Abstract
Purpose
The ratio of notochordal (NC) cells to mature nucleus pulposus (MNP) cells in the nucleus pulposus varies with species, age and health. Studies suggest that loss of NC cells is a key component of intervertebral disc degeneration. However, few studies have examined the phenotypes of these two cell populations. Therefore, this study aimed to isolate NC and MNP cells from the same intervertebral disc and study phenotypic differences in extracellular matrix production and cell morphology in 3D culture over 7 days.
Methods
Sequential mechanical dissociation and enzymatic digestion were used to isolate NC cell clusters and single MNP cells from bovine caudal discs. Cells were cultured in alginate beads and subsequently analysed for viability, cytokeratin-8 expression, GAG production and extracellular matrix gene expression.
Results
Mechanical dissociation allowed NC cells to be extracted as intact cell clusters. NC cells represented 8 % of the NP cell population and expressed both cytokeratin-8 and vimentin. MNP cells expressed vimentin, only. Both cells types were viable for 7 days. In addition to morphological differences, NC cells produced up to 30 times more total proteoglycan than MNP cells. NC cells had significantly higher aggrecan and brachyury expression.
Conclusions
NC and MNP cells can be isolated from the same bovine disc and maintain their distinct phenotypes in 3D culture.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The mammalian spinal column develops from the embryonic notochord and its surrounding somites [17, 21]. The nucleus pulposus (NP) contains two distinct populations of cells, notochordal (NC) and chondrocyte-like mature nucleus pulposus (MNP) cells. NC cells are the remnants of the embryonic notochord; these large vacuolated cells form dense clusters/islands within the NP matrix [10, 11, 21, 22]. In some animals such as rats and mice, the NP is composed of exclusively notochordal cells [17]. However, in animals such as cattle, and in humans, the NC cell population gradually decreases throughout life [25]. MNP cells are small, chondrocyte-like cells that are sparsely distributed throughout the NP. The origin of MNP cells is currently contentious; histological studies have suggested that they are of mesenchymal origin and infiltrate the NP from the surrounding annulus and/or the cartilaginous endplates [14]. The alternative hypothesis is that these cells are derived from the NC cell population [17]. A review by Rodrigues-Pinto et al. [26] provides an in-depth discussion around this debate.
Species that retain a full complement of NC cells throughout life maintain a proteoglycan-rich NP and generally do not develop disc degeneration [8]. Conversely, species that show a reduction in the NC cell population with age show a gradual loss of proteoglycan within the NP becoming more fibrous and prone to structural failure [6, 8, 25]. This correlation suggests that the reduction in NC cells with age is the underlying cause of disc degeneration. However, the exact role that each of these cell types play in both the physiology and pathology of the intervertebral disc is poorly understood.
Until recently, there has been a lack of specific markers for NC cells. Instead cells are classified by morphology/size alone (e.g. flow cytometry) or on species assumptions (e.g. non-chondrodystrophoid dogs) [3, 23]. Recent proteomic and transcriptional studies identified cytokeratin-8 (CK8) as the most robust NC-specific marker [7, 18], suggesting that NC cells are of non-mesenchymal lineage [28]. Healthy human adult discs have been shown to have up to 50 % CK8-positive cells, with the amount being inversely proportional to disc degeneration grade [29, 30]. These findings show that NC cells persist into adulthood, but it is still unclear whether NC loss is a cause or consequence of disc degeneration.
The morphological differences between NC and MNP cells in situ have been well described [10–12]. However, the few studies that have examined NC and MNP cell function and gene expression profiles have used cells from different species, e.g. porcine NC cells and bovine MNP cells [3, 6, 9, 23]. Furthermore, flow cytometry and 2D culture methods drastically alter the microenvironment and architecture of the cells, a caveat when studying substrate-dependant connective tissue cells [4].
The hypothesis that NC cells are responsible for proteoglycan-rich discs still remains difficult to prove. The challenge arises from the variability between species and age of the animals, as well as the varying culture conditions used between studies (10, 20, 21). We believe that to understand the function of these cells, NC cells should be compared to MNP cells from the same animal and examined in their native cluster formation. Therefore, in this study, we present a procedure that isolates NC cells as intact clusters together with individual MNP cells from the same disc, and a characterisation of their phenotypes.
