Abstract
Tortella rigens occurs in the European Baltic region and at a few localities in North America. At least 95 % of all localities are found in southern Sweden. It is morphologically similar to the more widespread T. bambergeri, which was found in Sweden during this study. Haplotype networks, based on ITS or the chloroplast markers atbB-rbcL and rps4, for T. rigens, T. bambergeri, and closely related species display significant reticulation and NeighborNet and Jacknife analyses are used to explore the species’ relationships. Significant incongruence is found between the two molecular data sets or between these and the morphologically defined species, and several species appear polyphyletic or paraphyletic. Sporophytes of T. rigens are described and appear to be a result of hybridisation, which could partly explain the reticulation. Among the four South Swedish T. rigens populations, those of Gotland (GTL) and the Stockholm archipelago (SAR) are not differentiated molecularly from each other and there are signs of significant migration between them. The ones in Västergötland (VAS) and Öland (OEL) are distinct from each other and from GTL and SAR. The haplotype pattern of GTL displays signs of population increase whereas a decrease is found for VAS. Although the time scale is uncertain, this decrease is of conservation concern, considering that VAS has one unique haplotype. OEL has both the highest number of unique haplotypes (4) and the highest haplotype diversity of all four populations and has therefore the highest importance for conservation. Tortella rigens and T. bambergeri can be differentiated by atpB-rbcL.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
In southern Scandinavia and nearby areas around the Baltic Sea, many bryophyte species are more or less restricted to regions with large surfaces of calcareous rock or soil. Compared with their surroundings, these regions are relatively sparsely covered by more competitive vascular plants. Such regions occur especially in the Swedish provinces Västergötland, Öland, Gotland, the Stockholm archipelago with adjacent mainland, and in western Estonia. Among the southern Scandinavian species that are almost restricted to these regions we find both some that occur also in the Scandinavian mountain range (Hedenäs 2014) and those that are known only from the lowlands. One of the latter is Tortella rigens Alberts.
Tortella rigens was described from southern Sweden in the middle of the 20th century and has for a long time been considered endemic to the European Baltic area, where it occurs in all five regions mentioned above (Albertson 1946; Nyholm 1991; Vellak and Ingerpuu 2012). In addition, the species was recently found in the westernmost portion of the Åland archipelago in Finland (Huttunen et al. 2014) and in eastern North America (Eckel 1998, 2007). Earlier reports of this species from Central Europe (Düll 1984; Pilous 1965) were erroneous (H. Köckinger, personal communication, 2013). Only a few tens of localities for T. rigens have so far been reported from outside Sweden. Based on the almost 400 registered collections in the Swedish herbaria (http://www.herbarium-ume.se/virtuella_herbariet/index.html; checked 18 March 2014), and own observations of the abundance of the species in the Swedish regions where it occurs, Sweden probably harbours more than 95 % of the localities of T. rigens. Sweden therefore has a special responsibility for this species’ long-term survival.
The genus Tortella includes several taxa in which origin, status, circumscription, or distribution are presently unclear. For example, Tortella rigens has been suggested to be of hybrid origin (Albertson 1946; Eckel 1998; Persson 1947) and T. bambergeri (Schimp.) Broth. was only recently found in Britain and Norway, despite that both areas are well-studied bryologically (Bosanquet 2006; Hassel and Høitomt 2013). Tortella bambergeri is similar to T. rigens in habit, especially when moist. Considering the restricted distribution and global rarity of T. rigens it is crucial to know whether some or numerous of the Scandinavian occurrences of this species belong to T. bambergeri. In the latter case, T. rigens could be even rarer than so far thought.
Because Sweden has a great responsibility for T. rigens, it is essential to understand both the species’ origin and how its intraspecific diversity and variation are distributed among the four main Swedish regional populations. Only with such knowledge it is possible to decide whether all regions require conservation management to secure the species’ long-term survival. Since T. rigens is almost completely restricted to the four-mentioned regions, T. rigens is also interesting as a model species for studying intraspecific variation in species with small and isolated distribution areas (Frankham et al. 2002). We also need to know how to differentiate T. rigens molecularly from the morphologically similar T. bambergeri. The following questions are addressed in this study: (1) which species are most closely related to T. rigens? Can this explain its morphological features? (2) How is the intraspecific variation partitioned among the four main Swedish regional populations? Are the populations stable and can we find evidence of migration among the populations. (3) Can the molecular markers used to reconstruct the phylogenetic relationships of T. rigens also serve as molecular tools, barcodes, to distinguish it from T. bambergeri?
Materials and methods
Study species and material
Tortella rigens grows in small to large and relatively dense cushions, and is often intermixed with other acrocarpous mosses, including other Tortella species. Individual shoots range from less than one to several centimetres tall. The leaves are lanceolate and narrow gradually towards the apex. When moist the leaves are straight and erect or erect-spreading, and when dry they are twisted and crisped, but not as strongly as in T. tortuosa. The margin is plane, and the costa has long, narrow cells on both the ad- and abaxial sides. The median and distal leaf lamina cells are strongly papillose and mostly 11–14 µm wide, and the border towards the long, thin-walled and hyaline basal cells is V-shaped. The species is dioicous and when this investigation started sporophytes were unknown in the species (Eckel 1998; Nyholm 1991). The species grows in habitats with abundant calcareous rock and soil, typically flat or slightly sloping and with at most sparse vascular plant vegetation.
Tortella bambergeri differs from T. rigens in having smaller median and distal leaf lamina cells, and the costa is covered by short and papillose cells both adaxially and at least mostly in its distal, abaxial portion (Bosanquet 2006; Eckel 2010). The occurrence of T. bambergeri in Sweden was not confirmed when this study started (Nyholm 1991), but its known distribution includes large portions of Europe, from Caucasus to the British Isles and Norway, as well as eastern North America (Bosanquet 2006; Eckel 2010; Hassel and Høitomt 2013). Considering the Norwegian finds, its occurrence in Sweden was expected and the species was found at a few localities in connection with this study. Tortella bambergeri can grow in similar habitats as T. rigens, but in addition in escarpments and on less calcareous rocks, such as base-rich gneiss.
To place T. rigens in a wider phylogenetic context, and to explore whether molecular differences exist between T. rigens and T. bambergeri, sequences were generated for 74 samples of T. rigens, 11 of T. bambergeri, and 20 of other Tortella species that were placed close to T. rigens in earlier phylogenetic studies (Grundmann et al. 2006; Inoue et al. 2012; Werner et al. 2005). In addition, four such Tortella specimens, in which sequences were available on GenBank were included. As outgroup, three samples of Tortella humilis (one newly generated sequence, two from GenBank), two samples of Oxystegus tenuirostris and two of T. crispulum were used. These three species are successively more distantly related to the clade with Tortella species around T. rigens according to overviews of the Trichostomoideae based on ITS by Werner et al. (2005), rps4 by Inoue et al. (2012) and several markers by Grundmann et al. (2006). All included samples are listed in “Appendix 1”, where authors of the Latin names are also found.
The other aim of this study was to explore whether genetic differences exist among the four Swedish, the Estonian, Åland, and North American regional populations of T. rigens. Since the populations in Sweden are most abundant and represent most of the species’ global occurrences, the focus is on these. Of the 74 T. rigens specimens, 16 were from Öland (OEL), 15 from Gotland (GTL), 13 from Västergötland (VAS), 18 from the Stockholm archipelago with adjacent mainland (SAR), 4 from Estonia (EST), 3 from Åland, Finland (FIN) and 5 from Ontario, Canada (CAN) (Appendix 1).
Molecular methods
Total DNA was extracted using the DNeasy® Plant Mini Kit for DNA isolation from plant tissue (QIAGEN). Double-stranded DNA templates were prepared by polymerase chain reaction (PCR). PCR was performed using IllustraTM Hot Start Mix RTG (GE Healthcare) in a 25 µl reaction volume according to the manufacturer’s instructions.
Initially, variation in the nuclear internal transcribed spacers 1 and 2 (ITS) and the plastid atpB-rbcL spacer (atbB-rbcL), rpl16 G2 intron (rpl16), the rps4 gene + trnS-rps4 spacer (rps4) and trnLUAA-trnFGAA spacer (trnL-trnF) were explored for five specimens of T. rigens, four of T. bambergeri, and two for each of. T. fragilis and T. tortuosa. The three most variable ones, ITS, atpB-rbcL, and rps4 were selected for the investigation. For the three used molecular markers, the PCR programs given below were initiated by a denaturation step of 5 min at 95 ºC and were followed by a final extension period of 10 min at 72 ºC. For ITS the PCR programme employed was 40 cycles of 30 s at 95 ºC, 30 s at 52 ºC, and 1 min at 72 ºC, with the primers ‘ITS4-bryo’ and ‘ITS5-bryo’ (Stech 1999). In a few cases, the internal primers ‘5.8SC’ and ‘5.8SN’ (Bartish et al. 2005) were used. For atpB-rbcL, 40 cycles of 30 s at 94 ºC, 30 s at 52 ºC, and 40 s at 72 ºC were employed, with the primers ‘ATPB-1’ and ‘RBCL-1’ (Chiang et al. 1998). For rps4, 40 cycles of 30 s at 94 ºC, 30 s at 52 ºC, and 40 s at 72 ºC were employed, with the primers ‘rps5F’ (Nadot et al. 1994) and ‘trna5R’ (‘trnS’ in Souza-Chies et al. 1997).
