Abstract
As the role of exosomes in physiological and pathological processes has been properly perceived, harvesting them and their internal components is critical for subsequent applications. This study is a debut of intermittent lysis, which has been integrated into a simple and easy-to-operate procedure on a single paper-based device to extract exosomal nucleic acid biomarkers for downstream analysis. Exosomes from biological samples were captured by anti-CD63-modified papers before being intermittently lysed by high-temperature, short-time treatment with double-distilled water to release their internal components. Exosomal nucleic acids were finally adsorbed by sol–gel silica for downstream analysis. Empirical trials not only revealed that sporadically dropping 95 °C ddH2O onto the anti-CD63-modified papers every 5 min for 6 times optimized the exosomal nucleic acids extracted by the anti-CD63 paper but also verified that the whole deployed procedure is applicable for point-of-care testing (POCT) in low-resource areas and for both in vitro (culture media) and in vivo (plasma and chronic lesion) samples. Importantly, downstream analysis of exosomal miR-21 extracted by the paper-based procedure integrated with this novel technique discovered that the content of exosomal miR-21 in chronic lesions related to their stages and the levels of exosomal carcinoembryonic antigen originated from colorectal cancer cells correlated to their exosomal miR-21.
Graphical Abstract

Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Existing in virtually all body fluids, exosomes are currently one of the most studied subjects in liquid biopsy to understand their roles in intercellular communication and disease pathogenesis. They are nano-sized extracellular vesicles (EVs, 30–200 nm) enveloped by a phospholipid membrane and secreted by living cells, carrying diverse functional molecules like proteins, nucleic acids, and lipids, which are abundant sources of biomarkers for numerous diseases, especially cancers [1,2,3,4]. For instance, microRNA-21 (miR-21) plays a critical role in a plethora of diseases, including cancer, wound healing, and inflammation [5,6,7,8] while CEA (carcinoembryonic antigen, a serologic protein) is universally ratified and is a sensitive, specific, and accurately categorical biomarker for the diagnosis and prognosis of colorectal cancer [9,10,11]. Therefore, profiling exosome-delivered nucleic acids and proteins for early diagnosis and intervention of intractable chronic diseases has become a highly invested field with a variety of samples from in vitro (cell culture supernatants) to clinics (plasma, saliva, urine, etc.) [12]. However, harvesting exosomal components for downstream analyses and applications is hindered not only due to the ultrasmall sizes and diverse origins of exosomes [13] but also by the absence of widely recognized and standardized lysis, although plentiful methods and buffers have been introduced for this purpose, including urea-thiourea, radioimmunoprecipitation assay (RIPA), phenol/guanidine-based buffer, Tris-based buffer, and so on [14,15,16,17,18,19,20,21]. These commercial buffers are composed of complex chemicals with organic substances and proteins that interfere with proteomic analyses and falsify nucleic acid assessments [22, 23].
This study leverages (1) the immunoaffinity technique, which has emerged as an advantageous tool to obtain exosomes by targeting their membrane proteins (especially the tetraspanin family) [24, 25], to capture exosomes; (2) double-distilled water (ddH2O), a highly purified and easily accessible source, for exosome lysis; and (3) sol–gel silica, to adsorbed nucleic acids released from captured exosomes. Silica, after undergoing a sol–gel process involving hydrolysis and polycondensation, will form an integrated network: the condensation (or polymerization) under high temperature between the silanol groups or between silanol groups and ethoxy groups will dehydrate the silica surfaces and create siloxane bridges (Si–O–Si) that form the entire silica structure [26]. Nucleic acid adsorption on the surface of silica particles is influenced by simultaneously various forces, which has been discussed in previous literatures [27,28,29]. The silanol groups generated are, therefore, divergent according to the environment, and their pH values vary in the range of 5.0–8.0 [30]. Regarding the increase of pH value, the level of dissociation and deprotonation of OH groups on the surface of silica is enhanced, accreting negative surface charges. Haynes et al. proposed the following mechanisms for manipulating the adsorption of nucleic acids on the silica surface [31]: (1) intermolecular hydrogen bond between silica and nucleic acid, (2) electrostatic shielding effect between silica and nucleic acid molecules, (3) dehydration and interaction between silica and the nucleic acid surface, and (4) the cationic salt bridge effect between silica and nucleic acids.
