphrynium placentarium and S. latifolium, all of which had 2n = 2x = 26 (Fig. 1e, n, o). It occurred also in the five studied species of Maranta, which had 2n = 4x = 52 (Fig. 1j), in five of the six species of genus Marantochloa (M. conferta, M. leucantha, M. mannii, M. purpurea), which had 2n = 2x = 26 (Fig. 1k, m) and 2n = 5x = 65 (only M. congensis; Fig. 1l). x = 14 occurred only in Goeppertia picturata, which had 2n = 2x = 28 (Fig. 1f).
Infrageneric variation in basic chromosome numbers was encountered in Ctenanthe, Goeppertia and Marantochloa. Species of Ctenanthe had x = 9 or x = 10 and of Goeppertia x = 13 or x = 14 (see above). Although x = 13 was the most common basic number in Marantochloa, M. cordifolia had x = 9 with 2n = 4x = 36.
Karyotypes
In the karyograms of Figs. 2, 3 and 4 usually one or two chromosome pairs of the complements show secondary constrictions and/or are almost twice as large as the other chromosomes. They were designated in the following as satellite chromosomes (SAT). Two of such pairs were present in three Asian accessions of the x = 11 diploids Phrynium and Phacelophrynium as well as in African x = 9 tetraploid Marantochloa cordifolia. One pair was present in x = 11 Asian diploid Donax canniformis, African triploid Thalia geniculata, x = 13 Asian diploids Stachyphrynium placentarium and S. latifolium, American x = 9 tetraploid Ctenanthe setosa, x = 11 tetraploid Stromanthe sanguinea, x = 13 diploids of Goeppertia and African Halopegia azurea, African x = 9 tetraploid Megaphrynium macrostachyum, x = 10 diploid Thaumatococcus danielii as well as x = 11 diploid Trachyphrynium brauneanum and Hypselodelphys sp. In the karyograms of all other accession, no chromosomes with clearly visible satellites could be verified with the method used.
Karyotype analyses with detailed measurements are presented in Table 1. Chromosomes of Marantaceae are small. Mean chromosome lengths (MCL) varied from 0.8 µm in 4x Maranta arundinacea to 1.9 µm in 4x Megaphrynium macrostachyum. The total monoploid lengths (TML) ranged from 10.0 μm in 4x Maranta arundinacea to 22.8 μm in 2x Stachyphrynium latifolium. Mean centromeric asymmetry (MCA) varied from 12.4 in 3x Thalia geniculata (symmetrical chromosomes) to 26.8 in 4x Marantochloa cordifolia with more asymmetrical chromosomes. Interchromosomal asymmetry (CVCL), indicating the variation of chromosome lengths within the whole complement, was lowest in Sarcophrynium schweinfurthianum (CVCL 11.5) and highest in Goeppertia ornata (CVCL 38.6).
Regression analysis
In Fig. 5, the results for the linear regression models are presented as scatter plots, which refer to the dependent (MCL, TML, MCA, CVCL) and independent variables (2n, pl, x). Calculations are listed under each plot with the following parameters: an estimate of the regression slope, its standard error, the t test statistic and the two-sided significance level for the null hypothesis of regression slopes equal to zero. Solid lines in A, B, F and I show significant linear regression, and dotted lines in the other plots represent not significant correlations. The four significant regression models in A, B, F and I had p values < 0.0008 that were significant after adjusting the p value for multiple testing, e.g. by the conservative Bonferroni correction, which divides the p value by the number of comparisons (0.05/12 = 0.004).
