Abstract
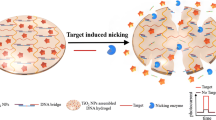
A novel photoelectrochemical (PEC) biosensor based on an enzyme-free nucleic acid dual-amplification strategy combined with a mimic enzyme to catalyze the deposition of a quencher is reported for the ultrasensitive detection of miRNA-21. A limited amount of target miRNA-21 can trigger the formation of long DNA duplexes on the electrode, owing to the synergistic effect of the enzyme-free nucleic acid dual-amplification strategy of entropy-driven strand displacement reaction (ESDR) amplification and hybridization chain reaction (HCR) amplification. The embedded manganese porphyrin (MnPP) in the long DNA duplexes acts as a horseradish peroxidase (HRP)–mimicking enzyme to catalyze the transformation of benzo-4-chlorohexadienone on the electrode surface, resulting in a significant reduction in photocurrent intensity. As a photosensitive material, BiOCl-BiOI is used as a tag to provide strong initial PEC signals. Based on the cascade integration of the enzyme-free nucleic acid dual-amplification strategy and the mimic enzyme-catalyzed precipitation reaction, the current PEC biosensor exhibits outstanding performance for miRNA-21 detection with an ultralow detection limit (33 aM) and a wide quantification range (from 100 aM to 1 nM). This work provides a new avenue toward the ultrasensitive detection of miRNAs, and is expected to be used for clinical and biochemical samples.
Graphical abstract
A unique PEC biosensor with the BiOCl-BiOI composite, as the photosensitive material, has been developed for ultrasensitive miRNA-21 determination based on the combination of an enzyme-free nucleic acid dual-amplification strategy and mimic enzyme catalytic precipitation reaction.

Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
microRNAs (miRNAs), endogenous non-coding small RNAs, play significant roles in cell proliferation, differentiation, and apoptosis by regulating the expression of genes and translation of proteins [1,2,3,4,5]. miRNAs start as pri-miRNAs, which are processed to become pre-miRNAs, as well as miRNA precursors, with a length of about 70–90 nucleotides; pre-miRNAs are cleaved by corresponding enzymes to become mature miRNAs with 18–25 nucleotides in length [6, 7]. Abnormal expression of mature miRNAs is closely related to the occurrence and development of diverse diseases, especially cancers [8,9,10]. However, high-performance detection of miRNAs faces severe challenges owing to the low abundance, short sequence, and sequence similarity of miRNAs [11, 12]. There is an urgent need to develop high-performance biosensors for the ultrasensitive detection of mature miRNAs.
In recent years, photoelectrochemical (PEC) biosensors have drawn extensive attention owing to their low background and high sensitivity [13,14,15], compared with counterparts based on electrochemistry [16, 17], electrochemiluminescence [18], and fluorescence [19]. Noteworthy, turn-off PEC sensing systems have been widely applied in the detection of miRNAs with the advantages of low cost, handy operation, and fast response speed [20, 21]. It is well accepted that the sensitivity of signal-off PEC biosensor primarily depends on the degree of reduction of photocurrent after target recognition, which is closely related to the intensity of initial PEC signal and the avenue to reduce the photocurrent [22, 23]. Excellent photoelectronic active materials play critical roles in offering strong initial PEC signals [24]. As a late-model photoactive material, BiOCl-BiOI has been applied in many areas for its excellent photoelectric properties, such as photocatalysis for photodegradation of organic contaminants and water decomposition and reduction of carbon dioxide [25, 26]. Here, BiOCl-BiOI is expected to become a promising photosensitive material which can provide a strong initial PEC signal for PEC sensors for miRNA detection.
