Abstract
Purpose
Mechanical loading is an important parameter that alters the homeostasis of the intervertebral disc (IVD). Studies have demonstrated the role of compression in altering the cellular metabolism, anabolic and catabolic events of the disc, but little is known how complex loading such as torsion–compression affects the IVD cell metabolism and matrix homeostasis. Studying how the duration of torsion affects disc matrix turnover could provide guidelines to prevent overuse injury to the disc and suggest possible beneficial effect of torsion. The aim of the study was to evaluate the biological response of the IVD to different durations of torsional loading.
Methods
Intact bovine caudal IVD were isolated for organ culture in a bioreactor. Different daily durations of torsion were applied over 7 days at a physiological magnitude (±2°) in combination with 0.2 MPa compression, at a frequency of 1 Hz.
Results
Nucleus pulpous (NP) cell viability and total disc volume decreased with 8 h of torsion–compression per day. Gene expression analysis suggested a down-regulated MMP13 with increased time of torsion. 1 and 4 h per day torsion–compression tended to increase the glycosaminoglycans/hydroxyproline ratio in the NP tissue group.
Conclusions
Our result suggests that load duration thresholds exist in both torsion and compression with an optimal load duration capable of promoting matrix synthesis and overloading can be harmful to disc cells. Future research is required to evaluate the specific mechanisms for these observed effects.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The intervertebral disc (IVD) is subjected to complex mechanical loading, depending on daily activity. With ageing and degeneration, the disc matrix composition is altered, water content is reduced and its ability to distribute loading is compromised. Disc degeneration is a biochemical process characterized partially by cell apoptosis and dedifferentiation and hence altered matrix homeostasis. Loading has been identified as one of the risk factors for disc degeneration, which affects the overall water and oxygen content, fixed-charged density and pH of the disc. In addition, the complex nature of disc loading, including compression, bending, shearing and torsion (twisting) applies variable direct mechanical stimulation to the cells.
It is well explored how static and dynamic compression affect the mechanobiology of disc cells and its relation to disc degeneration and possibly low back pain. In short, dynamic and static compression influence disc cell viability, matrix turnover and water content in a magnitude- and duration-dependent manner [5, 33]. It has also been demonstrated both in vitro and in vivo that static and dynamic loading affect the homeostasis of the disc matrix by the changes in matrix gene expression over different loading durations [8, 12, 21]. For instance, Lotz et al. found a magnitude- and duration-dependent cell death caused by static compression over 1–7 days in an in vivo rat tail model [19]. Maclean et al. [21] and Wuertz et al. [38] found that 4 and 8 h per day dynamic compression induced catabolic event of the IVD. However, how complex loading affects disc matrix homeostasis and its consequences for disc degeneration or low back pain still needs further investigation. Different spinal loadings are associated with variable risk of disc injury [2, 9], for example, repetitive flexion–extension combined with compression increased the susceptibility to disc herniation in a study using porcine spinal specimens [4]. Compression with bending has resulted in annulus fibrosus (AF) delamination and increased cell death therefore increasing the susceptibility to disc degeneration [36]. Torsional loading has been identified as a risk factor for disc injury and low back pain [17, 26]; clinical evidence indicates that cricket and golf players have a higher incidence of disc herniation [10, 16]. Therefore, understanding the mechanobiology of the IVD in response to complex loading has a significant implication for preventing low back injury by defining healthy and harmful spinal loading levels and for developing possible motion-based physiotherapies to relieve back pain [29].
The influence of increasing magnitude of torsion loading has been evaluated in an organ culture model, which revealed a positive effect on the cell viability when dynamic torsion was applied to the IVD at a physiological magnitude (±2° at a frequency of 0.1 Hz) [6]. Nonetheless, an 8 h per day dynamic complex torsion–compression loading induced substantial cell death even at a physiological magnitude and frequency [6]. In this study, it was hypothesized that torsion can be beneficial and harmful to the disc at a physiological magnitude, depending on the duration of the torsion applied. We aimed to investigate the biological response of the disc to a physiological magnitude of torsion, as a function of the duration of applied torsion.