Inspired by all the above findings, we developed a procedure for extracting exosomal nucleic acids that was utterly performed on a single paper-based device using ddH2O, avoiding complicated ultracentrifugation and transition stages and unwanted proteins in commercial buffers which potentially contaminate the harvested products, to improve the efficiency of downstream analysis. The procedure has been conducted to meet the ASSURED criteria for point-of-care testing (POCT) and to be applicable to diverse clinical samples for early-stage diagnosis of malignancies and chronic diseases. In our paper-based procedure, we delineate a novel tactic for exosomes to be lysed by watering the exosome-captured zones of the anti-CD63 papers at high temperature (HT, 95 °C) within a short time (ST), so-called intermittent lysis or HTST lysis, instead of immersing these devices into hot water within a long operating time, since exosomes were reportedly susceptible to damage in HTST (90–95 °C) [32]. Nucleic acids released from exosomes were then absorbed by sol–gel silica for downstream analysis. The top half verified and optimized the regime of intermittent lysis and its integrated paper-based procedure in extracting exosomal nucleic acids (Fig. 1) while the bottom half applied it to extract exosomal nucleic acids from biological samples for downstream analysis. Empirical data demonstrated not only the outstanding features of the deployed system in extracting exosomal nucleic acids from various biological samples (commercial exosomes, culture media, human plasma, and chronic wound) with superior efficiency compared to commercial kit and stable operability of the paper-based device but also the clinical applicability with successful applications on human-derived samples (plasma and chronic wounds). Moreover, pioneered evaluation on exosomal proteins was achieved when CEA in exosomes purified from HCT116 cell culture media (HCT116 CCM) was successfully determined by enzyme-linked immunosorbent assay (ELISA). The paper-based procedure, which completely relied on a single paper, not only overcomes the limitations in investigating exosomes caused by their ultrasmall size but also exhibits outstanding properties (economy, simple operation, small input requirement…) that facilitate the POCT and open further research potential on exosome-transported materials.
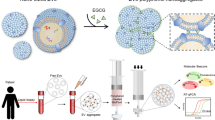
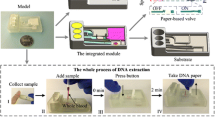
Extracting exosomal nucleic acids by paper-based intermittent (HTST) lysis procedure. Exosomes in biological samples were captured by anti-CD63-modified papers before being intermittently lysed to release their internal components. Exosomal nucleic acids were finally adsorbed by sol–gel silica, and their miR-21 content was analyzed as a representative target
Materials and methods
Materials
Sigma-Aldrich delivered Whatman chromatography papers grade 1, mesoporous silica (0.5 µm particles and 2 nm pores), (3-Mercaptopropyl)trimethoxysilane (MPTMS), and bovine serum albumin (BSA). 99.9% ethanol was from Jingming Chemical, whereas Thermo Fisher Scientific was the supplier of 4-Maleimidobutyric acid N-hydroxysuccinimide ester (GMBS), NeutrAvidin, ddH2O, phosphate-buffered saline (PBS), quantitative reverse transcription polymerase chain reaction (RT-qPCR) kit, and mirVana miRNA isolation kit. Lyophilized exosome standards (LESs) from HCT116 and biotin anti-human CD63 antibody were obtained from HansaBioMed and BioLegend, respectively. The remainder of the materials included the miRNeasy Mini kit (Qiagen) and the CEA (Human) ELISA kit (Abnova).
Fabricating paper-based devices
Paper-based devices were fabricated by exploiting the procedure used in our recent publication to immobilize the anti-CD63 antibodies onto the Whatman papers [33]. To do this, circular pieces of 6-mm Whatman paper were incubated in 50 µL of 4% MPTMS (in 99.9% ethanol) and 0.01 µmol GMBS (in 99.9% ethanol) for 30 min and 15 min, respectively. These pieces were then exposed to 50 µL NeutrAvidin (10 µg/mL in PBS) at 4 °C for 60 min. In the next step, 20 µL of 1% BSA in PBS (w/v) was dropped onto the papers for 3 times every 10 min to block the unreacted sites and prevent nonspecific absorption. Lastly, 20 µL of 20 µg/mL biotinylated anti-CD63 antibody (in PBS containing 1% w/v BSA) was dispensed onto the modified zones for 3 times every 10 min, and the unbound molecules were disposed of by PBS droplets for 3 times. All steps were performed at room temperature, unless otherwise stated.
Collecting biological samples
Commercial exosomes
LESs from HCT116 were stored at 4 °C and reconstituted with proper volumes of deionized water in accordance with the desired concentrations before use.
HCT116 cell culture media
HCT1116 CCM was centrifuged to eliminate cells, cell debris, and unwanted contaminants (Fig. S1). The settings of culturing HCT116 cells and centrifugation used in this study were identical to the ones described and employed in a recent publication on exosome extraction by our paper-based devices [33]. The samples of concentrated HCT116 exosomes (HCT116-exo) were then diluted in PBS at different ratios (1:6, 1:10, 1:20, 1:40) to vary their concentration.
Platelet-poor plasma and chronic lesions
Samples of platelet-poor plasma (PPP) and exudate from chronic wounds (or chronic wound fluid—CWF) with their information about sampling time were provided by Dr. Shin-Chen Pan at the National Cheng Kung University Hospital and stored at − 20 °C until used. Informed consent was written by all patients, and the study protocol was conducted in accordance with the Declaration of Helsinki and approved by the Ethics Committee of National Cheng Kung University Hospital (IRB no. B-ER-110–506). All experiments were performed in accordance with relevant procedures and regulations.
PPP was subjected through a membrane system consisting of 200- and 30-nm-layers as a preliminary filtration process to sequester exosomes (Fig. S2). The samples retained between the two filters (Retentate) and the samples passed through the second filter (Filtrate) were then resuspended in PBS. All three plasma-derived samples (original PPP, Retentate, and Filtrate) were analyzed by NTA before experiencing exosomal nucleic acid extraction (Fig. 1).