Twenty micro litres of each amplified fragment was cleaned using a mixture of 20 units of Exonuclease I, E.coli and 4 units of FastAP TM Thermosensitive Alkaline Phosphatase (Fermentas LIFE SCIENCE), mixed and incubated at 37 ºC for 30 min and inactivated at 80 ºC for 15 min. Cycle sequencing was performed using the ABI BigDye Terminator Kit (Applied Biosystems) according to the instructions on the kit (BDT ver. 3.1), and the sequencing products were cleaned using the DyeEx® 96 Kit (QIAGEN). The same primers as for the initial PCR were used. Sequencing products were resolved on an ABI3130xl automated sequencer. Double-stranded sequencing was performed.
Sequence editing and analysis
Nucleotide sequence fragments were edited and assembled for each DNA region using PhyDE® 0.9971 (http://www.phyde.de/index.html). The assembled sequences were manually aligned in PhyDE®. Regions of partially incomplete data in the beginning and end of the sequences were identified and excluded from subsequent analyses. The codable gaps, when coded as present or absent, provided additional evidence to distinguish haplotypes and the analyses were thus performed with the insertions and deletions coded as single informative characters independent of their length. The sequence alignments used in the analyses are available on request. GenBank accession numbers are listed in “Appendix 1”.
Paralogous ITS haplotypes are rarely encountered in bryophytes (but see Košnar et al. 2012). However, the ITS chromatograms generated in this study did not show ‘messy’ patterns or noise that could suggest paralogy, and the 5.8S gene was invariable among the samples (cf. Feliner and Rosselló 2007; Shaw et al. 2002). The revealed ITS variation is thus interpreted as being among homologous haplotypes.
The programme TCS (Clement et al. 2000) was first used to evaluate relationships among specimens in a haplotype context. For the data with several Tortella species, reticulation was revealed in the haplotype networks based on either ITS or chloroplast data (not shown). Therefore, the statistical support for potential recombination in ITS was tested by the Φw statistic (Bruen et al. 2006) as implemented in SplitsTree 4.12.6 (Huson and Bryant 2006). Because of the occurrence of abundant reticulation, a split network was computed with the NeighborNet (NN) method as implemented in SplitsTree 4.12.6 (Huson and Bryant 2006) to visualize similarities or relationships among samples. A Jacknife analysis (1,000 replications) was performed with the programme TNT (Goloboff et al. 2003) to test whether supported lineages exist among the studied Tortella species in a phylogenetic tree context. ITS and chloroplast data were analysed separately, since both visual inspection of the split networks and Jacknife trees, and the ILD test (Farris et al. 1995; 400 replicates, p = 0.0025) indicated that the two are incongruent. Since neither the ITS nor chloroplast data yielded results that agree with a monophyletic T. rigens, a maximum likelihood (ML) analysis was made for the two data sets, both without and with T. rigens constrained as monophyletic, employing GARLI (Zwickl 2006), a programme suitable for analysing large data sets. The models GTR + G (ITS) and TPM2uf + I + G (chloroplast data) were selected by jModelTest2 using the AIC criterion (Darriba et al. 2012; Guindon and Gascuel 2003). Ten or, if this was not sufficient, 20 searches were run with GARLI to get at least four equal shortest trees for both the unconstrained and constrained ITS and chloroplast data. For each of the two data sets, PAUP 4.0 (Swofford 2002) was employed to compare the best unconstrained with the corresponding constrained result to see if these were significantly different from each other, using the Shimodaira–Hasegawa test with 1,000 replicates (Shimodaira and Hasegawa 1999).
To investigate patterns of haplotype variation among the four Swedish populations of T. rigens, an analysis of molecular variance (AMOVA) was performed with GENALEX 6.5 (Peakall and Smouse 2006, 2012), using haplotypes based on all markers together and identified with TCS. The null hypothesis in this and the following analysis is that there exist no differences in haplotype composition among the studied populations. Pair-wise PhiPT (an analogue of F ST, i.e., genetic diversity among populations) was also estimated with GENALEX 6.5, and the same programme was used to calculate the effective number of haplotypes (Ne) and the haplotype diversity (H; the probability that two random haplotypes are different) for each population.
To decide whether the Swedish T. rigens populations are stable in size, expanding or decreasing, Tajima’s D test of selective neutrality was employed (Tajima 1989). Tajima’s D test was preferred over Fu’s F S test (Fu 1997), because it has been shown that the latter should not be used when recombination levels are unknown (Ramírez-Soriano et al., 2008). Tajima’s D test was run in Arlequin ver. 3.5.1.3 (Excoffier and Lischer 2010). Estimations of immigration rates, M = m/µ (chance for a lineage to immigrate as rate per generation/neutral mutation rate per site per generation), into each of the four Swedish populations from every other population were made using the programme LAMARC (Kuhner 2006). Because dating points for bryophytes are very scarce, absolute mutation rates are not possible to apply. Found migration events are therefore assumed to have taken place during the postglacial period. The likelihood-based approach of LAMARC was employed, and the analysis was initially run three times with the default settings of the programme. The results of these initial analyses were unproblematic and this was therefore followed by two runs with the following settings: ten initial chains, each with 50,000 steps (after 8,000 steps were first discarded in each), and two final chains with 1,500,000 (75,000) steps each. The conditions for the analysis were met, including requirements concerning potential directional changes in Θ and Data lnL among the chains and that the Posterior lnL was considerably lower than 2–3 times the number of estimated parameters. Since the results were similar in the last two runs, only the results of the second run were retained.
Finally, morphological observations on T. rigens spore capsules were made to explore whether the reticulate patterns and incongruent relationships of T. rigens specimens in the two molecular data sets could be a result of hybridisation. Tortella rigens was found with sporophytes for the first time ever during the fieldwork for the present study (cf. Eckel 1998), in three locations in SAR and seven in GTL. In several specimens only immature sporophytes or old setae remained, but in a few some capsules were in a condition where significant morphological observations could be made. These observations of capsule morphology were combined with a search for male T. rigens plants (so-far unknown; Eckel 1998) in five shoots in each of 68 collections, 340 shoots in total, including those specimens with sporophytes (Appendix 2).
Results
The total number of aligned ITS sites in the 109 studied Tortella ingroup specimens and the outgroup of three T. humilis, two Oxystegus, and two Trichostomum ones, after deletion of regions at the beginnings and ends that were incomplete for some specimens, was 904 and when only T. rigens was included the length was 806. This included 123 (33 in the Tortella ingroup, 12 in T. rigens) sites with base substitutions and 31 (10, 9) coded indels. For atpB-rbcL the corresponding values were 565 and 524, with 46 (19, 12) base substitutions and 8 (8, 4) coded indels. For rps4 the values were 605 and 601, with 39 (15, 7) base substitutions and 3 (1, 0) indels. The number of parsimony informative sites including indels was 96 (29, 21) for ITS, 34 (21, 11) for atpB-rbcL, and 24 (11, 4) for rps4. The sequence lengths for the species were for: Tortella arctica (n = 1): 800 (ITS), 518 (atpB-rbcL), 601 (rps4); T. bambergeri (n = 11): 795–803, 517–527, 601; T. densa (3): 794–801, 523, 601; T. fragilis (5): 785–800, 515–517, 601–602; T. humilis (3): 793–833, 487–511, 601; T. inclinata (5): 789–801, 518, 601; T. rigens (74): 793–800, 515–523, 601; T. tortuosa (10): 796–800, 508–553, 601; Oxystegus tenuirostris (2): 794–795, 484, 603; Trichostomum crispulum (2): 767–785, 503–520, 602.
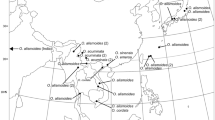
The NN split networks and the Jacknife trees with branches having support values of 60 or higher are shown in Figs. 1 and 2. Several of the morphologically defined species, including T. rigens and T. bambergeri, appeared polyphyletic or paraphyletic and their relationships differed between the nuclear and plastid networks and trees, respectively. As an example, both data sets placed most of the 74 T. rigens specimens in a single main clade, or on one side of a main split, together with a few specimens of other species. These specimens included a clear majority of the SAR, GTL, OEL and CAN specimens, whereas in the ITS data the majority of the VAS specimens appeared elsewhere and with the chloroplast data two of the four EST specimens were found outside the main group of T. rigens. The Shimodaira–Hasegawa test rejected the hypothesis that the unconstrained trees and the ones constrained to make T. rigens monophyletic are the same (p < 0.001).