Materials and methods
Bovine IVD isolation and organ culture
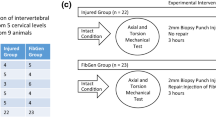
A total of 40 bovine caudal discs with intact vertebral endplates were isolated from six animals, 12–18-month-old cows obtained from a local slaughterhouse, as previously described [6]. Five discs per animal were harvested, one was used for day 0 control, and four were used for culturing under dynamic torsion for 7 days. Disc explants were placed in the centre of the custom-made specimen chambers that contained two serrated, porous titanium plates for securing the disc samples during the load application. The disc samples were provided with 40 mL Dulbecco’s Modified Eagle’s Medium (DMEM, 4.5 g/L glucose) containing 5 % foetal calf serum (FCS), 100 U/mL penicillin, 100 μg/mL streptomycin (all from Gibco, Basel, Switzerland) and cultured at 37 °C and 5 % CO2 for 8 days with media change every 2–3 days.
Mechanical loading
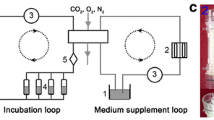
Using a custom-built bioreactor (Fig. 1), dynamic axial rotation of ±2° was applied at a frequency of 1 Hz, superimposed on a static compressive load of 0.2 MPa during the torsion loading period. The compression loading was adjusted to keep a pressure of 0.2 MPa, a secondary low-amplitude dynamic compression resulted along with the dynamic rotation (variation <1 % of nominal load). The changes in load magnitude and torque were recorded during the loading. Figure 2 shows an example of the changes in torque and compression force during the dynamic loading. Discs were assigned to four groups of different durations of torsion with static compression: 0, 1, 4, 8 h/day. After loading for the defined period, discs were allowed to rest in a free-swelling condition (therefore 24, 23, 20 and 16 h/day) before the next loading cycle. Disc diameter (d) and disc height (h) were measured using a digital caliper (Tesa, Renens, Switzerland) before the experiment to calculate the required compression force magnitude, based on disc area and desired pressure. Two diameters were measured on the outer annulus: at the observed largest width of the disc (d 1) and the perpendicular diameter (d 2). The disc surface area (A) was calculated as A = 0.25 × π × d 1 × d 2. Immediately after the final day of loading without resting time, sample dimensions were measured again and disc volume was calculated as V = 0.25 × h × π × d 1 × d 2. For biochemical analysis, NP and AF tissues were cut out using a 6-mm biopsy punch, and were further divided for cell viability, metabolic activity, gene expression and matrix content analysis, n = 6 for each group.
Schematic mechanical design of one loading station. The fluidic muscle moves the support structure, which is mounted on a sledge allowing movements in the x-direction, upwards and therefore compressed the specimen. A contact-free travel sensor records the movements. The servomotor including the gears and the encoder for applying torque on the specimen are also mounted on the support structure and therefore moves upwards with muscle contraction
Cell viability
NP and AF tissues were incubated in DMEM containing 10 µmol/L calcein AM and 1 µmol/L ethidium homodimer-1 (Sigma–Aldrich, Buchs, Switzerland) at 37 °C for 3 h. Stained samples were visualized on an inverted confocal laser-scanning microscope (LSM710, Carl Zeiss, Germany). Stacks were split into single colour channel images, and cell viability was quantified using imageJ software [13].
Metabolic activity
Cellular activity of the disc tissue was measured using resazurin sodium salt (Sigma–Aldrich, Buchs, Switzerland). The dissected tissues were incubated in culture medium with 50 μM resazurin sodium salt for 5 h. Relative fluorescence unit (RFU) was measured at an excitation wavelength of 547 nm and an emission wavelength of 582 nm using a microplate reader (SpectraMax M5, Molecular device, Switzerland). RFU measured for each tissue was normalized with the tissue dry weight.
