Abstract
Mycoheterotrophic plants (MHPs) rely on their mycorrhizal fungus for carbon and nutrient supply, thus a shift in mycobionts may play a crucial role in speciation. This study aims to explore the mycorrhizal diversity of two closely related and sympatric fully MHPs, Monotropastrum humile var. humile (Mhh) and M. humile var. glaberrimum (Mhg), and determine their mycorrhizal associations. A total of 1,108,710 and 1,119,071 ectomycorrhizal fungal reads were obtained from 31 Mhh and 31 Mhg, and these were finally assigned to 227 and 202 operational taxonomic units, respectively. Results show that sympatric Mhh and Mhg are predominantly associated with different fungal genera in Russulaceae. Mhh is consistently associated with members of Russula, whereas Mhg is associated with members of Lactarius. Associating with different mycobionts and limited sharing of fungal partners might reduce the competition and contribute to their coexistence. The ectomycorrhizal fungal communities are significantly different among the five forests in both Mhh and Mhg. The distinct mycorrhizal specificity between Mhh and Mhg suggests the possibility of different mycobionts triggered ecological speciation between sympatric species.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Mycoheterotrophic plants (MHPs) lack chlorophyll and depend on mycorrhizal fungi for carbon sources and nutrients (Leake 1994). These plants are categorized into three trophic groups: initial, partial, and fully mycoheterotrophic, based on their reliance on mycorrhizal fungi in their life cycle. The fully MHPs, in particular, depend on fungal partners throughout their lifecycle and are considered the closest symbionts of mycorrhizal fungi (Merckx 2013). Approximately 580 achlorophyllous species have been identified, constituting a small subset of land plants (Jacquemyn and Merckx 2019). The transition from autotrophy to full mycoheterotrophy is estimated to have occurred independently at least 40 times during plant diversification (Jacquemyn and Merckx 2019).
The evolutionary trajectory of fully MHP is closely tied to their associated fungal partners (Ogura-Tsujita et al. 2012; Yamato et al. 2014). The adaptation and specialization of mycorrhizal fungi may play a role in the diversification or speciation of MHPs (Jacquemyn et al. 2023). Fully MHPs often form connections with phylogenetically restricted fungi, compared to their autotrophic relatives (Bidartondo and Bruns 2001; Grubisha et al. 2014; Dowie et al. 2017).
The subfamily Monotropoideae (family Ericaceae) is distributed throughout the Northern Hemisphere and is notable for its fully mycoheterotrophic characteristics (Bidartondo and Bruns 2001, 2002, 2005). Members within Monotropoideae, are associated with phylogenetically restricted fungi, such as Russulaceae, Tricholoma, and Rhizopogon (Bidartondo and Bruns 2001; Grubisha et al. 2014; Dowie et al. 2017). Phylogenetic congruence between Monotropoideae and ectomycorrhizal (ECM) fungi has been observed, indicating complex relationships between Monotropoideae plants and their fungal partners (Yang and Pfister 2006). Each of the five related plant lineages within the Monotropoideae is specialized to one of five distantly related ECM fungal lineages (Bidartondo and Bruns 2001, 2002, 2005), suggesting intricate and multifaceted interactions. Additionally, a shift in fungal partners may play a crucial role in the evolution and diversification of MHPs (Hynson and Bruns 2009; Ogura-Tsujita et al. 2012; Yamato et al. 2014; Suetsugu et al. 2020; Jacquemyn et al. 2023).
Monotropastrum humile often classified into two varieties, var. humile and var. glaberrimum (Hara 1965), exhibits distinct distribution patterns. M. humile var. humile (Mhh) is distributed in eastern Asia, from the Himalayas to Japan (Wallace 1975), while M. humile var. glaberrimum (Mhg) is only found in Taiwan and China. Chou and Zhou (1990) and Hsu et al. (1998) treated these varieties as different species. Mhg can be easily distinguished from Mhh by the absence of trichomes on the floral organs and differences in the shape and color of the floral disc (Tsukaya et al. 2008). Phylogenetic analysis based on ITS2 sequences shows that the Mhh formed a monophyletic group. However, Mhg was not the sister taxon of Mhh, but formed a monophyletic group with Monotropa uniflora (Tsukaya et al. 2008). Tsukaya et al. (2008) suggested that Mhg should be considered a distinct species.
Some studies reveal the existence of two closely related varieties of M. humile that establish symbiotic relationships with distinct ECM fungal families. Mhh forms associations with fungi from the Russulaceae family (Bidartondo and Bruns 2001), whereas Mhg is connected with the Thelephoraceae family (Yokoyama et al. 2005). Notably, the symbiosis between Mhg and its affiliated ECM fungi marks a significant deviation in the pattern of mycorrhizal associations observed within the subfamily Monotropoideae (Yokoyama et al. 2005). However, these differences have been observed in limited samples (2 Mhg and 1 Mhh) within a single site (Yokoyama et al. 2005), without considering the spatial structure in the associated fungal community.