Regression analysis of chromosome data (2n chromosome number, ploidy level, basic chromosome number x) versus karyomorphology data (MCL mean chromosome length, TML total length of monoploid chromosome set, MCA mean centromeric asymmetry, CVCL interchromosomal asymmetry) in Marantaceae taxa. Parameter: circle an estimate of regression slope, star standard error, trianglet test statistic, square two-sided significance level for null hypothesis of regression slopes equal to zero. Regular lines significant-, dotted lines not significant linear regression
Discussion
Variation of chromosome numbers in Marantaceae
Chromosome counts in 43 accessions, along with information on previous reports for 51 taxa (15 of them were the same species; Table 1), reveal consistencies in eight species. In seven species, we found deviating chromosome numbers. In our study, a consistent somatic chromosome number occurred in 32 out of the 43 investigated accessions. Aberrations were rather the exception than the rule. Variation of chromosome numbers was found in only eleven accessions. In nine of them, two to four different chromosome numbers were observed (Table 1). In these instances, we consider the most frequently occurring somatic number of the respective taxon to establish the monoploid chromosome number and ploidy level (2x, 3x, 4x). Plants with different somatic chromosome numbers in root tip cells often have uniform euploid chromosome numbers at meiosis (Joachimiak et al. 2001) and mostly form bivalents during meiotic prophase (Armstrong 1981, 1984 and references therein). Possibly due to the rarity of flowering in green house cultures of Marantaceae species, reliable information on meiosis is scarce and the literature records are restricted to only a few species of the genera Calathea, Stromanthe, Maranta (Bisson et al. 1968; Mahanty 1970). In the light of our findings, we suppose that the chromosome numbers are rather stable at least within the generative tissues.
Somatic variation in chromosome numbers has been observed in root tips of many angiosperm species (e.g. Fedorov 1969; Kula 1999; Winterfeld et al. 2015). It was suggested that it is either a result of chromosome rearrangements such as fusion or split after recent polyploidization (Smulders et al. 1994; Mishiba and Mii 2000) or a result of defective chromosome segregation as a consequence of meiotic disturbances caused by somatic irregularities, namely lagging and late separation of chromosomes (polysomaty and aneusomaty; Kula 1999; Mishiba and Mii 2000; Orr et al. 2015).
Our study did not verify far-reaching inconsistencies of chromosome numbers in Marantaceae species that were reported by previous investigations (Table 1; Sharma and Bhattacharyya 1958; Bisson et al. 1968). Hence, we suppose that such findings are due to artefacts from preparation, misidentification of the specimens studied or the use of outdated genus and species concepts, and will allow far-reaching quantification of chromosome number variation (Peruzzi et al. 2014) and to test their phylogenetic and adaptional effects (Carta et al. 2018) in the future.
Dysploidy and polyploidy caused the extensive chromosome number variation
Our study validates the occurrence of considerable somatic chromosome number variation in Marantaceae. The presumed basic chromosome numbers and the inferred ploidy levels of the studied taxa are specified in Table 1. Our chromosomal survey revealed eleven different somatic chromosome numbers, namely 2n = 20, 22, 24, 26, 28, 33, 36, 44, 52, 65 and 72, which are considered as multiples of x = 9, 10, 11, 12, 13 and 14 as the possibly genuine basic chromosome numbers of the taxa investigated. Consequently, plants with these particular somatic chromosome numbers described above may correspond to diploids, triploids, tetraploids, pentaploids and octoploids, respectively.
Bisson et al. (1968) provided an explanation for the wide variation of chromosome numbers in the family Marantaceae by invoking frequent hybridization of taxa with different basic numbers. He presented a network spanning nearly all arithmetically possible chromosome numbers. However, it should be considered that in general only species with similar karyotypes are easily capable to produce hybrids, whereas species with different karyotypes are usually not able to cross (Baltisberger and Hörandl 2016). Despite this, hybridization is rather frequent in Marantaceae as shown by recent phylogenetic (interspecific) and phylogeographic (intraspecific) studies (Ley and Claßen-Bockhoff 2011; Ley and Hardy 2013, 2014, 2017). However, due to the facts that species of Marantaceae (1) propagate mainly by vegetative means, (2) seed setting after flowering is very rare (Sharma and Bhattacharyya 1958) and (3) the origin of such chromosomal biotypes through sexual reproduction is fairly impossible (Mahanty 1970), we suppose that hybridization plays also, but only an ancillary role in the chromosome evolution of Marantaceae species in our investigation. Few verifiable instances form the triploid Thalia geniculata and pentaploid Marantochloa congensis, in which meiotic disturbances cannot be excluded. For the taxa of Marantaceae, it is more likely that dysploidy, through gains and losses of single chromosomes or fission and/or fusion of chromosome segments, is the most common mechanism of karyotypic change in the family. Polyploidy, due to the initiation of reproductive isolation between diploids and the established polyploids, is considered the most common mechanism of sympatric plant speciation (Stace 2000; Husband and Sabara 2003; Bolnick and Fitzpatrick 2007; Rieseberg and Willis 2007; Wood et al. 2009; Ramsey 2011). It played seemingly a significant, but not dominant, role in the evolution and diversification of Marantaceae (e.g. potentially in the separation of Marantochloa congensis, from its sister species M. sulphurea and M. monophylla; see Ley and Hardy 2014).