Real-time polymerase chain reaction
RNA from disc tissues was extracted using TRI reagent (Molecular Research Center, Cincinnati, OH, USA) and purification kit with DNA digestion (Sigma–Aldrich, Buchs, Switzerland). Reverse transcription was performed using the iScript© cDNA synthesis kit (Bio-Rad, Reinach, Switzerland). Primers (Table 1) (Microsynth, Balgach, Switzerland) were newly designed with Beacon Designer™ software (Premier Biosoft, Palo Alto, USA) based on nucleotide sequences from GenBank. Real-time RT-PCR was run on an IQ5 cycler (Bio-Rad, Reinach, Switzerland), followed by melting curve analysis for specific amplification. C t values of the genes were normalized to the reference gene ribosomal 18S RNA (∆C t). ∆∆C t values were calculated by normalizing the loading groups to the no loading control group and then transformed using the 2−∆∆Ct-method [18].
Quantification of GAG, HYP and DNA content
Tissues were dried at 60 °C overnight and then digested with papain overnight at 60 °C. The papain-digested samples were used for glycosaminoglycan (GAG) and DNA measurement. The part of the papain-digested samples was hydrolysed with 6 M HCl at 96 °C for 24 h to release hydroxyproline (HYP). The GAG content was measured by the modified dimethylmethylene blue (DMB) assay (pH 1.5) [11]. HYP content was measured to determine the total collagen content. Colour reaction was prepared using chloramine T and 4-dimethylamino-benzaldehyde (DABA). The amount of DNA in the sample was measured with bisbenzimidol fluorescent dye (Hoechst 33258). Absorbance for GAG and HYP measurement was read at 595 and 560 nm, respectively, DNA fluorescence was read with the same microplate reader (SpectraMax M5, Molecular device, Switzerland). GAG, HYP and DNA concentrations were calculated from a standard curve obtained with chondroitin sulphate, HYP and calf thymus DNA (all chemicals from Sigma–Aldrich, Buchs, Switzerland), respectively.
Statistics
The statistical analysis was performed using PRISM software (GraphPad, La Jolla, USA, version 6.0e). The Friedman test for non-parametric and matched data with Dunn’s multiple comparison post hoc test was used to test for statistically significant differences between groups. A significance value of p < 0.05 was specified for significance.
Result
Cell viability and activity
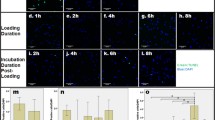
Nucleus pulposus cell (NPC) viability remained above 85 % after torsion–compression loading for 0, 1 or 4 h per day. Cell viability was statistical significantly reduced to below 70 % when torsion was applied for 8 h per day (p = 0.022) (Fig. 3). The daily duration of torsional loading did not affect the AF cell viability (>80 % for all loading durations). No difference in cell activity was noted between groups (Fig. 4).
Percentage cell viability of the disc tissue measured by fluorescence live/dead stain and confocal microscopy. There was a significant cell death in the nucleus pulposus (NP) after 8 h of torsion as compared to 1 h. Cell viability in the annulus fibrosus (AF) maintained stable for all groups. Plot of min to max, midline indicates median, + indicates mean. *p < 0.05. N = 6
Disc volume
A significant decrease in disc volume was observed in the 8 h group (−12.01 ± 8.74 %, p = 0.005) as compared to non-loaded controls (+20.55 ± 8.51 %) (Fig. 5).
Change in disc volume after torsion with static loading. Disc cultured without loading has swelled for about 20 % as compared to day 0, where disc geometry decreased with increasing duration of torsion which reflex the decrease in water content in the disc. Disc volume has reduced 20 % after 8 h of torsion with 0.2 MPa static loading. Plot of min to max, midline indicates median, + indicates mean. **Indicates p < 0.01
Gene expression
To monitor the matrix synthesis and degradation of the disc tissue related to the duration of torsion, a group of genes were monitored by real-time RT-PCR. A trend of collagen 2 and MMP-3 gene up-regulation with an increasing duration of torsion was observed in NP (Fig. 6). MMP-13 gene expression in the NP of the 8 h group was significantly lower than the 1 h group. (p = 0.023) A trend of increased COX-2 gene expression on the NP of the 4 h group than the 1 h group was observed (p = 0.05) (Fig. 6).