Microscopic characteristics of the mycorrhizal fungal sheath of Mhh were examined by Yamada et al. (2008). They categorized 78 samples of adult M. humile var. humile individuals into 37 root mycorrhizal morphotypes, with 24 types identified as Russula or Lactarius fungal taxa within the Russulaceae family. However, the remaining 13 types were left unidentified, suggesting the potential association of non-Russulaceae fungi with Mhh. While microscopic evidence is valuable for assessing these ECM fungal mycorrhizal formations, the limited characteristics make it challenging to comprehensively understand their ECM fungal diversity. This limitation may result in underestimating ECM fungi that deviate from the classic mycorrhizal structure and exhibit non-dominant colonization patterns.
Over the past decades, advancements in culture-independent approaches, particularly high-throughput sequencing, have significantly expanded our comprehension of the global diversity of root mycobionts. These cutting-edge techniques have provided a more comprehensive understanding of the intricate relationships between plants and mycorrhizal fungi. This progress underscores the need to revisit and reevaluate previous findings, especially in the context of mycorrhizal associations. In Taiwan, Mhh and Mhg often coexist in the same geographic area, providing a valuable model to investigate the mycorrhizal fungal community of these closely related MHPs in sympatry.
This study aims to assess the diversity of mycorrhizal fungi associated with Mhh and Mhg in Taiwan. Five sites were selected to identify variations in fungal preferences of these co-occurring MHPs by addressing the following research questions: (1) What is the identity and structure of the fungal communities associated with two closely related MHPs? (2) Do the co-occurring Mhh and Mhg overlap in their respective ECM fungi within their roots? (3) Do populations of Mhh or Mhg from different areas host distinct ECM fungal communities?
Materials and methods
Study site and sampling procedure
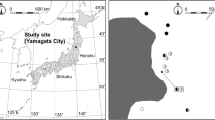
Throughout the flowering periods of MHP from 2017 to 2020, we collected samples of Mhh and Mhg from five locations across Taiwan, including Jailishan (JL), Henglingshan (HL), Hehuanshan (HH), Sun Link Sea (SLS), and Jinshuiying (JS). These sites are situated at elevations ranging from 1,430 to 3,030 m above sea level. The diverse habitats sampled include broadleaved forests with ectomycorrhizal (ECM) trees from the Fagaceae family, arbuscular mycorrhizal (AM) trees from the Symplocaceae family, and ericoid mycorrhizal (ERM) trees from the Ericaceae family; mixed conifer-broadleaved forests with ECM trees (Fagaceae and Pinaceae), AM trees (Lauraceae and Cupressaceae), and ERM trees (Ericaceae); and conifer forests dominated by Abies kawakamii and Tsuga chinensis, as detailed in Table 1. In the course of our fieldwork, individual plants were identified morphologically as one of the two MHP varieties. Five to nine individuals per MHP variety were sampled in each site, with a distance of more than one meter between specimens to prevent sampling the same plant population (Matsuda et al. 2011). The distance between MHPs at the HH site ranged from 30.9 to 234.1 m; at the HL site, the range was from 7.8 to 1,241.6 m; the distance at the JL site spanned from 8.5 to 62.9 m; at the JS site, it varied from 17.3 to 419.5 m; and, finally, at the SLS site, the MHPs were separated by distances ranging from 36.8 to 239.8 m.
A total of 31 Mhh and 31 Mhg individuals were collected. All samples were stored at 4 °C and processed within 48 h for DNA extraction. To confirm the taxonomic identity of these two MHPs, we meticulously selected multiple samples from both Mhh and Mhg. These samples were subjected to ITS sequence amplification and subsequent alignment with ITS sequences of other monotropoids, which are available in the GenBank international DNA database (Bidartondo and Bruns 2001).
Root Processing
The root structure within the rootball of two varieties of Monotropastrum humile differs significantly from that of autotrophic plants (Fig. S1). This distinction allows us to differentiate them and avoid mistakenly taking them based on their root morphology. The rootball was carefully rinsed in tap water using forceps to eliminate soil particles, tree roots, and debris. For each MHP specimen, we randomly selected at least 15 root tips (approximately 1 cm in length, totaling around 1 gram), spanning from the inner to the outer sections of the rootball. This approach was adopted to guarantee a thorough and representative sample.
To eliminate any surface contaminants or non-mycorrhizal fungi present, the mycorrhizal roots were subjected to a surface sterilization procedure. This involved the use of a 1% sodium hypochlorite (NaClO) solution, followed by thorough rinses in sterile deionized water (Suetsugu et al. 2021). This step is crucial for ensuring that the observed mycorrhizal associations are not influenced by external microbial or fungal populations.
