Abstract
The early diagnosis of canine leptospirosis is still a challenge for clinicians that in most cases only have basic routine tests as diagnostic tools. This study aimed to isolate and characterize leptospires from the blood of dogs with clinical signs suggestive of leptospirosis and to evaluate the results of some laboratory tests in hamsters experimentally infected with these isolates. The blood of 22 animals was submitted to bacteriological culturing, and the isolates recovered were characterized by serogrouping, partial DNA sequencing (secY), and multiple locus variable-number tandem repeat analysis (MLVA). Golden Syrian hamsters were infected with isolated leptospires and submitted to serology using microscopic agglutination test (MAT), polymerase chain reaction (PCR), complete blood counts, and serum biochemistry. Three isolates (13.63%) were obtained, and all were assigned to the serogroup Icterohaemorrhagiae. The DNA sequencing results showed that all the isolates belonged to the species Leptospira interrogans, and the MLVA results evidenced that the profile of the strains was compatible with Copenhageni/Icterohaemorrhagiae serovars. Regarding laboratory tests performed on hamsters infected with these isolated strains, marked leukocytosis by neutrophilia and monocytosis was observed, associated with an accentuated increase in renal and hepatic indicators. In addition, a titer of 800 was detected by MAT for only the serogroup Icterohaemorrhagiae, and blood, liver, kidneys, and lung samples were PCR positive. The presence of these isolates suggests that the dogs were infected with strains carried by rodents, and significant changes were found in routine laboratory tests, which, when correlated, can help veterinarians with early diagnosis.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Leptospirosis in dogs is a globally reported disease and is characterized by important morbidity and mortality in these animals (Klassen and Adler 2015). Furthermore, as it is caused by zoonotic organism, it can be transmitted to humans, especially when humans come in contact with the contaminated urine of asymptomatic dogs (Miotto et al. 2018a).
Several clinical symptoms have been reported in dogs diagnosed with leptospirosis, such as anorexia, vomiting, diarrhea, jaundice, dehydration, red to brownish urine, and fever, which occur regardless of the infecting Leptospira serogroup (Geisen et al. 2007). However, as the clinical signs are not disease-specific, the diagnosis of leptospirosis is not simple and should be differentiated from other clinical syndromes involving kidney, liver, lung, and gastrointestinal injuries (Shuller et al. 2015).
One of the most robust and cheapest diagnostic methods for leptospirosis is the microscopic agglutination test (MAT), which is still a reality in many laboratories worldwide (Guedes et al. 2021a). The great advantage of this test is that it can predict the most likely infecting serogroup of Leptospira, as well as the magnitude of the titers, especially the expressive increase when paired samples are performed at an ideal interval of 10 to 14 days (Levett 2001). Other methods, such as enzyme-linked immunosorbent assay (ELISA) (Ambily et al. 2019) and indirect fluorescent antibody tests (IFA), are also reported for the serological diagnosis of canine leptospirosis; nonetheless, they may not be as sensitive or specific as MAT (Silva and Riedemann 2007).
Isolation of Leptospira from dogs has low sensitivity (Silva and Riedemann 2007). Associated with this, prolonged incubation time and culture contamination are limiting factors in bacteriological methods (Levett 2001; Faine et al. 1999). Despite that, obtaining leptospires is still essential for many studies, whether for epidemiological investigation (Miotto et al. 2016; Guedes et al. 2021c) or to assess the pathophysiological behavior of the disease agent through experimental infection in laboratory animals (Gomes-Solecki 2017). Importantly, isolates are also necessary for vaccine production (Zarantonelli et al. 2018).
In this way, the detection of bacterial DNA from clinical samples by polymerase chain reaction (PCR) has been used as an essential tool in the direct diagnosis of leptospirosis in dogs and can be particularly useful in diagnosing the disease in acute cases (Miotto et al. 2018b). Moreover, laboratory tests that are readily available in most veterinary clinics are blood count and serum biochemistry, which together provide information valuable to a diagnosis of leptospirosis, with emphasis on leukocytosis and liver and kidney alterations (Miotto et al. 2018b; Constantinescu et al. 2015).