Relative gene expression quantified by real-time RT-PCR. Anabolic, catabolic and inflammatory genes were analysed in the NP and AF tissues. ADAMTS-4 (a disintegrin and metalloproteinase with thrombospondin motifs 4), MMP-3 and MMP-13 (matrix metalloproteinase-3, 13), COX-2 (prostaglandin-endoperoxide synthase 2), MCP-1 (monocyte chemotatic protein-1), NGF (nerve growth factor), IL-1b, 8 (interleukin 1b, 8). Plot of mean ± SD. *Statistically significant, p < 0.05. N = 6
DNA, GAG and HYP content
To assess influence of torsion to the disc at protein level, GAG and HYP content of the disc tissue after torsional loading were measured. There was no difference in absolute DNA, GAG and HYP content between groups. GAG/HYP ratio in the AF was not different between all groups, but a trend of higher GAG/HYP ratio in the NP of 1 h (29.57 ± 8.37) and 4 h (27.49 ± 6.95) torsion than 0 h (18.59 ± 2.583) and 8 h (19.81 ± 8.223) torsion was observed (p = 0.06) (Fig. 7).
Discussion
Spinal rotation has been identified as a risk factor for disc injury and low back pain [17, 26]. Long-duration torsion is possible in athletes who undergo extensive daily training, e.g. in tennis and golf. This study demonstrated the adverse effect of extended torsion that 8 h of torsion at physiological magnitude ±2° at a frequency of 1 Hz can cause increased cell death in the NP. This accumulated cell death might be a predisposing risk factor for disc degeneration in the long term. This indicates that different loading imposes different effects on the disc cells, where 8 h of physiological dynamic compression is not harmful but dynamic torsion is. This finding also explains partially the results of the previous study, i.e., an extended period (8 h/day) of complex dynamic torsion–compression loading has caused a substantial cell death in the NP [7].
It was counter-intuitive to find that torsion was more harmful to the NP cells than AF cells. It is assumed that axial rotation imposes a higher strain on the AF cells than NP cells since the NP is closer to the centre of rotation, therefore, the matrix surrounding NP cells experienced less distortion than the outer AF cells [32]. Pure torsion at this magnitude should result in zero disc volume change, thereby not promoting convective transport, in contrast to dynamic compression, which provides a pumping action to enhance solute transport across the disc. In contrast to dynamic compression loading, dynamic torsion–compression caused increased NP cell death due to the lowered glucose levels and increased accumulated lactic acid in the disc, resulting from a slower diffusion rate caused by the decreased tissue porosity following torsion–compression. As analysed in a recent in silico study, the accumulation of lactic acid was found to be the major cause of NP cell death during dynamic torsion–compression loading [31]. In addition, we found a reduced disc height and total disc volume only in the 8 h group. The tight packing of collagen fibrils during dehydration resulted in enhanced mechanical rigidity of the collagen and cells are suspected to experience higher stress and shear under loading within a dehydrated NP matrix.
Load magnitude, frequency, duration as well as loading type affect disc fluid flow and therefore water content and its implication on disc cell viability and metabolism [5, 25]. The result from this study should be interpreted carefully, since different durations of static compression also alter fluid flow and hence disc water content, disc height, nutrients and waste content [23] that can influence cell viability and matrix composition. Duration of static compression has been shown to influence cell viability and matrix gene expression in the AF and to a lesser extent in the NP [5]. Chen and colleagues found an increased AF matrix gene expression after 30 h of static compression, but not in the NP [8, 20]. Wang et al. found that 6 h of static loading caused a reduced collagen and aggrecan gene expression but elevated IL-1b and TNFa gene expression [37]. Paul et al. indicated that, after 7 days of dynamic physiological loading and 16 h of high static loading (0.6 MPa), there was no difference in NP cell viability but AF cell viability was 20 % lower in static loading than dynamic physiological loading [28]. In a rat tail disc in vitro culture model, Ramakrishnan et al. demonstrated that a compression of 0.25 MPa for 12 h per day did not alter the disc cell viability after 7 days [30]. We also found no change in cell viability with a static loading of 0.2 MPa for 8 h per day after 7 days in our pilot study. The drop in NP cell viability after 8 h of torsion–compression was attributed to the combinational effect of dynamic torsion and static compression but not only due to the static compression.