Preparing sol–gel silica
Sol–gel silica was obtained from a 2-step preparation. First, 0.5 g silica powder was dispersed in 15 mL DI water at 60 °C for 10 min. Second, 1 g silica powder and 50 µL 70% HNO3 were co-dispensed into the mixture and heated at 80 °C for 30 min.
Capturing exosomes from biological samples
Exosomes from biological samples (reconstituted LESs, concentrated HCT116-exo, PPP samples, and CWF) were captured by incubating the active zones of the paper-based devices with a volume of 20 µL analyte for 20 min before the unwanted materials were removed by PBS.
Extracting exosomal nucleic acids by intermittent lysis on paper-based devices
LESs reconstituted at the desired concentration were captured by paper-based devices before being intermittently lysed for their nucleic acids to be released and adsorbed by silica particles. This process was experimented with by slowly dropping 95 °C ddH2O onto the capturing-exosome zones at different volumes and time interval following with a 20 µL silica droplet to identify the regime which can maximize the content of extracted nucleic acids. 20 µL ddH2O was then dropped onto the experimental devices for 3 times to remove unwanted materials before incubating the devices in ddH2O at 55 °C for 45 min for nucleic acid elution (the volume of ddH2O was equal to the originally used volume of the samples). The content of extracted nucleic acids was finally estimated by RT-qPCR of miR-21 as a representative target. The settings of intermittent lysis were varied (Table 1) to establish the regime which could maximize the extracted amount of nucleic acids. The optimal regime was then identically employed for denaturing exosomes captured from different biological samples (concentrated HCT116-exo, PPP, and exudate of chronic lesions) and extracting their exosomal nucleic acids (Fig. 1).
SEM imaging
The paper-based devices were observed by S-3000H Scanning Electron Microscopes (Hitachi) after capturing LESs for their morphological analysis.
Quantifying captured LESs by ELISA on a paper-based device (P-ELISA)
After capturing LESs, both anti-CD63 papers and unmodified papers were alternately contacted with 5 µL of 1 µg/mL anti-CD9 (in PBS) for 1 min and 20 µL of PBS for 3 times. This step was repeated with 5 µL of HRP-conjugated anti-rabbit antibody before adding a 5 µL mixture of 3,3′,5,5′-tetramethylbenzidine (TMB) and hydrogen peroxide (1:1 v/v) for color development. All the paper pieces were finally photographed, and their color intensity was analyzed by ImageJ software.
Nanoparticle tracking analysis
All the PPP and CCM-derived samples were diluted for 120 times before being analyzed by NanoSight NS300 (Malvern Panalytical) equipped with a green laser and the NTA 3.4 software.
Analyzing miR-21 by RT-qPCR
Exosomal miR-21 extracted from all the samples (reconstituted LESs, concentrated HCT116-exo, PPP samples, and CWF) by the paper-based procedure was analyzed by RT-qPCR using identical probes and primers (Table S1), chemicals, and settings described in our recent publication [33].
Quantifying CEA by ELISA
This study employed two kinds of ELISA for the purposes of quantifying exosomal CEA and proving that the modified papers hardly capture free-circulating exosomes in bio-samples. The former was achieved using an Abnova CEA (Human) ELISA kit whereas P-ELISA on the anti-CD63 papers was designed for the latter.
On the one hand, paper pieces containing isolated exosomes were incubated in 50 µL lysis buffer from the mirVana miRNA Isolation kit for 5 min and an additional 10 µL Chloroform for 2–3 min to lyse exosomes for detection of exosomal CEA by the CEA (Human) ELISA kit. Briefly, 50 µL of samples were dispensed to the antibody-coated wells and incubated with 100 µL of enzyme conjugate reagent (goat anti-CEA antibody conjugated to horseradish peroxidase) for 60 min. One hundred microliter of TMB as a colorimetric reagent was added after the wells were emptied and rinsed 5 times with DI H2O. After incubating for 20 min, 100 µL of Stop solution (1N HCl) was finally added to the wells, and the color absorbance at 450 nm was evaluated by BioTek Synergy H1 Hybrid Multi-Mode Monochromator Fluorescence Microplate Readers (Thermo Fisher Scientific) within 15 min. The entire process was carried out at room temperature, as illustrated in Fig. S3.
On the other hand, different concentration of non-exosomal CEA solutions was dispensed to the anti-CD63-modified zones, and P-ELISA was performed by treating with anti-CEA-conjugated HRP (goat antibody) and TMB. The final state of the papers was captured to be analyzed by ImageJ software. All the reagents used in this ELISA were also supplied in a CEA (human) ELISA kit (Abnova).