The broad variation of presumed basic chromosome numbers and the relatively low proportion of polyploid taxa found in our study (Table 1, Figs. 2, 3, 4) support the evolutionary significance of karyotype changes through dysploidy, which may have comparatively longer-term persistence over evolutionary time than polyploid changes that fail in many cases to persist (Escudero et al. 2014). Our observations in Marantaceae are in contrast to the attention polyploidy and dysploidy usually received in the literature. While the evolutionary role of polyploidy has been stressed in many reviews, chromosomal change via dysploidy was less regarded. Only a few studies suggested the prevalence of dysploidy in species diversification among angiosperms so far (e.g. Ray and Chisaki 1957; Grant 1981; Lee and Namai 1993; Vickery 1995; Soltis et al. 2001; Church 2003; Mandakova and Lysak 2018; Winterfeld et al. 2018).
Chromosome structure and trends in chromosome evolution
Data on chromosome morphology are important to understand the variation in chromosome numbers and to identify potentially different genomes within a plant family. So far, karyotype studies of only a few species of Marantaceae have been published (Sharma and Bhattacharyya 1958; Mahanty 1970; Mukhopadhyay and Sharma 1987; Eksomtramage et al. 2007). Our detailed karyotype analyses reveal that the chromosomes are overall comparatively small (< 2 µm in length), and complement lengths (TML) are comparatively short. Mean centromeric asymmetry (MCA) and variation of chromosome lengths (CVCL) within the whole complements are variable. All in all, karyotypes of the studied taxa reveal considerable variation, but it is rather continuous.
Regression analyses of chromosome number (2n), ploidy level (pl) and basic chromosome number (x) versus karyotype data, such as mean chromosome length (MCL), total length of a monoploid chromosome set (TML), mean centromeric asymmetry (MCA) and interchromosomal asymmetry (CVCL), respectively, were conducted (Fig. 5). The following trends in karyotype evolution were retrieved: There is a linear correlation between 2n chromosome number and MCL/TML (Fig. 5a, b; significant in both), meaning that an increase in 2n is linked with a decrease in MCL/TML. An increase in 2n is coupled with polyploidy as seen in the ploidy-level graph (Fig. 5e, f; significant in f). There are no correlations between 2n and MCA (Fig. 5c) as well as between ploidy level and MCA (Fig. 5g). 2n and CVCL showed a weak interrelation (Fig. 5d), which is caused mainly by polyploidization (Fig. 5h). Thus, chromosomes of a complement seem to converge in their length after whole-genome duplication. Basic number x and MCL revealed a significant linear correlation (Fig. 5i). Some hypotheses were developed for the explanation of chromosome number variation in various angiosperm species (Fusion-, Fission-, Modal-Hypothesis; White 1973; Todd 1970, 1975; Matthey 1973) and to interpret the observed patterns in chromosome morphology and behaviour (e.g. Schubert and Lysak 2011; Lysak et al. 2006; Escudero et al. 2014).
However, the direction of dysploidy change cannot be fully determined in Marantaceae. There are two possibilities: either increasing dysploidy accompanied by decreasing chromosome length (MCL) or, alternatively, decreasing dysploidy accompanied by increasing chromosome length. The latter process leads to fewer and larger chromosomes and is a frequent pattern in monocots following polyploidization (Carta et al. 2018). To reveal its role in diploid and polyploidy Marantaceae and to determine the original basic number in the family, however, a comparison of chromosome data with a molecular phylogenetic framework would be essential, which is not yet available. There is no significant correlation or trend visible for change in x and TML, MCA and CVCL values (Fig. 5j, k, l).