Organ culture models provide a unique platform to bridge from simple cell culture models to in vivo models to study cell behaviour under the native cell microenvironment [12], but one of the limitations of organ culture studies is that the load distribution and the forces that are experienced by the disc cells are unknown. Usually, only the reaction force from the overall loading is recorded. To understand the difference in cell response in the NP and AF during axial rotation in depth, it is important to know the strains that are applied to the NP and AF during axial torsion. Further examination by mechanical experiments and mathematical models can estimate the stress and strain inside the disc. For example, the shear strain of the disc during loading could be measured by using a non-invasive magnetic resonance imaging (MRI) method [27]. In silico studies could also help not only to estimate the magnitude of stress and strain experienced by the cells, but also the distribution of nutrients and waste products in the disc [20, 22, 39].
Although extended torsion could be harmful to the disc, torsion could also be beneficial for matrix production. The duration of the loading stimulus applied to the IVD cells has a significant influence on their matrix production [5, 21]. A trend of increase in GAG to HYP ratio was observed in short torsion duration up to 4 h/day in our study, which may be induced by the mechanical stimulus applied to the disc cells during torsion. Similarly, 4 h/day physiological compression loading also restored the proteoglycan content in the trypsin-treated disc [14]. We did not observe a major change in disc matrix content in our short-term study, possibly a longer term of study is required to observe the difference at protein level.
NP cells experience both hydrostatic and shear stresses that could cause NP cells stretching. Matsumoto et al. [24] found an increased collagen synthesis by NP cells that when a 20 % strain was applied to NP cell monolayer by a Flexcell plate, the authors also suggested a synergistic influence of inflammation (in the presence of IL-1 or TNF) and stretching on the production of prostaglandin E2 (PGE2) by both NP and AF cells. AF cells also react stronger than NP cells if stimulated by stretching in an inflammatory environment. A recent study by Gawri et al. [15] also confirmed an increased synthesis of inflammatory cytokines TNF-α and neurotrophic factors when a 20 % stretching was applied to monolayer NP and AF cells. Although cell stretching is expected to be higher in the AF rather than NP during axial rotation, we found a tenfold increase in COX-2 in the NP after 4 and 8 h torsion, but not in the AF (Fig. 6). COX-2, a key enzyme to regulate PGE2 synthesis, is elevated in the degenerative disc [1], which induces inflammatory cytokines synthesis including interleukin-1 beta (IL-1 beta) and tumour necrosis factor-alpha (TNF-alpha). This implies that extended torsion could possibly induce the inflammatory response of the disc.
Lotz and Chin [19] found that the magnitude and the duration of static compressive force influenced the cell viability synergistically. They found that the percentage of dying cells followed the sum of the logarithmic transformations of compressive force and duration of compressive loading. We did not test both magnitude and duration at the same time, keeping the compression and the torsion magnitudes constant, and thus, we cannot test for combined effects of compression and torsion magnitude together with the duration of the loading stimulus. Instead, previous investigations have shown that magnitude of torsion affected cell viability, matrix turnover and inflammatory response of the IVD. Chan et al. [6] showed that 1 h/day torsion of ±2° increased inner disc cell viability, but the positive effect was lost with torsion >±5°. Barbir et al. [3] compared the magnitude of torsion (±5°, ±15° and ±30°) using an in vivo model and found increased inflammatory response at ±30° torsion. Similar to other types of loading, a synergistic influence on disc cell viability by magnitude and duration of torsion is speculated. It is relevant in future experiments to evaluate the synergistic effect of magnitude and duration of torsion. On the other hand, Sowa et al. [34, 35] suggested that a low-frequency stretching (0.1 Hz) had a beneficial anti-catabolic effect on AF cells after 4 h stretching. However, the authors also agreed that frequency is less important than magnitude and duration in altering the expression of catabolic factors [35]. Barbir et al. [3] performed mechanical testing on the rat tail disc from 0.01 to 10 Hz and suggested that torsion magnitude seemed to be more important than frequency in causing disc damage in their mechanical study, but duration was not tested. Frequency of torsion was fixed at 1 Hz in this study; it is relevant to test in a future study whether low-frequency torsion could provide an anti-catabolic effect and how high-frequency loading can cause damage to the disc from a mechano-biological point of view.