Results and discussion
Intermittent lysis to extract exosomal nucleic acids
In this study and our previous publication [33], high-temperature ddH2O was used for exosome lysis because it is an ultra-purified and conveniently accessible buffer, which not only meets the criteria for a POCT but also avoids product contamination and interference in subsequent analysis compared to protein-containing lysis buffers. However, in our previous publication [33], the paper-based devices after capturing exosomes were immersed in 95 °C ddH2O for 30 min, which is necessary to destabilize exosomes and their membrane proteins but also accidentally degrade their extracted components. To subdue this dilemma, we delineated a novel strategy for exosome lysis by sequentially dropping 95 °C ddH2O onto paper-based devices, so-called intermittent lysis, instead of immersing them in such a high temperature for a long operation time. There were 3 variables experimented in our survey to determine how they dominate the lysis, including the total volume of 95 °C ddH2O (30, 40, 45, and 60 µL), the interlude of dropping 95 °C ddH2O (5, 10, and 15 min), and the total lysis time (30 and 40 min) (Table 1). A glance at RT-qPCR data of miR-21 lysed by varied regimes (Fig. 2) demonstrated that lysing exosomes by periodically dropping 10 µL 95 °C ddH2O onto the paper-based devices every 5 min over the course of 30 min (10 × 6) provided the optimal condition for the release of nucleic acids with the most extracted miR-21 content (the lowest Ct value of 25.67 ± 0.25). Contacting exosome-captured papers with 15 µL HT water for 4 times over the course of 30 min (15 × 4) also can improve the extracting efficiency at a certain level, although it was still far from the result achieved by 10 × 6 regime. Since both regimes used the equal volume of 95 °C ddH2O (60 µL), it is feasible to conclude that while lengthening the operation time of the lysis process does not guarantee an improvement in the extracted amount of nucleic acids, raising the number that the HT water dropped onto the exosome-captured papers during this period is an efficient strategy for this aim. In addition to that, a comparison between two regimes (10 × 4 and 15 × 4) in which the exosome-captured papers were contacted with various volumes of HT water (40 µL and 60 µL) for identical 4 turns and total lysis times of 40 min revealed that using a greater volume of HT water also contributes to the lysis efficiency. These results inferred that the lysis process of exosomes can be optimized by maximizing the number of dropping HT water onto the exosome-captured papers and its volume, whereas lengthening the operation time is optional.
Since intermittent lysis at different regimes expressed the superiority in lysing exosomes captured by the paper-based devices, their impact on exosome morphology was further investigated and characterized by SEM (Fig. 3). A comparison among the images displaying the surfaces of exosome-captured papers before (Fig. 3A) and after being treated with 95 °C ddH2O at different regimes (Fig. 3B–D) suggested that exposing exosomes to HT ddH2O altered their shape and dimension. To be more details, an initial observation in Fig. 3A discovered tiny particles with round shapes of about 100 nm, which belongs to the reported range of exosome dimensions [3, 4]. It is also evidence that the anti-CD63-immobilized papers succeeded in capturing exosomes since the samples used in this section were commercially purified exosomes. Their diameter was then remarkably bloated to the range of 200–600 nm (Fig. 3B, C) after being treated with 95 °C ddH2O at different regimes (Fig. 3B–D), with the maximally bloated dimension up to more than 800 nm after the paper was added 10 µL of 95 °C ddH2O 6 times every 5 min (Fig. 3D). These results were consistent with previously reported that exosomes were unstable when incubated for 30 min at 90 °C when membrane proteins were virtually degraded [32]. Considering this outcome and the one above, we hypothesize that exposing exosomes to HT ddH2O for a certain period denatured its lipid bilayer structure so that the internal components can be favorably released. Based on the available results, adding 10 µL 95 °C ddH2O 6 times every 5 min was chosen to intermittently lyse exosomes captured from biological samples to extract their internal nucleic acids.
Before applying the optimal setting of intermittent lysis to extract exosomal components of biological samples, it is crucial to verify that the RT-qPCR data were exclusively from exosomal miR-21. To do this, (1) LESs captured by the paper-based devices were quantified by ELISA (P-ELISA) and (2) free-circulating miR-21 which was non-specifically adsorbed by the paper-based devices was determined by RT-qPCR. On the one hand, the Δgrey values of the unmodified papers performed in Fig. S4A after the ELISA procedures were approximately equal, indicating that these papers barely captured exosomes despite being incubated with them at different levels. In contrast, the Δgrey outputted by ELISA for anti-CD63-coated papers capturing various amounts of exosomes not only obviously discriminated from that value of the control samples (no exosome particles) but also linearly intensified with the increasing amounts of exosomes (Fig. S4B), demonstrating the presence of exosomes on the modified papers. These data also presented a significant distinguishability between non-modified and antibody-modified papers. Given the above results, different exosome concentration could be distinguished, and exosome concentration in unknown samples could be estimated. On the other hand, a comparison among Ct values of 1 pM miR-21, 1 pM miR-21 and several DNase-, RNase-, and exosome-free solutions (1 × PBS, 6 ng/mL CEA, and non-template control) as negative controls after undergoing the described exosomal nucleic acid extraction process endorsed that most of the nucleic acids obtained from our extraction process were originated from exosomes (Fig. S5). Not only was the Ct value of the 1 pM miR-21 that underwent the paper-based extraction process considerably higher than that value of the untreated sample (15.57 ± 0.55), which was consistent with the one reported in the literature [34] but also it was on par with those of the exosome-free solutions and NTC (no template control), supporting the argument that the antibody-immobilized papers hardly captured free nucleic acids. However, it is also worth noting that the nonspecific adsorption of nucleic acids onto the paper-based devices is not trivial, since there was a visible distinction between the Ct value of the miR-21 sample and negative control samples (PBS, CEA, and NTC) after all of them underwent the whole procedure on the papers. This adsorbate was eluted in the subsequent steps and resulted in a visibly lower Ct value of miR-21 samples compared to the non-miR-21 ones.