Materials and methods
Culture medium
Low glucose (5 mM) DMEM (Sigma-Aldrich, Auckland, New Zealand) supplemented with 1 % penicillin/streptomycin (Life Technologies, Auckland New Zealand), 0.5 % fungizone (Life Technologies), 85 µM ascorbate (Sigma-Aldrich), 1.85 g/L NaHCO3 (Sigma-Aldrich), and 10 % foetal bovine serum (Sigma-Aldrich). To give a final osmolality of 400 mOsm, 19 g/L mannitol (Sigma-Aldrich) was added; finally pH was adjusted to 7.2 with HCL.
Extraction of isolated nucleus pulposus cell populations
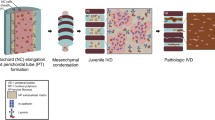
To determine the number of NC and MNP cells in the bovine nucleus pulposus, cells were isolated and separated using a combination of mechanical dissociation and enzymatic digestion. The schematic in Fig. 1 shows the procedures used to generate three different groups: full NP cell population, NC-rich cell fraction and MNP-rich cell fraction. Single-cell suspensions of each group were assessed for DNA and cytokeratin-8 expression.
Schematic illustrating the mechanical dissociation and enzymatic digestion procedures to generate three different populations from ccy4-5 and ccy5-6 bovine discs; total Nucleus Pulposus (NP), Notochordal (NC) and Mature Nucleus Pulposus (MNP) cells. A total of 16 discs were examined from a total of 8 separate animals
Bovine tails were obtained from a local abattoir and the two most proximal discs (ccy4-5 and ccy5-6) were removed. Nucleus pulposus tissue was separated from the surrounding annulus and finely minced. The tissue was enzymatically digested in pronase solution (8 units/mL; Sigma-Aldrich) for 45 min, then in collagenase (245 units/mL; Life Technologies) for 2.5 h at 37 °C to generate a single-cell suspension of the total NP cell population (n = 8 discs from 4 animals). To isolate NC and MNP cell-rich fractions, at total of eight NPs from four animals were finely minced and mechanically dissociated by manually triturating the tissue in a large volume of media to separate the NC cell clusters from the surrounding tissue. The mixture was then strained through a stainless steel sieve (pore size 0.5 mm) to separate liquid from undigested tissue. To allow the NC cells to be easily labelled and counted, the strained medium was enzymatically digested as above to yield a single-cell suspension. The remaining tissue was enzymatically digested to yield a single-cell suspension of MNP cells. The three preparations of isolated cells were each washed, centrifuged at 2,000g for 5 min and resuspended in 1 mL of medium.
Cell yield by DNA assay
Aliquots of cell suspension were mixed 1:5 with papain digest buffer (PBS, 150 mM NaCl, 55 mM sodium citrate, 5 mM EDTA, 5 mM cysteine hydrochloride and 0.6units/ml papain) and digested overnight at 60 °C. DNA was extracted by isopropanol precipitation and quantified using a SYBR green (Life Technologies) microplate assay [16, 20]. Calf thymus DNA (Sigma-Aldrich) was used to generate a standard curve. Given one bovine cell contains 7 pg DNA [15]; the number of cells per sample was calculated.
To determine fraction purity, cells were first seeded in alginate beads for immunocytochemistry. An equal volume of cell suspension was mixed with 2.5 % w/v alginate solution and expelled drop-wise through a 22G needle into a 100 mM CaCl2 solution. Beads were polymerised for 10 min at 37 °C, washed twice in serum-free medium and incubated overnight at 37 °C, 5 % CO2 to equilibrate.
Fluorescent immunocytochemistry
Following overnight culture, alginate beads were cross-linked in 100 mM BaCl2 for 10 min and fixed in 4 %w/v paraformaldehyde for 20 min at 37 °C. Beads were permeabilized in 0.5 % v/v Triton (Sigma Aldrich) for 10 min, washed in PBS containing 0.1 % (w/v) bovine serum albumin and 10 mM CaCl2, blocked in 5 % donkey serum (Sigma Aldrich) for 30 min and incubated in goat polyclonal vimentin primary antibody (1:200; Santa Cruz Biotechnology Inc) overnight at 4 °C. The following day beads were washed, incubated at room temperature for 2.5 h in donkey anti-goat Alexa-488 (1:500; Life Technologies), washed again, blocked in 5 % goat serum (Sigma Aldrich) and incubated with mouse monoclonal cytokeratin-8 primary antibody (1:200; Acris Germany). The following day beads were washed and incubated in goat anti-mouse Alexa-594. Finally beads were counterstained with Hoechst (1:500; Sigma-Aldrich) for 15 min, washed and mounted by firmly pressing onto a slide under a coverslip with Prolong Gold (Life Technologies). A minimum of 200 cells per condition were imaged using a Leica DMR fluorescence microscope.