To further improve the sensitivity of PEC biosensors for miRNA detection, enzyme-free amplification strategies, such as hybridization chain reaction (HCR) and entropy-driven strand displacement reaction (ESDR), have been widely used in PEC biosensor [27,28,29,30]. However, HCR amplification strategies can only convert one target into one output target mimic (long DNA duplexes), resulting in finite amplification efficiency [31, 32]. Noteworthy, ESDR can convert a target into an output, which can be used as a catalyst for other amplification reactions (HCR, for instance) to achieve higher amplification efficiency [33, 34]. Moreover, ESDR is an entropy-driven process with simple design, high reaction efficiency, outstanding amplification ability, and robust thermal stability [35, 36]. Hence, the combination of HCR and ESDR is an excellent alternative to gain higher signal amplification efficiency for highly sensitive detection of miRNA since more target mimics can participate in the HCR process. Besides, manganese porphyrin (MnPP), which can be inserted into DNA duplexes, has been extensively used as horseradish peroxidase (HRP) mimics to catalyze the transformation of certain reducing reagents into quenchers for reducing photocurrent by blocking electronic transmission [37, 38]. Embedding MnPP in HCR products will largely reduce the photocurrent signal. Therefore, the detection limits of miRNA-21 could be improved through the integration of an enzyme-free nucleic acid dual-amplification strategy with a mimicked enzyme catalytic precipitation reaction in PEC sensors.
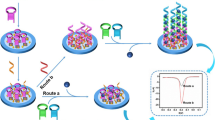
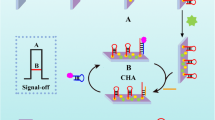
Herein, based on the enzyme-free nucleic acid dual-amplification strategy of HCR and ESDR amplification, along with the use of mimic enzyme catalytic reaction products as an efficient quencher, a novel PEC biosensor was designed for ultrasensitive detection of miRNA-21. As shown in Scheme 1, owing to the high photoelectric conversion efficiency of BiOCl-BiOI, the BiOCl-BiOI-modified glassy carbon electrode (GCE) could generate a strong photocurrent signal. Meanwhile, by modifying magnetic beads with the hybrid products of DNA S0 and S1, a capture DNA was constructed. Then, the target miRNA-21 could activate toehold-mediated ESDR and double-output amplification process by hybridizing with the capture DNA and S2, which would convert a small amount of target RNA into abundant output DNA (S1). The output S1 could further hybridize with hairpin H1 immobilized on the electrode to trigger HCR with the participation of hairpin H2 and H3, thereby obtaining a large amount of long DNA duplex that could be loaded with the abundant mimic enzyme MnPP. Finally, in the presence of H2O2, the embedded MnPP could catalyze 4-CN to benzo-4-chlorohexadienone (4-CD) and deposit it onto the surface of electrodes. The deposited 4-CD could be used as an effective quencher to significantly reduce the photocurrent signal by blocking the electronic transmission between the electrolyte and the photoactive substances. Thus, based on the above amplification principles, this PEC biosensor could be employed for the detection of low abundance miRNA-21. Moreover, the magnetic beads could effectively eliminate by-products under magnetic field and avoid the interference of S2 on subsequent HCR, which is conducive to improving the selectivity of the PEC biosensor. The experimental results confirmed that the PEC biosensor could quantitatively monitor miRNA-21 with ultra-sensitivity and high selectivity. This enzyme-free nucleic acid dual-amplification strategy with mimic enzyme catalytic precipitation reaction for the design of PEC sensors provides a simple but powerful protocol for ultrasensitive detection of miRNAs.
Experimental section
Interfacial modification of Au@Fe3O4
Au@Fe3O4 was prepared according to the protocols reported previously [39]. The prepared Au@Fe3O4 nanoparticles were dispersed in 2 mL of PBS and stored at 4 °C.
Synthesis of BiOCl-BiOI nanocomposite
BiOI powder was prepared by a simple hydrothermal method. A total of 1 mmol of KI was dissolved in 17 mL of deionized water. Subsequently, the above solution was dripped to 17 mL of ethanol containing 1 mmol Bi(NO3)3·5H2O and magnetically stirred at room temperature for 30 min to make all reagents uniformly mixed. The resultant precursor suspension was transferred to a Teflon-lined stainless autoclave and maintained at 150 °C for 8 h. After cooling to room temperature, a red precipitate was obtained by filtering and washing with water, and dried at 80 °C.
A total of 0.5 mmol of the prepared BiOI was dissolved in 17 mL of ultrapure water, followed by adding 1.2 mmol of NaCl with stirring. After stirring for 30 min, 17 mL of ethanol containing 1.2 mmol Bi(NO3)3·5H2O was added. Then, the mixture was transferred to autoclave, reacted at 100 °C for 18 h, and subsequently cooled to room temperature. The resultant products were cleaned with distilled water and then dried in air to obtain BiOCl-BiOI nanocomposite. In addition, pure BiOCl was prepared under the same conditions without the addition of BiOI.