Operability of the paper-based procedure in extracting exosomal nucleic acids
This section is a further examination of the workability of the whole system by testing the function of the fabricated paper-based devices after being preserved at different conditions in terms of time and temperature, as well as comparing its working efficiency with miRNeasy Mini kit, a popular commercial miRNA extraction tool which is universally applicable for various of culture media and cell types (humans, plants, and animals) (Fig. 4). On the one hand, the former was implemented by employing the anti-CD63 papers, after storing them under room temperature at 4 °C for 1, 2, 3, and 4 weeks to extract the nucleic acids derived from the same amount of LES samples. It is obvious to realize from the Ct values of the extracted miR-21 that storing the fabricated papers at ambient temperature rapidly deteriorates their function within less than 7 days. Conversely, the immunoaffinity of the paper-based devices will be consistent for more than a fortnight if they are maintained under cold condition of 4 °C (Fig. 4A), which is very promising for mass-production and transportation to the areas far from the manufacturing site or specialized laboratory. On the other hand, the latter was completed by evaluating Ct values of miR-21 in the nucleic acid solution extracted from the LESs using the novel strategy developed in this study and miRNeasy Mini kit. Empirical data indicated that extracting nucleic acids by our method yielded higher content of miR-21 despite consuming only half of the sample required by the kit (Fig. 4B), feasibly commercialized to replace this kit as a low-cost and environmental-friendly for miRNA extraction. In summary, it is possible to conclude from the results in this section and the previous section that the technology developed in this study is applicable for point-of-care applications since it fulfills the ASSURED criteria (affordable, sensitive, specific, user-friendly, rapid and robust, equipment-free, and deliverable to end users) proposed by the World Health Organization (WHO) for the development of POCT applications in areas with limited resources.
Extracting exosomal nucleic acids from biological samples by paper-based intermittent lysis
HCT116 cell culture media
After optimizing the intermittent lysis and verifying the workability of the designed procedure on commercial exosomes, the whole system was deployed to extract exosomal nucleic acids from CCM, PPP, and CWF as biological samples, which consist of not only exosomes but also miscellaneous components. Initially, exosomes secreted by HCT116 were concentrated by centrifugation to remove cells, cell debris, and unwanted contaminants, before being diluted to different ratios of 1:6, 1:10, 1:20, and 1:40 in PBS. These diluted samples were then subjected to the developed system for extracting exosomal biomarkers. Our previous publication has demonstrated that the immunoaffinity paper-based devices are capable of capturing and quantifying exosomes from cells cultured in different environments [33]. In this study, the particle concentration and particle size distribution of these HCT116-ex samples were determined prior to their exosomal biomarker contents (CEA and miR-21) being evaluated (Table S2 and Fig. 5). Obviously, a majority of the particles detected in Fig. 5A belonged to the reported range of exosome dimensions [3, 4] with high concentrations (Table S2), essentially guaranteeing that the concentrated exosomes were not damaged by the pretreatment steps, and they were intact for exosomal biomarker extraction, as well as the data used to evaluate exosomal biomarker contents (CEA and miR-21) were from the HCT116 exosomes. It also can be inferred from the calibration line in Fig. 5B, which presented an inversely proportional relationship between the particle concentration of captured exosomes and corresponding Ct values of miR-21, that the extracted exosomal miR-21 was proportional to the captured HCT116-exo. In addition to those, we also conducted quantitative assays for exosome-derived proteins and investigated the correlation between exosomal protein, with exosomal CEA as a representative target and exosomal miR-21 as an extension for downstream analysis. Since CEA has been ratified as a biomarker for colorectal cancer [9,10,11] and miR-21 has been found to increase in patients with this chronic disease [5, 6], discovering the correlation between these cancerous cell-derived exosomal biomarkers can benefit in diagnosing and monitoring it for appropriate intervention and treatment. A graph delineating the relationship between the exosomal CEA concentration (horizontal axis) and the exosomal miR-21 Ct value (vertical axis) performed an inversely proportional relationship between these values (Fig. 5C) with the concentration of CEA in exosomes derived from colorectal cancer cells is significantly higher than the normal concentration of CEA plasma (5 ng/mL) [9], despite low content of exosomes (0.758 × 1011 particles/mL), raising the question of whether exosomal CEA levels might become a more effective biomarker than free-circulating CEA in plasma for cancer diagnosis and prognosis. These discoveries were concreted by the fact that the non-specific CEA were hardly retained on the paper surface after washing steps, as the Δgrey values analyzed by ImageJ indicated that there was no difference between the papers incubated with 3, 6, and 12 ng/mL CEA, whereas the paper incubated with 30 ng/mL without washing exhibited a superior intensity of Δgrey compared to the paper which was washed after incubating with 30 ng/mL CEA (Fig. 5D). Since the interest of this experiment is exosome-derived CEA, it is critical to prove that the anti-CD63 papers did not nonspecifically adsorb free-circulating CEA in the samples. In summary, these results confirmed that our deployed system not only could harvest exosomal nucleic acids from exosomes released by HCT116 cells at various concentration but also supports an in-depth analysis of the relationship between exosome-originated components.