Although canine leptospirosis has been well studied, difficulty still persists in clinicians making the diagnosis, due to financial factors limiting access to advanced laboratory methods, or due to the super acute course of the disease, which culminates in rapid death of affected animals. Therefore, to advance the diagnosis of leptospirosis, this study aimed to isolate and characterize leptospires from the blood of dogs with clinical signs suggestive of leptospirosis and to evaluate the results of serology, PCR, complete blood count, and serum biochemistry from hamsters experimentally infected with these bacteria.
Materials and methods
Animals and bacteriological culturing for Leptospira
During 2019 and 2020, blood from 22 dogs treated at a veterinary clinic in the metropolitan region of São Paulo, Brazil, was received for the isolation of leptospires. These animals were of both sexes and of different ages and breeds and had some of the clinical signs suggestive of leptospirosis, such as fever, anorexia, vomiting, jaundice, diarrhea, and/or abdominal pain. No information was available on any prior vaccination of the animals against leptospirosis.
Blood samples were collected in tubes containing the anticoagulant EDTA (ethylenediaminetetraacetic acid) and sent as soon as possible for bacterial isolation. Two to three drops of blood were seeded into liquid and semisolid EMJH medium (Ellighausen, McCullough, Johnson, and Harris) enriched 10% with improved supplement for Leptospira (Guedes et al. 2021b) added with 100 mg/L fluorouracil. The samples were incubated in a bacteriological incubator at 30 °C for 6 weeks, and the presence of leptospires was determined by observation under dark field microscopy and later confirmed by 16S PCR (Mérien et al. 1992).
Serological and molecular characterization of the isolates
The obtained isolates were submitted to serogrouping via a microscopic agglutination test (MAT) according to Faine et al. [9] using a panel of 26 polyclonal antisera against Leptospira spp., which represented 18 distinct serogroups. The serogroup was determined by detection of the highest titer found for any of the 26 antisera used.
For molecular characterization, extraction and purification of DNA from the isolates was performed using a PureLink® Genomic DNA Mini Kit (Invitrogen) following the manufacturer protocol.
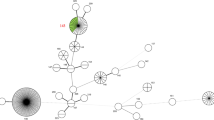
PCR was carried out using a primer pair that amplified a 549-bp region of the secY gene (Ahmed et al. 2006). The amplicons were sequenced by the Sanger method with BigDye Terminator v.3.1 chemistry (Applied Biosystems) and an ABI-3500 automatic sequencer (Applied Biosystems) according to the manufacturer’s instructions. The sequences were assembled using BioEdit Sequence Alignment Editor (Hall 1999). A phylogenetic tree was built using homologous sequences retrieved from the GenBank database (accession numbers in Fig. 1) with the neighbor-joining method, the Tamura-3-parameter model, and 1000 bootstrap replicates in MEGA 7 (Kumar et al. 2016).
For typing of isolates by multiple locus variable number tandem repeat analysis (MLVA) were used the five discriminatory markers for the VNTR loci 4, 7, 10, Lb4, and Lb5 for L. interrogans, L. kirschneri, and L. borgpetersenii according to Salaün et al. (2006).
Experimental infection in hamsters and hematobiochemical analyses
The isolates were grown in liquid EMJH medium until they reached a concentration of ~ 2 × 108 leptospires/ml. Then, a 0.5-ml sample was inoculated intraperitoneally into a golden Syrian hamsters (Mesocricetus auratus) weighing between 60 and 100 g. The animals were divided into groups considering the number of isolates, and each group consisted of two animals, one of which was used as a control (without infection).
The animals were kept under observation for a period of 7 days or until the manifestation of clinical signs of leptospirosis (bristly hair, prostration, photophobia, jaundice, dyspnea), and the presence of these symptoms was the determining factor for conducting euthanasia, thus avoiding prolonged stress and natural death. The hamsters were anesthetized, and blood collection was conducted for MAT, PCR, hematological, and biochemical analyses. Immediately after blood collection, euthanasia and necropsy of the animals were performed, and macroscopic lesions in organs were recorded when seen. Fragments of the liver, lung, and kidney were obtained and macerated in sterile Sorensen buffered saline (1:10) and submitted to PCR for detection of leptospiral DNA (Mérien et al. 1992).