Biogeographic differentiation
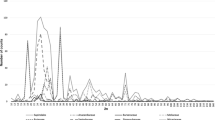
Variation of chromosome and karyotype characters was analysed in relation to the main distribution areas (America, Africa, Asia) and their species diversity (Fig. 6). All in all, the strongest variation occurred in the taxa of tropical America. They displayed eight different 2n chromosome numbers, three ploidy levels and six different basic chromosome numbers. We sampled 17 of about 450 species in this region. Lower variation with six different 2n numbers, two ploidy levels (except for triploid and pentaploid hybrids) and four different basic numbers were characteristic of tropical Africa, a region with ca. 50 species, of which we sampled 14 species. The lowest variation occurred in the Asian taxa, of which we studied eight of about 50 species. They had only two different chromosome numbers, two different ploidy levels and were consistently diploid. Mean centromeric asymmetry (MCA) was similar in taxa from all continents. African taxa stood out by low interchromosomal asymmetry (CVCL). Interestingly, high rates of chromosomal variation are characteristic of taxa with low chromosome sizes, both MCL and TML, as typically found in the American taxa (MCL 1.2 µm, TML 14.5 µm; Fig. 6). The lowest chromosomal variation occurred in the Asian taxa having the largest chromosomes (MCL 1.6 µm, TML 18.5 µm). African taxa were intermediate (MCL 1.3 µm, TML 15.5 µm). One explanation would be the fact that smaller genomes might have improved evolvability through benefiting from raising general genome flexibility (Puttick et al. 2015).
Relations between chromosome and karyotype data and species diversity in American, African and Asian Marantaceae. Number of different 2n chromosome numbers, pl ploidy levels, x basic chromosome numbers and species number are on the left-hand side. Mean values of karyotype parameters: MCL mean chromosome length, TML total length of monoploid chromosome set, MCA mean centromeric asymmetry, CVCL interchromosomal asymmetry, are on the right-hand side. The lines between the symbols are only ledger lines for better visualizing of common chromosome parameters from the three areas. The highest variability of chromosome data occurred in America (red), lower variability in Africa and the lowest variability in Asia, which corresponds widely with the respective species diversity
If we accept the suggestion of Africa as the original home of the family (Mahanty 1970; Andersson and Chase 2001; Prince and Kress 2006), a karyotype with x = 13 could possibly be the phylogenetically ancestral state as hypothesized by Mahanty (1970). Decreasing dysploidy would have led to basic numbers of x = 11, 10 and 9. Diploids prevail, and tetraploids occur sporadically such as triploid and pentaploid hybrids (Fig. 6). The relatively low species diversity in Africa is possibly due to higher extinction rates as a result of shrinking lowland tropical forests during the tertiary (Maley 1996; Prince and Kress 2006).
Following Prince and Kress (2006), the New World tropics were reached secondarily by dispersal events from Africa. The enormous species richness in America and higher chromosomal diversity (basic chromosome numbers of x = 9, 10, 11, 12, 13, 14; diploid, tetraploid and octoploid ploidy levels) thus would reflect a secondary radiation and not indicate the original home of the family in America as proposed by Holttum (1951).
The Asian tropics were considered under all scenarios to be colonized secondarily by Marantaceae. On the basis of the present knowledge, the species have a low variation of chromosome features and chromosomes are larger than in the other regions, which might support their status as phylogenetically rather derived.