ESDR amplification process
First, 20 µL of S0 (2 µM) was incubated with 45 µL of S1 (2 µM) at 37 °C for 2 h to obtain double-stranded DNA (dsDNA). The above dsDNA and 100 µL of Au@Fe3O4 stock solution were added into the centrifuge tube and mixed overnight on a shaking table at 4 °C in order to obtain dsDNA-Au@Fe3O4 as a capture DNA by Au–S bonding. Subsequently, after magnetic separation, the capture DNA, miRNA-21 in different concentrations, and 20 µL of S2 (2 µM) were incubated together at 37 °C for 2 h to trigger the ESDR amplification process. Finally, the supernatant was collected by magnetic separation to obtain the output DNA (S1) for participation in HCR process.
Fabrication of the proposed PEC biosensor
Prior to modification, the bare GCE was burnished with alumina powder (0.3 µm, 0.05 µm) and then thoroughly cleaned to attain a mirror-like surface. A total of 10 µL of BiOCl-BiOI (1 mg/mL) was modified onto the cleaned GCE surface, and then dried at 37 °C. Subsequently, the modified electrode was bathed in HAuCl4 solution (1%) for electrochemical deposition to obtain the Au nanoparticles layer (DepAu) on the surface of BiOCl-BiOI/GCE (deposition time, 15 s; constant potential, − 0.2 V). Next, 10 µL of thiol-modified H1 (2 µM) was dropped on the surface of DepAu/BiOCl-BiOI/GCE and allowed to incubate at 4 °C for 12 h. The H1 was immobilized on the surface of the electrode via Au–S binding, and then, 10 µL of hexanethiol (HT, 0.1 mM) was dropped on the electrode surface to block nonspecific sites at room temperature for 30 min. Afterwards, 10 µL of the output DNA was dropped on the surface of electrode at 37 °C for 2 h. And then, 7.5 µL of H2 (2 µM) and 7.5 µL of H3 (2 µM) were added and kept at 37 °C for 2 h to trigger the HCR. After that, 5 µL of MnPP (100 µM) was added to the electrode surface and incubated at 37 °C for 1 h to make MnPP embedded into the HCR products to be an HRP mimic enzyme. Finally, H2O2 (0.2 mM) and 4-CN (1 mM) were incubated on the synthesized electrode for 15 min to induce the enzymatic biocatalytic precipitation reaction.
PEC measurements
PEC biosensor measurements were performed in 5 mL of PBS (0.1 M, pH = 7.0) with the addition of H2O2 (50 μL, 30%) as electron donor. In this measurement system, a light-emitting diode lamp was used as the excitation light source with excitation wavelength of 365 nm, switched off–on-off for 10–20-10 s by the applied potential of 0 V (vs. reference electrode).
Statistical analysis
All data are presented as the mean ± SD. Unless stated otherwise, the experiments were performed in triplicate. The significance of the difference was determined through one-way analysis of variance (ANOVA) by *p < 0.05, **p < 0.01, ***p < 0.001, and ns p ≥ 0.05.
Analysis of real samples
Total RNA samples were extracted from fixed number of MCF-7 or HeLa cells (obtained after precise calculation and dilution). Like the previous “Fabrication of the proposed PEC biosensor” process, after the preparation of HT/H1/DepAu/BiOCl-BiOI/GCE, 10 µL of the output DNA along with total RNA samples was dropped on the surface of the electrode at 37 °C for 2 h. The trigged HCR and added MnPP were as described in the previous steps, with all processing times and assay detection conditions as before.
Results and discussion
Characterization of BiOCl-BiOI composites
The morphology of resultant products was characterized by scanning electron microscopy (SEM) and transmission electron microscopy (TEM). As observed in Fig. 1A and B, pure BiOI and BiOCl exhibited a flake-like shape with a smooth surface and a nanoflower structure, respectively. The BiOCl nanoflowers were successfully assembled on the surface of BiOI nanoflakes, as characterized by the SEM image (Fig. 1C) and the TEM image (Fig. 1E). Meanwhile, the uniform distribution of Bi, O, Cl, and I in BiOCl-BiOI was confirmed by energy-dispersive X-ray spectrometry (EDS) mapping (Fig. 1D). In addition, the lattice spacing of 0.335 nm and 0.301 nm in the high-resolution TEM (HRTEM) image can be ascribed to the (101) plane of BiOCl and the (012) plane of BiOI (Fig. 1F), respectively, indicating the successful assembly of BiOCl-BiOI composites.