A Size distribution of particles in HCT116-exo 1:40. B Calibration line presented a linear relationship between exosome concentration and Ct values of exosomal miR-21 extracted by paper-based intermittent lysis procedure from different HCT116-exo samples, y = 30x − 0.8. C Calibration line presented a linear relationship between exosomal CEA concentration and Ct values of exosomal miR-21 extracted by paper-based intermittent lysis procedure from different HCT116-exo samples, y = 31.6x − 0.1. D Quantification of non-exosomal CEA non-specifically absorbed on anti-CD63 papers by ELISA (P-ELISA) after incubating the modified papers with different CEA concentration
Platelet-poor plasma
Encouraged by a successful trial on cell culture samples under the assistance of centrifugation, we applied the intermittent lysis to extract exosomal nucleic acids from human blood, which is not only the most popular sample in clinical diagnosis but also the source containing the most exosomes. In this experiment, PPP was first undergone a dual-layer membrane system constituted of 200- and 30-nm layers to collect Retentate and Filtrate samples, which theoretically retained exosomes and filtered plasma proteins and nucleic acids, respectively. On the one hand, the whole procedure was then applied to extract exosomal nucleic acids from three plasma-derived samples (PPP, Retentate, and Filtrate), and the Ct values of these samples were evaluated with the non-template control (NTC) (Fig. 6A). Obviously, there was a considerable distance between the Ct data of two groups, with the values of PPP and Retentate samples were significantly lower than the ones of Filtrate and NCT samples, evidencing a more abundant amount of miR-21 in the former group than in the latter group. It is also observed from this figure that the Ct value of the PPP was slightly lower than the Retentate sample, and so are the Ct values of Filtrate and NTC. Although these phenomena were supposedly a consequence of non-specific adsorption of plasma miR-21 onto the papers during the extraction process, which has been noted in the “Intermittent lysis to extract exosomal nucleic acids” section, they were also a demonstration that the whole procedure could independently extract exosomal nucleic acids without sample pretreatment. This conclusion was, on the other hand, also concreted by the NTA data, which was used to measure the particle concentration and outputted approximately equal numbers between PPP and Retentate (Table S3).
Chronic lesions
In this section, the established procedure was further employed to extract exosomal nucleic acids from CWF as a clinical without pretreatment by centrifugation nor membrane. Since miR-21 has been shown to be intimately involved in multiple stages of wound healing, such as playing a role of an inhibitor of phosphatase and tensin homolog (PTEN) and an activator of many cell-survival and proliferative pathways [5, 6, 34], monitoring and determining the expression of this biomarker is crucial to evaluate the treatment efficiency. On the one hand, there was no difference among Ct values of the samples simultaneously collected from distinct positions of a patient (who was coincidentally injured in the left leg, right leg, and left elbow), suggesting that the amount of exosomal miR-21 in one person is independent from the injured location with the same physiologic and treatment conditions (Fig. 6B). On the other hand, the exudate from chronic lesions of several patients (ciphered as W58, W87, W103, W121, W134) was harvested before and after a fortnight to extract the exosomal miR-21 for treatment evaluation (Table S4). It can be inferred from Fig. 6C that the miR-21 level in the fluid exudated from the same chronic wounds greatly reduced after a 2-week treatment since their Ct values remarkably rose. This finding was not only in line with the previously mentioned that exosomal miR-21 plays an important role in the wound healing process [7, 8] but also in agreement with our recent discovery that exosomal miR-21 level starts depleting after 1-week treatment [33]. Notably, its depletion was more substantial in the wounds which were intervened for less than 2 months (2 and 7 weeks) than in the wounds with longer treatment (6, 7, and 11 months). Based on these discoveries, we conjecture that exosomal miR-21 content experienced 3 phases during the healing process: (1) an initial rise to trigger the process, (2) a dramatic decline in the next stage, and (3) a gradual depletion with the prolongation of the healing process. This hypothesis proposes that miR-21-targeted therapy is potentially applicable in the treatment of chronic wounds.
In summary, downstream analysis of exosomal miR-21 contents extracted from HCT116 cell culture media, PPP, and chronic wounds demonstrated the capability of our strategy in independently extracting exosomal nucleic acids from various biological samples, both independently and in combination with other exosome isolation methods (centrifugation and ultrafiltration), for diagnostics and therapeutics without elaborate sample pretreatments. Challenges to widely apply it in clinical settings include regulatory approval and standardization of the whole procedure, and solutions to develop sample-to-result paper-based systems (closed systems) for exosomal nucleic acid biomarker quantification.