NeighborNet split network for Tortella rigens and related species, using Trichostomum crispulum, Oxystegus tenuirostris, and Tortella humilis as outgroup, based on the nuclear ITS (a) and the chloroplast markers atpB-rbcL and rps4 (b). Specimen numbers correspond with those in “Appendix 1”
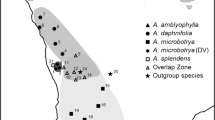
Jacknife tree for Tortella rigens and related species, using Trichostomum crispulum, Oxystegus tenuirostris, and Tortella humilis as outgroup, based on the nuclear ITS (a) and the chloroplast markers atpB-rbcL and rps4 (b). Branches with Jacknife support values of at least 60 are resolved. Specimen numbers correspond with those in “Appendix 1”
A combination of three atpB-rbcL positions correctly identifies all studied Tortella rigens and T. bambergeri specimens, as these are morphologically defined (Table 1; Online Resource 1). With rps4, 89 % of the specimens can be correctly assigned to morpho-species, but when the unequal numbers of specimens were considered, it separated 96 % of the 74 T. rigens specimens from T. bambergeri but only 45 % of the 11 T. bambergeri ones from T. rigens. The corresponding figures for ITS were 85, 84 and 91 % (see Online Resource 1).
Morphological studies of T. rigens sporophytes showed that these were of two kinds (Fig. 3a vs. b–d). The SAR capsule (Fig. 3a) comes from a cushion that grew close to a cushion of T. tortuosa. The capsule was not quite mature and was longer, ca. 4 mm with lid, than those from GTL. It had a tall peristome that was red, twisted, papillose, and had transverse ridges in its uppermost portion. Two of the GTL capsules came from cushions growing close to T. inclinata (Fig. 3b, d). The adjoining T. inclinata had male plants and in one case sporophytes (Fig. 3e). Although no other Tortella species was associated with the third specimen having developed sporophytes (Fig. 3c), T. inclinata occurred also at that locality. In all GTL specimens the capsules were shorter than in SAR, 2.3-2.7 mm long, and had a short peristome that was hyaline or yellowish, straight or very weakly twisted, smooth, and lacked transverse ridges. In the GTL specimens no spores were found in the capsules and the lids were still attached despite that the capsules appeared old (possibly aborted). The illustrated T. inclinata sporophyte was similar to those of the GTL T. rigens capsules, 2.5 mm long, with a short peristome that was pale orange-red, weakly twisted (almost straight), finely papillose, and lacked transverse ridges. Contrary to the similarly looking T. rigens capsules, this capsule had young spores that appeared viable (Fig. 3e). Out of the 340 shoots in 68 collections, 22 bore archegonia and 318 did not express sex. No male expressing T. rigens was found.
Sporophytes found on Tortella rigens (a–d) and T. inclinata (e), all from Sweden. a Stockholm archipelago (S; Reg. No. B196687), a not quite mature capsule from a plant that grew close to T. tortuosa. b Gotland (S; Reg. No. B196947), a capsule that appeared to be aborted or from the previous season, without spores, that grew close to T. inclinata (with male plants). c Gotland (S; Reg. No. B196942), a capsule that appeared to be aborted or from the previous season, without spores, from a plant in a pure T. rigens specimen, but with T. inclinata growing nearby. d, e Gotland (S; Reg. No. B196941), an old and aborted T. rigens capsule (d) and a young T. inclinata one (e) from a specimen where both species grew close to each other and males were present in T. inclinata
The statistical parsimony network for T. rigens, based on ITS and chloroplast data combined, is shown in Fig. 4a. Haplotypes 2–6 differ from haplotype 1 in 1–3 mutations, whereas haplotypes 8-9 are separated from Haplotype 1 in 11–12 mutations and haplotypes 7, 10, and 11 are even more distantly related. Of the eleven haplotypes revealed, two occur only in CAN and one only in EST. No statistical support for recombination was found for the ITS data set with all Tortella species (p = 0.4238), according to the Φw statistic and no reticulation was found among the T. rigens specimens when these were analysed on their own (Fig. 4a).
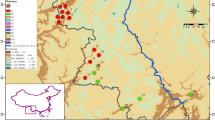
a Haplotype network for the 74 studied Tortella rigens specimens, based on all markers (ITS, atpB-rbcL, rps4). Circle size is proportional to the number of sampled populations of a certain haplotype and haplotype numbers are used throughout the paper. Circles connected by a line differ in a single mutational difference; dots indicate’missing’ haplotypes; numbers associated with the dots indicate the number of missing haplotypes when there are more than one. Different line and dot styles indicate which markers support differences between haplotypes. b Distribution of haplotypes among the regions Öland (OEL), Gotland (GTL), Västergötland (VAS), and the Stockholm archipelago with adjacent mainland (SAR), all in southern Sweden, and western Estonia (EST), Åland, Finland (FIN), and Ontario, Canada (CAN). Circle size is proportional to the number of sampled populations in a region
The geographical distributions of the T. rigens haplotypes are shown in Fig. 4b. The highest number of haplotypes was found on OEL, which had also got more unique haplotypes than any other area. The number of specimens, number of haplotypes, effective number of haplotypes, and haplotype diversity are compared among the four focal regional Swedish populations in Table 2. OEL had a markedly higher number of effective haplotypes (Ne) and higher haplotype diversity (H) than the other three populations. Tajima’s D test yielded a negative value below −1.791 for GTL (Tajima’s D: −2.304, n = 15) and a positive value above 1.976 for VAS (2.391, n = 13). This is significantly different from 0 (Table 2 in Tajima 1989), suggesting that the GTL population is increasing and/or subject to purifying selection and the VAS one is decreasing and/or subject to balancing selection (Tajima 1989). The OEL and SAR values did not differ from 0 (−1.732, n = 16; 0.488, n = 18). Out of the total haplotype variation, 78 % could be referred to within population variation, whereas 22 % was due to variation among the populations (Table 3). Pair-wise PhiPT values for the four Swedish populations distinguished all except GTL and SAR from each other (Table 4).
The migration analysis by LAMARC indicated that immigration occurred mainly into GTL from SAR and to SAR from GTL (Table 5). More limited immigration was revealed into OEL from VAS, into GTL from OEL, and into VAS from SAR.
Discussion
Relationships among Tortella rigens and related species
The frequent reticulation, as well as the incongruence among the data sets, appear to affect several ingroup Tortella species, although the extent of this is most evident for T. rigens and T. bambergeri, where numerous specimens were studied. Actually, several investigation have shown that suggested relationships among relatively closely related moss species, or individual specimens within the species, are incongruent between nuclear and chloroplast data sets, or that molecular relationships and morphologically defined species sometimes disagree (e.g., Draper and Hedenäs 2009; Draper et al. 2007; Hedenäs 2009; Hedenäs and Eldenäs 2007; Hedenäs and Rosborg 2009; Sotiaux et al. 2009). Neither ITS nor chloroplast data support a morphologically defined T. rigens as monophyletic, since a few specimens appear in positions far from the main group of samples. In view of the conflicting molecular evidence it is evident that molecular data are not necessarily more reliable than the morphological information when inferring species circumscriptions and relationships. All available information must therefore be evaluated.
The majority of the T. bambergeri and T. rigens specimens are separated according to the two sides of the main left–right split in both the ITS and chloroplast split networks (Fig. 1), which suggests that these two species are not closely related despite a similar general habit. The majority (ITS), or all T. bambergeri specimens are nested among the other included Tortella species in the right portion of the networks, whereas T. rigens and a few other specimens are found far to the left. On the other hand the molecular results suggest relationships between T. rigens and either T. inclinata and, to some degree, T. densa, or with T. fragilis. This is also what was suggested based on morphology when the species was originally described (Albertson 1946) and in the recent North American revision (Eckel 1998), where especially a relationship with T. inclinata was suggested. In the ITS network, T. inclinata, and one specimen of each of T. bambergeri and T. tortuosa are found together with T. rigens, with T. densa in between these and the remaining members of Tortella. In the chloroplast network, the specimens of T. fragilis and a possible T. tortuosa with a habit reminding of T. rigens, but with smaller leaf lamina cells and small cells that partly to entirely cover the adaxial costa, are instead found with most of the T. rigens specimens. Relatively few specimens of T. tortuosa were included, and significantly more specimens of this variable species need to be studied to clarify the background of its variation. For the recognition of T. rigens as a separate species, it is significant that it does not occur in Central Europe and the Alps, where all other European Tortella species are found. If it would only be a minor variant or deviant phenotype of some other species, one would expect that it should occur also outside its present geographical range. Neither of the two data sets support the idea that T. inclinata and T. densa are more closely related to each other than to several other Tortella species (Figs. 1, 2), as was suggested by, for example, Eckel (1998) and Hill et al. (2006). As for T. bambergeri, most of the specimens appear in two separate branches in both networks, suggesting that this species deserves a closer study (H. Köckinger et al., in preperation).