References
Al-Gharaibeh MM (2017) Seed germination and genetic structure of two Salvia species in response to environmental variables among phytogeographic regions in Jordan (Part I) and phylogeny of the pan-tropical family Marantaceae (Part II). PhD Thesis, Martin Luther University, Halle-Wittenberg
Andersson L (1998) Marantaceae. In: Kubitzki K (ed) The families and genera of vascular plants, vol 4. Springer, Berlin, pp 278–293
Andersson L, Chase MW (2001) Phylogeny and classification of Marantaceae. Bot J Linn Soc 135:275–287
Armstrong KC (1981) The evolution of Bromus inermis and related species of Bromus sect. Pnigma Bot Jahrb Syst 102:427–443
Armstrong KC (1984) Chromosome pairing affinities between Old and New World species of Bromus section Pnigma. Canad J Bot 62:581–585
Baack E, Melo MC, Rieseberg LH, Ortiz-Barrientos D (2015) The origins of reproductive isolation in plants. New Phytol 207:968–984. https://doi.org/10.1111/nph.13424
Baltisberger M, Hörandl E (2016) Karyotype evolution supports the molecular phylogeny in the genus Ranunculus (Ranunculaceae). Perspect Pl Ecol Syst 18:1–14
Bharathan G, Lambert G, Galbraith DW (1994) Nuclear DNA content of monocotyledons and related taxa. Amer J Bot 81:381–386
Bisson S, Guillemet S, Hamel JL (1968) Contribution a l’étude caryo-taxonomique des Scitaminées. Mém Mus Natl Hist Nat, B Bot 18:59–133
Bolnick DI, Fitzpatrick BM (2007) Sympatric speciation: models and empirical evidence. Annual Rev Ecol Evol Syst 381:459–487
Carta A, Bedini G, Peruzzi L (2018) Unscrambling phylogenetic effects and ecological determinants of chromosome numbers in major angiosperms clades. Sci Rep 8:14258. https://doi.org/10.1038/s41598-018-32515-x
Cave MS (1960) Index to plant chromosome numbers for 1959. California Botanical Society, Berkeley
Church SA (2003) Molecular phylogenetics of Houstonia (Rubiaceae): descending aneuploidy and breeding system evolution in the radiation of the lineage across North America. Molec Phylogen Evol 27:223–238
Dhetchuvi JB (1996) Taxonomie et phytogéographie des Marantaceae et des Zingiberaceae de l’Afrique Centrale (Gabon, Congo, Zaire, Rwanda et Brundi). PhD Thesis Université Libre de Bruxelles, Bruxelles
Eksomtramage L, Jornead S, Decharun S, Jansone A, Tanpho S (2007) Chromosome numbers of some angiosperm plants in Thailand Songklanakarin. J Sci Technol 29:61–72
Escudero M, Martín-Bravo S, Itay Mayrose I, Fernández-Mazuecos M, Fiz-Palacios O, Hipp AL, Pimentel M, Jiménez-Mejías P, Valcárcel V, Vargas P, Luceño M (2014) Karyotypic changes through dysploidy persist longer over evolutionary time than polyploid changes. PLoS ONE 9:e85266
Fedorov A (1969) Chromosome numbers of flowering Plants. Nauka, Leningrad
Gadella TW (1982) IOPB chromosome number reports LXXVI. Taxon 31:595–596
Grabowska-Joachimiak A, Kula A, Gernand-Kliefoth D, Joachimiak AJ (2015) Karyotype structure and chromosome fragility in the grass Phleum echinatum Host. Protoplasma 252:301–306. https://doi.org/10.1007/s00709-014-0681-5
Grant V (1981) Plant speciation, 2nd edn. Columbia University Press, New York
Hanson L, Leitch IJ, Bennett MD (1999) Unpublished values from the Jodrell Laboratory, Royal Botanic Gardens, Kew. Original references for DNA C-values listed in the Plant DNA C-values database (release 5.0 Dec. 2010)
Holttum RE (1951) The Marantaceae of Malaya. Gard Bull Singapore 13:254–296
Husband BC, Sabara HA (2003) Reproductive isolation between autotetraploids and their diploid progenitors in fireweed, Chamerion angustifolium (Onagraceae). New Phytol 161:703–713