DNA extraction, PCR, and high-throughput sequencing
Each root sample was pooled with 15 root tips and extracted DNA using the cetyltrimethylammonium bromide (CTAB) method (Doyle and Doyle 1987). DNA purity and concentration were determined with a NanoDrop spectrophotometer. Fungal-specific primers ITS1F (Gardes and Bruns 1993) and ITS2 (White et al. 1990) were used to amplify the nuclear internal transcribed spacer region (Boeraeve et al. 2018; Truong et al. 2019). PCR was performed using FastStart polymerase buffer and Taq DNA Polymerase (Roche, Germany) with the following PCR conditions, an initial denaturation for 5 min at 95°C, followed by 35 cycles of 30 sec at 95°C, 55°C and 72°C, and a final extension for 5 min at 72°C. Secondary PCR was performed using the following forward (5’-[8-mer NS]-[ITS1F]-3’) and reverse (5’-[8-mer NS]-[ITS2]-3’) sequencing primers fused with an 8-mer barcode as follows: an initial denaturation for 3 min at 95 °C, followed by 5 cycles of 20 s at 98 °C, 57.5 °C and 72 °C; and a final extension for 3 min at 72 °C. The Illumina MiSeq platform processed the high-throughput sequencing libraries (Tri-I Biotech Inc, Taiwan).
Bioinformatics
Raw data underwent quality control analysis using CLC Genomics Workbench v10. Mothur v. 1.35.1 was employed to screen reads with a Phred score > 20, aligning quality-filtered sequences against the UNITE database. Bayesian classifier with the UNITE training database was utilized to classify sequences, and USEARCH 7.0 clustered these sequences into OTUs at 97% similarity. OTU sequences comprising less than 0.05% of the total reads in any sample were removed (Mujic et al., 2023). Taxonomy assignment at the generic level (identity > 95%), family level (identity > 90%), and order level (identity > 80%) was performed (Tedersoo et al. 2015; Nilsson et al. 2019). The functional group of fungal OTUs was recognized using FUNGuild (Nguyen et al. 2016) and research articles (Rinaldi et al. 2008; Tedersoo and Smith 2013; Tedersoo et al. 2014). The ECM guild taxa were used for further analysis.
Statistical analysis
Sequencing depth adequacy was measured by rarefaction analysis using Past 4 software version 4.03 (Hammer et al. 2001). R software version 4.1.2 was used for statistical procedures. T-tests and ANOVA compared alpha diversity between MHPs and among sites, respectively. The GUniFrac package normalized reads for each sample (Chen and Zhang 2021). Multidimensional scaling (MDS) plot was used to visualize the ECM fungal community pattern based on Hellinger-based distances (Pérez-Izquierdo et al. 2020) by Primer 6 software (Clarke et al. 2014). PERMANOVA tested the difference in ECM fungal communities between two MHPs or among sites. Venn diagram was generated using InteractiVenn (http://www.interactivenn.net) (Heberle et al. 2015).
Results
Overall fungal community composition
A total of 62 DNA samples from plant roots were amplified and sequenced, with the effective tags generated by high-throughput sequencing aggregated at 97% sequence similarity, yielding 1,942 fungal OTUs (2,861,545 sequencing reads). After analysis, 77.9% of the sequences (2,227,781 sequencing reads, 408 OTUs) were assigned to putative ECM fungi (Table S1). Notably, the ECM guild represented the predominant fungal group associated with the MHPs (Table S2, Table S3). The rarefaction curves (Fig. S2) constructed for these samples indicated that the OTU diversity reached near-saturation, suggesting that our sequencing depth captured the majority of the fungal diversity present in each sample.
ECM fungal diversity associated with M. humile var. humile and M. humile var. glaberrimum
The totals of ECM fungal OTUs in 31 samples of Monotropastrum humile var. humile (Mhh) and 31 samples of M. humile var. glaberrimum (Mhg) were 227 (with 1,108,710 sequencing reads) and 202 (with 1,119,071 reads), respectively. Among these, only 21 ECM OTUs were communal between the two MHPs. A comparison of OTU richness showed that Mhh and Mhg had a similar number of ECM fungal OTUs, with no significant difference (t = -0.67057, p = 0.5051) in the number of ECM fungal OTU between Mhh (13.1 ± 7.9; mean ± SD) and Mhg (11.7 ± 7.7; mean ± SD). On average, there were 35,764.8 ± 9,881.6 reads and 36,099.1 ± 10,735.1 reads per individual associated with Mhh and Mhg, respectively (t = 0.12754, p = 0.8989). Table S2 compared ECM fungal OTU numbers detected in Mhh and Mhg from the five study sites. The number of ECM fungal OTUs of Mhh ranged from 8.2 ± 3.6 to 18.4 ± 14, which was not significantly different among the five sites (ANOVA, F = 1.42, p = 0.255). In contrast, the number of ECM fungal OTUs of Mhg ranged from 6.8 ± 4.4 to 21.6 ± 4.9, significantly different among the five sites (ANOVA, F = 4.529, p = 0.007). The highest ECM fungal OTUs were detected in the HL site (Table S2).