X-ray photoelectron spectroscopy (XPS) was further applied to confirm the successful preparation of BiOCl-BiOI. As shown in Fig. S1A, all peaks matched well with the corresponding elements of Bi, O, Cl, and I, thus confirming their existence (C element comes from the indeterminate carbon in the instrument). Moreover, two main peaks at 164.74 and 159.40 eV could be observed in the high-resolution XPS spectrum of Bi 4f (Fig. S1B), corresponding to the 4f5/2 and 4f7/2 peaks of Bi, respectively. For O 1 s spectrum (Fig. S1C), one signal appeared at 530.26 eV, and the other peak at 531.66 eV was ascribed to the surface hydroxyl groups [25]. As shown in Fig. S1D, the two peaks at binding energies 198.26 and 199.90 eV were attributed to Cl 2p3/2 and Cl 2p1/2, respectively, indicating the existence of Cl element. Additionally, the XPS characteristic peaks at 631.03 and 619.62 eV corresponding to I 3d3/2 and I 3d5/2 respectively could verify the existence of I element (Fig. S1E). All above results strongly confirmed the successful fabrication of BiOCl-BiOI.
Electrochemical characterization of the proposed PEC biosensor
To further confirm the feasibility of the designed PEC biosensor, electrochemical impedance spectroscopy (EIS) and cyclic voltammetry (CV) measurements were carried out. As displayed in Fig. 2A, compared to bare GCE (curve (a)), BiOCl-BiOI-modified GCE (curve (b)) showed an increased charge transfer resistance (Ret) because of the steric hindrance effect that blocked electronic transformation. After electrodeposition of AuNPs on the BiOCl-BiOI/GCE electrode surface, the outstanding conductivity of AuNPs caused the obvious decrease of Ret (curve (c)). When H1, HT, and output DNA were stepwise decorated, the Ret value maintained a continuous increase (curves (d) to (f)) due to the electrostatic repulsion of the negatively charged DNA and closure of HT for nonspecific binding sites. Subsequently, the mixture of H2, H3, and MnPP was dropped onto the surface of the electrode (curve (g)), and the Ret was diminished due to the large number of free electrons on the large p-conjugated structure of MnPP [38]. Notably, the Ret markedly increased in the presence of H2O2 and 4-CN (curve (h)) because the MnPP inserted in HCR products can catalyze 4-CN to form 4-CD precipitation, leading to a significant increase in steric hindrance and effective suppression of electron transfer.
EIS (A) and CV responses (B) of bare GCE (a), BiOCl-BiOI/GCE (b), DepAu/BiOCl-BiOI/GCE (c), H1/DepAu/BiOCl-BiOI/GCE (d), HT/H1/DepAu/BiOCl-BiOI/GCE (e), output DNA/HT/H1/DepAu/BiOCl-BiOI/GCE (f), H2 + H3 + MnPP/output DNA/HT/H1/DepAu/BiOCl-BiOI/GCE (g), and H2O2 + 4-CN/H2 + H3 + MnPP/output DNA/HT/H1/DepAu/BiOCl-BiOI/GCE (h) in 0.1 M PBS with 5 mM [Fe(CN)6]3−/4−
Simultaneously, the CV results also confirmed the successful modification of the electrode. As displayed in Fig. 2A, the bare GCE (curve (a)) showed a pair of reversible redox peaks. With the BiOCl-BiOI nanocomposites deposited on the electrode surface, a decreased peak appeared as a result of the steric hindrance effect (curve (b)). When AuNPs were deposited on the above composite electrode surface (curve (c)), the peak current increased because of the preeminent conductivity of AuNPs. Then, the peak current declined with the incubation of H1, HT, and output S1 (curves (d)–(f)), which was attributed to electronic repulsion and poor conductivity. The peak intensity increased again with the addition of H2, H3, and MnPP (curve (g)), which was attributed to the preeminent conductivity of MnPP. Finally, the peak intensity decreased with the treatment of H2O2 and 4-CN (curve (h)), indicating that electron transfer was inhibited by the precipitation (4-CD) generated from the catalytic reaction of MnPP.