The reticulation and incongruence between molecular partitions can have several causes, including insufficient data, rapid diversification, horizontal chloroplast transfer, horizontal gene transfer, hybridization, incomplete lineage sorting, convergence caused by natural selection, and variation in evolutionary rate (Harris 2008; Stegemann et al. 2012; Wendel and Doyle 1998). Phylogenetic analyses can often not decide which of these causes explain a particular case unless additional evidence is at hand (Wendel and Doyle 1998). For firm sequence-based evidence of hybridization and introgression, extensive studies based on numerous (up to 24–48) unlinked co-dominant markers are required (Twyford and Ennos 2012), and the present three markers are thus not sufficient for this. For T. rigens, a possible hybrid origin has been discussed several times based on morphological evidence, mostly suggesting that T. inclinata and T. fragilis are the likely parental species (Albertson 1946; Eckel 1998; Persson 1947). From a morphological point of view, a hybrid origin of T. rigens appears plausible. Both T. rigens and T. fragilis have erect and upwards gradually narrowed leaves that are fragile and have smooth, elongate marginal lamina cells. However, all these characteristics are less strongly developed in T. rigens than in T. fragilis. Short-leaved modifications of T. rigens can also resemble T. inclinata with relatively longly acuminate leaves, and very small shoots or small leaves of T. rigens may sometimes have somewhat cucullate leaf apices. In addition, both the latter species have smooth, elongate adaxial costa cells. Two kinds of sporophytes have been found in T. rigens, similar to those in either T. tortuosa or T. inclinata. Since male plants are unknown in T. rigens, one of the other two species were growing close to T. rigens (male plants were found in T. inclinata), and spore production was failing in several capsules, this suggests that the found sporophytes are actually of hybrid origin. If viable spores are occasionally produced by such hybrid sporophytes, this could potentially at least to some degree explain both the reticulation, the incongruence among the data sets, and the morphological similarities between T. rigens and the two potential parental species (Albertson 1946; Eckel 1998). Successful spore production could lead to introgression and potential transfer of one or a few morphological traits from one of the male parental species to T. rigens, and this could also explain why not only morphological traits of the parental species suggested above, but also such that remind of T. tortuosa are sometimes found in T. rigens (Eckel 1998). Recently another case of likely hybrid origin in Tortella was reported (Werner et al. 2014), suggesting that hybridisation is important in the evolution within this genus, despite that the morphology of the species appears stable in most respects.
Reticulation and incongruent molecular data sets appear to be frequent among closely related morphologically defined moss species (Draper and Hedenäs 2009; Draper et al. 2007; Harris 2008; Hedenäs 2011; Shaw et al. 2005), and is problematic for the development of molecular identification tools, such as barcodes. To base circumscriptions of biological entities and studies of their biology and history entirely on evidence from non-coding markers, as was done in an otherwise elegant biogeographic study of Tetraplodon by Lewis et al. (2014), disregards the vast amount of evidence present in morphology and habitat specializations that depends on coding genes. For T. rigens and T. bambergeri, only the chloroplast marker atpB-rbcL places all included specimens in the correct morphologically defined species (Table 1). Even so, since a smaller proportion of the total variation in T. bambergeri may be included among the fewer (11) studied specimens than in T. rigens (74) some caution regarding the reliability of this particular marker is still required. Of the three studied markers, rps4 displayed a strongly unbalanced distinguishing potential, since despite that 89 % of all specimens were correctly identified, 96 % of the morphologically defined T. rigens specimens were correctly distinguished from T. bambergeri whereas only 45 % of the latter were correctly separated from T. rigens. This aspect is important when developing barcodes.
Molecular relationships among Swedish Tortella rigens populations
The four focal Swedish regional populations of T. rigens display a wide variation in haplotype diversity, with the highest values in OEL and much lower in the other three areas. The OEL values are similar to those found in most of the populations of Rhytidium rugosum (Hedw.) Kindb., a species that occurs in OEL and VAS, and in addition in the Scandinavian mountain range and adjoining regions (Hedenäs 2014). However, the most isolated of the studied R. rugosum populations, that on OEL, displayed as low haplotype diversity as T. rigens in GTL, VAS, and SAR. The explanations for the low T. rigens haplotype diversity in the three latter areas probably differ. While the studied molecular markers are either non-coding or a chloroplast gene (rps4), it seems unlikely that they are subject to purifying or balancing selection in the studied populations (Tajima 1989), and the significantly negative and positive values of Tajima’s D, therefore, likely reflect changing population sizes. The population in GTL is increasing, suggesting that larger surfaces of suitable habitat have recently become available, possibly in connection with a lowered grazing pressure or changes in climate (Högström 1997). Considering that hybrid spore capsules are produced in this area, this could eventually lead to a higher diversity if viable spores are occasionally produced. On the other hand, the population in VAS is decreasing, which suggests a loss of diversity due to deterioration of the species’ habitat in this area. It is difficult to estimate the time frame of population changes since we lack reliable dating points or rates of molecular evolution in Tortella. However, it is evident that the area of suitable habitat for T. rigens in Kinnekulle in VAS has decreased since Albertson (1940, 1946) worked there, due to closing shrub and tree vegetation (Hedenäs, personal observation). Finally, the population in SAR, which has slightly higher haplotype diversity than those in GTL and VAS, is stable.
Based on the morphological variation that Albertson (1946) found in T. rigens, he correctly suspected that the species is genetically variable. Among population variation is 22 % of the total, and the species occurs in relatively isolated populations as evidenced by the relatively high and significant PhiPT values in most of the pair-wise comparisons among populations. However, GTL and SAR could not be distinguished from each other, possibly as a result of high interpopulation migration rates. Obviously these two populations are highly interconnected, and seem to function as a single larger population, probably together with the FIN and possibly the EST populations.
The CAN population differs from those in Europe, but although its haplotypes are unique within the species they are not more different from the most common T. rigens haplotype than some other European haplotypes (Fig. 4). The differences found between North America and Europe are therefore not large enough to, on their own, suggest that the two originated separately, for example through separate hybridization events.
The high haplotype diversity of T. rigens in OEL suggests that this population may be closest to where the species survived during the last glacial period or, alternatively, closest to the area where it originated through hybridisation. When the ice retreated, more base-rich conditions and conditions with little competition from vascular plants were widespread in southern Scandinavia (Andersen 1994; Berglund et al. 2008; Kuneš et al. 2011), and if the origin was through hybridisation this could at that time have occurred at more places than today in this area. Under both scenarios regarding its postglacial origin, the species may have been widespread in southern Scandinavia during a period of several 1,000 years until acidification (usually relatively fast; Matthews 1992) and closing vegetation allowed more limited populations to survive only in the four focal regions, besides small other areas around the Baltic Sea. Thus, founder effects or isolation and gradual elimination of haplotypes in small populations could contribute to explain why OEL, VAS, GTL-SAR-FIN, and EST, have all got their unique haplotypes.
Implications for the conservation of Tortella rigens
Tortella bambergeri occurs in Sweden. However, the vast majority of the T. rigens specimens are correctly identified and there is no reason to revise our ideas regarding its frequency. In southern Sweden we have three entities of T. rigens that should be considered in a conservation context. The largest consists of the GTL plus SAR populations, which together form a single unit probably thanks to frequent migration. The GTL one is presently increasing and the SAR one is stable, and the GTL-SAR population therefore presently does not appear to require special management to ensure its survival, despite that two of the haplotypes are only found here within Sweden. The VAS population, on the other hand, has a low haplotype diversity, is decreasing as a result of observed habitat changes, and one of its haplotypes is unique among the focal regions. Unless the species’ survival in this area is secured, one of the species eleven haplotypes (i.e., 9 %) or nine European haplotypes (11 %) may be lost. However, overall OEL is the population that is most important to manage wisely for the future, since this is stable, has the highest haplotype diversity of the three entities, and four of the haplotypes (36/44 %) are only found here. The very distinct North American population may require special management for its survival, but since it was only recently recognized a primary aim must instead be to explore how common, widespread, and variable the species is on this continent.