Histology and immunohistochemistry on intact bovine IVD
Whole bovine caudal IVDs were fresh-frozen in liquid nitrogen and cryo-sectioned to give 14 µm transverse sections. The sections were stained with haematoxylin and eosin or fluorescently labelled using antibodies against cytokeratin-8 and vimentin.
Phenotypic characterisation of NC cell clusters and MNP cells in culture over 7 days
In a separate set of experiments, NP tissue from IVDs was removed from tails as described above. The tissue was mechanically dissociated and strained to isolate intact NC cell clusters. Note, NC cell clusters were not enzymatically digested. The remaining tissue was incubated in pronase solution (8 units/mL; Sigma-Aldrich,) for 45 min, then in collagenase (245 units/mL; Life Technologies) for 2.5 h at 37 °C to isolate MNP cells. For each time point (day 1 and day 7) within an individual experiment, a total of six discs from three tails were examined and cell fractions from the six discs were pooled. The experiment was conducted a total of three times (N = 3). The MNP cell suspension volume was adjusted to give a concentration of 8 × 106 cells/ml. The NC fraction was then resuspended in the same volume to maintain in situ cell ratios. Cell fractions were mixed with an equal volume of 2.5 %w/v alginate solution, formed into beads and cultured for 1 and 7 days in 5 % CO2 at 37 °C in DMEM plus 10 % FBS. On days 1 and 7, beads were assayed for cell viability, DNA and GAG content and gene expression.
Cell viability
Cell viability was assessed using an Apoptotic, Necrotic and Healthy Cells Quantification Kit (Huntingtree Bioscience Supplies). Beads were labelled with FITC-Annexin antibody, Ethidium homodimer and Hoechst according to manufacture instructions. Beads were firmly pressed down onto a microscope slide under a coverslip and imaged immediately on a Nikon TE2000 widefield fluorescence microscope using a 20× objective. Cells from three fields of view per group were examined.
DNA and GAG quantification
Two beads per condition were digested in 1 mL of papain buffer at 60 °C overnight. The 1,9-dimethylmethylene blue (DMMB) dye binding assay was used to measure total sulphated GAG content [5]. Bovine chondroitin sulphate (Sigma Aldrich) was used to generate a standard curve. Since alginate is a polyanionic polysaccharide that can react with DMMB, digests of cell-free beads were used for background subtraction. GAG content within each sample was normalised to DNA content, data expressed as μg GAG/ng DNA per µL of digested sample.
Quantitative RT-PCR
At each time point, ~25 beads per group were placed in 500 μL RNA Later (Life Technologies) and stored at −20 °C. To extract RNA, beads were dissolved in ice-cold buffer containing 55 mM sodium citrate, 30 mM EDTA and 150 mM NaCl, mixed by inversion for 10 min at 4 °C then centrifuged at 2000 g for 10 min. The resultant cell pellet was washed in ice-cold PBS, homogenised in 1 mL Trizol using a 22G needle and frozen overnight at −80 °C, to ensure cell lysis. RNA was isolated using chloroform extraction and isopropanol precipitation, according to the manufacturer’s instructions (Life Technologies). DNAse treatment was performed using “Turbo DNAse kit” (Life Technologies) followed by reverse transcription and cDNA synthesis using Superscript III kit (Life Technologies).
Gene expression levels of notochordal marker, brachyury; anabolic matrix proteins, type II collagen and aggrecan; and catabolic matrix enzymes, matrix metalloprotease-3 and ADAMTS5 were assessed using Taqman gene expression assays (Life Technologies; Table 1). The tyrosine activation protein, YWHAZ, and ribosomal 18S were used as reference genes. Expression was calculated relative to mean expression of r18S and YWHAZ in each sample. If gene expression was not detected, the Ct value was set to 40 to conduct statistical analysis.
Statistics
All data were expressed as mean and standard deviation (SD) from a minimum of three biological repeats. All molecular assays were conducted with ≥2 technical repeats. Statistical significance within and between cell types was determined using two tailed t-tests with Holm-sidak post hoc correction and a p value of <0.05 was considered significant.