The following hematological parameters were evaluated: hematocrit (HCT), hemoglobin (HGB), red blood cells (RBC), morphological evaluation of red blood cells, mean cellular volume (MCV), mean cellular hemoglobin (MCH), mean cellular hemoglobin concentration (MCHC), white blood cells (WBC), leukocyte differential, platelet count, and plasma protein (PP) (Hervey 2001; Trhall et al. 2015). Serum biochemistry parameters evaluated included urea, creatinine, alanine aminotransferase (ALT), and alkaline phosphatase (ALP) as indicators of renal and liver function.
The MAT was carried out applying a panel with 24 serovars of Leptospira spp. used in the routine serological diagnosis of leptospirosis by the laboratory (Supplementary material 1) (Faine et al. 1999).
All analyses were performed on the animals before inoculation (pre-infection) and after inoculation (post-infection).
Results
Of the 22 blood samples submitted for bacteriological culturing, three isolates were recovered (13.63%). Leptospire growth was seen only in semisolid EMJH medium and confirmed by 16S PCR. In the serogrouping test, all isolates were reactive only for the serogroup Icterohaemorrhagiae, with an observed titer of 12,800. Regarding molecular characterization, the isolates were assigned to the species Leptospira interrogans by DNA sequencing (GenBank accession numbers: OL408066, OL408067, and OL408068) (Fig. 1). The MLVA results showed that the VNTR loci 4, 7, and 10 profile of all the isolates was 2–1-7 compatible with serovars Copenhageni/ Icterohaemorrhagiae, including the Fiocruz L1-130 strain, autochthonous in Brazil (Jaeger et al. 2018).
The isolates were cataloged in the laboratory bank of national Leptospira isolates and coded as M24/19, M60/19, and M12/20.
With respect to experimental infection of hamsters, because the serological and molecular profiles of the isolates were identical, two strains were randomly selected for infection of the animals (M60/19 and M12/20). Infected animals began to show symptoms such as bristly hair, prostration, photophobia, jaundice, dyspnea, and conjunctivitis between the fifth and sixth days post-infection and were immediately euthanized, whereas the control group did not develop any clinical signs and were euthanized on the seventh day.
The laboratory test results pre- and post-infection are shown in Table 1. At necropsy of the infected animals, the macroscopic lesions observed included jaundice, marked discoloration of the kidneys and liver, and hemorrhages in the form of petechial in the lungs.
Discussion
In canine leptospirosis, the species L. interrogans and serogroups Canicola and Icterohaemorrhagiae are seen most often. This is likely because dogs are reported to be natural hosts of serogroup Canicola, whereas infection by Icterohaemorrhagiae occurs through contact with rodent urine, since these animals can harbor leptospires belonging to this serogroup (Levett 2001; Faine et al. 1999; Pinto et al. 2016; Santos et al. 2021). Nevertheless, dogs can be infected by other species such as L. borgpetersenii, L. kirschneri, and L. kmetyi (Rahman et al. 2021), involving a variety of serogroups that can cause high mortality (Koizumi et al. 2013) and even atypical serogroups for dogs such as Sejroe (Miotto et al. 2016).
Strains of Leptospira isolated from rodents and dogs in Brazil revealed identical characteristics themselves and to strain Fiocruz L1-130, characterized as L. interrogans serogroup Icterohaemorrhagiae. This sample may have exceptional zoonotic potential, considering that it is the major agent of human leptospirosis in the country and may be widely distributed in Brazil in different hosts (Jaeger et al. 2018). The three strains isolated from dogs (M24/19, M60/19, and M12/20) had the VNTR profile 2–1-7, the same described for Fiocruz L1-130 strain, suggesting that they may also belong to the clonal subpopulation of this strain.
Golden Syrian hamsters (Mesocricetus auratus) are the most used animal model in studies of leptospirosis as they develop the acute phase of the disease, which can be interpreted as the severe form of leptospirosis (Haake 2006). In this study, experimental animals infected with isolated leptospires showed classic clinical symptoms of canine leptospirosis, highlighting prostration and jaundice. However, other clinical signs drew attention, such as dyspnea and conjunctivitis, which are not common, but they should not be neglected. This is reinforced by the observation of lesions in the lungs of infected hamsters, likewise the detection of bacteria in this organ. Conjunctivitis is also a genuine symptom in dogs with leptospirosis (Van de Maele et al. 2008).