References
Albertson N (1940) Rhytidium rugosum (Hedw.) Lindb. i Fennoscandia. Svensk Bot Tidskr 34:77–100
Albertson N (1946) Österplana hed. Ett alvarområde på Kinnekulle. Acta Phytogeogr Suec 20:I–XII, 1–267
Andersen ST (1994) History of the terrestrial environment in the quaternary of Denmark. Bull Geol Soc Denmark 41:219–228
Bartish IV, Swenson U, Munzinger J, Anderberg AA (2005) Phylogenetic relationships among New Caledonian Sapotaceae (Ericales): molecular evidence for generic polyphyly and repeated dispersal. Amer J Bot 92:667–773
Berglund BE, Persson T, Björkman L (2008) Late quaternary landscape and vegetation diversity in a North European perspective. Quaternary Int 184:187–194
Bosanquet SDS (2006) Tortella bambergeri (Schimp.) Broth. in the British Isles. J Bryol 28:5–10
Bruen TC, Hervé P, Bryant D (2006) A simple and robust statistical test for detecting the presence of recombination. Genetics 172:2665–2681
Chiang TI, Schaal BA, Peng C-I (1998) Universal primers for amplification and sequencing a noncoding spacer between the atpB and rbcL genes of chloroplast DNA. Bot Bull Acad Sin 39:245–250
Clement M, Posada D, Crandall KA (2000) TCS: a computer program to estimate gene genealogies. Molec Ecol 9:1657–1659
Darriba D, Taboada GL, Doallo R, Posada D (2012) jModelTest 2: more models, new heuristics and parallel computing. Nat Meth 9:772
Draper I, Hedenäs L (2009) Circumscription of European taxa within the Sciuro-hypnum reflexum complex (Brachytheciaceae, Bryophyta), based on molecular and morphological data. Taxon 58:572–584
Draper I, Hedenäs L, Grimm GW (2007) Molecular and morphological incongruence in European species of Isothecium (Bryophyta). Molec Phylogen Evol 42:700–716
Düll R (1984) Distribution of the European and Macaronesian mosses (Bryophytina). Part I. Bryol Beitr 4:1–113
Eckel PM (1998) Re-evaluation of Tortella (Musci, Pottiaceae) in conterminous USA and Canada with a treatment of the European species Tortella nitida. Bull Buffalo Soc Nat Sci 36:117–191
Eckel PM (2007) Tortella. In: FoNAE Committee (ed) Flora of North America north of Mexico, vol 27., Bryophyta, part 1, Oxford University Press, New York, pp 498–511
Eckel PM (2010) Tortella bambergeri in North America and an evaluation of its taxonomy. Bull Buffalo Soc Nat Sci 39:1–10
Excoffier L, Lischer HEL (2010) Arlequin suite ver 3.5: a new series of programs to perform population genetics analyses under Linux and Windows. Mol Ecol Resour 10:564–567
Farris JS, Källersjö M, Kluge AG, Bult C (1995) Testing significance of incongruence. Cladistics 10:315–319
Feliner GN, Rosselló JA (2007) Better the devil you know? Guidelines for insightful utilization of nrDNA ITS in species-level evolutionary studies in plants. Molec Phylogen Evol 44:911–919
Frankham R, Ballou JD, Briscoe DA (2002) Introduction to conservation genetics. Cambridge University Press, Cambridge
Fu Y (1997) Statistical tests of neutrality of mutations against population growth, hitchhiking and background selection. Genetics 147:915–925
Goloboff P, Farris J, Nixon K (2003) Tree analysis using new technology. www.zmuc.dk/public/phylogeny. Accessed 5 Mar 2014
Grundmann M, Schneider H, Russell SJ, Vogel JC (2006) Phylogenetic relationships of the moss genus Pleurochaete Lindb. (Bryales: Pottiaceae) based on chloroplast and nuclear genomic markers. Org Divers Evol 6:33–45
Guindon S, Gascuel O (2003) A simple, fast, and accurate algorithm to estimate large phylogenies by maximum likelihood. Syst Biol 52:696–704
Harris ESJ (2008) Paraphyly and multiple causes of phylogenetic incongruence in the moss genus Plagiomnium (Mniaceae). Taxon 57:417–433
Hassel K, Høitomt T (2013) Tortella vrimoseslekta i Norge, nye arter og arter vi kan være på utkikk etter. Blyttia 71:215–224
Hedenäs L (2009) Relationships among Arctic and non-Arctic haplotypes of the moss species Scorpidium cossonii and Scorpidium scorpioides (Calliergonaceae). Pl Syst Evol 277:217–231
Hedenäs L (2011) Incongruence among morphological species circumscriptions and two molecular data sets in Sarmentypnum (Bryophyta: Calliergonaceae). Taxon 60:1596–1606
Hedenäs L (2014) Southern Scandinavian lowland populations of Rhytidium rugosum (Bryophyta, Rhytidiaceae) differ significantly from those in the mountains. J Bryol 36:1–14
Hedenäs L, Eldenäs P (2007) Cryptic speciation, habitat differentiation, and geography in Hamatocaulis vernicosus (Calliergonaceae, Bryophyta). Pl Syst Evol 268:131–145
Hedenäs L, Rosborg C (2009) Pseudocalliergon is nested within Drepanocladus (Bryophyta: Amblystegiaceae). Lindbergia 33:67–74
Hill MO et al (2006) An annotated checklist of the mosses of Europe and Macaronesia. J Bryol 28:198–267
Högström S (1997) Habitats and increase of Sphagnum in the Baltic Sea island Gotland, Sweden. Lindbergia 22:69–74
Huson DH, Bryant D (2006) Application of phylogenetic networks in evolutionary studies. Molec Biol Evol 23:254–267
Huttunen S, Ahonen I, Bisang I, Hedenäs L, Laaka-Lindberg S, Vänni J (2014) Sammalretki Ahvenanmaalle keväällä 2013. Bryobrotherella 17:114–135
Inoue Y, Tsubota H, Sato H, Yamaguchi T (2012) Phylogenetic note on Pachyneuropsis miyagii T. Yamag. (Pottiaceae, Bryophyta). Hikobia 16:221–228
Košnar J, Herbstová M, Kolář F, Koutecký P, Kučera J (2012) A case of intragenomic ITS variation in bryophytes: assessment of gene flow and role of plyploidy in the origin of European taxa of the Tortula muralis (Musci: Pottiaceae) complex. Taxon 61:709–720
Kuhner MK (2006) LAMARC 2.0: maximum likelihood and Bayesian estimation of population parameters. Bioinformatics 22:768–770
Kuneš P, Odgaard BV, Gaillard M-J (2011) Soil phosphorus as a control of productivity and openness in temperate interglacial forest systems. J Biogeogr 38:2150–2164
Lewis LR, Rozzi R, Goffinet B (2014) Direct long-distance dispersal shapes a New World amphitropical disjunction in the dispersal-limited dung moss Tetraplodon (Bryopsida: splachnaceae). J Biogeogr. doi:10.1111/jbi.12385
Matthews JA (1992) The ecology of recently deglaciated terrain: a geoecological approach to glacier forelands and primary succession. Cambridge University Press, Cambridge
Nadot S, Bajon R, Lejeune B (1994) The chloroplast gene rps4 as a tool for the study of Poaceae phylogeny. Pl Syst Evol 191:27–38
Nyholm E (1991) Illustrated flora of Nordic mosses. Fasc. 2 Pottiaceae-Splachnaceae-Schistostegaceae. Nordic Bryological Society, Copenhagen and Lund
Peakall R, Smouse PE (2006) GENALEX 6: genetic analysis in Excel. Population genetic software for teching and research. Molec Ecol Notes 6:288–295
Peakall R, Smouse PE (2012) GenAlEx 6.5: genetic analysis in excel. Population genetic software for teaching and research—an update. Bioinformatics 28:2537–2539
Persson H (1947) Bryum arcticum och några andra mossfynd från Stora Karlsö. Svensk Bot Tidskr 41:141–150
Pilous Z (1965) Fragmenta bryologica 56. Tortella rigens Albertson und Tortella densa Crund. et Nyh., zwei neue tschechoslowakische Moose. Preslia 37:20–22
Shaw AJ, McDaniel SF, Werner O, Ros RM (2002) New frontiers in bryology and lichenology. Phylogeography and phylodemography. Bryologist 105:373–383
Shaw AJ, Melosik I, Cox CJ, Boles SB (2005) Divergent and reticulate evolution in closely related species of Sphagnum section Subsecunda. Bryologist 108:363–376
Shimodaira H, Hasegawa M (1999) Multiple comparisons of loglikelihoods with applications to phylogenetic inference. Molec Biol Evol 16:1114–1116
Sotiaux A, Enroth J, Olsson S, Quandt D, Vanderpoorten A (2009) When morphology and molecules tell us different stories: a case-in-point with Leptodon corsicus, a new and unique endemic moss species from Corsica. J Bryol 31:186–196
Souza-Chies TT, Bittar G, Nadot S, Carter L, Besin E, Lejeune B (1997) Phylogenetic analysis of lridaceae with parsimony and distance methods using the plastid gene rps4. Pl Syst Evol 204:109–123
Stech M (1999) Molekulare Systematik haplolepider Laubmoose (Dicranaceae, Bryopsida). PhD Thesis, Freie Universität Berlin, Berlin
Stegemann S, Keuthe M, Greiner S, Bock R (2012) Horizontal transfer of chloroplast genomes between plant species. Proc Natl Acad Sci USA 109:2434–2438
Swofford DL (2002) PAUP*. Phylogenetic analysis using parismony (*and other methods), version 4.0b10. Sinauer Association, Sunderland, Massachusetts
Tajima F (1989) Statistical method for testing the neutral mutation hypothesis by DNA polymorphism. Genetics 123:585–595
Twyford AD, Ennos RA (2012) Next-generation hybridization and introgression. Heredity 108:179–189
Vellak K, Ingerpuu N (2012) The state of bryophyte conservation in Estonia. Stud Bot Hung 43:59–68
Wendel JF, Doyle JJ (1998) Phylogenetic incongruence: window into genome history and molecular evolution. In: Soltis DE, Soltis PS, Doyle JJ (eds) Molecular systematics of plants II. DNA sequencing. Chapman and Hall, New York, pp 265–296
Werner O, Ros RM, Grundmann M (2005) Molecular phylogeny of Trichostomoideae (Pottiaceae, Bryophyta) based on nrITS sequence data. Taxon 54:361–368
Werner O, Köckinger H, Magdy M, Ros RM (2014) On the systematic position of Tortella arctica and Trichostomum arcticum (Bryophyta, Pottiaceae). Nova Hedwigia 98:273–293
Zwickl DJ (2006) Genetic algorithm approaches for the phylogenetic analysis of large biological sequence datasets under the maximum likelihood criterion. PhD Thesis, The University of Texas, Austin
Acknowledgments
I thank Keyvan Mirbakhsh for her efficient laboratory work, Tomas Hallingbäck and Heribert Köckinger for discussions and for providing important material, and the curators of CANM, NY, and TU for loans. Two reviewers provided valuable input on an earlier version of this paper. This study was funded by the Swedish Environmental Protection Agency, Naturvårdsverket (No. NV-10009-12).