Results
Isolation of nucleus pulposus cell sub-populations
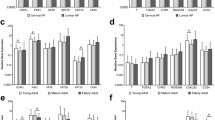
The mean number of nucleus pulposus cells per disc was 684 × 103 (Fig. 2a). The MNP cell-rich fraction yielded 597 × 103 per disc and the NC cell-rich fraction yielded 40 × 103 NC cells per disc. This gives a ratio of 15 MNP to 1 NC cell (approximately 7 % of cells were notochordal) (Fig. 2a). This proportion was confirmed using immunocytochemistry, where 9 ± 2 % of the total NP cells expressed the notochordal marker, cytokeratin-8 (Fig. 2b). The purity of the separated cell fractions was verified using this approach, which showed that 3 ± 1.9 % of cells in the MNP-rich fraction and 89 ± 1.4 % were cytokeratin-8 positive (Fig. 2b).
Nucleus pulposus cell sub-populations. a Cell yield: number of cells per disc for each of the cell fractions; total NP, MNP and NC. b Percentage cytokeratin 8 (CK8)-positive cells in each fraction. Each bar represents the mean ± standard deviation from eight discs acquired from four separate animals
Figure 3 shows that in situ CK8 expressing NC cells were located in dense clusters/islands of cells that appeared loosely attached to the NP matrix. Mechanical extraction and 3D culture preserved the cluster arrangement in vitro (Fig. 4). Individual NC cells had an elongated fibroblast-like morphology but were encapsulated in a gel-like matrix (arrow; Fig. 4) and expressed both vimentin and CK8. In contrast, in situ MNP cells were singular cells embedded in NP matrix and expressed only vimentin (Fig. 3). In vitro, they showed a spherical morphology with no visible pericellular matrix, and did not express CK8 (Fig. 4).
Histology and immunohistochemistry on sequential sections of a bovine nucleus pulposus (NP). a, b H&E stained NP tissue showing islands of NC cells under ×50 and ×100 magnification. c, d Immunohistochemical labelling against CK8 (red) and vimentin (green), showing the CK8 expressing NC cells in situ
Immunocytochemistry on primary MNP and NC cells in vitro (day 1). Cells were labelled against both vimentin (green) and CK8 (red) and counterstained with Hoechst (blue). Phase contrast images show the morphology of each cell type in vitro is similar to observed in situ. Arrow and dotted line indicates pericellular material. Scale 20 µm
Viability
Both MNP and NC cell-rich fractions showed ≥80 % viability for up to 7 days (Fig. 5a). There was no significant difference between MNP and NC cell viability at either time point or within cell type. Cell death was mainly due to necrosis with no differences across cell types (data not shown).
a Percentage viable cells within alginate beads after 1 and 7 days in culture. Cell viability was determined by live cell staining with ethidium homodimer and Hoechst. b DNA content expressed as cells per bead. Data shown represents the mean ± standard deviation of three experiments, with a total of six discs from three animals per timepoint
DNA content
MNP cells were seeded at 4 × 106 cells/ml, with an approximate bead volume of 10 µL. Figure 5b shows that MNP beads contained 41,000 and 45,000 cells on day 1 and 7, respectively. NC beads contained 3,600 and 3,500 cells, respectively, indicating no significant change in DNA content over time for either cell type.
Real-time PCR
NC cells expressed brachury (T) at significantly greater levels than MNP cells on both days (p = 0.015 and p = 0.028, Fig. 6a). Both MNP and NC cells expressed high levels of type II collagen and aggrecan. On day 1, collagen II expression was approximately equal for MNP and NC cells. There was a non-significant increase in collagen II expression on day 7 for MNP cells (Fig. 6b). In contrast, NC cells expressed significantly more aggrecan than MNP cells on day 1 (p = 0.0036). However, by day 7, aggrecan expression in NC cells was dramatically reduced (p = 0.014); while MNP aggrecan expression increased, leaving no measurable difference between cell types (Fig. 6c). MMP3 and ADAMTS5 were expressed at very low levels in both cell types (Fig. 6d, e). However, there was a significant increase in MMP3 expression at day 7 in MNP cells (p = 0.008).