Conclusion
This study engineered the intermittent lysis method to extract exosome-originated nucleic acids for downstream analysis and applied it to biological samples from cancer cell culture media, human plasma, and wound exudate. Our novel strategy has been evidenced to specifically extract exosomal nucleic acids both in vitro (culture media) and in vivo samples (plasma and wound fluids) without absorbing unwanted impurities. The technique and the paper-based procedure presented in this study not only overcomes the barriers in exosome purification due to their ultrasmall size but also possesses outstanding features (low budget, easy-to-operate steps, small volume sample, better performance than conventional kit, and applicable at low resource setting areas and locations far from specialized laboratories) that facilitate the POCT and pave the way to further analysis of exosome-transported genetic materials. Additionally, experimental results on exosome-derived components manifested that exosomal miR-21 levels in chronic wound fluid are strongly associated with wound recovery and have enormous potential as a therapeutic target for chronic injuries discovered a relevance between the content of exosomal miR-21 and exosomal CEA in colorectal cancer cells, as well as proposed a more pertinent role of exosomal CEA in this chronic disease.
Data availability
Data will be made available on reasonable request.
References
Makler A, Asghar W (2020) Exosomal biomarkers for cancer diagnosis and patient monitoring. Expert Rev Mol Diagn 20:387–400. https://doi.org/10.1080/14737159.2020.1731308
Hong C-H, Chen Y-C (2019) Clinical significance of exosomes as potential biomarkers in cancer. World J Clin Cases 7:171–190https://doi.org/10.12998/wjcc.v7.i2.171
Cunnane EM, Weinbaum JS, O’Brien FJ, Vorp DA (2018) Future perspectives on the role of stem cells and extracellular vesicles in vascular tissue regeneration. Front Cardiovasc Med 5:86. https://doi.org/10.3389/fcvm.2018.00086
Cheng Y, Qu X, Dong Z et al (2020) Comparison of serum exosome isolation methods on co-precipitated free microRNAs. PeerJ 8:e9434.https://doi.org/10.7717/peerj.9434
Raposo G, Stoorvogel W (2013) Extracellular vesicles: exosomes, microvesicles, and friends. J Cell Biol 200:373–383 (www.jcb.org/cgi/doi/10.1083/jcb.201211138)
Xie Y, Dang W, Zhang S et al (2019) The role of exosomal noncoding RNAs in cancer. Mol Cancer 18:37.https://doi.org/10.1186/s12943-019-0984-4
You C, Jin L, Xu Q, Shen B, Jiao X, Huang X (2019) Expression of miR-21 and miR-138 in colon cancer and its effect on cell proliferation and prognosis. Oncol Lett 17:2271–2277. https://doi.org/10.3892/ol.2018.9864
Wang T, Feng Y, Sun H et al (2012) miR-21 regulates skin wound healing by targeting multiple aspects of the healing process. Am J Pathol 181:1911–1920. https://doi.org/10.1016/j.ajpath.2012.08.022
Yang X, Wang J, Guo S-L et al (2011) miR-21 promotes keratinocyte migration and re-epithelialization during wound healing. Int J Biol Sci 7:685–690. https://doi.org/10.7150/ijbs.7.685
Goldstein MJ, Mitchell EP (2005) Carcinoembryonic antigen in the staging and follow-up of patients with colorectal cancer. Cancer Invest 23:338–351. https://doi.org/10.1081/CNV-58878
Björkman K, Jalkanen S, Salmi M et al (2021) A prognostic model for colorectal cancer based on CEA and a 48-multiplex serum biomarker panel. Sci Rep 11:4287. https://doi.org/10.1038/s41598-020-80785-1
Yokoyama S, Takeuchi A, Yamaguchi S et al (2017) Clinical implications of carcinoembryonic antigen distribution in serum exosomal fraction—measurement by ELISA. PLoS ONE 12:e0183337. https://doi.org/10.1371/journal.pone.0183337
Colombo M, Raposo G, Théry C (2014) Biogenesis, secretion, and intercellular interactions of exosomes and other extracellular vesicles. Annu Rev Cell Dev Biol 30:255–289. https://doi.org/10.1146/annurev-cellbio-101512-122326
Patel GK, Khan MA, Zubair H et al (2019) Comparative analysis of exosome isolation methods using culture supernatant for optimum yield, purity and downstream applications. Sci Rep 9:5335. https://doi.org/10.1038/s41598-019-41800-2
Duarte M, Subedi P, Yilmaz E, Marcus K, Laurell T, Ekström S (2018) Molecularly imprinted polymers synthesized via template immobilization on fumed silica nanoparticles for the enrichment of phosphopeptides. J Mol Recognit 31:e2677. https://doi.org/10.1002/jmr.2677
Kuo WP, Jia S (2017) Methods in molecular biology 1660: extracellular vesicles. Humana, New York. https://doi.org/10.1007/978-1-4939-7253-1
Channavajjhala SK, Rossato M, Morandini F et al (2014) Optimizing the purification and analysis of miRNAs from urinary exosomes. Clin Chem Lab Med 52:345–354. https://doi.org/10.1515/cclm-2013-0562
Eldh M, Lötvall J, Malmhäll C, Ekström K (2012) Importance of RNA isolation methods for analysis of exosomal RNA: evaluation of different methods. Mol Immunol 50:278–286. https://doi.org/10.1016/j.molimm.2012.02.001