Joachimiak A, Kula A, Śliwínka E, Sobieszczanska A (2001) C-banding and nuclear DNA amount in six Bromus species. Acta Biol Cracov, Ser Bot 43:105–115
Jongkind CCH (2008) Two new species of Hypselodelphys (Marantaceae) from West Africa. Adansonia Ser 3 30:57–62
Kennedy H (1978a) Systematics and Pollination of the “Closed-Flowered” Species of Calathea (Marantaceae). Constancea 71. University of California Press, Berkeley
Kennedy H (1978b) Notes on Central American Marantaceae: 3. New species of Calathea from Costa Rica and Panama. Brenesia 14:349–356
Kennedy H (2000) Diversification in pollination mechanisms in the Marantaceae. In: Wilson KL, Morrison DA (eds) Monocots: systematics and evolution. CSIRO, Melbourne
Knight A, Molinari NA, Petrov DA (2005) The large genome constraint hypothesis: evolution, ecology and phenotype. Ann Bot (Oxford) 95:177–190
Kula A (1999) Cytogenetic studies in the cultivated form of Bromus carinatus (Poaceae). Fragm Florist Geobot Polon 7:101–106
Lee KH, Namai H (1993) Cytogenetic and morphological characteristics of new types of diploids (2n = 22, 24, 40) derived from consecutive selfing of aneuploids in Brassica crops. Euphytica 1–2:15–22
Ley AC, Claßen-Bockhoff R (2011) Evolution in African Marantaceae—evidence from phylogenetic, ecological and morphological studies. Syst Bot 36:1–14
Ley AC, Hardy OJ (2013) Improving AFLP analysis of large-scale patterns of genetic variation—a case study with the Central African lianas Haumania spp (Marantaceae) showing interspecific gene flow. Molec Ecol 22:1984–1997
Ley AC, Hardy OJ (2014) Contrasting patterns of gene flow between sister plant species in the understorey of African moist forests—the case of sympatric and parapatric Marantaceae species. Molec Phylogen Evol 77:264–274
Ley AC, Hardy OJ (2017) Hybridization and asymmetric introgression after secondary contact in two tropical African climber species Haumania danckelmaniana and H. liebrechtsiana (Marantaceae). Int J Pl Sci 178:421–430
Lowry DB, Willis JH (2010) A widespread chromosomal inversion polymorphism contributes to a major life-history transition, local adaptation, and reproductive isolation. PLoS Biol 8:e1000500. https://doi.org/10.1371/journal.pbio.1000500
Lysak MA, Berr A, Pecinka A, Schmidt R, McBreen K, Schubert I (2006) Mechanisms of chromosome number reduction in Arabidopsis thaliana and related Brassicaceae species. Proc Natl Acad Sci USA 103:5224–5229. https://doi.org/10.1073/pnas.0510791103
Mahanty HK (1970) A cytological study of the Zingiberales with special reference to their taxonomy. Cytologia 35:13–48
Maley J (1996) The African rainforest—main characteristics of changes in vegetation and climate from the Upper Cretaceous to the Quaternary. Proc Roy Soc Edinburgh B Biol 104B:31–73
Mandakova T, Lysak MA (2018) Post-polyploid diploidization and diversification through dysploid changes. Curr Opin Pl Biol 42:55–65
Mangenot S, Mangenot G (1957) Nombres chromosomiques nouveaux chez diverses Dicotylédones et Monocotylédones d’Afrique occidentale. Bull Jard Bot État Bruxelles 27:639–654
Mangenot S, Mangenot G (1958) Deuxième liste de nombres chromosomiques nouveaux chez diverses Dicotylédones et Monocotylédones d’Afrique occidentale. Bull Jard Bot État Bruxelles 28:315–329
Mangenot S, Mangenot G (1962) Enquête sur les nombres chromosomiques dans une collection d’esp èces tropicales. Acta Bot Gallica 109:411–447. https://doi.org/10.1080/00378941.1962.10838117
Matthey R (1973) The chromosome formulae of eutherian mammals in cytotaxonomy and vertebrate evolution. In: Chiarelli AB, Capanna E (eds) Cytotaxonomy and vertebrate evolution. Academic Press, London, pp 531–616