Relative gene expression of a T (brachyury), b Col2a1 (collagen type II), c ACAN (aggrecan), d matrix metalloprotease 3 (MMP3), and e ADAMTS5. Gene expression normalised to averaged expression of the reference genes r18 s and YWHAZ. Data shown represents the mean ± standard deviation of three experiments, with a total of six discs from three animals were at each timepoint. *p ≤ 0.05, **p ≤ 0.01, ***p ≤ 0.001. Hash within cell type, asterisk between cell types
GAG
Figure 7 shows that NC cell cultures had significantly higher GAG/DNA ratios than MNP cell-rich cultures on day 1 (p = 0.009) and day 7 (p = 0.0126), respectively. There was an increase in GAG content for NC beads over time (p = 0.042), but no change in MNP GAG levels over the culture period.
GAG content in MNP and NC cell-rich beads after 1 and 7 days in culture. GAG content (µg) was normalised to DNA content (ng) for each sample. Data shown represents the mean ± standard deviation of three experiments, with a total of six discs from three animals were at each timepoint. **p ≤ 0.01. Hash within cell type, asterisk between cell types
Immunocytochemistry
Both MNPs and NCs maintained their distinctive morphologies over 7 days (Figs. 4, 8). MNP cells remained as singular spherical cells and expressed vimentin (green) but not CK8. NC cells in clusters had an elongated morphology (white arrow) and expressed both vimentin and CK8. Pericellular material was often present surrounding the cell clusters (Fig. 4, dashed line).
Cell morphology and phenotype on day 7. Upper panels show the phase contrast images of MNP and NC cells in vitro. Lower panels show MNP and NC cells double labelled against vimentin (green) and cytokeratin 8 (red) and counter stained with Hoechst (blue). Scale bar 20 µm, arrow indicates the fibroblast-like shape of the NC cells
Discussion
The phenotypic differences between NC and MNP cells have to date been poorly understood, with many studies giving conflicting results. Aguiar et al. [1] compared cells from bovine and canine discs. The paper shows a gelatinous canine NP, and a fibrous bovine NP. However, they found that MNPs from bovine tissue produced higher levels of proteoglycan than the canine NC cells. In contrast, work by Cappello et al. [2] found cells from non-chondrodystrophoid dogs (NC cell rich) to produce a more proteoglycan-rich matrix compared to cells from chondrodystrophoid dogs (MNP cell rich). Furthermore Miyazaki et al. [19] found that rat NC cells produced more proteoglycan per cell than rabbit NC cells in vitro, even though both cells were isolated from a gelatinous NC cell-rich NP tissue. Such studies illustrate that the critical importance of studying cells from the same species.
Studies by Chen et al. and Kim et al., examined isolated MNP and NC cells from the same tissue but were hindered by a historical lack of NC markers to confirm cell populations [3, 13]. These studies also dramatically altered the micro-environment of the cells by digesting the clusters before phenotypic characterisation. Given that the micro-environment of the NC cells has been shown to be a major determinant of cell behaviour [24], we aimed to examine the phenotypes of NC and MNP cells from the same species, to maintain the NC cells as native clusters and to culture cells in the ratio they are found in situ.
We first mechanically dissociated the nucleus pulposus tissue, which allowed “islands” of NC cells to be released into culture medium, whereas MNP cells required enzymatic digestion to be released from the remaining fibrous matrix. Following separation, we were able to assess cell yield per disc and the relative purity of each cell fraction. In agreement with Stosiek et al. and Gilson et al. [7, 28], we found that NC cells expressed both CK8 and vimentin, while MNP cells expressed vimentin only. Additionally, confirming the work by Gilson et al. [7], we found that CK8-expressing NC cells account for 7–9 % of the total bovine NP cell population.
Phenotypic differences between these cell types were maintained in vitro for 7 days. Expression of the notochordal marker brachyury was significantly greater in NC cells compared to MNPs throughout the culture period. Immunohistochemical expression of CK8 was also maintained within the NC clusters for the 7 days, suggesting that this approach is suitable for in vitro culture and for the phenotypic stability of NC cells.
Using cells from the same disc ensures that cells are at the same age and have similar mechanical/loading histories. We found that NC cells by themselves produce 20 times more GAG and have 4-fold higher aggrecan expression levels compared to MNP cells on day 1, showing a clear relationship between NC phenotype and proteoglycan-rich tissue. On day 7, the cumulative GAG content had increased yet the aggrecan expression was reduced. This was also seen by Spillekom et al. [27], who found a steady accumulation of GAG over 28 days, despite decreasing aggrecan expression levels. This study also found that brachyury gene expression decreased initially but returned to day 1 levels with long-term culture [27]. Thus, it is likely that the slight reduction in gene expression levels in this study is a response to culture conditions and not a true phenotypic shift.