PEC characterization of the biosensing interface
Stepwise PEC characterization was carried out to confirm the successful construction of PEC biosensor interface. As exhibited in Fig. 3, almost no photocurrent response was observed in the naked GCE (curve (a)). The photocurrent response increased with BiOCl-BiOI coated on the surface of GCE (curve (b)), owing to the remarkable photoelectric activity. The electrodeposition of AuNPs on the BiOCl-BiOI/GCE electrode surface corresponded to the highest photocurrent response (curve (c)), which was ascribed to the outstanding conductivity of AuNPs. When H1, HT, and output DNA were dropped on the modified electrode in a stepwise manner, the photocurrent response gradually decreased (curves (d)–(f)), which resulted from the poor charge transport of the DNA skeleton and small organic molecules [40]. Notably, PEC response decreased significantly after incubation with H3, MnPP, H2O2, and 4-CN (curve (g)). With the addition of H2O2, MnPP acted as the enzyme mimic to catalyze the generation of 4-CD precipitation, thereby hindering electron transfer. These experimental results demonstrated the successful construction of the designed sensor.
Analytical performance of PEC biosensor
The analytical performance of this designed PEC biosensor was evaluated using different concentrations of miRNA-21. As depicted in Fig. 4A and B, the photocurrent responses of PEC biosensor gradually decreased as the concentration of miRNA-21 increased. Good linearity was obtained with common logarithm (lg) of miRNA-21 concentrations ranging from lg100 aM to lg1 nM (equation: I = − 0.1236lgc + 1.152, R2 = 0.9962, where I and c represent as PEC response and concentration of miRNA-21, respectively). The detection limit of the PEC biosensor was determined to be 33 aM (S/N = 3). As shown in Tab. S2, compared with the reported biosensors for miRNA-21 detection, this elaborated PEC biosensor exhibited better analytical performance in terms of sensitivity and linear range. This excellent performance could be ascribed to the effective integration of the enzyme-free nucleic acid dual-amplification strategy and mimic enzyme catalytic precipitation reaction that can cause a significant reduction of photocurrent signal.
A Photocurrent responses of biosensor with incubation of different concentrations of miRNA-21 (100 aM, 1 fM, 10 fM, 100 fM, 1 pM, 10 pM, 100 pM, and 1 nM) with excitation wavelength of 365 nm, switched off–on-off for 10–20-10 s by the applied potential of 0 V (vs. reference electrode). B Calibration curve for miRNA-21 detection with common logarithm (lg) of miRNA-21 concentrations ranging from lg100 aM to lg1 nM
Selectivity and stability of constructed PEC biosensor
Compared to other small molecules (glucose, amino acids) and large molecules (proteins), various miRNAs with base complementary ligands were most likely to act as substances to influence the PEC responses. Thus, under the same experimental conditions, selectivity of the as-prepared PEC biosensor was examined by incubation of the PEC biosensor with different miRNAs. As indicated in Fig. 5A, the treatment of coexisting miRNAs with PEC biosensor exhibited no obvious influence on photocurrent response. Notably, target miRNA-21 and the mixture containing miRNA-21 had a significant quenching effect on photocurrent response, indicating excellent specificity of the designed PEC biosensor toward miRNA-21. Moreover, to evaluate the stability of the biosensor, the PEC response toward miRNA-21 was explored by continuous light processing for 10 cycles in the “off–on-off” mode. As displayed in Fig. 5B, the relative standard deviation (RSD) of 10 cycles was calculated to be 1.8%, which demonstrated good stability of the fabricated biosensor. Moreover, the analysis of this experiment showed that the measured analyte, which irreversibly adsorbed onto the electrode, exhibited excellent sensor stability, so there was no carryover effect upon successive measurements.
Preliminary assessment of the biosensor in cancer cells
Studies have shown that MCF-7 cells and HeLa cells have high and low expression levels of miRNA-21, respectively [18]. In order to assess the feasibility of the constructed biosensor in real samples, the expression level of miRNA-21 in cancer cells was measured by the response of PEC to lysates from HeLa cells and MCF-7 cells. As exhibited in Fig. 6, the photocurrent response showed almost no significant change with the number of HeLa cells increased from 10 to 105. However, the photocurrent response decreased rapidly after incubating the designed biosensor with the same concentration of MCF-7 cell lysates, which resulted from the increase in total content of miRNA-21 from MCF-7 cell lysates as the number of MCF-7 cells increased. The above results were consistent with present research [41, 42], indicating that the proposed PEC biosensor could monitor miRNA expression level in cancer cells.