Miege J (1960) Nombres chromosomiques de plantes d’Afrique Occidentale. Rev Cytol Biol Vég 21:373–384
Mishiba K, Mii M (2000) Polysomaty analysis in diploid and tetraploid Portulaca grandiflora. Pl Sci 156:213–219. https://doi.org/10.1016/S0168-9452(00)00257-0
Mukherjee S (1981) Cytomorphological studies on Datura metel Linn. from different ecological habitats. Proc Indian Sci Congr Assoc 68 (Sect. VI): 84
Mukhopadhyay S, Sharma AK (1987) Karyomorphological analysis of different species and varieties of Calathea, Maranta and Stromanthe of Marantaceae. Cytologia 52:821–831
Orr B, Godek KM, Compton D (2015) Aneuploidy. Curr Biol 25:523–548
Ouyang Y, Zhang Q (2013) Understanding reproductive isolation based on the rice model. Annual Rev Pl Biol 64:111–135
Paszko B (2006) A critical review and a new proposal of karyotype asymmetry indices. Pl Syst Evol 258:39–48
Peruzzi L (2013) “x” is not a bias, but a number with real biological significance. Pl Biosyst 147:1238–1241. https://doi.org/10.1080/11263504.2013.86.1533
Peruzzi L, Eroğlu H (2013) Karyotype asymmetry: again, how to measure and what to measure? Comp Cytogen 7:1–9
Peruzzi L, Goralski G, Joachimiak AJ, Bedini G (2012) Does actually mean chromosome number increase with latitude in vascular plants? An answer from the comparison of Italian, Slovak and Polish floras. Comp Cytogen 6:371–377
Peruzzi L, Caparelli KF, Bedini G (2014) A new index for the quantification of chromosome number variation: an application to selected animal and plant groups. J Theor Biol 353:55–60
Prince LM, Kress WJ (2006) Biogeography of the prayer plant family: getting to the root problem in Marantaceae. In: Columbus JT, Friar EA, Porter JM, Prince LM, Simpson MG (eds) Monocots: comparative biology and evolution. Rancho Santa Ana Botanic Garden, Claremont, pp 643–657
Puttick MN, Clark J, Donoghue PCJ (2015) Size is not everything: rates of genome size evolution, not C-value, correlate with speciation in angiosperms. Proc Roy Soc B 282:20152289. https://doi.org/10.1098/rspb.2015.2289
Ramsey J (2011) Polyploidy and ecological adaptation in wild yarrow. Proc Natl Acad Sci USA 108:7096–7101
Ray PM, Chisaki HF (1957) Studies on Amsinckia. II. Relationships among the primitive species. Amer J Bot 44:529–536
Ricklefs RE, Starck JM (1996) Applications of phylogenetically independent contrasts: a mixed progress report. Oikos 77:167–172
Rieseberg LH, Willis JH (2007) Plant speciation. Science 317:910–914
Sato D (1948) Karyotype and systematics of Zingiberales. Jap J Genet 23:44–45
Sato D (1960) The karyotype analysis in Zingiberales with special reference to the protokaryotype and stable karyotype. Sci Pap Coll Gen Educ Univ Tokyo 10:225–243
Schubert I, Lysak MA (2011) Interpretation of karyotype evolution should consider chromosome structural constraints. Trends Genet 27:207–216. https://doi.org/10.1016/j.tig.2011.03.004
Schwarzacher T, Ambros P, Schweizer D (1980) Application of Giemsa banding to orchid karyotype analysis. Pl Syst Evol 134:293–297
Sharma AK, Bhattacharyya NK (1958) Inconstancy in chromosome complements in species of Maranta and Calathea. Proc Natl Inst Sci India 24B:101–117
Sharma AK, Mukhopadhyay S (1984) Feulgen microspetrophotometric estimation of nuclear DNA of species and varieties of three different genera of Marantaceae. Proc Indian Acad Sci 93:337–347
Simmonds NW (1954) Chromosome behaviour in some tropical plants. Heredity 8:139–146
Smulders MJM, Rus-Kortekaas W, Gilissen LJW (1994) Development of polysomaty during differentiation in diploid and tetraploid tomato (Lycopersicon esculentum) plants. Pl Sci 97:53–60. https://doi.org/10.1016/0168-9452(94)90107-4