The main change found in the blood analyses was intense leukocytosis due to neutrophilia and monocytosis, which is not unusual in infected dogs (Constantinescu et al. 2015; Miller et al. 2007; Aswathanarayanappa et al. 2019). Additionally, in the leukocyte differential, a moderate number of granulocytes with a ring-shaped nucleus were observed (data not shown), which represent newly released and not fully matured neutrophils in mice (Ostanin et al. 2012). This situation points to the presence of a left shift that can sometimes be present in the blood counts of animals with leptospirosis (Schuller et al. 2015).
In severe cases of leptospirosis in dogs, thrombocytopenia is one of the most common alterations seen in blood counts, which is also associated with signs of lung damage (Kohn et al. 2010). This change was not remarked in hamsters infected with the strains isolated in this research, a parameter that seems to be influenced by the infecting serogroup, since the serogroup Pomona may be more related to thrombocytopenia than other serogroups (Goldstein et al. 2006). The pathological behavior of different serogroups of L. interrogans was reported in experimental infection in Wistar rats, which showed changes in different organs independently (Tonin et al. 2012). The serogroup Icterohaemorrhagiae can also cause thrombocytopenia (Andrade et al. 2020), which does not exclude the hypothesis of regional variability in strains influenced by climatic and environmental issues (Plank and Dean 2020).
Increased values of kidney and liver parameters in serum biochemistry analysis are a principal predictor of infection by Leptospira in dogs (Miotto et al. 2018a; Constantinescu et al. 2015; Hsu et al. 2018). In the present study, a significant increase in the values of urea and creatinine was discerned, which, together with the macroscopic lesions seen in the kidneys of infected hamsters, indicates the capacity of these strains to cause kidney injury. The serogroup Icterohaemorrhagiae has been described as a cause of considerable azotemia in dogs with acute leptospirosis (Paz et al. 2021). A factor that must be emphasized was the elevation in the enzyme alkaline phosphatase (ALP), indicating liver damage due to bacterial endotoxins, including a possible cholestatic disease (Constantinescu et al. 2015; Miller et al. 2007).
Concerning MAT, a titer of 800 was detected only for the serogroup Icterohaemorrhagiae. This confirms the serogroup to which the sample belongs. A titer ≥ 800 observed in MAT in a single serum sample can be considered for the diagnosis of the disease in dogs, especially when clinical signs are manifested (Geisen et al. 2007; Santos et al. 2021). However, MAT-negative samples can occur in early cases of the disease, and the antibodies can be detected only after the first sample is analyzed (Geisen et al. 2007). This is a limitation of this technique, as it is recommended to analyze paired samples with an approximate interval of 10 to 14 days; nonetheless, some patients may die even before seroconversion occurs (Levett 2001). This was a worrying condition; afterward, the leptospires recovered in this research were from dogs that died about 3 days after entering the hospital (data not shown).
PCR results showed that infected hamsters were in the acute phase of the disease, when leptospires were detected in the liver and blood. Interestingly, the kidneys were also positive, which means that the elimination of bacteria in the urine occurred since the acute phase. Urine should also be analyzed as a material of choice for the molecular diagnosis of leptospirosis in dogs in the acute phase because it may reveal positive animals that were negative in blood and serology by MAT (Santos et al. 2021).
Conclusion
Leptospires isolated from the blood of dogs suggest that these animals were infected with strains maintained by rodents. In addition, the possibility of these samples belonging to a clonal subpopulation of an autochthonous strain in Brazil responsible for cases of human leptospirosis in the country is not ruled out. Consequently, dogs can be victims just like humans in the urban environment. The routine laboratory tests used indicated critical changes, which, when analyzed together, can help veterinarians diagnose leptospirosis early.