Mutschelknaus L, Azimzadeh O, Heider T et al (2017) Radiation alters the cargo of exosomes released from squamous head and neck cancer cells to promote migration of recipient cells. Sci Rep 7:12423. https://doi.org/10.1038/s41598-017-12403-6
Hernandez-Valladares M, Aasebø E, Mjaavatten O et al (2016) Reliable FASP-based procedures for optimal quantitative proteomic and phosphoproteomic analysis on samples from acute myeloid leukemia patients. Biol Proced Online 18:13. https://doi.org/10.1186/s12575-016-0043-0
Yang M, Song D, Cao X et al (2017) Comparative proteomic analysis of milk-derived exosomes in human and bovine colostrum and mature milk samples by iTRAQ-coupled LC-MS/MS. Int Food Res J 92:17–25. https://doi.org/10.1016/j.foodres.2016.11.041
Subedi P, Schneider M, Philipp J et al (2019) Comparison of methods to isolate proteins from extracellular vesicles for mass spectrometry-based proteomic analyses. Anal Biochem 584:113390. https://doi.org/10.1016/j.ab.2019.113390
Schrader C, Schielke A, Ellerbroek L, Johne R (2012) PCR inhibitors – occurrence, properties and removal. J Appl Microbiol 113:1014–1026. https://doi.org/10.1111/j.1365-2672.2012.05384.x
Chugh PE, Sin S-H, Ozgur S et al (2013) Systemically circulating viral and tumor-derived microRNAs in KSHV-associated malignancies. PLoS Pathog 9:e1003484. https://doi.org/10.1371/journal.ppat.1003484
Clayton A, Court J, Navabi H et al (2001) Analysis of antigen presenting cell derived exosomes, based on immuno-magnetic isolation and flow cytometry. J Immunol Methods 247:163–174. https://doi.org/10.1016/S0022-1759(00)00321-5
Davtyan SP, Berlin A, Agabekov V, Lekishvili N (2012) Synthesis, properties, and applications of polymeric nanocomposites. J Nanomater 2012:215094. https://doi.org/10.1155/2012/215094
Sahoo SL, Liu C-H (2015) Adsorption behaviors of DNA by modified magnetic nanoparticles: effect of spacer and salt. Colloids Surf A Physicochem Eng Asp 482:184–194. https://doi.org/10.1016/j.colsurfa.2015.05.010
Castro-Smirnov FA, Pietrement O, Aranda P et al (2016) Physical interactions between DNA and sepiolite nanofibers, and potential application for DNA transfer into mammalian cells. Sci Rep 6:36341. https://doi.org/10.1038/srep36341
Li X, Zhang J, Gu H (2012) Study on the adsorption mechanism of DNA with mesoporous silica nanoparticles in aqueous solution. Langmuir 28:2827–2834. https://doi.org/10.1021/la204443j
Hair ML, Hertl W (1970) Acidity of surface hydroxyl groups. J Phys Chem 74:91–94 https://doi.org/10.1021/j100696a016
Melzak KA, Sherwood CS, Turner RF, Haynes CA (1996) Driving forces for DNA adsorption to silica in perchlorate solutions. J Colloid Interface Sci 181:635–644. https://doi.org/10.1006/jcis.1996.0421
Lee M, Ban J-J, Im W, Kim M (2016) Influence of storage condition on exosome recovery. biotechnol bioprocess Eng 21:299-304. https://doi.org/10.1007/s12257-015-0781-x
Lai C-H, Lee C-L, Vu C-A et al (2022) Paper-based devices for capturing exosomes and exosomal nucleic acids from biological samples. Front Bioeng Biotechnol 10:836082. https://doi.org/10.3389/fbioe.2022.836082
Hu W-P, Chen Y-C, Chen W-Y (2020) Improve sample preparation process for miRNA isolation from the culture cells by using silica fiber membrane. Sci Rep 10:21132. https://doi.org/10.1038/s41598-020-78202-8
Acknowledgements
The authors acknowledge the financial support from the Taiwan Ministry of Science and Technology (grant no. 109-2811-E-008-518 and 111-2314-B-006-093) for this research.
Funding
This work was supported by the Taiwan Ministry of Science and Technology (grant no. 109–2811-E-008–518 and 111–2314-B-006–093).
Author information
Authors and Affiliations
Contributions
V.-T.V.: data curation, formal analysis, visualization, investigation, methodology, project administration, validation, and writing—original draft. C.-A.V.: data curation, formal analysis, visualization, investigation, methodology, supervision, validation, and writing—review and editing. C.-J.H.: formal analysis, resources, methodology, supervision, and validation. C.-M.C.: formal analysis, resources, methodology, supervision, and validation. S.-C.P.: formal analysis, funding acquisition, methodology, resources, supervision, and validation. W.-Y.C.: formal analysis, funding acquisition, methodology, project administration, resources, supervision, validation, and writing—review and editing.
Corresponding authors
Ethics declarations
Ethics approval
This study was performed in line with the principles of the Declaration of Helsinki. Approval was granted by the Institutional Review Board at National Cheng Kung University Hospital (no. B-ER-110–506).
Consent for publication
The authors affirm that human research participants provided informed consent for publication.
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Vu, VT., Vu, CA., Huang, CJ. et al. Intermittent lysis on a single paper-based device to extract exosomal nucleic acid biomarkers from biological samples for downstream analysis. Microchim Acta 191, 501 (2024). https://doi.org/10.1007/s00604-024-06566-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00604-024-06566-z