Soltis DE, Tago-Nakazawa M, Xiang Q-Y, Kawanon S, Murata J et al (2001) Phylogenetic relationships and evolution in Chrysosplenium (Saxifragaceae) based on matK sequence data. Amer J Bot 88:883–893
Stace CA (2000) Cytology and cytogenetics as a fundamental resource for the 20th and 21th centuries. Taxon 49:451–477
Stevens PF (2001 onwards) Angiosperm Phylogeny Website. Version 14, July 2017. Available at: http://www.mobot.org/MOBOT/research/APweb/
Suessenguth K (1921) Bemerkungen zur meiotischen und somatischen Kernteilung bei einigen Monokotylen. Flora 114:313–328
Suksathan P, Borchsenius F (2005) Nomenclatural synopsis of the Marantaceae in Thailand. Taxon 54:1083–1090
R Core Team (2014) R: a language and environment for statistical computing, version 3.1.0. Available at: www.r-project.org. Accessed 16 Jun 2014
Todd NB (1970) Karyotypic fissioning and canid phylogeny. J Theor Biol 26:445–480
Todd NB (1975) Chromosomal mechanisms in the evolution of artiodactyls. Paleobiology 1:175–188
Venkatasubban KR (1946) A preliminary survey of chromosome numbers in Scitamineae of Bentham and Hooker. Proc Indian Acad Sci B 23:281–300
Vickery RK (1995) Speciation by aneuploidy and polyploidy in Mimulus (Plantaginaceae). Great Basin Nat 55:174–176
Vovides AP, Lascurain M (1995) Numeros cromosomicos de cuatro especies de Costus (Costaceae), una de Calathea, una de Maranta y una de Stromanthe (Marantaceae). Acta Bot Mex 33:81–86
White MJD (1973) Animal cytology and evolution. Cambridge University Press, Cambridge
Winterfeld G, Perner K, Röser M (2014) Polyploidy and hybridization as main factors of speciation: complex reticulate evolution within the grass genus Helictochloa. Cytogenet Genome Res 142:204–225
Winterfeld G, Schneider J, Becher H, Dickie J, Röser M (2015) Karyosystematics of the Australasian stipoid grass Austrostipa and related genera: chromosome sizes, ploidy, chromosome base numbers, and phylogeny. Austral Syst Bot 28:145–159
Winterfeld G, Wölk A, Röser M (2016) Genome evolution in alpine oat-like grasses through homoploid hybridization and polyploidy. AoB PLANTS 8:plw039. https://doi.org/10.1093/aobpla/plw039
Winterfeld G, Becher H, Voshell S, Hilu K, Röser M (2018) Karyotype evolution in Phalaris (Poaceae): the role of reductional dysploidy, polyploidy and chromosome alteration in a wide-spread and diverse genus. PLoS ONE 13:e0192869. https://doi.org/10.1371/journal
Wölk A, Winterfeld G, Röser M (2015) Genome evolution in a Mediterranean species complex: phylogeny and cytogenetics of Helictotrichon (Poaceae) allopolyploids based on nuclear DNA sequences (rDNA, topoisomerase gene) and FISH. Syst Biodivers 13:326–345. https://doi.org/10.1080/14772000.2015.1023867
Wood TE, Takebayashi N, Barker MS, Mayrose I, Greenspoon PB, Rieseberg LH (2009) The frequency of polyploid speciation in vascular plants. Proc Natl Acad Sci USA 106:13875–13879
Acknowledgements
Open Access funding provided by Projekt DEAL. We thank Mirko Hause (Botanical Garden of the University Halle-Wittenberg) for taking care of the living plant collection and Denise Marx (Herbarium HAL) for preparing of the herbarium vouchers. We are grateful to Ina Reichelt for assistance in laboratory work.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Handling Editor: Martin A. Lysak.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Winterfeld, G., Ley, A., Hoffmann, M.H. et al. Dysploidy and polyploidy trigger strong variation of chromosome numbers in the prayer-plant family (Marantaceae). Plant Syst Evol 306, 36 (2020). https://doi.org/10.1007/s00606-020-01663-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00606-020-01663-x