References
Ahmed N, Devi SM, Valverde MDLA, Vijayachari P, Machang RS, Ellis WA, Hartskeerl RA (2006) Multilocus sequence typing method for identification and genotypic classification of pathogenic Leptospira species. Ann Clin Microbiol Antimicrob. https://doi.org/10.1186/1476-0711-5-28
Ambily R, Mini M, Siju J, Vamshikrishna S, Abhinay G, Gleeja VL, Sunanda C (2019) Utility of recombinant LipL41 based IgM and IgG ELISA in diagnosis of canine leptospirosis in an endemic area - a study from Kerala, India. Trop Biomed 36:654–663
Andrade MC, Oliveira LB, Santos AF, Moreira MVL, Pierezan F, Ecco R (2020) Differential diagnoses in 83 dogs with icterus. Pesq Vet Bras 40:451–465. https://doi.org/10.1590/1678-5150-PVB-6482
Aswathanarayanappa V, Kamal H, Shankaregowda, Shivakumar V, Vismitha V (2019) Haematobiochemical alterations in canine leptospirosis in Hassan: retrospective study. J Pharm Innov 8: 466–467. https://doi.org/10.13140/RG.2.2.20284.59529
Constantinescu R, Crivineanu V, Goran G, Danes D, Togoe D, Codreanu MD (2015) Correlation between clinical signs and different laboratory investigations in dogs diagnosed with leptospirosis. Scientific Works. Series C Vet Medic LXI 2015
Faine S, Adler B, Bolin C, Perolat P (1999) Leptospira and leptospirosis, second ed, Medi Sci Melbourne
Geisen V, Stengel C, Brem S, Múller W, Greenez C, Hartmann K (2007) Canine leptospirosis infections – clinical signs and outcome with different suspected Leptospira serogroups (42 cases). J Small Anim Pract 48:324–328. https://doi.org/10.1111/j.1748-5827.2007.00324.x
Goldstein RE, Lin RC, Langston CE, Scrivani PV, Erb HN, Barr SC (2006) Influence of infecting serogroup on clinical features of leptospirosis in dogs. J Vet Intern Med 20:489–494
Gomes-Solecki M, Santecchia I, Werts C (2017) Animal models of leptospirosis: of mice and hamsters. Front Immunol 58:1–20. https://doi.org/10.3389/fimmu.2017.00058
Guedes IB, Souza OS, Castro JFP, Cavalini MB, Souza Filho AFSF, Heinemann MB (2021a) Usefulness of the ranking technique in the microscopic agglutination test (MAT) to predict the most likely infecting serogroup of Leptospira. Front Vet Sci. https://doi.org/10.3389/fvets.2021.654034
Guedes IB, Souza OS, Castro JFP, Cavalini MB, Souza Filho AF, Aizawa J, Cortez A, Heinemann MB (2021b) Improvement of the enrichment used in the EMJH medium (Ellinghausen-McCullough-Johnson-Harris) for the cultivation of Leptospira spp. Rev Argent Microbiol. https://doi.org/10.1016/j.ram.2021.03.002
Guedes IB, Souza OS, Rocha KS, Cavalini MB, Damasceno Neto MS, Castro JFP, Souza Filho AF, Negrão MP, Cortez A, Moraes CCG, Heinemann MB (2021c) Leptospira strains isolated from cattle in the Amazon region, Brazil, evidence of a variety of species and serogroups with a high frequency of the Sejroe serogroup. Comp Immunol Microbiol Infect Dis. https://doi.org/10.1016/j.cimid.2020.101579
Haake DA (2006) Hamster Model of Leptospirosis. Curr Protoc Microbiol. https://doi.org/10.1002/9780471729259.mc12e02s02
Hall TA (1999) BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic Acids Symp Ser 41:95–98
Heatley JJ, Harris MC (2009) Hamsters and gerbils. In: Mitchell, Tully, (Eds.), Manual of exotic pet practice Elsevier
Hervey JW (2001) Atlas of veterinary hematology – blood and bone marrow of domestic animals, 1st edn. Sounder Elsevier, Philadelphia
Hsu CH, Liu IL, Liu CC, Liu BH, Pan MJ, Lin CS (2018) Seroepidemiologic survey of canine leptospirosis in northern Taiwan during 2008–2015. Taiwan Vet J 44:141–149. https://doi.org/10.1142/S1682648518500038
Jaeger LH, Pestana CP, Carvalho-Costa FA, Medeiros MA, Lilenbaum W (2018) Characterization of the clonal subpopulation Fiocruz L1–130 of Leptospira interrogans in rats and dogs from Brazil. J Med Microbiol 67:1361–1367. https://doi.org/10.1099/jmm.0.000806