Author information
Authors and Affiliations
Corresponding author
Additional information
Handling editor: Jochen Heinrichs.
Electronic supplementary material
Below is the link to the electronic supplementary material.
606_2014_1159_MOESM1_ESM.pdf
Online Resource 1 Aligned sequences for atpB-rbcL, rps4, and ITS for Tortella rigens and T. bambergeri. For each species the different haplotypes, based on the respective markers, are displayed. (PDF 36 kb)
Appendices
Appendix 1
Specimen data and GenBank accession numbers for the studied specimens. Data format: Sample No.: [for Tortella rigens: REGIONAL POPULATION (see Fig. 4b)]; Locality; Coll. Year, Collector (collector’s No.) (LH = L. Hedenäs); Herbarium acronym: registration No.; GenBank Accession numbers for ITS, atpB-rbcL, rps4. Sample numbers for specimens which sequences were downloaded from GenBank begin with ‘GB’
Tortella arctica (Arnell) Crundw. & Nyholm: P138: Sweden. Jämtland, Åre, 2010, LH; S: B182517; KM020640, KM020530, KM020750. Tortella bambergeri (Schimp.) Broth.: P091: Sweden. Dalsland, Skållerud, 1985, T. Hallingbäck 44376; S: B184928; KM020617, KM020506, KM020726. P095: Austria. Oberoesterreich, Koppenwinkel Alm, 1962, J. Froehlich; S: B196096; KM020611, KM020500, KM020720. P096: Austria. Kärnten, Kanziani-Berg, 1975, J. Poelt (Pl. Graec., Bryoph. 2); S: B196097; KM020612, KM020501, KM020721. P097: Austria. Steiermark, Lavanttal, 1988, H. Pittoni & J. Poelt (Pl. Graec., Bryoph. 145); S: B196098; KM020613, KM020502, KM020722. P104: Sweden. Västergötland, Djupadalen, 1974, T. Hallingbäck 44375; S: B184932; KM020620, KM020509, KM020729. P129: Hungary. (no locality provided), 1936, A. Boros; S: B186073; KM020614, KM020503, KM020723. P130: United Kingdom. Scotland. Gleann Beag, 1963, A.C. Crundwell; S: B196677; KM020615, KM020504, KM020724. P131: Austria. Nideroesterreich, Kuhschneeberg, 1951, J. Froehlich; S: B196678; KM020616, KM020505, KM020725. P151: Sweden. Dalsland, Steneby, 1986, T. Hallingbäck 44374; herb. Hallingbäck; KM020618, KM020507, KM020727. P153: Sweden. Dalsland, Skållerud, 2011, T. Hallingbäck 5755; herb. Hallingbäck; KM020619, KM020508, KM020728. P200: Sweden. Södermanland, Utö, 2013, LH; S: B198239; KM020621, KM020510, KM020730. Tortella densa (Lorentz & Molendo) Crundw. & Nyholm: P136: Sweden. Gotland, Boge, 1997, LH; S: B42917; KM020638, KM020528, KM020748. P137: Norway. Hordaland, Stord, 1990, N. Hakelier; S: B184679; KM020639, KM020529, KM020749. GB1de: Ireland. Claire, 1992, E. Wiltshire; BM; AY854412, AY950332, AY950377. Tortella fragilis (Hook. & Wilson) Limpr.: P098: Sweden. Södermanland, Utö, 2010, LH; S: B176012; KM020622, KM020511, KM020731. P099: Sweden. Pite Lappmark, Arjeplog, 2006, LH et al.; S: B114557; KM020623, KM020512, KM020732. P132: Norway. Finnmark, Söröysund, 2001, LH; S: B59844; KM020624, KM020513, KM020733. GB1fr: Greenland, F.J.A. Daniels; MSUN: 0795-869.5; AY854417, AY950337, AY950382. GB2fr: Russia. Gorno-Altai, 1991, M. Ignatov; BM; AY854416, AY950336, AY950381. Tortella inclinata (R.Hedw.) Limpr.: P113: Sweden. Södermanland, Mörkö, 2003, T. Hallingbäck 39628; S: B184931; KM020634, KM020524, KM020744. P117: Sweden. Södermanland, Mörkö, 1984, LH; S: B196111; KM020635, KM020525, KM020745. P134: Sweden, Värmland, Filipstad, 2010, LH & G. Odelvik; S: B179320; KM020636, KM020526, KM020746. P135: Sweden. Västmanland, Sala, 2011, LH et al.; S: B185169; KM020637, KM020527, KM020747. GB1in: Germany. Northrhine-Westphalia, C. Schmidt; BM: 000824494; AY854420, AY950340, AY950385. Tortella rigens Alberts.: P089: VAS; Sweden. Västergötland, Dala, 1991, N. Hakelier; S: B195852; KM020536, KM020426, KM020646. P090: VAS; Sweden. Västergötland, Högstena, 1974, T. Hallingbäck 44373; S: B184934; KM020537, KM020427, KM020647. P092: OEL; Sweden. Öland, Sandby, 2005, P. Lundquist 1198842; S: B148156; KM020549, KM020439, KM020659. P093: OEL; Sweden. Öland, Kastlösa, 2005, M. Arnesson 1198767; S: B148179; KM020550, KM020440, KM020660. P094: SAR; Sweden. Uppland, Djurö, 2011, LH; S: B187057; KM020565, KM020455, KM020675. P102: VAS; Sweden. Västergötland, Högstena, 1985, N. Hakelier; S: B195851; KM020538, KM020428, KM020648. P103: VAS; Sweden. Västergötland, Österplana, 1961, N. Hakelier; S: B195850; KM020539, KM020429, KM020649. P105: VAS; Sweden. Västergötland, Högstena alvar, 1974, T. Hallingbäck 42384; S: B184933; KM020540, KM020430, KM020650. P106: VAS; Sweden. Västergötland, Österplana hed, 1975, T. Hallingbäck 44377; S: B184936; KM020541, KM020431, KM020651. P107: VAS; Sweden. Västergötland, Dala, 1974, E. Nyholm; S: B184787; KM020542, KM020432, KM020652. P108: VAS; Sweden. Västergötland, Österplana, 1975, G. Bråvander; S: B81291; KM020543, KM020433, KM020653. P109: VAS; Sweden. Västergötland, Valtorp, 1942, A. Hülphers; S: B184759; KM020544, KM020434, KM020654. P110: VAS; Sweden. Västergötland, Vilske-Kleva, 1942, A. Hülphers; S: B184785; KM020545, KM020435, KM020655. P111: VAS; Sweden. Västergötland, Södra Kyrketorp, 1946, N. Albertson; S: B184767; KM020546, KM020436, KM020656. P112: OEL; Sweden. Öland, Gräsgård, 2005, P. Stjernfeldt 1198769; S: B148190; KM020551, KM020441, KM020661. P114: SAR; Sweden. Södermanland, Utö, 1983, LH; S: B196110; KM020566, KM020456, KM020676. P115: SAR; Sweden. Södermanland, Mörkö, 1985, LH; S: B184796; KM020567, KM020457, KM020677. P116: SAR; Sweden. Södermanland, Ornö, 1983, LH; S: B196109; KM020568, KM020458, KM020678. P118: OEL; Sweden. Öland, Södra Möckleby, 2005, P. Lundquist 1198752; S: B148185; KM020552, KM020442, KM020662. P119: OEL; Sweden. Öland, Böda, 1962, A.C. Crundwell & E. Nyholm; S: B77839; KM020553, KM020443, KM020663. P120: OEL; Sweden: Öland, Föra, 1960, A.C. Crundwell & E. Nyholm 60-4; S: B184705; KM020554, KM020444, KM020664. P121: OEL; Sweden. Öland, Ås, 2005, M. Beckman 1198759; S: B148148; KM020555, KM020445, KM020665. P122: OEL; Sweden. Öland, Vickleby, 2005, M. Aronsson 1198735; S: B148163; KM020556, KM020446, KM020666. P123: OEL; Sweden. Öland, Södra Möckleby, 2005, P. Lundquist 1198751; S: B148184; KM020557, KM020447, KM020667. P124: OEL; Sweden. Öland, Ventlinge, 2005, M. Aronsson; S: B148292; KM020558, KM020448, KM020668. P125: OEL; Sweden. Öland, Hulterstad, 2005, M. Miller 1198766; S: B148149; KM020559, KM020449, KM020669. P126: OEL; Sweden. Öland, Resmo, 2005, M. Beckman 1198764; S: B148168; KM020560, KM020450, KM020670. P127: OEL; Sweden. Öland, Bårby, 2006, T. Hallingbäck & N. Lönnell 43866; S: B184937; KM020561, KM020451, KM020671. P128: OEL; Sweden. Öland, Möckelmossen, 2006, T. Hallingbäck & N. Lönnell 43898; S: B184939; KM020562, KM020452, KM020672. P140 (Originally identified as Tortella humilis by P. M. Eckel in 1987): SAR; Sweden. Uppland, Runmarö, 1984, J.T. Johansson; S: B184798; KM020569, KM020459, KM020679. P154: VAS; Sweden. Västergötland, Västerplana, 2012, T. Hallingbäck 6183; herb. Hallingbäck; KM020547, KM020437, KM020657. P155: VAS; Sweden. Västergötland, Södra Kyrketorp, 2003, T. Hallingbäck 39536; herb. Hallingbäck; KM020548, KM020438, KM020658. P156: CAN; Canada. Ontario, Renfrew, 1987, R.R. Ireland 22737; NY: 00134127; KM020598, KM020488, KM020708. P157: CAN; Canada. Ontario, Renfrew, 1987, R.R. Ireland 22735; CANM: 304175; KM020599, KM020489, KM020709. P158: CAN; Canada. Ontario, Algoma, 1989, R.R. Ireland 24437; CANM: 314373; KM020610, KM020499, KM020719. P159: CAN; Canada. Ontario, Carleton, 1978, J. Reddoch; CANM: 260653; KM020600, KM020490, KM020710. P160: CAN; Canada. Ontario, Ottawa-Carleton, 1988, D.F. Brunton 8639; CANM: 308477; KM020601, KM020491, KM020711. P161: EST; Estonia. Saare mk, Kärla, 2012, K. Vellak; TU: 168632; KM020602, KM020521, KM020742. P162: EST; Estonia. Harju Co., Väiki-Pakri, 2008, N. Ingerpuu; TU: 162045; KM020603, KM020492, KM020712. P163: EST; Estonia. Saare Co., Kärla, 2008, K. Vellak; TU: 168765; KM020604, KM020493, KM020713. P164: EST; Estonia. Saaremaa, Lümanda, 2010, S. Pihu; TU: 168246; KM020605, KM020494, KM020714. P166: SAR; Sweden. Södermanland, Nämdö, 2013, LH; S: B196865; KM020570, KM020460, KM020680. P167: SAR; Sweden. Södermanland, Nämdö, 2013, LH; S: B196870; KM020571, KM020461, KM020681. P168: SAR; Sweden. Södermanland, Trosa-Vagnhärad, 2013, LH & I. Bisang; S: B196734; KM020572, KM020462, KM020682. P169: SAR; Sweden. Södermanland, Trosa-Vagnhärad, 2013, LH & I. Bisang; S: B196736; KM020573, KM020463, KM020683. P170: SAR; Sweden. Uppland, Djurö, 2013, LH; S: B196684; KM020574, KM020464, KM020684. P171: SAR; Sweden. Uppland, Djurö, 2013, LH; S: B196694; KM020575, KM020465, KM020685. P172: SAR; Sweden. Uppland, Djurö, 2013, LH; S: B196688; KM020576, KM020466, KM020686. P173: SAR; Sweden. Södermanland, Nämdö, 2013, LH; S: B196885; KM020577, KM020467, KM020687. P174: FIN; Finland. Åland, Eckerö, 2013, LH; S: B197025; KM020606, KM020495, KM020715. P175: FIN; Finland. Åland, Eckerö, 2013, LH & I. Bisang; S: B197029; KM020607, KM020496, KM020716. P176: FIN; Finland. Åland, Eckerö, 2013, LH & I. Bisang; S: B197037; KM020608, KM020497, KM020717. P177: GTL; Sweden. Gotland, Fleringe, 2013, LH & I. Bisang; S: B196893; KM020583, KM020473, KM020693. P178: GTL; Sweden. Gotland, Bunge, 2013, LH & I. Bisang; S: B196898; KM020584, KM020474, KM020694. P179: GTL; Sweden. Gotland, Fleringe, 2013, LH & I. Bisang; S: B196900; KM020585, KM020475, KM020695. P181: GTL; Sweden. Gotland, Hellvi, 2013, LH; S: B196908; KM020586, KM020476, KM020696. P182: GTL; Sweden. Gotland, Hellvi, 2013, LH; S: B196910; KM020587, KM020477, KM020697. P183: GTL; Sweden. Gotland, Hellvi, 2013, LH; S: B196912; KM020588, KM020478, KM020698. P184: GTL; Sweden. Gotland, Hellvi, 2013, LH; S: B196917; KM020589, KM020479, KM020699. P185: GTL; Sweden. Gotland, Hellvi, 2013, LH; S: B196923; KM020590, KM020480, KM020700. P186: GTL; Sweden. Gotland, Hangvar, 2013, LH; S: B196929; KM020591, KM020481, KM020701. P187: GTL; Sweden. Gotland, Hangvar, 2013, LH; S: B196934; KM020592, KM020482, KM020702. P188: GTL; Sweden. Gotland, Stenkyrka, 2013, LH; S: B196937; KM020593, KM020483, KM020703. P189: GTL; Sweden. Gotland, Stenkyrka, 2013, LH; S: B196945; KM020594, KM020484, KM020704. P190: GTL; Sweden. Gotland, Hangvar, 2013, LH; S: B196950; KM020595, KM020485, KM020705. P191: OEL; Sweden. Öland, Mörbylånga, 2005, M. Beckman 1198738; S: B148169; KM020563, KM020453, KM020673. P192: OEL; Sweden. Öland, Gårdby, 2005, M. Miller 1198732; S: B148152; KM020564, KM020454, KM020674. P193: GTL; Sweden. Gotland, Hellvi, 2013, LH; S: B196915; KM020596, KM020486, KM020706. P194: GTL; Sweden. Gotland, Endre, 1989, LH G89-36; S: B197080; KM020597, KM020487, KM020707. P195: SAR; Sweden. Uppland, Djurö, 2013, LH; S: B196689; KM020578, KM020468, KM020688. P196: SAR; Sweden. Uppland, Djurö, 2013, LH; S: B196691; KM020579, KM020469, KM020689. P197: SAR; Sweden. Södermanland, Trosa-Vagnhärad, 2013, LH & I. Bisang; S: B196737; KM020580, KM020470, KM020690. P198: SAR; Sweden. Södermanland, Nämdö, 2013, LH; S: B196873; KM020581, KM020471, KM020691. P199: SAR; Sweden. Södermanland, Nämdö, 2013, LH; S: B196876; KM020582, KM020472, KM020692. Tortella tortuosa (Hedw.) Limpr.: P100: Sweden. Gotland, Stenkyrka, 2001, LH; S: B62094; KM020625, KM020514, KM020734. P101: Sweden. Medelpad, Borgsjö, 2006, LH; S: B116614; KM020626, KM020515, KM020735. P133: Sweden. Pite Lappmark, Arjeplog, 2006, LH et al.; S: B116150; KM020627, KM020516, KM020736. P139: Canada. NWT (Nunavut). Bathurst Island, 1973, R.R. Ireland & I. Brodo 16585; S: B196676; KM020609, KM020498, KM020718. P146: Austria. Styria, Eisenerzer Alpen, 2012, H. Köckinger 14932; S: B196697; KM020628, KM020517, KM020737. P147: Austria. Carinthia, Karawanken, 