ECM fungal composition among MHP varieties and sites
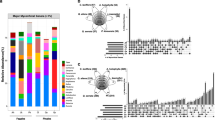
To examine whether the ECM fungal OTU composition varied between two MHP varieties, ECM fungal OTU matrices were selected for MDS and PERMANOVA. The OTU-level MDS plot illustrated distinct ECM fungal communities between Mhh and Mhg (Fig. 1), confirmed by PERMANOVA (pseudo-F = 5.2018, p = 0.001). There were thirteen ECM fungal genera and one Boletaceae unclassified genus associated with Mhh. Members of Russula were most abundant accounting for 92.3% of total ECM guild reads, followed by Lactarius (3.4%), Sebacina (2.0%), and Chloridium (1.6%). Members of Amanita, Cortinarius, Elaphomyces, Entoloma, Inocybe, Phylloporus, Piloderma, Tomentella, Tuber, and one Boletaceae unclassified genus represented < 1% relative read abundance (Fig. 2). Fifteen ECM fungal genera were associated with Mhg. Members of Lactarius were most abundant accounting for 80.2%, followed by Russula (14.0%), Sebacina (2.2%), Elaphomyces (1.3%), and Chloridium (1.1%). Members of Amanita, Amaurodon, Cortinarius, Entoloma, Hydnum, Inocybe, Lactifluus, Piloderma, Thelephora and Tomentella represented < 1% relative read abundance (Fig. 2).
Variation in ECM fungal communities (OTU-level) associating with two MHPs. MDS graph displays variation in ECM fungal community composition between individuals of M. humile var. humile (Mhh) and M. humile var. glaberrimum (Mhg) sampled at Hehuanshan (HH), Jailishan (JL), Henglingshan (HL) Sun Link Sea (SLS) and Jinshuiying (JS)
Generic level of ECM fungal communities observed in the coexisting populations of M. humile var. humile (Mhh) and M. humile var. glaberrimum (Mhg). Bar charts representing the cumulative proportions of sequences belonging to different ectomycorrhizal fungal genera observed in the sampled Mhh and Mhg populations from Hehuanshan (HH), Jailishan (JL), Henglingshan (HL) Sun Link Sea (SLS) and Jinshuiying (JS)
The top 100 ECM fungal OTUs were used to compare the dominant ECM fungal composition and relative read abundance between two MHP varieties (Fig. 3). Each MHP was predominantly associated with one to four ECM fungal OTUs at each site. Two MHP varieties did not share dominant ECM fungal OTUs in three sites (HL, JL, and SLS; Fig. 3b, c and e). In contrast, five ECM fungal OTUs were shared between two MHP varieties in the HH site (Fig. 3a), and one ECM fungal OTU was detected in both MHP varieties in JS (Fig. 3d). These limited ECM fungal OTUs sharing between two MHPs had lower relative read abundance. In the HH site, Lactarius OTU00007 was detected in both roots of MHPs but was only predominantly associated with Mhg (Fig. 3a). At the OTU level, we also found two MHPs had differences in ECM fungal associations in the co-occurring site.
PERMANOVA test revealed that sampling sites significantly influenced the ECM fungal communities associated with MHPs (pseudo-F = 3.526, p = 0.001). MDS plot showed that ECM fungal communities associated with Mhh were grouped according to sampling sites (Fig. 4a) and the ANOSIM test show that the ECM fungal communities were significantly different among sites (p = 0.001). Only ECM fungal communities associated with Mhh from SLS and HL were not significantly different. The Venn diagram revealed that 11, 10, 5, 1, and 1 OTUs were shared between Mhh populations from SLS-JL, SLS-JS, SLS-HL, HH-JL, and HH-JS, respectively (Fig. 4c). Similar site effects were also found in ECM fungal communities associated with Mhg (Fig. 4b, p = 0.001). The Venn diagram revealed that 1 OTU was shared among Mhg populations from JL, SLS, and HL. Eleven, seven, four, and one OTUs were shared between Mhg populations from JS-JL, JL-SLS, HL-SLS, and SLS-JS, respectively (Fig. 4d).
Variation in ECM fungal communities (OTU-level) associating with Mhh (a) and Mhg (b) among the five sites. The MDS graph displayed variation in ECM fungal community composition between individuals of Mhh (a) and Mhg (b) sampled at five sites. A Venn diagram showing the number of ECM fungal OTUs shared among Mhh (c) or Mhg (d) populations from Hehuanshan (HH), Jailishan (JL), Henglingshan (HL) Sun Link Sea (SLS) and Jinshuiying (JS)
Discussion
This study investigated the ECM fungal community of monotropoid roots through Illumina sequencing of the ITS1 region. Our findings reveal a high ECM fungal diversity associated with Monotropastrum humile var. humile (Mhh) and M. humile var. glaberrimum (Mhg). The dominant association pattern emerged, with Mhh having a consistent link to Russulaceae fungi, particularly members of Russula, while Mhg preferred members of Lactarius. Additionally, our study also identified a few undominant ECM taxa, such as Sebacina, Chloridium, Elaphomyces and Thelephora associated with Mhh and Mhg. Roots of M. humile were found to have a highly specialized association with Russulaceae (Yokoyama et al. 2005; Yamada et al. 2008; Matsuda et al. 2011). However, Thelephoraceae (Yokoyama et al. 2005; Matsuda et al. 2011), Phellodon sp. and Gymnopilus aff. penetrans (Shen et al. 2012) were found in the rootball of M. humile. Contrary to prior research (Bidartondo and Bruns 2001; Yokoyama et al. 2005; Yamada et al. 2008; Matsuda et al. 2011; Shen et al. 2012), our findings suggest that MHPs can host a diverse array of ECM fungal species, with certain fungi potentially outcompeting others or being preferentially selected by the host, leading to the dominance of specific ECM fungi while others remain less abundant.