This study has shown that an extended duration of torsion reduces the survival of NP cells within the IVD in organ culture, on the other hand hinting that shorter durations of dynamic torsion could alter the matrix content and hence the proteoglycan/collagen ratio. We suggest that load duration thresholds exist in both torsion and compression with an optimal load duration capable of promoting matrix synthesis, and overloading can be harmful to disc cells. Nevertheless, further experimentation is required to evaluate the specific mechanisms for these observed effects.
References
Alimasi W, Sawaji Y, Endo K, Yorifuji M, Suzuki H, Kosaka T, Shishido T, Yamamoto K (2013) Regulation of nerve growth factor by anti-inflammatory drugs, a steroid, and a selective cyclooxygenase 2 inhibitor in human intervertebral disc cells stimulated with interleukin-1. Spine (Phila Pa 1976) 38(17):1466–1472. doi:10.1097/BRS.0b013e318294edb1
Aultman CD, Drake JD, Callaghan JP, McGill SM (2004) The effect of static torsion on the compressive strength of the spine: an in vitro analysis using a porcine spine model. Spine (Phila Pa 1976) 29(15):E304–E309
Barbir A, Godburn KE, Michalek AJ, Lai A, Monsey RD, Iatridis JC (2010) Effects of torsion on intervertebral disc gene expression and biomechanics, using a rat tail model. Spine (Phila Pa 1976) 36(8):607–614. doi:10.1097/BRS.0b013e3181d9b58b
Callaghan JP, McGill SM (2001) Intervertebral disc herniation: studies on a porcine model exposed to highly repetitive flexion/extension motion with compressive force. Clin Biomech (Bristol, Avon) 16(1):28–37. doi:10.1016/S0268-0033(00)00063-2
Chan SC, Ferguson SJ, Gantenbein-Ritter B (2011) The effects of dynamic loading on the intervertebral disc. Eur Spine J 20(11):1796–1812. doi:10.1007/s00586-011-1827-1
Chan SC, Ferguson SJ, Wuertz K, Gantenbein-Ritter B (2011) Biological response of the intervertebral disc to repetitive short term cyclic torsion. Spine (Phila Pa 1976) 36(24):2021–2030. doi:10.1097/BRS.0b013e318203aea5
Chan SCW, Walser J, Käppeli P, Shamsollahi MJ, Ferguson SJ, Gantenbein-Ritter B (2013) Region specific response of intervertebral disc cells to complex dynamic loading: an organ culture study using a dynamic torsion-compression bioreactor. PLoS One 8(8):e72489. doi:10.1371/journal.pone.0072489
Chen J, Yan W, Setton LA (2004) Static compression induces zonal-specific changes in gene expression for extracellular matrix and cytoskeletal proteins in intervertebral disc cells in vitro. Matrix Biol 22(7):573–583. doi:10.1016/j.matbio.2003.11.008
Drake JD, Aultman CD, McGill SM, Callaghan JP (2005) The influence of static axial torque in combined loading on intervertebral joint failure mechanics using a porcine model. Clin Biomech (Bristol, Avon) 20(10):1038–1045. doi:10.1016/j.clinbiomech.2005.06.007
Elliott B, Khangure M (2002) Disk degeneration and fast bowling in cricket: an intervention study. Med Sci Sports Exerc 34(11):1714–1718. doi:10.1249/01.MSS.0000036863.74140.90
Farndale RW, Buttle DJ, Barrett AJ (1986) Improved quantitation and discrimination of sulphated glycosaminoglycans by use of dimethylmethylene blue. Biochim Biophys Acta 883(2):173–177. doi:10.1016/0304-4165(86)90306-5
Gantenbein B, Illien-Jünger S, Chan SC, Walser J, Haglund L, Ferguson SJ, Iatridis JC, Grad S (2015) Organ culture bioreactors-platforms to study human intervertebral disc degeneration and regenerative therapy. Curr Stem Cell Res Ther 10(4):339. doi:10.2174/1574888X10666150312102948