Conclusions
In this work, based on the combination of enzyme-free nucleic acid dual-amplification strategy and mimic enzyme catalytic precipitation reaction, a unique PEC biosensor using BiOCl-BiOI composites as the photosensitive material was developed for the ultrasensitive miRNA-21 determination. BiOCl-BiOI was employed as a tag to offer the strong initial PEC signal which was effectively quenched by a large number of precipitations induced by the low abundance of miRNA-21. Experimental results demonstrated that the constructed PEC biosensor was ultrasensitive with wide linear range and excellent selectivity and stability. Importantly, the PEC biosensor could also monitor the expression level of miRNA-21 in real samples. This work provides a distinctive platform for quantitative determination of miRNA, which could find broad application prospects in biological analysis and early clinical diagnosis. Noteworthy, this biosensor was not reversible in practical applications because of the single recognition ability and the irreversible adsorption of the electrode of target miRNA-21. The following plan was to optimize its design and construct a superior performance enzyme-free nucleic acid dual-amplification strategy–based miRNA sensing platform.
References
Ma W, Fu P, Sun M, Xu L, Kuang H, Xu C (2017) Dual quantification of microRNAs and telomerase in living cells. J Am Chem Soc 139:11752–11759
Wang H, Tang H, Yang C, Li Y (2019) Selective single molecule nanopore sensing of microRNA using PNA functionalized magnetic core–shell Fe3O4–Au nanoparticles. Anal Chem 91:7965–7970
Calin GA, Croce CM (2006) MicroRNA signatures in human cancers. Nat Rev Cancer 6:857–866
Dong H, Lei J, Ding L, Wen Y, Ju H, Zhang X (2013) microRNA: function, detection, and bioanalysis. Chem Rev 113:6207–6233
Yi J-T, Chen T-T, Huo J, Chu X (2017) Nanoscale zeolitic imidazolate framework-8 for ratiometric fluorescence imaging of microRNA in living cells. Anal Chem 89:12351–12359
Greco F, Inferrera A, La Rocca R, Navarra M, Casciaro M, Grosso G, Gangemi S, Ficarra V, Mirone V (2019) The potential role of microRNAs as biomarkers in benign prostatic hyperplasia: a systematic review and meta-analysis. Eur Urol Focus 5:497–507
Zhao T, Zhang N, Zhang Y, Ren J, Xu P, Liu Z, Cheng L, Hu Y (2017) A novel method to identify pre-microRNA in various species knowledge base on various species. J Biomedical Semantics 8:30
Bejerano T, Etzion S, Elyagon S, Etzion Y, Cohen S (2018) Nanoparticle delivery of miRNA-21 mimic to cardiac macrophages improves myocardial remodeling after myocardial infarction. Nano Lett 18:5885–5891
Li M-X, Zhao W, Wang H, Li X-L, Xu C-H, Chen H-Y, Xu J-J (2018) Dynamic single molecular rulers: toward quantitative detection of microRNA-21 in living cells. Anal Chem 90:14255–14259
Zhang P, Lin Z, Zhuo Y, Yuan R, Chai Y (2017) Dual microRNAs-fueled DNA nanogears: a case of regenerated strategy for multiple electrochemiluminescence detection of microRNAs with single luminophore. Anal Chem 89:1338–1345
Xia Y, Zhang R, Wang Z, Tian J, Chen X (2017) Recent advances in high-performance fluorescent and bioluminescent RNA imaging probes. Chem Soc Rev 46:2824–2843
van Rooij E, Sutherland Lillian B, Qi X, Richardson James A, Hill J, Olson Eric N (2007) Control of stress-dependent cardiac growth and gene expression by a microRNA. Science 316:575–579
Zhao W-W, Xu J-J, Chen H-Y (2014) Photoelectrochemical DNA biosensors. Chem Rev 114:7421–7441
Haddour N, Chauvin J, Gondran C, Cosnier S (2006) Photoelectrochemical immunosensor for label-free detection and quantification of anti-cholera toxin antibody. J Am Chem Soc 128:9693–9698