Our results align with prior research suggesting that plant phylogenetic constraints play a crucial role in shaping mycorrhizal communities in MHP species. Jacquemyn et al. (2011) and Xing et al. (2020) proposed that phylogenetic constraints influence the specificity levels of dominant mycorrhizal partners. Our study observed diversified mycobionts in the roots of Mhh and Mhg, with the changes in specificity levels (i.e., phylogenetic breadth) of dominant mycobionts influenced by phylogenetic constraints. Previous studies showed that plant lineages are specifically dependent on different lineages of fungi in the monotropoid mycorrhizal symbiosis and that MHP plants in the clades of Monotropastrum and Monotropa are constraint associated with Russulaceae fungi (Bidartondo and Bruns 2001). However, the identity of the fungal symbiotic partners of the Monotropastrum used limited samples. The present study has the largest sample, numerically, taxonomically and geographically.
The phylogenetic tree for the subfamily Monotropoideae was constructed and presented in Fig. S3. This phylogenetic tree indicates that M. humile var. humile (Mhh) samples form a monophyletic group. In contrast, M. humile var. glaberrimum (Mhg) samples form a monophyletic group closely related to Monotropa uniflora. This finding corroborated the study by Tsukaya et al. (2008), disclosing that these two M. humile varieties and Monotropa uniflora, despite belonging to different genera, form a monophyletic clade. Therefore, our results support the reclassification of Monotropastrum humile var. glaberrimum as a separate species.
Additionally, our research uncovers that Mhh and Mhg are associated with distinct dominant ECM fungi. Monotropastrum humile var. humile (Bidartondo and Bruns 2001; Suetsugu et al. 2023), Monotropastrum kirishimense (Suetsugu et al. 2023), and Monotropa uniflora (Kong et al. 2015) are predominantly associated with Russula fungi. Monotropastrum humile var. glaberrimum (Mhg) samples form a monophyletic group closely related to Monotropa uniflora (Fig. S3), and they preferred to be associated with members of Lactarius. Monotropa brittonii is closely related to Monotropa uniflora (Fig. S3) and Keesling et al. (2021) found that Monotropa brittonii is only associated with Lactifluus which is a member of the Russulaceae family. This demonstrates that the symbiotic associations between the MHPs and fungi are dictated by the plants’ phylogenetic relationships, with a predilection for specific ECM genera in the Russulaceae family. A thorough examination of the ECM community and taxonomies of these MHPs in the Monotropastrum-Monotropa clade is necessary to uncover their ECM association and shed light on this complicated evolutionary process.
Our results revealed that coexisting Mhh and Mhg are associated with distinct sets of fungal partners. This observation aligns with existing studies suggesting that co-occurring MHP species prefer distinct mycorrhizal fungi (Waterman et al. 2011; Jacquemyn et al. 2012, 2014). In sympatric habitats, MHP species tend to engage with different fungal partners, minimizing overlap, and reducing competition (Gomes et al. 2017). The limited sharing of fungal partners between Mhh and Mhg might reduce competition and allow coexistence.
The significant role of habitat in shaping ECM fungal communities associated with both Mhh and Mhg was observed in this study, thereby demonstrating the profound influence of site effects on such communities. Bidartondo and Bruns (2001) conducted an expansive investigation into the phylogenetic patterns of Ericaceae from North America and Eurasia, thereby inferring the existence of geographical patterns of specificity. Our research provides further substantiating findings, suggesting that Monotropastrum plant species have affiliations with local fungal partners that are area-specific. Although approximately 200 ECM fungal OTUs may establish associations with MHPs, typically only 12–13 OTUs emerge as dominant. Across different sites, 2–4 ECM fungal OTUs tend to dominate, with some regions hosting three dominant ECM fungi. For instance, HH site harbors unique autotrophic host plant species, resulting in a distinct ECM fungal pool. The Venn diagram illustrates that ECM fungi associated with Mhh/Mhg from the HH site do not overlap with those from the other four sites. The presence of varied ECM fungi in different habitats provides MHPs with ample colonization opportunities across diverse elevations and forest types. The composition of the ECM fungal pool is influenced by the forest type, thus impacting the associations between MHPs and ECM fungi.