2012, H. Köckinger 14933; S: B196698; KM020629, KM020518, KM020738. P148: Austria. Carinthia, Gailtaler Alpen, 2008, H. Köckinger 14934; S: B196699; KM020630, KM020519, KM020739. P149: Austria. Carinthia, Gurktal, 2003, H. Köckinger 12334; S: B196700; KM020631, KM020520, KM020740. P150: United Kingdom. Northern Ireland, Fermanagh, 1993, T. Hallingbäck 42536; herb. Hallingbäck; KM020632, KM020522, KM020741. P152: Sweden. Dalsland, Skållerud, 1986, T. Hallingbäck 1452; herb. Hallingbäck; KM020633, KM020523, KM020743. OUTGROUP: Oxystegus tenuirostris (Hook. & Taylor) A.J.E.Sm.: P144: Sweden. Dalarna, By, 2011, LH et al.; S: B185165; KM020644, KM020534, KM020754. P145: Sweden. Närke, Fellingsbro, 1994, N. Hakelier; S: B196695; KM020645, KM020535, KM020755. Tortella humilis (Hedw.) Jenn.: P141: Spain. Murcia, Bullas, 2002, T. Hallingbäck 39035; herb. Hallingbäck; KM020641, KM020531, KM020751. GB1hu: Paraguay. Guaira, E. Zardini & P. Aquino 32386; BM; AY854419, AY950339, AY950384. GB2hu: United States. Arkansas, 1992, P.L. Redfearn Jr.; BM; AY854418, AY950338, AY950383. Trichostomum crispulum Bruch: P142: Sweden. Västergötland, Dala, 1984, N. Hakelier; S: B196696; KM020642, KM020532, KM020752. P143: Sweden. Gotland, Langs hage, 2002, T. Hallingbäck 38746; S: B185015; KM020643, KM020533, KM020753.
Appendix 2
The 68 specimens of Tortella rigens that were searched for male plants. Five arbitrarily chosen shoots were studied in each plant. All specimens are from Sweden or Finland (Åland) and were collected by Lars Hedenäs (LH), sometimes together with Irene Bisang (IB). The specimens are in S, and the ‘B’ numbers are S registration numbers
Gotland: Bunge: ca. 1 km W of Bunn, 12 May 2013, LH & IB: B196898; ca. 500 m E of Strå, 11 May 2013, LH & IB: B196886. Fleringe: ca. 1.2 km WSW of Tvärlingsmyr, 12 May 2013, LH & IB: B196900; ca. 1.6 km WSW of Tvärlingsmyr, 12 May 2013, LH & IB: B196907; ca. 400 m SW of Hau farm, 11 May 2013, LH & IB: B196889; ca. 800 m WSW of Hau farm, 11 May 2013, LH & IB: B196893; W end of Kyrkgatmyr, S side, 11 May 2013, LH & IB: B196896. Hangvar: central portion of Hajdhagen, 14 May 2013, LH: B196933; S portion of Hajdhagen, 14 May 2013, LH: B196934; Skarphagen, 14 May 2013, LH: B196950; Skarphagen, 14 May 2013, LH: B196950; W portion of Hajdhagen, 14 May 2013, LH: B196929. Hellvi: ca. 1 km SE of Sudergårde, 13 May 2013, LH: B196921; ca. 1,5 km SSE of Sudergårde, 13 May 2013, LH: B196923; ca. 200 m N of Lörge, 13 May 2013, LH: B196917; ca. 750 m WNW of Lörge, 13 May 2013, LH: B196920; just SW of Kyllaj, 13 May 2013, LH: B196915; NW end of Kyllajhajdar, 13 May 2013, LH: B196908; NW end of Kyllajhajdar, 13 May 2013, LH: B196909; NW end of Kyllajhajdar, 13 May 2013, LH: B196910; S end of Kyllajhajdar, 13 May 2013, LH: B196912. Lärbro: ca. 750 S of Stora Ire, 13 May 2013, LH: B196924. Stenkyrka: ca. 1 km N of Ekebys, 14 May 2013, LH: B196941; ca. 1.2 km N of Ekebys, 14 May 2013, LH: B196942; ca. 1.6 km N of Ekebys, 14 May 2013, LH: B196945; ca. 2 km NNW of Ekebys, 14 May 2013, LH: B196946; ca. 2 km NNW of Ekebys, 14 May 2013, LH: B196947; ca. 750 m NW of Bromyr, 14 May 2013, LH: B196949; Myrskogen, ca. 600 m W of Kvie, 14 May 2013, LH: B196937. Öland: Borgholm: N of Borgholm, 21 May 1980, LH: B198240. Södermanland: Nämdö:between Klippudden and Östanvik, 8 May 2013, LH: B196873; between Sandviksudden and Klippudden, 8 May 2013, LH: B196872; just ENE of Nämdö church, 8 May 2013, LH: B196885; Klockberget, 8 May 2013, LH: B196876; Kyrknäset, 8 May 2013, LH: B196877; Nämdö Böte, just inside N margin of summer house area, 8 May 2013, LH: B196865; Nämdö Nature Reserve, SW of Östanvik, 8 May 2013, LH: B196875; Sandviksudden, 8 May 2013, LH: B196868; SW of Sandviksudden, 8 May 2013, LH: B196870; Västanvik, 8 May 2013, LH: B196880; Västanvik, 8 May 2013, LH: B196883. Ornö: Sundby Nature Reserve, E of S end of Lake Långträsk, 29 July 2013, LH: B198246; Sundby Nature Reserve, SW end of peninsula E of Västra Varpet, 29 July 2013, LH: B198244. Trosa-Vagnhärad: SSE of Fänsåker, 1 May 2013, LH & IB: B196734; SSE of Fänsåker, 1 May 2013, LH & IB: B196735; SSE of Fänsåker, 1 May 2013, LH & IB: B196736; SSE of Fänsåker, 1 May 2013, LH & IB: B196737. Utö: Ängsbergen, 24 July 2013, LH: B198241; Ängsbergen, 24 July 2013, LH: B198242; Källviksberget, 24 July 2013, LH: B198243. Uppland: Djurö: Runmarö, between Noreträsk and Viträsk, 24 April 2013, LH: B196692; Runmarö, between Noreträsk and Viträsk, 24 April 2013, LH: B196693; Runmarö, between Noreträsk and Viträsk, 24 April 2013, LH: B196694; Runmarö, between Storsten and Nore, 24 April 2013, LH: B196688; Runmarö, between Storsten and Nore, 24 April 2013, LH: B196689; Runmarö, between Storsten and Nore, 24 April 2013, LH: B196690; Runmarö, N of Storsten, 24 April 2013, LH: B196684; Runmarö, N of Storsten, 24 April 2013, LH: B196685; Runmarö, N of Storsten, 24 April 2013, LH: B196686; Runmarö, N of Storsten (10 m from B196682), 24 April 2013, LH: B196683; Runmarö, N of Storsten (10 m from B196683), 24 April 2013, LH: B196682; Runmarö, W of Byholmen, 24 April 2013, LH: B196681; Runmarö, WNW of Nore, 24 April 2013, LH: B196691; Runmarö, WNW of Storsten, 24 April 2013, LH: B196687. Åland: Eckerö: Enskär, 350 m SE of old Coast Guard Station, 27 May 2013, LH: B197025; Signilskär, along small brooklet 300 m S of Mjölkarskatan, 25 May 2013, LH & IB: B197029; Signilskär, N of Bird Station, 24 May 2013, LH & IB: B197028; Signilskär, Söderskatan, 26 May 2013, LH & IB: B197037.
Rights and permissions
About this article
Cite this article
Hedenäs, L. Tortella rigens (Bryophyta, Pottiaceae): relationships, regional variation, and conservation aspects. Plant Syst Evol 301, 1361–1375 (2015). https://doi.org/10.1007/s00606-014-1159-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00606-014-1159-9