Wen G, Ju H (2016) Enhanced photoelectrochemical proximity assay for highly selective protein detection in biological matrixes. Anal Chem 88:8339–8345
Hu Q, Han D, Gan S, Bao Y, Niu L (2018) Surface-initiated-reversible-addition–fragmentation-chain-transfer polymerization for electrochemical DNA biosensing. Anal Chem 90:12207–12213
Ge L, Wang W, Li F (2017) Electro-grafted electrode with graphene-oxide-like DNA affinity for ratiometric homogeneous electrochemical biosensing of microRNA. Anal Chem 89:11560–11567
Zhang Y, Chai Y, Wang H, Yuan R (2019) Target-induced 3d DNA network structure as a novel signal amplifier for ultrasensitive electrochemiluminescence detection of microRNAs. Anal Chem 91:14368–14374
Chen P, Wu P, Zhang Y, Chen J, Jiang X, Zheng C, Hou X (2016) Strand displacement-induced enzyme-free amplification for label-free and separation-free ultrasensitive atomic fluorescence spectrometric detection of nucleic acids and proteins. Anal Chem 88:12386–12392
Zhang N, Shi X-M, Guo H-Q, Zhao X-Z, Zhao W-W, Xu J-J, Chen H-Y (2018) Gold nanoparticle couples with entropy-driven toehold-mediated DNA strand displacement reaction on magnetic beads: toward ultrasensitive energy-transfer-based photoelectrochemical detection of miRNA-141 in Real Blood Sample. Anal Chem 90:11892–11898
Xia L-Y, Li M-J, Wang H-J, Yuan R, Chai Y-Q (2020) Novel single-enzyme-assisted dual recycle amplification strategy for sensitive photoelectrochemical microRNA assay. Anal Chem 92:14550–14557
Huang L, Zhang L, Yang L, Yuan R, Yuan Y (2018) Manganese porphyrin decorated on DNA networks as quencher and mimicking enzyme for construction of ultrasensitive photoelectrochemistry Aptasensor. Biosens Bioelectron 104:21–26
Kong W, Xiang M-H, Xia L, Zhang M, Kong R-M, Qu F (2020) In-situ synthesis of 3D Cu2O@Cu-based Mof nanobelt arrays with improved conductivity for sensitive photoelectrochemical detection of vascular endothelial growth factor 165. Biosens Bioelectron 167:112481
Li L, Zhang Y, Yan Z, Chen M, Zhang L, Zhao P, Yu J (2020) Ultrasensitive photoelectrochemical detection of microRNA on paper by combining a cascade nanozyme-engineered biocatalytic precipitation reaction and target-triggerable DNA motor. ACS Sensors 5:1482–1490
Sun L, Xiang L, Zhao X, Jia C-J, Yang J, Jin Z, Cheng X, Fan W (2015) Enhanced visible-light photocatalytic activity of BIOI/BIOCl heterojunctions: key role of crystal facet combination. ACS Catal 5:3540–3551
Su X, Yang J, Yu X, Zhu Y, Zhang Y (2018) In situ grown hierarchical 50%Biocl/Bioi hollow flowerlike microspheres on reduced graphene oxide nanosheets for enhanced visible-light photocatalytic degradation of rhodamine B. Appl Surf Sci 433:502–512
Yi W, Cai R, Xiang D, Wang Y, Zhang M, Ma Q, Cui Y, Bian X (2019) A novel photoelectrochemical strategy based on an integrative photoactive heterojunction nanomaterial and a redox cycling amplification system for ultrasensitive determination of microRNA in cells. Biosens Bioelectron 143:111614
Gao X, Niu S, Ge J, Luan Q, Jie G (2020) 3D DNA nanosphere-based photoelectrochemical biosensor combined with multiple enzyme-free amplification for ultrasensitive detection of cancer biomarkers. Biosens Bioelectron 147:111778
Chu Y, Deng A-P, Wang W, Zhu J-J (2019) Concatenated catalytic hairpin assembly/hyperbranched hybridization chain reaction based enzyme-free signal amplification for the sensitive photoelectrochemical detection of human telomerase RNA. Anal Chem 91:3619–3627
Li H, Wang J, Wang X, Lin H, Li F (2019) Perylene-based photoactive material as a double-stranded DNA intercalating probe for ultrasensitive photoelectrochemical biosensing. ACS Appl Mater Interfaces 11:16958–16964