Additionally, the selective symbiotic relationship between Mhh/Mhg and their fungal symbionts likely evolved over time. For example, Johansson et al. (2017) demonstrated that Hypopitys monotropa, during its seed germination stage, can form associations with various genera of ECM fungi, but as it progresses to later stages, it exhibits heightened mycorrhizal specificity. This specialization could potentially enhance the efficiency of carbon acquisition (Leake and Cameron 2010). While dominant ECM fungi may contribute to carbon acquisition efficiency, further research is required to fully elucidate this relationship.
Data availability
No datasets were generated or analysed during the current study.
References
Bidartondo MI, Bruns TD (2001) Extreme specificity in epiparasitic Monotropoideae (Ericaceae): widespread phylogenetic and geographical structure. Mol Ecol 10:2285–2295. https://doi.org/10.1046/j.1365-294X.2001.01358.x
Bidartondo MI, Bruns TD (2002) Fine-level mycorrhizal specificity in the Monotropoideae (Ericaceae): specificity for fungal species groups. Mol Ecol 11:557–569. https://doi.org/10.1046/j.0962-1083.2001.01443.x
Bidartondo MI, Bruns TD (2005) On the origins of extreme mycorrhizal specificity in the Monotropoideae (Ericaceae): performance trade-offs during seed germination and seedling development. Mol Ecol 14:1549–1560. https://doi.org/10.1111/j.1365-294X.2005.02503.x
Boeraeve M, Honnay O, Jacquemyn H (2018) Effects of host species, environmental filtering and forest age on community assembly of ectomycorrhizal fungi in fragmented forests. Fungal Ecol 36:89–98. https://doi.org/10.1016/j.funeco.2018.08.003
Chen J, Zhang X (2021) D-MANOVA: fast distance-based multivariate analysis of variance for large-scale microbiome association studies. Bioinformatics 38:286–288. https://doi.org/10.1093/bioinformatics/btab498
Chou Y-L, Zhou R-C (1990) Pyrolaceae. In: Fang WP, Hu WK (eds) Flora Reipublicae Popularis Sinica. Science, Beijing, pp 158–216
Clarke K, Gorley R, Somerfield P, Warwick R (2014) Change in marine communities: an approach to statistical analysis and interpretation. Primer-E Ltd, Plymouth
Dowie NJ, Grubisha LC, Burton BA et al (2017) Development of anonymous nuclear loci for Pterospora andromedea (Monotropoideae) using Illumina and Ion Torrent sequencing data. Conserv Genet Resour 9:371–373. https://doi.org/10.1007/s12686-017-0686-4
Doyle JJ, Doyle JL (1987) A rapid DNA isolation procedure for small quantities of fresh leaf tissue. Phytochem Bull 19:11–15
Gardes M, Bruns TD (1993) ITS primers with enhanced specificity for basidiomycetes - application to the identification of mycorrhizae and rusts. Mol Ecol 2:113–118. https://doi.org/10.1111/j.1365-294X.1993.tb00005.x
Gomes SI, Merckx VS, Saavedra S (2017) Fungal-host diversity among mycoheterotrophic plants increases proportionally to their fungal‐host overlap. Ecol Evol 7:3623–3630. https://doi.org/10.1002/ece3.2974
Grubisha LC, Dowie NJ, Miller SL et al (2014) Rhizopogon kretzerae sp. nov.: the rare fungal symbiont in the tripartite system with Pterospora andromedea and Pinus strobus. Botany 92:527–534. https://doi.org/10.1139/cjb-2013-0309
Hammer Ø, Harper DAT, Ryan PD (2001) PAST: paleontological statistics software package for education and data analysis
Hara H (1965) New or noteworthy flowering plants from Eastern Himalaya (4). Shokubutsu Kenkyu Zasshi 40:97–103. https://doi.org/10.51033/jjapbot.40_4_5239
Heberle H, Meirelles GV, da Silva FR et al (2015) InteractiVenn: a web-based tool for the analysis of sets through Venn diagrams. BMC Bioinform 16:169. https://doi.org/10.1186/s12859-015-0611-3
Hsu T-W, Kuoh C-S, Hsieh C-F (1998) Cheilotheca. In: Huang T-C, Boufford DE, Lowry PP, Ohashi H, Peng C-I (eds) Flora of Taiwan, Dept. Bot. National Taiwan Univ., Taipei, pp 5–6
Hynson NA, Bruns TD (2009) Evidence of a myco-heterotroph in the plant family Ericaceae that lacks mycorrhizal specificity. Proc Royal Soc B 276:4053–4059. https://doi.org/10.1098/rspb.2009.1190
Jacquemyn H, Merckx V (2019) Mycorrhizal symbioses and the evolution of trophic modes in plants. J Ecol 107:1567–1581. https://doi.org/10.1111/1365-2745.13165