Gantenbein-Ritter B, Potier E, Zeiter S, van der Werf M, Sprecher CM, Ito K (2008) Accuracy of three techniques to determine cell viability in 3D tissues or scaffolds. Tissue Eng Part C Methods 14(4):353–358. doi:10.1089/ten.tec.2008.0313
Gawri R, Moir J, Ouellet J, Beckman L, Steffen T, Roughley P, Haglund L (2014) Physiological loading can restore the proteoglycan content in a model of early IVD degeneration. PLoS One 9(7):e101233. doi:10.1371/journal.pone.0101233
Gawri R, Rosenzweig DH, Krock E, Ouellet JA, Stone LS, Quinn TM, Haglund L (2014) High mechanical strain of primary intervertebral disc cells promotes secretion of inflammatory factors associated with disc degeneration and pain. Arthritis Res Ther 16(1):R21. doi:10.1186/ar4449
Gluck GS, Bendo JA, Spivak JM (2008) The lumbar spine and low back pain in golf: a literature review of swing biomechanics and injury prevention. Spine J 8(5):778–788. doi:10.1016/j.spinee.2007.07.388
Hangai M, Kaneoka K, Hinotsu S, Shimizu K, Okubo Y, Miyakawa S, Mukai N, Sakane M, Ochiai N (2009) Lumbar intervertebral disk degeneration in athletes. Am J Sports Med 37(1):149–155. doi:10.1177/0363546508323252
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods 25(4):402–408. doi:10.1006/meth.2001.1262
Lotz JC, Chin JR (2000) Intervertebral disc cell death is dependent on the magnitude and duration of spinal loading. Spine (Phila Pa 1976) 25(12):1477–1483
Lotz JC, Colliou OK, Chin JR, Duncan NA, Liebenberg E (1998) Compression-induced degeneration of the intervertebral disc: an in vivo mouse model and finite-element study. Spine (Phila Pa 1976) 23(23):2493–2506
MacLean JJ, Lee CR, Alini M, Iatridis JC (2005) The effects of short-term load duration on anabolic and catabolic gene expression in the rat tail intervertebral disc. J Orthop Res 23(5):1120–1127. doi:10.1016/j.orthres.2005.01.020
Malandrino A, Noailly J, Lacroix D (2011) The effect of sustained compression on oxygen metabolic transport in the intervertebral disc decreases with degenerative changes. PLoS Comput Biol 7(8):e1002112. doi:10.1371/journal.pcbi.1002112
Masuoka K, Michalek AJ, MacLean JJ, Stokes IA, Iatridis JC (2007) Different effects of static versus cyclic compressive loading on rat intervertebral disc height and water loss in vitro. Spine (Phila Pa 1976) 32(18):1974–1979. doi:10.1097/BRS.0b013e318133d591
Matsumoto T, Kawakami M, Kuribayashi K, Takenaka T, Tamaki T (1999) Cyclic mechanical stretch stress increases the growth rate and collagen synthesis of nucleus pulposus cells in vitro. Spine (Phila Pa 1976) 24(4):315–s319
McMillan DW, Garbutt G, Adams MA (1996) Effect of sustained loading on the water content of intervertebral discs: implications for disc metabolism. Ann Rheum Dis 55(12):880–887. doi:10.1136/ard.55.12.880
Mundt DJ, Kelsey JL, Golden AL, Panjabi MM, Pastides H, Berg AT, Sklar J, Hosea T (1993) An epidemiologic study of sports and weight lifting as possible risk factors for herniated lumbar and cervical discs. The Northeast Collaborative Group on Low Back Pain. Am J Sports Med 21(6):854–860
O’Connell GD, Johannessen W, Vresilovic EJ, Elliott DM (2007) Human internal disc strains in axial compression measured noninvasively using magnetic resonance imaging. Spine (Phila Pa 1976) 32(25):2860–2868. doi:10.1097/BRS.0b013e31815b75fb