Yuan Y, Hu T, Zhong X, Zhu M, Chai Y, Yuan R (2020) Highly sensitive photoelectrochemical biosensor based on quantum dots sensitizing Bi2te3 nanosheets and DNA-amplifying strategies. ACS Appl Mater Interfaces 12:22624–22629
Liu P, Yang X, Sun S, Wang Q, Wang K, Huang J, Liu J, He L (2013) Enzyme-free colorimetric detection of DNA by using gold nanoparticles and hybridization chain reaction amplification. Anal Chem 85:7689–7695
Chu Y, Wu R, Fan G-C, Deng A-P, Zhu J-J (2018) Enzyme-free photoelectrochemical biosensor based on the co-sensitization effect coupled with dual cascade toehold-mediated strand displacement amplification for the sensitive detection of microRNA-21. ACS Sustain Chem Eng 6:11633–11641
Li S-K, Liu Z-T, Li J-Y, Chen A-Y, Chai Y-Q, Yuan R, Zhuo Y (2018) Enzyme-free target recycling and double-output amplification system for electrochemiluminescent assay of mucin 1 with MoS2 nanoflowers as co-reaction accelerator. ACS Appl Mater Interfaces 10:14483–14490
Li J, Weng X, Mo F, Han M, Li H (2020) Superparamagnetic nanostructures coupled with an entropy-driven DNA circuit for elegant and robust photoelectrochemical biosensing. Anal Chem 92:15145–15151
Feng Q-M, Guo Y-H, Xu J-J, Chen H-Y (2017) Self-assembled DNA tetrahedral scaffolds for the construction of electrochemiluminescence biosensor with programmable DNA cyclic amplification. ACS Appl Mater Interfaces 9:17637–17644
Long D, Li M, Wang H, Wang H, Chai Y, Li Z, Yuan R (2020) Ultrasensitive photoelectrochemical assay for DNA detection based on a novel SnS2/Co3O4 sensitized structure. Anal Chem 92:14769–14774
Li M, Tian X, Liang W, Yuan R, Chai Y (2018) Ultrasensitive photoelectrochemical assay with PtB7-Th/CdTe quantum dots sensitized structure as signal tag and benzo-4-chlorohexadienone precipitate as efficient quencher. Anal Chem 90:14521–14526
Xie S, Dong Y, Yuan Y, Chai Y, Yuan R (2016) Ultrasensitive lipopolysaccharides detection based on doxorubicin conjugated N-(aminobutyl)-N-(ethylisoluminol) as electrochemiluminescence indicator and self-assembled tetrahedron DNA dendrimers as nanocarriers. Anal Chem 88:5218–5224
Li M, Xiong C, Zheng Y, Liang W, Yuan R, Chai Y (2018) Ultrasensitive photoelectrochemical biosensor based on DNA tetrahedron as nanocarrier for efficient immobilization of CdTe Qds-methylene blue as signal probe with near-zero background noise. Anal Chem 90:8211–8216
Ye S, Li X, Wang M, Tang B (2017) Fluorescence and SERS imaging for the simultaneous absolute quantification of multiple miRNAs in living cells. Anal Chem 89:5124–5130
Si ML, Zhu S, Wu H, Lu Z, Wu F, Mo YY (2007) MIR-21-mediated tumor growth. Oncogene 26:2799–2803
Funding
This work was financially supported by the National Natural Science Foundation of China (22064010, 51862014, 21775089, 22074080), Changjiang Scholar Program of the Ministry of Education of China (Q2019258), and the Taishan Scholar Program of Shandong Province (tsqn201909106).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
604_2022_5345_MOESM1_ESM.docx
Supplementary file1 Materials and reagents, DNA and RNA sequence, optimization of experimental conditions, mechanism of the electron transformation, and additional data. (DOCX 286 KB)
Rights and permissions
About this article
Cite this article
Zhao, Y., Li, X., Xiang, MH. et al. Enzyme-free nucleic acid dual-amplification strategy combined with mimic enzyme catalytic precipitation reaction for the photoelectrochemical detection of microRNA-21. Microchim Acta 189, 249 (2022). https://doi.org/10.1007/s00604-022-05345-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00604-022-05345-y