Klassen HE, Adler B (2015) Recent advances in canine leptospirosis: focus on vaccine development. Vet Med (auckl) 6:245–260. https://doi.org/10.2147/VMRR.S59521
Kohn B, Steinicke K, Arndt G, Gruber AD, Guerra B, Jansen A et al (2010) Pulmonary abnormalities in dogs with leptospirosis. J Vet Intern Med 24:1277–1282. https://doi.org/10.1111/j.1939-1676.2010.0585.x
Koizumi N, Muto MM, Akachi S, Okano S, Yamamoto S, Horikawa K et al (2013) Molecular and serological investigation of Leptospira and leptospirosis in dogs in Japan. J Med Microbiol 62:630–636. https://doi.org/10.1099/jmm.0.050039-0
Kumar S, Stecher G, Tamura K (2016) MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol Biol Evol 33:1870–1874. https://doi.org/10.1093/molbev/msw054
Levett PN (2001) Leptospirosis. Clin Microbiol Rev 14:296–326. https://doi.org/10.1128/CMR.14.2.296-326.2001
Mérien F, Amouriaux P, Perolat P, Baranton G, Saint-Girons I (1992) Polymerase chain reaction goes detection of Leptospira spp. in clinical samples. J Clin Microbiol 30:2219–2224
Miller RI, Ross SP, Sullivan ND, Perkins NR (2007) Clinical and epidemiological features of canine leptospirosis in North Queensland. Aust Vet J 85:13–19. https://doi.org/10.1111/j.1751-0813.2006.00089.x
Miotto BA, Moreno LZ, Guilloux AGA, Sousa GO, Loureiro AP, Moreno AM, Lilenbaum W, Vasconcellos SA, Heinemann MB, Hagiwara MK (2016) Molecular and serological characterization of the first Leptospira santarosai strain isolated from a dog. Acta Trop 162:1–4. https://doi.org/10.1016/j.actatropica.2016.06.007
Miotto BA, Moreno LZ, Guilloux AGA, Tozzi BF, Moreno LZ, Hora AS et al (2018a) Prospective study of canine leptospirosis in shelter and stray dog populations: identification of chronic carriers and different Leptospira species infecting dogs. PLoS ONE 13:1–23. https://doi.org/10.1371/journal.pone.0200384
Miotto BA, Tozzi BF, Penteado MS, Guilloux AGA, Moreno LZ, Heinemann MB, Souza Filho AF, Lilenbaum W, Hagiwara MK (2018b) Diagnosis of acute canine leptospirosis using multiple laboratory tests and characterization of the isolated strains. BMC Vet Res 14:1–9. https://doi.org/10.1186/s12917-018-1547-4
Ostanin DV, Kurmaeva E, Furr K, Bao R, Hoffman J, Berney S, Grisham MB (2012) Acquisition of antigen-presenting functions by neutrophils isolated from mice with chronic colitis. J Immunol 188:1491–1502. https://doi.org/10.4049/jimmunol.1102296
Paz LN, Dias CS, Almeida DS, Balassiano IT, Medeiros MA, Costa F, Silva DN, Reis JN, Estrela-Lima A, Hamond C, Pinna MH (2021) Multidisciplinary approach in the diagnosis of acute leptospirosis in dogs naturally infected by Leptospira interrogans serogroup Icterohaemorrhagiae: a prospective study. Comp Immunol Microbiol Infect Dis. https://doi.org/10.1016/j.cimid.2021.101664
Pinto PS, Libonati H, Lilenbaum W (2016) A systematic review of leptospirosis on dogs, pigs, and horses in Latin America. Trop Anim Health Prod. https://doi.org/10.1007/s11250-016-1201-8
Plank R, Dean D (2020) Overview of the epidemiology, microbiology, and pathogenesis of Leptospira spp. in humans. Microbes Infect 2:1265–1276. https://doi.org/10.1016/S1286-4579(00)01280-6
Rahman SA, Khor KH, Khairani-Bejo S, Lau SF, Mazlan M, Roslan A, Goh SH (2021) Detection and characterization of Leptospira spp. in dogs diagnosed with kidney and/or liver disease in Selangor. Malaysia J Vet Diagn Invest 33:834–843. https://doi.org/10.1177/10406387211024575
Salaün L, Mérien F, Gurianova S, Baranton G, Picardeau M (2006) Application of multilocus variable-number tandem-repeat analysis for molecular typing of the agent of leptospirosis. J Clin Microbiol 44:3954–3962. https://doi.org/10.1128/JCM.00336-06