Despite the different cell seeding densities, both cell types were viable over the culture period and there was no change in cell number (DNA content), showing that the extraction and culture environment does not induce cell death or changes in cell growth rate, making this model a closer approximation of in situ cell senescence.
Human studies show that NC cells persist into adulthood, and thus should not be ignored when studying IVD pathology [29, 30]. This study shows that even in an animal that is considered chondrodystrophic (MNP cell rich), NC cells are present and viable, and produce a more proteoglycan-rich matrix compared to their MNP cell neighbours. Therefore, this study shows that our extraction and culture methods are a suitable model to improve our understanding of the genetic and molecular signature of healthy NC cells and their suitability for future regenerative cell replacement therapy.
Conclusion
This study has shown that NC cells can be isolated as intact clusters from bovine caudal discs. The NC cells represented ~8 % of the total bovine NP cell population. The cell separation procedure detailed here maintained the NC cluster arrangement in vitro. Over the 7 day culture period, both cell types were over 80 % viable. In addition to being morphological different, NC cells were distinguishable by their expression of cytokeratin-8 and brachyury, as well significantly higher proteoglycan levels. The model described here provides a platform for studies into the ontogeny and functional differences between these two cell types.
References
Aguiar DJ, Johnson SL, Oegema TR (1999) Notochordal cells interact with nucleus pulposus cells: regulation of proteoglycan synthesis. Exp Cell Res 246:129–137
Cappello R, Bird JL, Pfeiffer D, Bayliss MT, Dudhia J (2006) Notochordal cell produce and assemble extracellular matrix in a distinct manner, which may be responsible for the maintenance of healthy nucleus pulposus. Spine (Phila Pa 1976) 31:873–882 discussion 883
Chen J, Yan W, Setton LA (2006) Molecular phenotypes of notochordal cells purified from immature nucleus pulposus. Eur Spine J 15(Suppl 3):S303–S311
Chou AI, Reza AT, Nicoll SB (2008) Distinct intervertebral disc cell populations adopt similar phenotypes in three-dimensional culture. Tissue Eng Part A 14:2079–2087
Farndale RW, Sayers CA, Barrett AJ (1982) A direct spectrophotometric microassay for sulfated glycosaminoglycans in cartilage cultures. Connect Tissue Res 9:247–248
Gantenbein-Ritter B, Chan SC (2011) The evolutionary importance of cell ratio between notochordal and nucleus pulposus cells: an experimental 3-D co-culture study. Eur Spine J 21:S819–S825
Gilson A, Dreger M, Urban JP (2010) Differential expression level of cytokeratin 8 in cells of the bovine nucleus pulposus complicates the search for specific intervertebral disc cell markers. Arthritis Res Ther 12:R24
Gruber HE, Hanley Jr EN (2002) Ultrastructure of the human intervertebral disc during aging and degeneration: comparison of surgical and control specimens. Spine (Phila Pa 1976) 27:798–805
Guehring T, Wilde G, Sumner M, Grunhagen T, Karney GB, Tirlapur UK, Urban JP (2009) Notochordal intervertebral disc cells: sensitivity to nutrient deprivation. Arthritis Rheum 60:1026–1034
Hunter CJ, Matyas JR, Duncan NA (2003) The three-dimensional architecture of the notochordal nucleus pulposus: novel observations on cell structures in the canine intervertebral disc. J Anat 202:279–291
Hunter CJ, Matyas JR, Duncan NA (2004) Cytomorphology of notochordal and chondrocytic cells from the nucleus pulposus: a species comparison. J Anat 205:357–362
Hunter CJ, Matyas JR, Duncan NA (2004) The functional significance of cell clusters in the notochordal nucleus pulposus: survival and signaling in the canine intervertebral disc. Spine (Phila Pa 1976) 29:1099–1104
Kim JH, Deasy BM, Seo HY, Studer RK, Vo NV, Georgescu HI, Sowa GA, Kang JD (2009) Differentiation of intervertebral notochordal cells through live automated cell imaging system in vitro. Spine (Phila Pa 1976) 34:2486–2493