Jacquemyn H, Merckx V, Brys R et al (2011) Analysis of network architecture reveals phylogenetic constraints on mycorrhizal specificity in the genus Orchis (Orchidaceae). New Phytol 192:518–528. https://doi.org/10.1111/j.1469-8137.2011.03796.x
Jacquemyn H, Deja A, De hert K et al (2012) Variation in mycorrhizal associations with tulasnelloid fungi among populations of five Dactylorhiza species. PLoS ONE 7:e42212. https://doi.org/10.1371/journal.pone.0042212
Jacquemyn H, Brys R, Merckx VSFT et al (2014) Coexisting orchid species have distinct mycorrhizal communities and display strong spatial segregation. New Phytol 202:616–627. https://doi.org/10.1111/nph.12640
Jacquemyn H, Suetsugu K, Merckx V (2023) The role of mycorrhizal fungi in driving ecotype formation in mycoheterotrophic plants. Nord J Bot e 04083. https://doi.org/10.1111/njb.04083
Johansson VA, Bahram M, Tedersoo L, Kõljalg U, Eriksson O (2017) Specificity of fungal associations of Pyroleae and Monotropa hypopitys during germination and seedling development. Mol Ecol 26:2591–2604. https://doi.org/10.1111/mec.14050
Keesling AR, Broe MB, Freudenstein JV (2021) Reevaluating the species status of the southern ghost pipe, Monotropa brittonii (Ericaceae). Syst Bot 46:1067–1079. https://doi.org/10.1600/036364421X16370109698722
Kong A, Cifuentes J, Estrada-Torres A et al (2015) Russulaceae associated with mycoheterotroph Monotropa uniflora (Ericaceae) in Tlaxcala, Mexico: a phylogenetic approach. Cryptogam Mycol 36:479–512. https://doi.org/10.7872/crym/v36.iss4.2015.479
Leake JR (1994) The biology of myco-heterotrophic (‘saprophytic’) plants. New Phytol 127:171–216. https://doi.org/10.1111/j.1469-8137.1994.tb04272.x
Leake JR, Cameron DD (2010) Physiological ecology of mycoheterotrophy. New Phytol 185:601–605. https://www.jstor.org/stable/25609647
Matsuda Y, Okochi S, Katayama T et al (2011) Mycorrhizal fungi associated with Monotropastrum humile (Ericaceae) in central Japan. Mycorrhiza 21:569–576. https://doi.org/10.1007/s00572-011-0365-3
Merckx VSFT (2013) Mycoheterotrophy: an introduction. In: Merckx V (ed) Mycoheterotrophy. Springer, New York, pp 1–17
Mujic AB, Policelli N, Nuñez MA, Truong C, Smith ME (2023) Co-invasive ectomycorrhizal fungi alter native soil fungal communities. Plant Soil 484:547–567. https://doi.org/10.1007/s11104-022-05820-8
Nguyen NH, Song Z, Bates ST et al (2016) FUNGuild: an open annotation tool for parsing fungal community datasets by ecological guild. Fungal Ecol 20:241–248. https://doi.org/10.1016/j.funeco.2015.06.006
Nilsson RH, Larsson K-H, Taylor AFS et al (2019) The UNITE database for molecular identification of fungi: handling dark taxa and parallel taxonomic classifications. Nucleic Acids Res 47:D259–D264. https://doi.org/10.1093/nar/gky1022
Ogura-Tsujita Y, Yokoyama J, Miyoshi K, Yukawa T (2012) Shifts in mycorrhizal fungi during the evolution of autotrophy to mycoheterotrophy in Cymbidium (Orchidaceae). Am J Bot 99:1158–1176. https://doi.org/10.3732/ajb.1100464
Pérez-Izquierdo L, Zabal-Aguirre M, Verdú M et al (2020) Ectomycorrhizal fungal diversity decreases in Mediterranean pine forests adapted to recurrent fires. Mol Ecol 29:2463–2476. https://doi.org/10.1111/mec.15493
Rinaldi AC, Comandini O, Kuyper TW (2008) Ectomycorrhizal fungal diversity: seperating the wheat from the chaff. Fungal Divers 33:1–45
Shen M, Zhang C-Q, Ma Y-P et al (2012) Mycorrhizal features and fungal partners of four mycoheterotrophic Monotropoideae (Ericaceae) species from Yunnan, China. Symbiosis 57:1–13. https://doi.org/10.1007/s13199-012-0180-4
Suetsugu K, Matsubayashi J, Tayasu I (2020) Some mycoheterotrophic orchids depend on carbon from dead wood: novel evidence from a radiocarbon approach. New Phytol 227:1519–1529. https://doi.org/10.1111/nph.16409
Suetsugu K, Matsuoka S, Shutoh K et al (2021) Mycorrhizal communities of two closely related species, Pyrola subaphylla and P. japonica, with contrasting degrees of mycoheterotrophy in a sympatric habitat. Mycorrhiza 31:219–229. https://doi.org/10.1007/s00572-020-01002-5
Suetsugu K, Hirota SK, Hsu TC et al (2023) Monotropastrum kirishimense (Ericaceae), a new mycoheterotrophic plant from Japan based on multifaceted evidence. J Plant Res 136:3–18. https://doi.org/10.1007/s10265-022-01422-8