Santos CM, Dias GCRS, Saldanha AVP, Esteves SB, Cortez A, Guedes IB, Heinemann MB, Gonçales AP, Miotto BA (2021) Molecular and serological characterization of pathogenic Leptospira spp. isolated from symptomatic dogs in a highly endemic area. Brazil BMC Vet Res 17:1–11. https://doi.org/10.1186/s12917-021-02930-w
Schuller S, Francey T, Hartmann K, Hugonnard M, Kohn B, Nally JE, Sykes J (2015) European consensus statement on leptospirosis in dogs and cats. J Small Anim Pract 56:159–179. https://doi.org/10.1111/jsap.12328
Silva RF, Riedemann S (2007) Seroprevalencia de leptospirosis canina en perros atendidos en clínicas veterinarias, mediante aglutinación microscópica y comparación con las técnicas de aislamiento e inmunofluorescencia indirect. Arch Med Vet 39:269–274. https://doi.org/10.4067/S0301732X2007000300011
Tonin AA, Da Silva AS, De Azevedo MI, França RT, Paim FC, Schaefer PC, Martins JLR, Badke MRT, Lopes STA (2012) Hematologic and biochemical alterations in Wistar rats experimentally infected by Leptospira interrogans. Com Clin Pathol 21:833–838. https://doi.org/10.1007/s00580-011-1186-7
Trhall AM, Weiser G, Allison RW, Campbell TW (2015) Hematologia e bioquímica – clínica veterinária. 2, nd. Guanabara Koogan, Rio de Janeiro
Van de Maele I, Claus A, Haesebrouck F, Daminet S (2008) Leptospirosis in dogs: a review with emphasis on clinical aspects. Vet Rec 163:409–413. https://doi.org/10.1136/vr.163.14.409
Zarantonelli L, Suanes A, Meny P, Buroni F, Nieves C, Salaberry X, Briano C, Ashfield N, Silveira CS, Dutra F, Easton C, Fraga M, Giannitti F, Hamond C, Macías-Rioseco M, Menéndez C, Mortola A, Picardeau M, Quintero J, Ríos C, Rodríguez V, Romero A, Varela G, Rivero R, Schelotto F, Riet-Correa F, Buschiazzo A (2018) Isolation of pathogenic Leptospira strains from naturally infected cattle in Uruguay reveals high serovar diversity, and uncovers a relevant risk for human leptospirosis. PLoS Negl Trop Dis 12:1–22. https://doi.org/10.1371/journal.pntd.0006694
Acknowledgements
MBH acknowledges CNPq (Conselho Nacional de Desenvolvimento Científico e Tecnológico) for the fellowship (CNPq 309146/2017-8). IBG and JFPC acknowledge CAPES (Coordenação de Aperfeiçoamento de Pessoal de Nível Superior) for the scholarship. DBN acknowledges Fapesp (Fundação de Amparo à Pesquisa do Estado de São Paulo) for the fellowship (2020/16634-2).
Author information
Authors and Affiliations
Contributions
IG, JC, DN, and GS: conceptualization, methodology, investigation, and writing. AE and JI: performed the sample collection. MH: accurately reviewed the manuscript. All authors have read and approved the final version of the manuscript.
Corresponding author
Ethics declarations
We have read and have abided by the statement of ethical standards for manuscripts submitted.
Funding
This study was financed in part by CAPES, Brasil (Finance Code 001) by CNPq (420110/2018–6), and by Fapesp (2019/09896–3).
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
This work was approved by the Ethics Committee on Animal Use of the School of Veterinary Medicine and Animal Science (Universidade de São Paulo) – CEUA/FMVZ nº 1,361,131,218.
Informed consent
None.
Consent for publication
All authors have approved the manuscript and agreed with publication.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Guedes, I.B., de Paula Castro, J.F., Estefanuto, A. et al. Characterization of Leptospira strains recovered from the blood of dogs and usefulness of laboratory tests in hamsters experimentally infected with these isolates. Comp Clin Pathol 32, 147–153 (2023). https://doi.org/10.1007/s00580-022-03424-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00580-022-03424-3