Kim KW, Lim TH, Kim JG, Jeong ST, Masuda K, An HS (2003) The origin of chondrocytes in the nucleus pulposus and histologic findings associated with the transition of a notochordal nucleus pulposus to a fibrocartilaginous nucleus pulposus in intact rabbit intervertebral discs. Spine (Phila Pa 1976) 28:982–990
Kim YJ, Sah RL, Doong JY, Grodzinsky AJ (1988) Fluorometric assay of DNA in cartilage explants using Hoechst 33258. Anal Biochem 174:168–176
Leggate J, Allain R, Isaac L, Blais BW (2006) Microplate fluorescence assay for the quantification of double stranded DNA using SYBR Green I dye. Biotechnol Lett 28:1587–1594
McCann MR, Tamplin OJ, Rossant J, Seguin CA (2012) Tracing notochord-derived cells using a Noto-cre mouse: implications for intervertebral disc development. Dis Model Mech 5:73–82
Minogue BM, Richardson SM, Zeef LA, Freemont AJ, Hoyland JA (2010) Transcriptional profiling of bovine intervertebral disc cells: implications for identification of normal and degenerate human intervertebral disc cell phenotypes. Arthritis Res Ther 12:R22
Miyazaki T, Kobayashi S, Takeno K, Meir A, Urban J, Baba H (2009) A phenotypic comparison of proteoglycan production of intervertebral disc cells isolated from rats, rabbits, and bovine tails; which animal model is most suitable to study tissue engineering and biological repair of human disc disorders? Tissue Eng Part A 15:3835–3846
Myers MA (1998) Direct measurement of cell numbers in microtitre plate cultures using the fluorescent dye SYBR green I. J Immunol Methods 212:99–103
Peacock A (1951) Observations on the prenatal development of the intervertebral disc in man. J Anat 85:260–274
Peacock A (1952) Observations on the postnatal structure of the intervertebral disc in man. J Anat 86:162–179
Potier E, Ito K (2014) Using notochordal cells of developmental origin to stimulate nucleus pulposus cells and bone marrow stromal cells for intervertebral disc regeneration. Eur Spine J 23:679–688
Rastogi A, Thakore P, Leung A, Benavides M, Machado M, Morschauser MA, Hsieh AH (2009) Environmental regulation of notochordal gene expression in nucleus pulposus cells. J Cell Physiol 220:698–705
Roberts S, Evans H, Trivedi J, Menage J (2006) Histology and pathology of the human intervertebral disc. J Bone Joint Surg Am 88(Suppl 2):10–14
Rodrigues-Pinto R, Richardson SM, Hoyland JA (2014) An understanding of intervertebral disc development, maturation and cell phenotype provides clues to direct cell-based tissue regeneration therapies for disc degeneration. Eur Spine J 23:1803–1814
Spillekom S, Smolders LA, Grinwis GC, Arkesteijn IT, Ito K, Meij BP, Tryfonidou MA (2014) Increased osmolarity and cell clustering preserve canine notochordal cell phenotype in culture. Tissue Eng Part C Methods 20:652–662
Stosiek P, Kasper M, Karsten U (1988) Expression of cytokeratin and vimentin in nucleus pulposus cells. Differentiation 39:78–81
Sun Z, Wang HQ, Liu ZH, Chang L, Chen YF, Zhang YZ, Zhang WL, Gao Y, Wan ZY, Che L, Liu X, Samartzis D, Luo ZJ (2013) Down-regulated CK8 expression in human intervertebral disc degeneration. Int J Med Sci 10:948–956
Weiler C, Nerlich AG, Schaaf R, Bachmeier BE, Wuertz K, Boos N (2010) Immunohistochemical identification of notochordal markers in cells in the aging human lumbar intervertebral disc. Eur Spine J 19:1761–1770
Acknowledgments
This work was supported by an Auckland Medical Research Foundation Doctoral Scholarship (TS) and the School of Medical Sciences, Faculty of Medical and Health Sciences, University of Auckland. Thanks to Satya Amirapu for histological processing, the Biomedical Imaging Research Unit, University of Auckland for use of their facilities and Dr. Kelly Wade for critical review of the manuscript.
Conflict of interest
None.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Saggese, T., Redey, P. & McGlashan, S.R. Same-species phenotypic comparison of notochordal and mature nucleus pulposus cells. Eur Spine J 24, 1976–1985 (2015). https://doi.org/10.1007/s00586-014-3697-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00586-014-3697-9