Tedersoo L, Smith ME (2013) Lineages of ectomycorrhizal fungi revisited: foraging strategies and novel lineages revealed by sequences from belowground. Fungal Biol Rev 27:83–99. https://doi.org/10.1016/j.fbr.2013.09.001
Tedersoo L, Bahram M, Põlme S et al (2014) Global diversity and geography of soil fungi. Science 346:1256688. https://doi.org/10.1126/science.1256688
Tedersoo L, Anslan S, Bahram M et al (2015) Shotgun metagenomes and multiple primer pair-barcode combinations of amplicons reveal biases in metabarcoding analyses of fungi. MycoKeys 10:1–43. https://doi.org/10.3897/mycokeys.10.4852
Truong C, Gabbarini LA, Corrales A et al (2019) Ectomycorrhizal fungi and soil enzymes exhibit contrasting patterns along elevation gradients in southern Patagonia. New Phytol 222:1936–1950. https://doi.org/10.1111/nph.15714
Tsukaya H, Yokoyama J, Imaichi R, Ohba H (2008) Taxonomic status of Monotropastrum humile, with special reference to M. humile var. glaberrimum (Ericaceae, Monotropoideae). J Plant Res 121:271–278. https://doi.org/10.1007/s10265-008-0157-9
Wallace G (1975) Studies of the Monotropoideae (Ericaceae) taxonomy and distribution. Wasmann J Biol 33:1–88
Waterman RJ, Bidartondo MI, Stofberg J et al (2011) The effects of above- and belowground mutualisms on orchid speciation and coexistence. Am Nat 177:E54–68. https://doi.org/10.1086/657955
White T, Bruns T, Lee S et al (1990) Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In: Pcr Protocols: a guide to methods and applications. pp 315–322
Xing X, Gao Y, Zhao Z et al (2020) Similarity in mycorrhizal communities associating with two widespread terrestrial orchids decays with distance. J Biogeogr 47:421–433. https://doi.org/10.1111/jbi.13728
Yamada A, Kitamura D, Setoguchi M et al (2008) Monotropastrum humile var. humile is associated with diverse ectomycorrhizal Russulaceae fungi in Japanese forests. Ecol Res 23:983–993. https://doi.org/10.1007/s11284-008-0463-7
Yamato M, Ogura-Tsujita Y, Takahashi H, Yukawa T (2014) Significant difference in mycorrhizal specificity between an autotrophic and its sister mycoheterotrophic plant species of Petrosaviaceae. J Plant Res 127:685–693. https://doi.org/10.1007/s10265-014-0661-z
Yang S, Pfister DH (2006) Monotropa uniflora plants of eastern Massachusetts form mycorrhizae with a diversity of russulacean fungi. Mycologia 98:535–540
Yokoyama J, Fukuda T, Tsukaya H (2005) Molecular identification of the mycorrhizal fungi of the epiparasitic plant Monotropastrum humile var. glaberrimum (Ericaceae). J Plant Res 118:53–56. https://doi.org/10.1007/s10265-004-0188-9
Acknowledgements
The work was supported by funding from the Ministry of Science and Technology, Taiwan (MOST 108-2621-B-080-001-MY2). The authors would like to express their gratitude to several organizations for granting permission to collect samples of two Monotropastrum taxa in Taiwan. The acknowledgments go to Taroko National Park Headquarters, Endemic Species Research Institute, Dongshih Forest District Office, Pingtung Forest District Office, and Sun-Link-Sea Resort. We also thank Dr. Yu-Ting Wu, Dr. Ko-Hsuan Chen for their critical reading, insightful discussions, and constructive comments greatly enhanced the quality of the research. This research was conducted by Ren-Cheng Liu as part of his doctoral dissertation, under the guidance and supervision of Dr. Wan-Rou Lin and Dr. Pi-Han Wang.
Funding
The work was supported by funding from the Ministry of Science and Technology, Taiwan (MOST 108-2621-B-080-001-MY2).
Author information
Authors and Affiliations
Contributions
R.C.L. conducted research, analyzed the data and contributed to writing the manuscript; P.H.W. designed the experiments, contributed to writing the manuscript and reviewed the manuscript; W.R.L. obtained funding, planned the experiments, analyzed the data, and wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Liu, RC., Lin, WR. & Wang, PH. Exploring mycorrhizal diversity in sympatric mycoheterotrophic plants: a comparative study of Monotropastrum humile var. humile and M. humile var. glaberrimum. Mycorrhiza 34, 283–292 (2024). https://doi.org/10.1007/s00572-024-01158-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00572-024-01158-4