Paul CP, Schoorl T, Zuiderbaan HA, Zandieh Doulabi B, van der Veen AJ, van de Ven PM, Smit TH, van Royen BJ, Helder MN, Mullender MG (2013) Dynamic and static overloading induce early degenerative processes in caprine lumbar intervertebral discs. PLoS One 8(4):e62411. doi:10.1371/journal.pone.0062411
Rainville J, Hartigan C, Martinez E, Limke J, Jouve C, Finno M (2004) Exercise as a treatment for chronic low back pain. Spine J 4(1):106–115. doi:10.1016/S1529-9430(03)00174-8
Ramakrishnan PS, Hong J, Martin JA, Kurriger GL, Buckwalter JA, Lim TH (2011) Biomechanical disc culture system: feasibility study using rat intervertebral discs. Proc Inst Mech Eng H 225(6):611–620
Reagh J, Vizel S, Wills CR, di Blasi T, Malandrino A, Loeser F, Chan SCW, Gantenbein B, Noaily J (2015) Multi-scale simulation of intervertebral disc biophysics under bioreactor conditions by coupling finite element and agent-based models. Czech Republic, Prague
Setton LA, Chen J (2004) Cell mechanics and mechanobiology in the intervertebral disc. Spine (Phila Pa 1976) 29(23):2710–2723
Setton LA, Chen J (2006) Mechanobiology of the intervertebral disc and relevance to disc degeneration. J Bone Joint Surg Am 88(Suppl 2):52–57. doi:10.2106/JBJS.F.00001
Sowa G, Agarwal S (2008) Cyclic tensile stress exerts a protective effect on intervertebral disc cells. Am J Phys Med Rehabil 87(7):537–544. doi:10.1097/PHM.0b013e31816197ee
Sowa G, Coelho P, Vo N, Bedison R, Chiao A, Davies C, Studer R, Kang J (2011) Determination of annulus fibrosus cell response to tensile strain as a function of duration, magnitude, and frequency. J Orthop Res 29(8):1275–1283. doi:10.1002/jor.21388
Walter BA, Korecki CL, Purmessur D, Roughley PJ, Michalek AJ, Iatridis JC (2011) Complex loading affects intervertebral disc mechanics and biology. Osteoarthr Cartil 19(8):1011–1018. doi:10.1016/j.joca.2011.04.005
Wang DL, Jiang SD, Dai LY (2007) Biologic response of the intervertebral disc to static and dynamic compression in vitro. Spine (Phila Pa 1976) 32(23):2521–2528. doi:10.1097/BRS.0b013e318158cb61
Wuertz K, Godburn K, MacLean JJ, Barbir A, Donnelly JS, Roughley PJ, Alini M, Iatridis JC (2009) In vivo remodeling of intervertebral discs in response to short- and long-term dynamic compression. J Orthop Res 27(9):1235–1242. doi:10.1002/jor.20867
Zhu Q, Jackson AR, Gu WY (2012) Cell viability in intervertebral disc under various nutritional and dynamic loading conditions: 3d finite element analysis. J Biomech 45(16):2769–2777. doi:10.1016/j.jbiomech.2012.08.044
Acknowledgments
This project was supported by funds from the Swiss National Science Foundation (project # 310030_153411), the Orthopedic Department of the Insel University Hospital of Bern and the Lindenhof foundation (project # 14-03-F). We thank Dr Stefan Bauer for his contribution on the digital data processing. The imaging part of this study was performed with the facility of the Microscopy Imaging Center (MIC), University of Bern.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
None.
Rights and permissions
About this article
Cite this article
Chan, S.C.W., Walser, J., Ferguson, S.J. et al. Duration-dependent influence of dynamic torsion on the intervertebral disc: an intact disc organ culture study. Eur Spine J 24, 2402–2410 (2015). https://doi.org/10.1007/s00586-015-4140-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00586-015-4140-6